Sex-Related Differences in Allelic Frequency of the Human Beta T Cell Receptor SNP rs1800907: A Retrospective Analysis from Milan Metropolitan Area
Abstract
1. Introduction
2. Materials and Methods
2.1. Studies Included for Data Aggregation
2.2. Study Participants
2.3. Allelic Frequency
- Forward primer: 5′-TAATTTTGAAATAAGGGAAGATGAC-3′
- Reverse primer: 5′-TTTTGTATCCACCCTATGGGTTGGC-3′
2.4. Statistical Analysis
3. Results
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Ober, C.; Loisel, D.A.; Gilad, Y. Sex-Specific Genetic Architecture of Human Disease. Nat. Rev. Genet. 2008, 9, 911–922. [Google Scholar] [CrossRef]
- Taneja, V. Sex Hormones Determine Immune Response. Front. Immunol. 2018, 9. [Google Scholar] [CrossRef]
- Lotter, H.; Altfeld, M. Sex Differences in Immunity. Semin. Immunopathol. 2019, 41, 133–135. [Google Scholar] [CrossRef]
- Mor, G.; Cardenas, I. The Immune System in Pregnancy: A Unique Complexity. Am. J. Reprod. Immunol. 2010, 63, 425–433. [Google Scholar] [CrossRef]
- Spagnolo, P.A.; Manson, J.E.; Joffe, H. Sex and Gender Differences in Health: What the COVID-19 Pandemic Can Teach Us. Ann. Intern. Med. 2020. [Google Scholar] [CrossRef] [PubMed]
- McCombe, P.A.; Greer, J.M.; Mackay, I.R. Sexual Dimorphism in Autoimmune Disease. Curr. Mol. Med. 2009, 9, 1058–1079. [Google Scholar] [CrossRef] [PubMed]
- Ngo, S.T.; Steyn, F.J.; McCombe, P.A. Gender Differences in Autoimmune Disease. Front. Neuroendocrinol. 2014, 35, 347–369. [Google Scholar] [CrossRef]
- Watson, C.T.; Para, A.E.; Lincoln, M.R.; Ramagopalan, S.V.; Orton, S.M.; Morrison, K.M.; Handunnetthi, L.; Handel, A.E.; Chao, M.J.; Morahan, J.; et al. Revisiting the T-Cell Receptor Alpha/Delta Locus and Possible Associations with Multiple Sclerosis. Genes Immun. 2011, 12, 59–66. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Khan, D.; Ansar Ahmed, S. The Immune System Is a Natural Target for Estrogen Action: Opposing Effects of Estrogen in Two Prototypical Autoimmune Diseases. Front. Immunol. 2016, 6. [Google Scholar] [CrossRef]
- Subrahmanyan, L.; Eberle, M.A.; Clark, A.G.; Kruglyak, L.; Nickerson, D.A. Sequence Variation and Linkage Disequilibrium in the Human T-Cell Receptor Beta (TCRB) Locus. Am. J. Hum. Genet. 2001, 69, 381–395. [Google Scholar] [CrossRef]
- Millward, B.A.; Welsh, K.I.; Leslie, R.D.; Pyke, D.A.; Demaine, A.G. T Cell Receptor Beta Chain Polymorphisms Are Associated with Insulin-Dependent Diabetes. Clin. Exp. Immunol. 1987, 70, 152–157. [Google Scholar]
- Manabe, K.; Hibberd, M.L.; Donaldson, P.T.; Underhill, J.A.; Doherty, D.G.; Demaine, A.G.; Mieli-Vergani, G.; Eddleston, A.L.W.F.; Williams, R. T-Cell Receptor Constant β Germline Gene Polymorphisms and Susceptibility to Autoimmune Hepatitis. Gastroenterology 1994, 106, 1321–1325. [Google Scholar] [CrossRef]
- Tatsuma, M.; Tsugu, H.; Murakami, M. HLA-DQ region and TCR gene polymorphism associated with primary IgA nephropathy in japanese children. Nihon. Jinzo. Gakkai. Shi. 1997, 39, 734–739. [Google Scholar] [CrossRef]
- Demaine, A.G.; Vaughan, R.W.; Taube, D.H.; Welsh, K.I. Association of Membranous Nephropathy with T-Cell Receptor Constant Beta Chain and Immunoglobulin Heavy Chain Switch Region Polymorphisms. Immunogenetics 1988, 27, 19–23. [Google Scholar] [CrossRef] [PubMed]
- Ito, M.; Tanimoto, M.; Kamura, H.; Yoneda, M.; Morishima, Y.; Yamauchi, K.; Itatsu, T.; Takatsuki, K.; Saito, H. Association of HLA Antigen and Restriction Fragment Length Polymorphism of T Cell Receptor Beta-Chain Gene with Graves’ Disease and Hashimoto’s Thyroiditis. J. Clin. Endocrinol. Metab. 1989, 69, 100–104. [Google Scholar] [CrossRef]
- Tripputi, P.; Graziani, D.; Alfano, R.M.; Cassani, B.; Coggi, G. A Common T/C Polymorphism in the Promoter Region of the Beta T-Cell Receptor Gene. Mol. Cell. Probes 2000, 14, 195–197. [Google Scholar] [CrossRef]
- Tripputi, P.; Cigognini, D.; Bianchi, S.; Fedele, L. Different Allelic Distribution of a Single SNP between Sexes in Humans. Biochem. Genet. 2008, 46, 733–736. [Google Scholar] [CrossRef] [PubMed]
- Tripputi, P.; Bianchi, S.; Fedele, L. A Possible Mechanism for Non-Replication of Allelic Association between a Single Nucleotide Polymorphism of the Human Beta T-Cell Receptor and Autoimmune Diseases. Int. J. Immunogenet. 2008, 35, 141–144. [Google Scholar] [CrossRef]
- Selmi, C.; Invernizzi, P.; Tripputi, P.; Battezzati, P.M.; Bignotto, M.; Zuin, M.; Crosignani, A.; Podda, M. T-Cell Receptor Polymorphism in Primary Biliary Cirrhosis. Ann. Ital. Med. Interna Organo Uff. Della Soc. Ital. Med. Interna 2003, 18, 149–153. [Google Scholar]
- Marschall, H.-U.; Henriksson, I.; Lindberg, S.; Söderdahl, F.; Thuresson, M.; Wahlin, S.; Ludvigsson, J.F. Incidence, Prevalence, and Outcome of Primary Biliary Cholangitis in a Nationwide Swedish Population-Based Cohort. Sci. Rep. 2019, 9, 11525. [Google Scholar] [CrossRef] [PubMed]
- Hirschhorn, J.N.; Lohmueller, K.; Byrne, E.; Hirschhorn, K. A Comprehensive Review of Genetic Association Studies. Genet. Med. Off. J. Am. Coll. Med. Genet. 2002, 4, 45–61. [Google Scholar] [CrossRef] [PubMed]
- Redden, D.T.; Allison, D.B. Nonreplication in Genetic Association Studies of Obesity and Diabetes Research. J. Nutr. 2003, 133, 3323–3326. [Google Scholar] [CrossRef] [PubMed]
- Tripputi, P.; Bignotto, M. Allele Frequency between Males and Females of a SNP of the Human Beta T Cell Receptor. Adv. Stud. Biol. 2012, 4, 521–528. [Google Scholar]
- Tripputi, P.; Guérin, S.L.; Moore, D.D. Two Mechanisms for the Extinction of Gene Expression in Hybrid Cells. Science 1988, 241, 1205–1207. [Google Scholar] [CrossRef] [PubMed]
- Tripputi, P.; Bianchi, P.; Fermo, E.; Bignotto, M.; Zanella, A. Chromosome 7q31.1 Deletion in Myeloid Neoplasms. Hum. Pathol. 2014, 45, 368–371. [Google Scholar] [CrossRef] [PubMed]
- Tripputi, P.; Bignotto, M.; Cigognini, D.; Bianchi, S.; Fedele, L. T-Cell Receptor Beta Polymorphism Is Not Associated with Endometriosis. J. Obstet. Gynaecol. Res. 2011, 37, 1405–1408. [Google Scholar] [CrossRef] [PubMed]
- Weiss, L.A.; Pan, L.; Abney, M.; Ober, C. The Sex-Specific Genetic Architecture of Quantitative Traits in Humans. Nat. Genet. 2006, 38, 218–222. [Google Scholar] [CrossRef]
- Klein, S.L.; Flanagan, K.L. Sex Differences in Immune Responses. Nat. Rev. Immunol. 2016, 16, 626–638. [Google Scholar] [CrossRef]
- Ortona, E.; Pierdominici, M.; Maselli, A.; Veroni, C.; Aloisi, F.; Shoenfeld, Y. Sex-Based Differences in Autoimmune Diseases. Ann. Ist. Super. Sanita 2016, 52, 205–212. [Google Scholar] [CrossRef]
- Roved, J.; Westerdahl, H.; Hasselquist, D. Sex Differences in Immune Responses: Hormonal Effects, Antagonistic Selection, and Evolutionary Consequences. Horm. Behav. 2017, 88, 95–105. [Google Scholar] [CrossRef]
- Hou, X.-L.; Wang, L.; Ding, Y.-L.; Xie, Q.; Diao, H.-Y. Current Status and Recent Advances of next Generation Sequencing Techniques in Immunological Repertoire. Genes Immun. 2016, 17, 153–164. [Google Scholar] [CrossRef] [PubMed]
- Matos, T.R.; de Rie, M.A.; Teunissen, M.B.M. Research Techniques Made Simple: High-Throughput Sequencing of the T-Cell Receptor. J. Invest. Dermatol. 2017, 137, e131–e138. [Google Scholar] [CrossRef] [PubMed]
- Clark, V.J.; Ptak, S.E.; Tiemann, I.; Qian, Y.; Coop, G.; Stone, A.C.; Przeworski, M.; Arnheim, N.; Di Rienzo, A. Combining Sperm Typing and Linkage Disequilibrium Analyses Reveals Differences in Selective Pressures or Recombination Rates across Human Populations. Genetics 2007, 175, 795–804. [Google Scholar] [CrossRef] [PubMed][Green Version]
- King, R.A.; Rotter, J.I.; Motulsky, A.G. The Genetic Basis of Common Diseases; Oxford University Press: Oxford, UK, 2002; ISBN 978-0-19-512582-5. [Google Scholar]
- Payami, H.; Zhu, M.; Montimurro, J.; Keefe, R.; McCulloch, C.C.; Moses, L. One Step Closer to Fixing Association Studies: Evidence for Age- and Gender-Specific Allele Frequency Variations and Deviations from Hardy-Weinberg Expectations in Controls. Hum. Genet. 2005, 118, 322–330. [Google Scholar] [CrossRef] [PubMed]
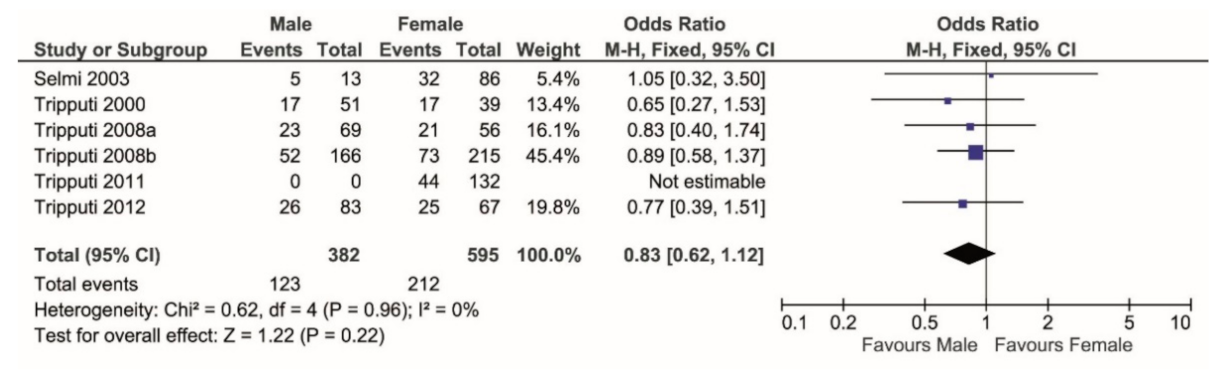

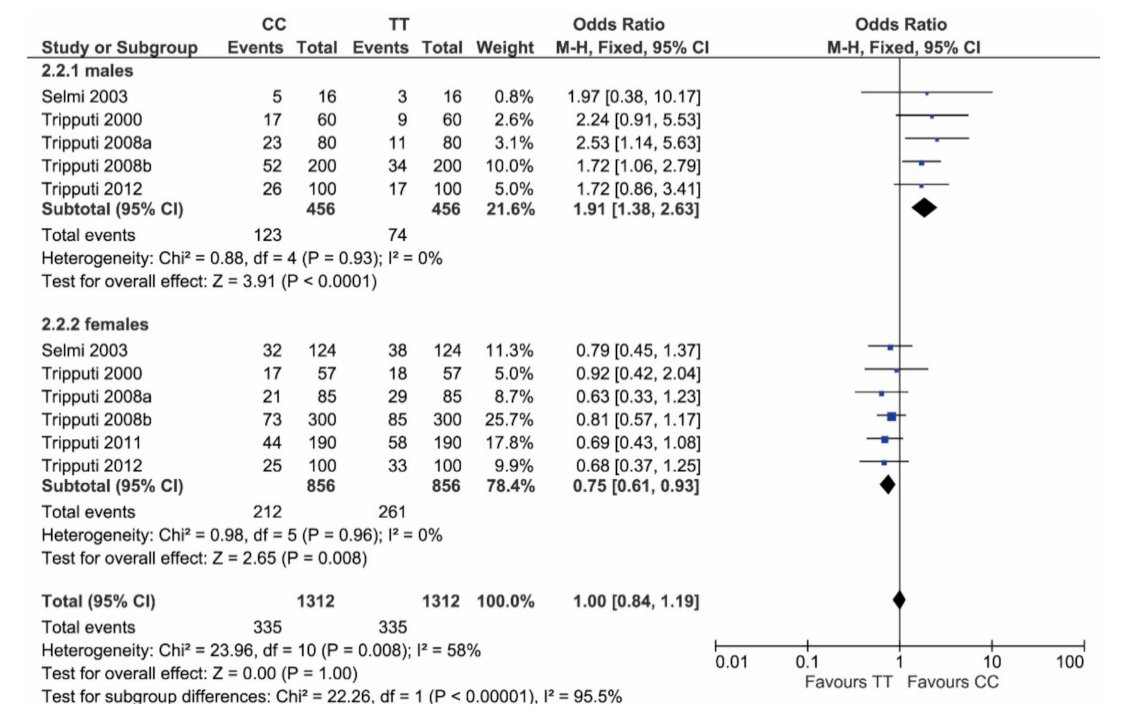
| Reference | Males | Females | Total Enrolled Individuals |
|---|---|---|---|
| Tripputi and Bignotto 2012 [23] | 100 | 100 | 200—only healthy controls |
| Tripputi et al., 2011 [26] | 0 | 190 | 190—70 cases and 120 controls |
| Tripputi et al., 2008 [17] | 80 | 85 | 165—only healthy controls |
| Tripputi et al., 2008 [18] | 200 | 300 | 500—only healthy controls |
| Selmi et al., 2003 [19] | 16 | 124 | 140—70 cases (62 females and 8 males) and 70 healthy controls (62 females and 8 males) |
| Tripputi et al., 2000 [16] | 60 | 57 | 117 (only healthy controls) |
| Total | 456 | 856 | 1312 (140 cases and 1172 healthy controls) |
| Individuals | Males (Number and %) | Females (Number and %) | Overall (Number and %) |
|---|---|---|---|
| T/C a | 259 (57%) | 383 (45%) | 642 (49%) |
| T/T b | 123 (27%) | 212 (25%) | 335 (25.5%) |
| C/C c | 74 (16%) | 261 (30%) | 335 (25.5%) |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Varoni, E.M.; Lodi, G.; Del Fabbro, M.; Sardella, A.; Carrassi, A.; Iriti, M.; Tripputi, P. Sex-Related Differences in Allelic Frequency of the Human Beta T Cell Receptor SNP rs1800907: A Retrospective Analysis from Milan Metropolitan Area. Vaccines 2021, 9, 333. https://doi.org/10.3390/vaccines9040333
Varoni EM, Lodi G, Del Fabbro M, Sardella A, Carrassi A, Iriti M, Tripputi P. Sex-Related Differences in Allelic Frequency of the Human Beta T Cell Receptor SNP rs1800907: A Retrospective Analysis from Milan Metropolitan Area. Vaccines. 2021; 9(4):333. https://doi.org/10.3390/vaccines9040333
Chicago/Turabian StyleVaroni, Elena M., Giovanni Lodi, Massimo Del Fabbro, Andrea Sardella, Antonio Carrassi, Marcello Iriti, and Pasquale Tripputi. 2021. "Sex-Related Differences in Allelic Frequency of the Human Beta T Cell Receptor SNP rs1800907: A Retrospective Analysis from Milan Metropolitan Area" Vaccines 9, no. 4: 333. https://doi.org/10.3390/vaccines9040333
APA StyleVaroni, E. M., Lodi, G., Del Fabbro, M., Sardella, A., Carrassi, A., Iriti, M., & Tripputi, P. (2021). Sex-Related Differences in Allelic Frequency of the Human Beta T Cell Receptor SNP rs1800907: A Retrospective Analysis from Milan Metropolitan Area. Vaccines, 9(4), 333. https://doi.org/10.3390/vaccines9040333









