Primary Human Dendritic Cells and Whole-Blood Based Assays to Evaluate Immuno-Modulatory Properties of Heat-Killed Commensal Bacteria
Abstract
1. Introduction
2. Materials and Methods
2.1. Growth of Commensal Bacteria
2.2. In Vitro Differentiation and Stimulation of Primary Human Monocyte-Derived Dendritic Cells (MoDC) with Live Bacteria
2.3. In Vitro Differentiation and Stimulation of Primary Human Monocyte-Derived Dendritic Cells (MoDC) with Heat-Killed Bacteria
2.4. Quantification of Adenosine Triphosphate (ATP) and Immune Mediators in Cell Culture Supernatant
2.5. Immuno-Staining and Flow Cytometry
2.6. Stimulation of Primary Human Whole-Blood with Heat-Killed Bacteria Using the TruCulture® Ex Vivo Whole-Blood Culture System
2.7. Quantification of Immune Mediators in Human Sera
2.8. Statistical Analysis
3. Results
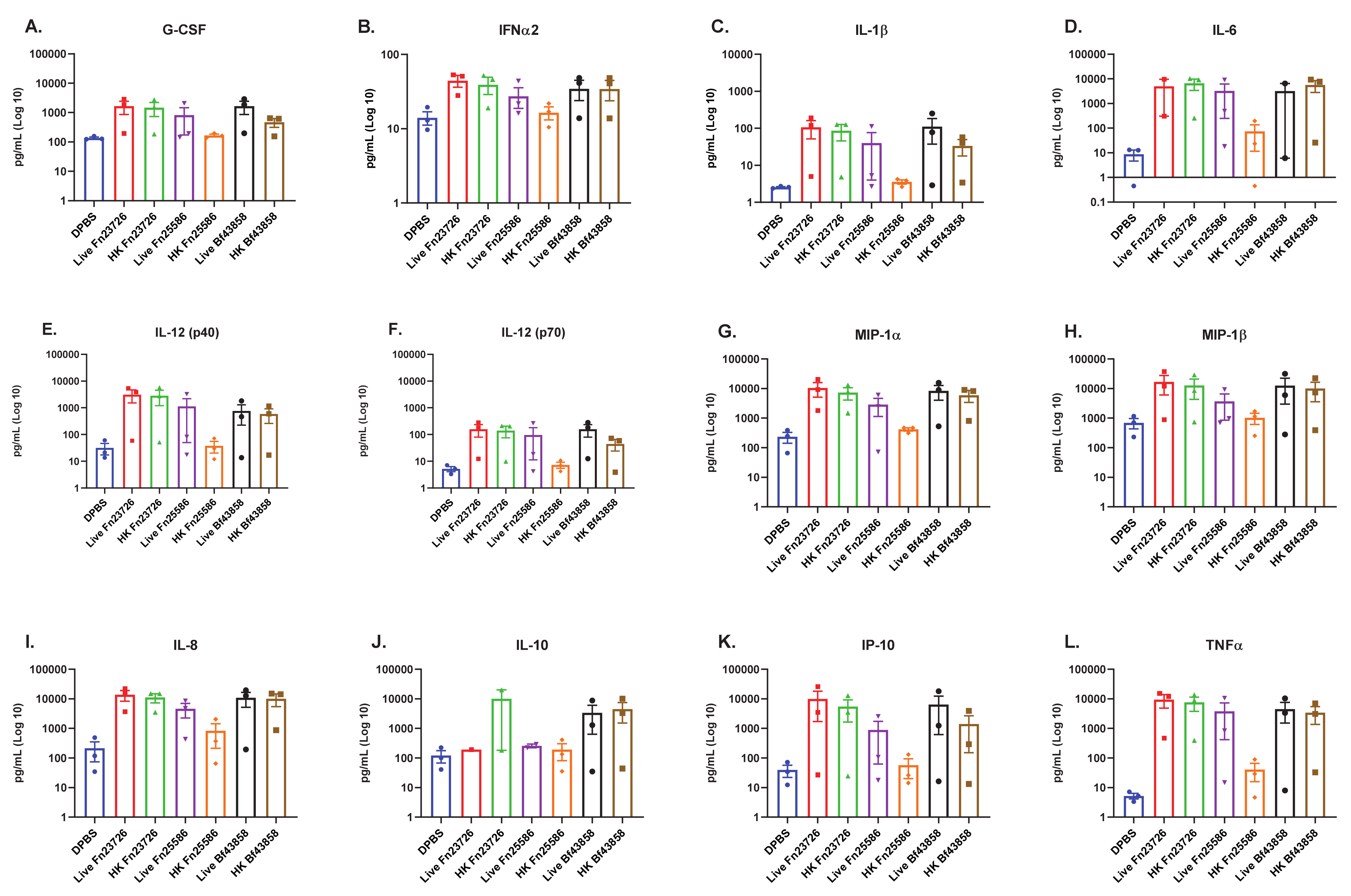
3.1. Primary Human MoDC Stimulated with Live and Heat-Killed Bacteria Demonstrate Comparable Levels of Immune Activation
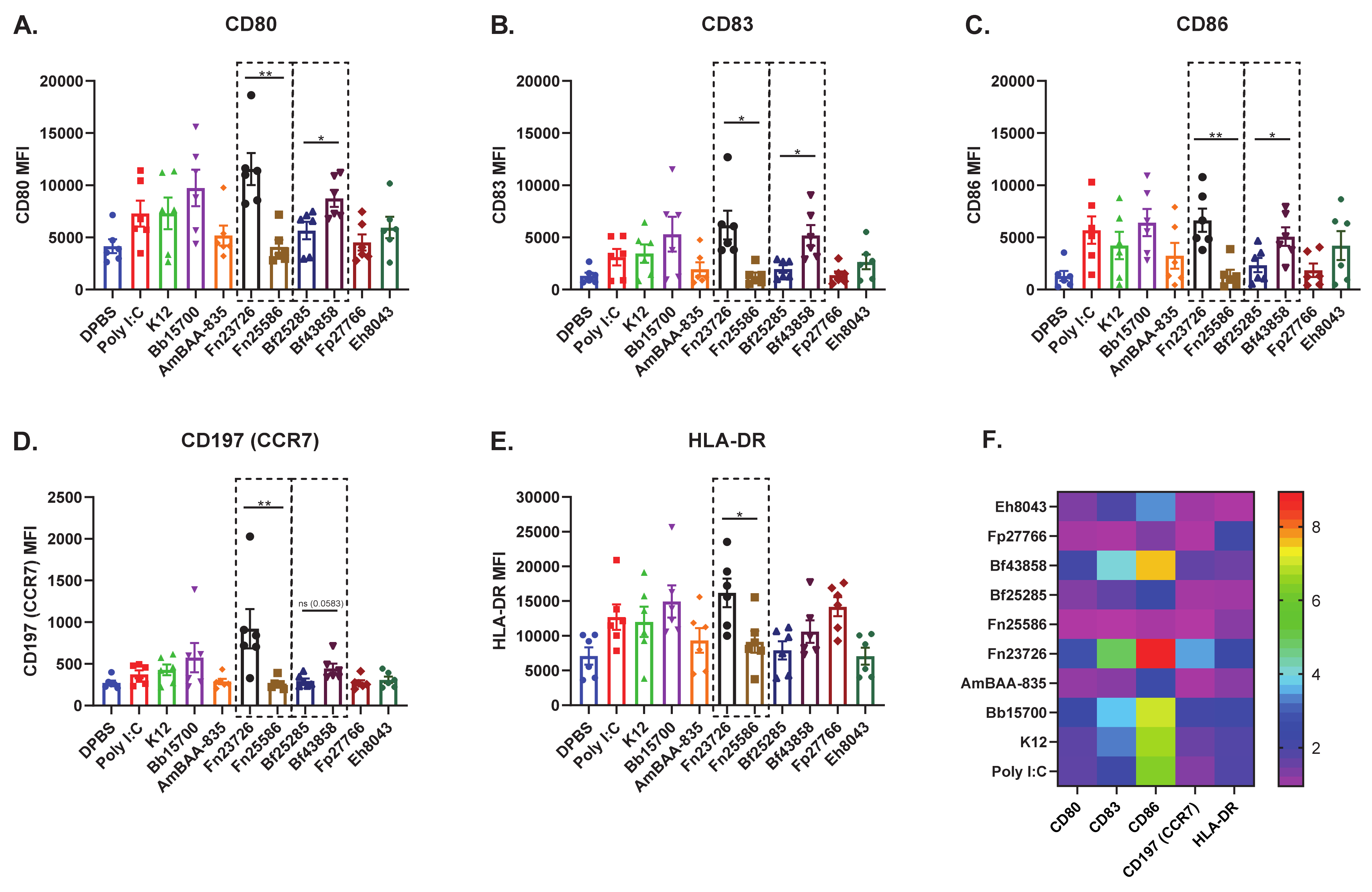
3.2. Primary Human MoDC Stimulated with Heat-Killed Bacteria Can Detect Strain-Based Differences in Immuno-Modulation
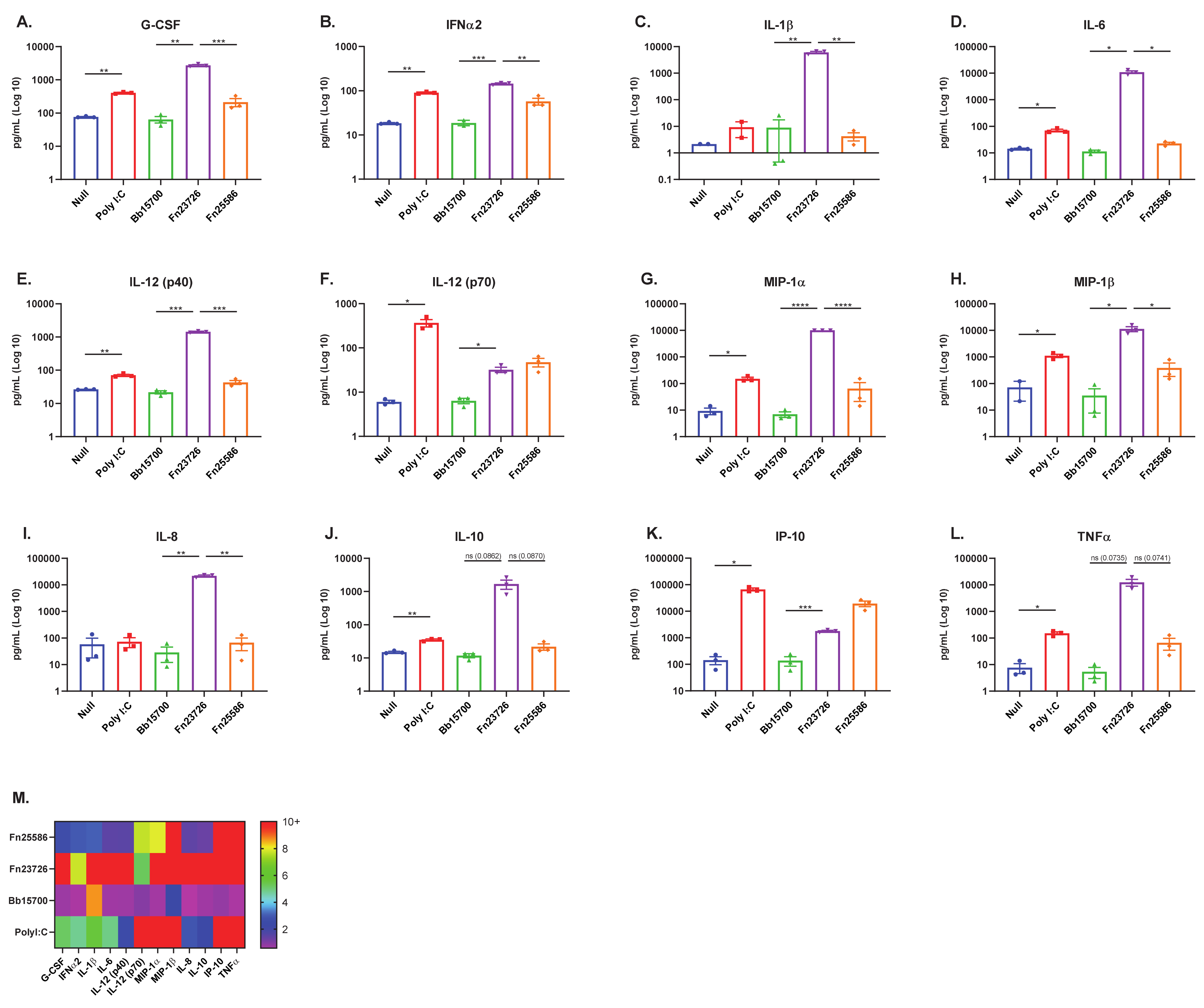
3.3. Primary Human Whole-Blood Stimulated with Heat-Killed Bacteria Can Detect Strain-Based Differences in Immuno-Modulation
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Informed Consent Statement
Acknowledgments
Conflicts of Interest
References
- Ostman, S.; Rask, C.; Wold, A.E.; Hultkrantz, S.; Telemo, E. Impaired regulatory T cell function in germ-free mice. Eur. J. Immunol. 2006, 36, 2336–2346. [Google Scholar] [CrossRef] [PubMed]
- Dimmit, R.A.; Staley, E.M.; Chuang, G.; Tanner, S.C.; Soltau, T.D.; Lorenz, R.G. The role of postnatal acquisition of the intestinal microbiome in the early development of immune function. J. Pediatr. Gastroenterol. Nutr. 2010, 51, 262–273. [Google Scholar] [CrossRef] [PubMed]
- Macpherson, A.J.; Harris, N.L. Interactions between commensal intestinal bacteria and the immune system. Nat. Rev. Immunol. 2004, 4, 478–485. [Google Scholar] [CrossRef] [PubMed]
- Lathrop, S.K.; Bloom, S.M.; Rao, S.M.; Nutsch, K.; Lio, K.W.; Santacruz, N.; Peterson, D.A.; Stappenback, T.S.; Hsieh, C.S. Peripheral education of the immune system by colonic commensal microbiota. Nature 2011, 478, 250–254. [Google Scholar] [CrossRef]
- Kuhn, K.A.; Stappenback, T.S. Peripheral education of the immune system by the colonic microbiota. Sem. Immunol. 2013, 25, 364–369. [Google Scholar] [CrossRef]
- Kim, D.; Kim, Y.G.; Seo, S.U.; Kim, D.J.; Kamada, N.; Prescott, D.; Chamaillard, M.; Philpott, D.J.; Rosenstiel, P.; Inohara, N.; et al. Nod2-mediated recognition of the microbiota is critical for mucosal adjuvant activity of cholera toxin. Nat. Med. 2016, 22, 524–530. [Google Scholar] [CrossRef]
- Zimmerman, P.; Curtis, N. The influence of the intestinal microbiome on vaccine responses. Vaccine 2018, 36, 4433–4439. [Google Scholar] [CrossRef]
- Valdez, Y.; Brown, E.M.; Finlay, B.B. Influence of the microbiota on vaccine effectiveness. Trends Immunol. 2014, 35, 526–537. [Google Scholar] [CrossRef]
- Hagan, T.; Cortese, M.; Rouphael, N.; Boudreau, C.; Linde, C.; Maddur, M.S.; Das, J.; Wang, H.; Guthmiller, J.; Zheng, N.Y.; et al. Antibiotics-driven gut microbiome perturbation alters immunity to vaccines in humans. Cell 2019, 178, 1313–1328. [Google Scholar] [CrossRef]
- Routy, B.; Le Chatelier, E.; Derosa, L.; Doung, C.P.M.; Alou, M.T.; Daillere, R.; Fluckiger, A.; Messaoudene, M.; Rauber, C.; Roberti, M.P.; et al. Gut microbiome influences efficacy of PD-1-ased immunotherapy against epithelial tumors. Science 2018, 359, 91–97. [Google Scholar] [CrossRef]
- Gopalakrishnan, V.; Spencer, C.N.; Nezi, L.; Reuben, A.; Andrews, M.C.; Karpinets, T.V.; Prieto, P.A.; Vicente, D.; Hoffman, K.; Wei, S.C.; et al. Gut microbiome modulates response to anti-PD-1 immunotherapy in melanoma patients. Science. [CrossRef] [PubMed]
- Zheng, Y.; Wang, T.; Tu, X.; Huang, Y.; Zhang, H.; Tan, D.; Jiang, W.; Cai, S.; Zhao, P.; Song, R.; et al. Gut microbiome affects the response to anti-PD-1 immunotherapy in patients with hepatocellular carcinoma. J. Immunother. Cancer 2019, 23, 1–7. [Google Scholar] [CrossRef]
- Chanput, W.; Mess, J.J.; Wichers, H.J. THP-1 cell line: An in vitro cell model for immune modulation. Int. Immunopharmacol. 2014, 23, 37–45. [Google Scholar] [CrossRef] [PubMed]
- Banchereau, J.; Briere, F.; Caux, C.; Davoust, J.; Lebecque, S.; Lui, Y.-J.; Pulendran, B.; Palucka, K. Immunobiology of dendritic cells. Annu. Rev. Immunol. 2000, 18, 767–811. [Google Scholar] [CrossRef]
- Worbs, T.; Hammerschmidt, S.I.; Forster, R. Dendritic cell migration in health and disease. Nat. Rev. Immunol. 2017, 17, 30–48. [Google Scholar] [CrossRef] [PubMed]
- Wculek, S.K.; Cueto, F.J.; Mujal, A.M.; Melero, I.; Krummel, M.F.; Sancho, D. Dendritic cells in cancer immunology and immunotherapy. Nat. Rev. Immunol. 2019, 20, 7–24. [Google Scholar] [CrossRef]
- Howard, C.J.; Charleston, B.; Stephens, S.A.; Sopp, P.; Hope, J.C. The role of dendritic cells in shaping the immune system. Anim. Health Res. Rev. 2004, 5, 191–195. [Google Scholar] [CrossRef]
- Perez, C.R.; De Palma, M. Engineering dendritic cells vaccines to improve cancer immunotherapy. Nat. Commun. 2019, 10, 1–10. [Google Scholar] [CrossRef]
- Blanco, P.; Palucka, A.K.; Pascual, V.; Banchereau, J. Dendritic cells and cytokines in human inflammatory and autoimmune diseases. Cytokine Growth Factor Rev. 2008, 19, 41–52. [Google Scholar] [CrossRef]
- Ziegler-Heitbrock, L. Blood monocytes and their subsets: Established features and open questions. Front. Immunol. 2015, 6, 1–5. [Google Scholar] [CrossRef] [PubMed]
- Fink, L.N.; Zeuthen, L.H.; Ferlazzo, G.; Frokiaer, H. Human antigen-presenting cells respond differently to gut-derived probiotic bacteria but mediate similar strain-dependent NK and T cell activation. FEMS Immunol. Med. Microbiol. 2007, 51, 535–546. [Google Scholar] [CrossRef]
- Bene, K.; Varga, Z.; Petrov, V.O.; Boyko, N.; Rajnavolgyi, E. Gut microbiota species can provoke both inflammatory and tolerogenic immune response in human dendritic cells mediated by retinoic acid receptor alpha ligation. Front. Immunol. 2017, 8, 1–17. [Google Scholar] [CrossRef]
- Ivanov, I.I.; Honda, K. Intestinal commensal microbes as immune modulators. Cell Host Microbe 2013, 12, 496–508. [Google Scholar] [CrossRef] [PubMed]
- Derrien, M.; Belzer, C.; de Vos, W.M. Akkermansia muciniphila and its role in regulating host functions. Microb. Pathog. 2017, 106, 171–181. [Google Scholar] [CrossRef] [PubMed]
- Leylabadlo, H.E.; Ghotaslou, R.; Feizabadi, M.M.; Farajina, S.; Moaddab, S.Y.; Ganbarov, K.; Khodadadi, E.; Tanomand, A.; Sheykhsaran, E.; Yousefi, B.; et al. The critical role of Faecalibacterium prausnitzii in human health: An overview. Microb. Pathog. 2020, 149, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Casasanta, M.A.; Yoo, C.C.; Udayasuryan, B.; Sanders, B.; Umana, A.; Zhang, Y.; Peng, H.; Duncan, A.J.; Wang, Y.; Li, L.; et al. Fusobacterium nucleatum host-cell binding and invasion induces IL-8 and CXCL1 secretion that drives colorectal cancer cell migration. Sci. Signal. 2020, 13, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Stappers, M.H.T.; Janssen, N.A.F.; Oosting, M.; Plantinga, T.S.; Arvis, P.; Mouton, J.W.; Joosten, L.A.B.; Netea, M.G.; Gyssens, I.C. A role for TLR1, TLR2, and NOD2 in cytokine induction by Bacteroides fragilis. Cytokine 2012, 60, 861–869. [Google Scholar] [CrossRef]
- Duffy, D.; Rouilly, V.; Bradeau, C.; Corbiere, V.; Djebali, R.; Ungeheuer, M.-N.; Josein, R.; LaBrie, S.T.; Lantz, O.; Louis, D.; et al. Standardized whole blood stimulation improves immunomonitoring of induced immune responses in multi-center study. Clin. Immunol. 2017, 183, 325–335. [Google Scholar] [CrossRef] [PubMed]
- Duffy, D.; Rouilly, V.; Libri, V.; Hasan, M.; Beitz, B.; David, M.; Urrutia, A.; Bisiaux, A.; LaBrie, S.T.; Dubois, A.; et al. Functional analysis via standardized whole-blood stimulation systems defines the boundaries of a healthy immune responses to complex stimuli. Immunity 2014, 40, 436–450. [Google Scholar] [CrossRef]
- Zamarareva, M.V.; Sabirov, R.Z.; Maeno, E.; Ando-Akatsuka, Y.; Bessonova, S.V.; Okada, Y. Cells die with increased cytosolic ATP during apoptosis: A bioluminescence study with intracellular luciferase. Cell Death Differ. 2005, 12, 1390–1397. [Google Scholar] [CrossRef] [PubMed]
- Goradel, N.H.; Heidarzadeh, S.; Jahangiri, S.; Farhood, B.; Mortezaee, K.; Khanlarkhani, N.; Negahdari, B. Fusobacterium nucleatum and colorectal cancer: A mechanistic overview. J. Cell. Physiol. 2019, 234, 2337–2344. [Google Scholar] [CrossRef] [PubMed]
- Kasper, S.H.; Morell-Perez, C.; Wyche, T.P.; Sana, T.R.; Lieberman, L.A.; Hett, E.C. Colorectal cancer-associated anaerobic bacteria proliferate in tumor spheroids and alter the microenvironment. Sci. Rep. 2020, 10, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Sanchez, B.; Gonzalez-Rodriguez, I.; Arboleya, S.; Lopez, P.; Suarez, A.; Ruas-Madiedo, P.; Margolles, A.; Gueimonde, M. The Effects of Bifidobacterium breve on Immune Mediators and Proteome of HT29 Cells Monolayers. Biomed Res. Int. 2015, 2015, 1–6. [Google Scholar] [CrossRef] [PubMed]
- He, F.; Morita, H.M.; Ouwehand, A.C.; Hosoda, M.; Hiramatsu, M.; Kurisaki, J.-I.; Isolauri, E.; Benno, Y.; Salminen, S. Stimulation of the secretion of pro-inflammatory cytokines by Biffidobacterium strains. Microbiol. Immunol. 2002, 46, 781–785. [Google Scholar] [CrossRef] [PubMed]
- Van Nood, E.; Vrieze, A.; Nieudorp, M.; Fuentes, S. Duodenal infusion of donor feces for recurrent Clostridium difficile. N. Engl. J. Med. 2013, 368, 407–415. [Google Scholar] [CrossRef]
- Rao, K.; Young, V.B. Fecal microbiota transplantation for the management of Clostridium difficile infection. Infect. Dis. Clin. N. Am. 2015, 29, 109–122. [Google Scholar] [CrossRef] [PubMed]
- Bermudez-Humaran, L.G.; Langella, P. Live bacterial biotherapeutics in the clinic. Nat. Biotechnol. 2018, 36, 816–818. [Google Scholar] [CrossRef]
- Tanoue, T.; Morita, S.; Plichta, D.R.; Skelly, A.N.; Suda, W.; Sugiura, Y.; Narushima, S.; Vlamakis, H.; Motoo, I.; Sugita, K.; et al. A defined commensal consortium elicits CD8 T cell and anti-cancer immunity. Nature 2019, 565, 600–605. [Google Scholar] [CrossRef]
- Furusawa, Y.; Obata, Y.; Fukuda, S.; Endo, T.A.; Nakato, G.; Takahashi, D.; Nakanishi, Y.; Uetake, C.; Kato, K.; Kato, T.; et al. Commensal microbe-derived butyrate induces the differentiation of colonic regulatory T cells. Nature 2013, 504, 446–450. [Google Scholar] [CrossRef]
- Mathewson, N.; Jenq, R.; Mathew, A.V.; Koenigsknecht, M.; Hanash, A.; Toubai, T.; Oravecz-Wilson, K.; Wu, S.-R.; Sun, Y.; Rossi, C.; et al. Gut microbiome-derived metabolites modulate intestinal epithelial cell damage and mitigate graft-versus-host disease. Nat. Immunol. 2016, 16, 505–513. [Google Scholar] [CrossRef]
- Guermonprez, P.; Valladeau, J.; Zitvogel, L.; Thery, C.; Amigorena, S. Antigen presentation and T cell stimulation by dendritic cells. Annu. Rev. Immunol. 2002, 20, 621–667. [Google Scholar] [CrossRef]
- Li, Z.; Ju, X.; Silveira, P.A.; Abadir, E.; Hsu, W.-H.; Har, D.N.J.; Clark, G.J. CD83: Activation marker for antigen presenting cells and its therapeutic potential. Front. Immunol. 2019, 10, 1–9. [Google Scholar] [CrossRef]
- Forester, R.; Davalos-Misslitz, A.C.; Rot, A. CCR7 and its ligands: Balancing immunity and tolerance. Nat. Rev. Immunol. 2008, 8, 362–371. [Google Scholar] [CrossRef]
- Pulendran, B.; Ahmed, R. Immunological mechanisms of vaccination. Front. Immunol. 2011, 12, 509–517. [Google Scholar] [CrossRef] [PubMed]
- Disis, M.L. Mechanism of action of immunotherapy. Semin. Onc. 2014, 41, S3–S13. [Google Scholar] [CrossRef]
- Hoarau, C.; Lagaraine, C.; Martin, L.; Velge-Roussel, F.; Lebranchu, Y. Supernatant of Bifidobacterium breve induces dendritic cell maturation, activation, and survival through Toll-like receptor 2 pathway. J. Allergy Clin. Immunol. 2006, 117, 696–702. [Google Scholar] [CrossRef] [PubMed]
- Huda, M.N.; Ahmad, S.M.; Alam, M.J.; Khanam, A.; Kalanetra, K.M.; Taft, D.H.; Raqib, R.; Underwood, M.A.; Mills, D.A.; Stephensen, C.B. Bifidobacterium abundance in early infancy and vaccine response at 2 years of age. Pediatrics 2019, 13, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Ma, Y.; Luo, Y.; Huang, X.; Song, F.; Liu, G. Construction of Bifidobacterium infantis as a live oral vaccine that expresses antigens of the major fimbrial subunit (CfaB) and the B subunit of heat-labile enterotoxin (LTB) from enterotoxigenic Escherichia coli. Microbiology 2011, 158, 498–504. [Google Scholar] [CrossRef]

| Bacteria | Strain Number | Source | Gram | Order | Aerobe/Anaerobe |
|---|---|---|---|---|---|
| Akkermansia muciniphila (Am) | BAA-835 | ATCC | Gram -ve | Verrucomicrobiales | Anaerobe |
| Fusobacterium nucleatum (Fn) | 23726 | ATCC | Gram -ve | Fusobacteriales | Anaerobe |
| Fusobacterium nucleatum (Fn) | 25586 | ATCC | Gram -ve | Fusobacteriales | Anaerobe |
| Bacteroides fragilis (Bf) | 25285 | ATCC | Gram -ve | Bacteroidales | Anaerobe |
| Bacteroides fragilis (Bf) | 43858 | ATCC | Gram -ve | Bacteroidales | Anaerobe |
| Escherichia coli K-12 (K12) | 10798 | ATCC | Gram -ve | Enterobacterales | Aerobe |
| Bifidobacterium breve (Bb) | 15700 | ATCC | Gram +ve | Bifidobacteriales | Anaerobe |
| Faecalibacterium prausnitzii (Fp) | 27766 | ATCC | Gram +ve | Clostridiales | Anaerobe |
| Enterococcus hirae (Eh) | 8043 | ATCC | Gram +ve | Lactobacillales | Facultative Anaerobe |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Norton, J.E., Jr.; Kommineni, S.; Akrivoulis, P.; Gutierrez, D.A.; Hazuda, D.J.; Swaminathan, G. Primary Human Dendritic Cells and Whole-Blood Based Assays to Evaluate Immuno-Modulatory Properties of Heat-Killed Commensal Bacteria. Vaccines 2021, 9, 225. https://doi.org/10.3390/vaccines9030225
Norton JE Jr., Kommineni S, Akrivoulis P, Gutierrez DA, Hazuda DJ, Swaminathan G. Primary Human Dendritic Cells and Whole-Blood Based Assays to Evaluate Immuno-Modulatory Properties of Heat-Killed Commensal Bacteria. Vaccines. 2021; 9(3):225. https://doi.org/10.3390/vaccines9030225
Chicago/Turabian StyleNorton, James E., Jr., Sushma Kommineni, Patricia Akrivoulis, Dario A. Gutierrez, Daria J. Hazuda, and Gokul Swaminathan. 2021. "Primary Human Dendritic Cells and Whole-Blood Based Assays to Evaluate Immuno-Modulatory Properties of Heat-Killed Commensal Bacteria" Vaccines 9, no. 3: 225. https://doi.org/10.3390/vaccines9030225
APA StyleNorton, J. E., Jr., Kommineni, S., Akrivoulis, P., Gutierrez, D. A., Hazuda, D. J., & Swaminathan, G. (2021). Primary Human Dendritic Cells and Whole-Blood Based Assays to Evaluate Immuno-Modulatory Properties of Heat-Killed Commensal Bacteria. Vaccines, 9(3), 225. https://doi.org/10.3390/vaccines9030225






