The Importance of RNA-Based Vaccines in the Fight against COVID-19: An Overview
Abstract
1. Introduction
2. A Brief History of RNA-Based Vaccines
2.1. RNA Vaccines against Cancer
2.2. RNA Vaccines against Non-Infectious Diseases
2.3. RNA Vaccines against Infectious Diseases
3. RNA Vaccines in the Context of COVID-19
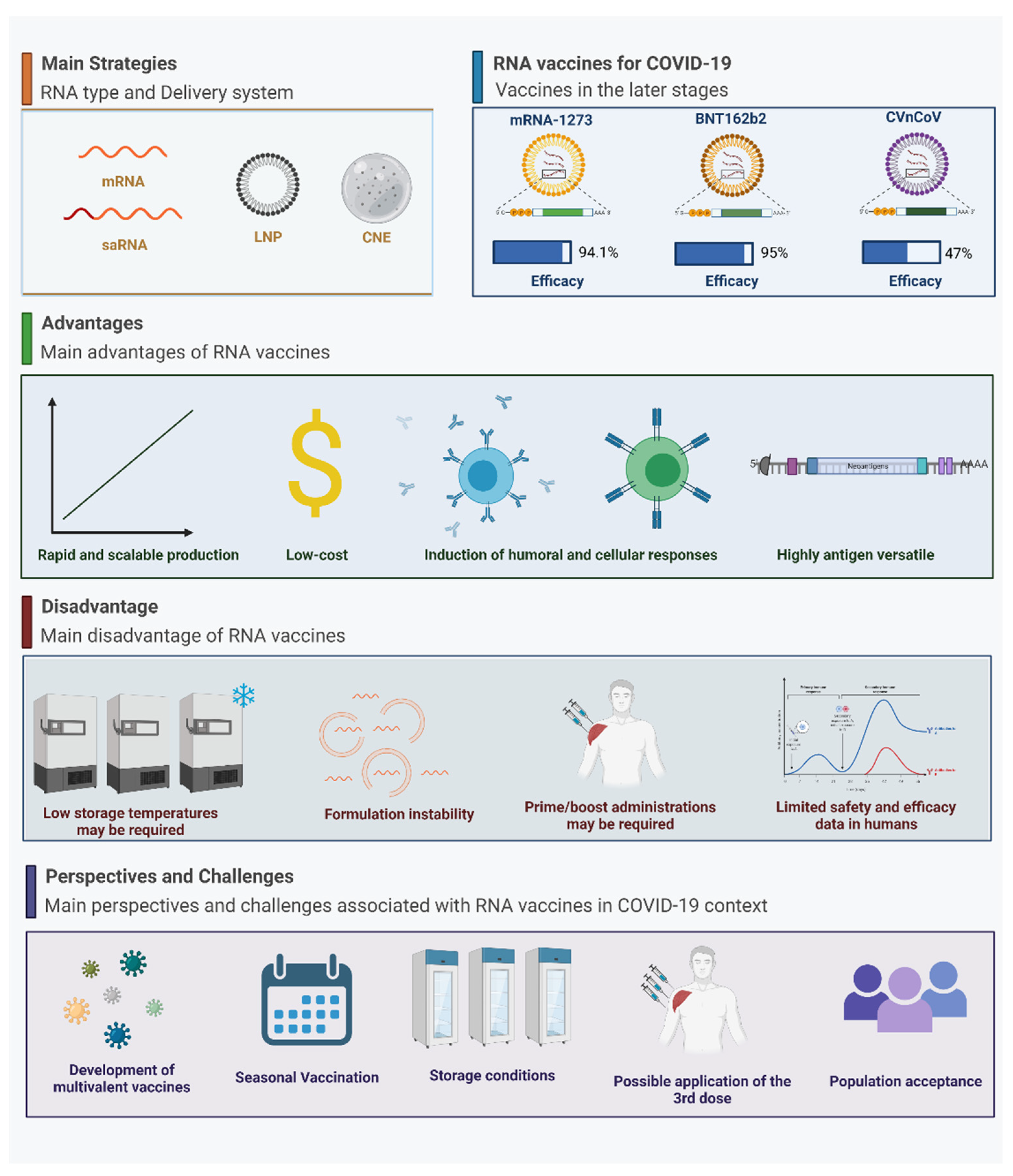
3.1. The Main RNA-Based Vaccines for COVID-19
| Developer | Vaccine Name | Active Substance (Antigen Type) and Formulation Details | Animal Model | Main Findings and/or Conclusions Considering | Reference |
|---|---|---|---|---|---|
| CureVac AG | CVnCoV | Lipid-nanoparticle-encapsulated mRNA that encodes full-length, prefusion-stabilized SARS-CoV-2 S protein | Mice (BALB/c) | Mice: innate immune response (systemic IL-6 and IFNα and robust cellular and humoral immune responses Hamsters: protected against challenge with wild-type SARS-CoV-2 NHP: those vaccinated with 8 µg were protected from challenge with wild-type SARS-CoV-2 (Victoria/1/2020) | [146] |
| Syrian Hamsters (Mesocricetus auratus) | |||||
| Rhesus macaques (Macaca mulatta) | [148] | ||||
| Arcturus Therapeutics | ARCT-021 (LUNAR-COV19) | Lipid-nanoparticle-encapsulated saRNA that encodes an alphavirus-based replicon and the SARS-CoV-2 full-length S glycoprotein (saRNA) | Mice (C57BL/6 (human ACE2 transgenic mouse model) and BALB/c) | Mice: single vaccination led to robust antibody responses Human ACE2 transgenic mice: protected from mortality and measurable infection following wild-type SARS-CoV-2 challenge | [149] |
| Imperial College London | COVAC1 (LNP-nCoVsaRNA) | saRNA encoding SARS-CoV-2 S protein encapsulated within a cationic liposom | Mice (BALB/c) | Mice: high cellular responses (IFN-γ production), specific IgG | [143] |
| Pfizer and BioNTech | BNT162b1 | Nucleoside-modified mRNA that encodes immunogens derived from the S protein of SARS-CoV-2 stabilized in the prefusion conformation, formulated in lipid nanoparticle | Mice (BALB/c) | Mice: one intramuscular dose of either elicited dose-dependent antibody responses, strong CD4+, and CD8+ T-cell response NHP: protected macaques against challenge with wild-type SARS-CoV-2 | [144] |
| BNT162b2 (known commercially as Comirnaty) | Rhesus macaques (Macaca mulatta) | ||||
| HDT Bio | HDT-301 (LION/repRNA-CoV2S) | saRNA that encodes SARS-CoV-2 full S protein formulated with alipid inorganic nanoparticle (LION) emulsion | Mice (BALB/c and C57BL/6) | Mice: single immunization triggered robust IgG and Th1 cells Aged mice: induced S-specific IgG and Th1 responses NHP: antibody responses persisted for at least 70 days, neutralized SARS-CoV-2 at titers comparable to those of convalescent plasma | [145] |
| Pigtail macaque (Macaca nemestrina) | |||||
| Moderna | mRNA-1273 (known commercially as Spikevax) | Nucleoside-modified mRNA that encodes prefusion-stabilized SARS-CoV-2 encapsulated in a lipid nanoparticle | Rhesus macaques (Macaca mulatta) | NHP: induced robust neutralizing activity against SARS-CoV-2, rapid protection in the upper and lower airways without pathological changes in the lung after challenge | [150] |
| PLA Academy of Military Sciences, Abogen and Walvax | ARCoV | Lipid-nanoparticle-encapsulated mRNA encoding the receptor-binding domain of SARS-CoV-2 S protein | Mice (BALB/c) | Mice: neutralizing antibodies against SARS-CoV-2, cellular responses (Th1 cells) Two doses conferred complete protection against the challenge (wild-type SARS-CoV-2) NHP: Th1 responses, neutralizing antibodies against SARS-CoV-2 Note: manufactured as a liquid formulation that can be stored at room temperature for at least 1 week | [147] |
| Cynomolgus monkeys (Macaca fascicularis) |
| Vaccine | Trial Phase | Location (NCT Number) | Doses | Dose Level | Main Immunogenicity and/or Efficacy Findings | Main Safety Findings | Reference |
|---|---|---|---|---|---|---|---|
| mRNA-1273 (Spikevax) | 1 | USA | 2 (28 days apart) | 25 μg, 100 μg, or 250 μg | Dose-dependent responses observed. Antibody GMT of 40,227, 109,209, and 213,526 in 25 μg, 100 μg, and 250 μg recipients, respectively. After second vaccination, titers increased to 299,751, 782,719, and 1,192,154 for 25 μg, 100 μg, or 250 μg, respectively. | AEs were commonly reported at the highest doses and after the second immunization. Systemic and local AEs occurred after both injections (fatigue, chills, headache, myalgia, and injection site pain). No AEs were noted; no prespecified halting rules were met. | [163] |
| 2 | USA | 50 μg or 100 μg | The 100 μg dose induced a > binding antibody concentrations than 50 μg. | The most common solicited AEs were injection site pain, headache, and fatigue. | [164] | ||
| 3 | USA | 100 μg | There was 94.1% efficacy (95% CI, 89.3 to 96.8%; p < 0.001) for the prevention of symptomatic SARS-CoV-2 infection within two weeks of second dose. | AEs were rare and occurred with the same incidence as the placebo. The most common treatment-related AEs were fatigue, headache, and injection-site pain. | [158] | ||
| BNT162b2 (Comirnaty) | 1/2 | Germany (NCT04380701) USA (NCT04380701) | 2 (21 days apart) | 10 µg, 20 µg, 30 µg | Dose-dependent nAbs > convalescent serum. (1.7 to 4.6-fold higher in 18–55-year-olds, 1.1–2.2-fold higher in 65-to-85-year-olds. SARS-CoV-2 antigen-specific CD4+ and CD8+ T cells. | Lower incidence and severity of systemic reactions, especially in older adults (65 to 85 years). Transient local reactions and systemic events were dose-dependent, greater after the second dose. | [165] |
| 3 | USA, Argentina, Brazil, Germany, South Africa, Turkey (NCT04368728) | 30 μg | There was 95% efficacy (95% CI 90.3–97.6) evaluated 7 days after second dose. Efficacy in participants reported comorbidity of 94.7%. | 16–55-year-olds experienced more systemic effects than >55-year-olds. Headache and fatigue were the most common systemic effects of local, injection-site pain. | [166] | ||
| CVnCoV | 1 | Germany (NCT04449276) | 2 (28 days apart) | 2 µg, 4 µg, 6 µg, 8 µg, or 12 µg | nAbs and IgG S protein or RBD-binding IgG two weeks after the second 12 μg dose comparable to convalescent plasma. | Dose-dependent increase in the frequency and severity of systemic AEs and local reactions; most were mild or moderate and transient. No vaccine-related SAEs were reported. | [167] |
| 2/3 | Argentina, Belgium, Colombia, Dominican Republic, Germany, Mexico, Netherlands, Panama, Peru, and Spain (NCT04652102) | 12 µg | Reported 47% efficacy versus disease. | Data not yet released. | [168] | ||
| ARCT-021 (LUNAR-COV19) | 1 | Singapore (NCT04480957) | 1 | 1 μg, 5 μg, 7.5 μg, and 10 μg | nAbs increased dose, similar following one 5 μg or 7.5 μg dose. S-binding IgG titers overlapped with levels in convalescent plasma (except1 μg dose). | The 10 μg dose was associated with more local and systemic solicited AEs; 7.5 μg was well tolerated. Fatigue, headache, myalgia, chills, and fever were the most common AEs. | [169] |
| 2 | 2 (28 days apart) | 3 μg and 5 μg | nAbs increased with dose. No increase in CD8 IFN-γ T cells was reported after the second dose. | Fatigue and arthralgia occurred in a single participant following the second 5 μg dose (56–80-year-old cohort). |
3.2. Efficacy of RNA-Based Vaccines against SARS-CoV-2 Variants
3.3. Composition of Main RNA-Based Vaccines for COVID-19
4. Perspectives and Challenges Associated with COVID-19 Immunization Based on RNA Vaccines
4.1. Perspectives and Challenges Associated with Widespread Production and Availability
4.2. Perspectives and Challenges Associated with Acceptance of COVID-19 Vaccines
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Plotkin, S.A. Vaccines: Past, present and future. Nat. Med. 2005, 11, S5–S11. [Google Scholar] [CrossRef] [PubMed]
- Zhang, N.; Li, C.; Zhu, Y.; Li, K.; Liang, J.; Wang, L.; Du, L.; Jiang, S.; Yang, F.; He, X.; et al. Current development of COVID-19 diagnostics, vaccines and therapeutics. Microbes Infect. 2019, 22, 231–235. [Google Scholar] [CrossRef]
- Zhu, N.; Zhang, D.; Wang, W.; Li, X.; Yang, B.; Song, J.; Zhao, X.; Huang, B.; Shi, W.; Lu, R.; et al. A Novel Coronavirus from Patients with Pneumonia in China, 2019. N. Engl. J. Med. 2020, 382, 727–733. [Google Scholar] [CrossRef] [PubMed]
- Dodd, R.H.; Pickles, K.; Nickel, B.; Cvejic, E.; Ayre, J.; Batcup, C.; Bonner, C.; Copp, T.; Cornell, S.; Dakin, T.; et al. Concerns and motivations about COVID-19 vaccination. Lancet Infect. Dis 2021, 21, 161–163. [Google Scholar] [CrossRef]
- Lu, H.; Stratton, C.W.; Tang, Y.-W. Outbreak of pneumonia of unknown etiology in Wuhan, China: The mystery and the miracle. J. Med. Virol. 2020, 92, 401–402. [Google Scholar] [CrossRef]
- Walker, P.G.T.; Whittaker, C.; Watson, O.J.; Baguelin, M.; Winskill, P.; Hamlet, A.; Djafaara, B.A.; Cucunubá, Z.; Mesa, D.O.; Green, W.; et al. The impact of COVID-19 and strategies for mitigation and suppression in low- and middle-income countries. Science 2020, 369, 413–422. [Google Scholar] [CrossRef]
- WHO. WHO Coronavirus Disease (COVID-19) Dashboard. Available online: https://covid19.who.int/ (accessed on 28 October 2021).
- Kampf, G.; Todt, D.; Pfaender, S.; Steinmann, E. Persistence of coronaviruses on inanimate surfaces and their inactivation with biocidal agents. J. Hosp. Infect. 2020, 104, 246–251. [Google Scholar] [CrossRef] [PubMed]
- Meselson, M. Droplets and Aerosols in the Transmission of SARS-CoV-2. N. Engl. J. Med. 2020, 382, 2063. [Google Scholar] [CrossRef]
- Arslan, M.; Xu, B.; El-Din, M.G. Transmission of SARS-CoV-2 via fecal-oral and aerosols–borne routes: Environmental dynamics and implications for wastewater management in underprivileged societies. Sci. Total Environ. 2020, 743, 140709. [Google Scholar] [CrossRef]
- Huang, C.; Wang, Y.; Li, X.; Ren, L.; Zhao, J.; Hu, Y.; Zhang, L.; Fan, G.; Xu, J.; Gu, X.; et al. Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Lancet 2020, 395, 497–506. [Google Scholar] [CrossRef]
- Yang, J.; Chen, X.; Deng, X.; Chen, Z.; Gong, H.; Yan, H.; Wu, Q.; Shi, H.; Lai, S.; Ajelli, M.; et al. Disease burden and clinical severity of the first pandemic wave of COVID-19 in Wuhan, China. Nat. Commun. 2020, 11, 5411. [Google Scholar] [CrossRef]
- Piroth, L.; Cottenet, J.; Mariet, A.-S.; Bonniaud, P.; Blot, M.; Tubert-Bitter, P.; Quantin, C. Comparison of the characteristics, morbidity, and mortality of COVID-19 and seasonal influenza: A nationwide, population-based retrospective cohort study. Lancet Respir. Med. 2021, 9, 251–259. [Google Scholar] [CrossRef]
- Ferrara, P.; Albano, L. COVID-19 and healthcare systems: What should we do next? Public Health 2020, 185, 1–2. [Google Scholar] [CrossRef] [PubMed]
- Chan, J.F.-W.; Kok, K.-H.; Zhu, Z.; Chu, H.; To, K.K.-W.; Yuan, S.; Yuen, K.-Y. Genomic characterization of the 2019 novel human-pathogenic coronavirus isolated from a patient with atypical pneumonia after visiting Wuhan. Emerg. Microbes Infect. 2020, 9, 221–236. [Google Scholar] [CrossRef]
- Wu, A.; Peng, Y.; Huang, B.; Ding, X.; Wang, X.; Niu, P.; Meng, J.; Zhu, Z.; Zhang, Z.; Wang, J.; et al. Genome Composition and Divergence of the Novel Coronavirus (2019-nCoV) Originating in China. Cell Host Microbe 2020, 27, 325–328. [Google Scholar] [CrossRef] [PubMed]
- Du, L.; He, Y.; Zhou, Y.; Liu, S.; Zheng, B.-J.; Jiang, S. The spike protein of SARS-CoV—A target for vaccine and therapeutic development. Nat. Rev. Microbiol. 2009, 7, 226–236. [Google Scholar] [CrossRef] [PubMed]
- Wan, Y.; Shang, J.; Graham, R.; Baric, R.S.; Li, F. Receptor Recognition by the Novel Coronavirus from Wuhan: An Analysis Based on Decade-Long Structural Studies of SARS Coronavirus. J. Virol. 2020, 94, e00127-20. [Google Scholar] [CrossRef]
- Malick, M.S.S.; Fernandes, H. The Genomic Landscape of Severe Acute Respiratory Syndrome Coronavirus 2. Adv. Mol. Pathol. 2021, 4, 231–235. [Google Scholar] [CrossRef]
- Wang, G.-L.; Wang, Z.-Y.; Duan, L.-J.; Meng, Q.-C.; Jiang, M.-D.; Cao, J.; Yao, L.; Zhu, K.-L.; Cao, W.-C.; Ma, M.-J. Susceptibility of Circulating SARS-CoV-2 Variants to Neutralization. N. Engl. J. Med. 2021, 384, 2354–2356. [Google Scholar] [CrossRef]
- Bernal, J.L.; Andrews, N.; Gower, C.; Gallagher, E.; Simmons, R.; Thelwall, S.; Stowe, J.; Tessier, E.; Groves, N.; Dabrera, G.; et al. Effectiveness of Covid-19 Vaccines against the B.1.617.2 (Delta) Variant. N. Engl. J. Med. 2021, 385, 585–594. [Google Scholar] [CrossRef]
- WHO Tracking SARS-CoV-2 Variants. Available online: https://www.who.int/en/activities/tracking-SARS-CoV-2-variants/ (accessed on 13 August 2021).
- Zhou, P.; Yang, X.-L.; Wang, X.-G.; Hu, B.; Zhang, L.; Zhang, W.; Si, H.-R.; Zhu, Y.; Li, B.; Huang, C.-L.; et al. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature 2020, 579, 270–273. [Google Scholar] [CrossRef]
- Andersen, K.G.; Rambaut, A.; Lipkin, W.I.; Holmes, E.C.; Garry, R.F. The proximal origin of SARS-CoV-2. Nat. Med. 2020, 26, 450–452. [Google Scholar] [CrossRef]
- Yuan, M.; Wu, N.C.; Zhu, X.; Lee, C.-C.D.; So, R.T.Y.; Lv, H.; Mok, C.K.P.; Wilson, I.A. A highly conserved cryptic epitope in the receptor binding domains of SARS-CoV-2 and SARS-CoV. Science 2020, 368, 630–633. [Google Scholar] [CrossRef]
- Seib, K.; Dougan, G.; Rappuoli, R. The Key Role of Genomics in Modern Vaccine and Drug Design for Emerging Infectious Diseases. PLoS Genet. 2009, 5, e1000612. [Google Scholar] [CrossRef]
- Chakraborty, S.; Mallajosyula, V.; Tato, C.M.; Tan, G.S.; Wang, T.T. SARS-CoV-2 vaccines in advanced clinical trials: Where do we stand? Adv. Drug Deliv. Rev. 2021, 172, 314–338. [Google Scholar] [CrossRef]
- Graham, B.S. Rapid COVID-19 vaccine development. Science 2020, 368, 945–946. [Google Scholar] [CrossRef]
- Wang, J.; Peng, Y.; Xu, H.; Cui, Z.; Williams, R. The COVID-19 Vaccine Race: Challenges and Opportunities in Vaccine Formulation. AAPS PharmSciTech 2020, 21, 225. [Google Scholar] [CrossRef] [PubMed]
- Allen, A. For Billion-Dollar COVID Vaccines, Basic Government-Funded Science Laid the Groundwork. Available online: https://www.scientificamerican.com/article/for-billion-dollar-covid-vaccines-basic-government-funded-science-laid-the-groundwork/ (accessed on 26 January 2021).
- Thanh Le, T.; Andreadakis, Z.; Kumar, A.; Gómez Román, R.; Tollefsen, S.; Saville, M.; Mayhew, S. The COVID-19 vaccine development landscape. Nat. Rev. Drug Discov. 2020, 19, 305–306. [Google Scholar] [CrossRef] [PubMed]
- Conte, C.; Sogni, F.; Affanni, P.; Veronesi, L.; Argentiero, A.; Esposito, S. Vaccines against Coronaviruses: The State of the Art. Vaccines 2020, 8, 309. [Google Scholar] [CrossRef] [PubMed]
- Bradley, E.S.; McNeel, D.G. DNA Vaccines. In Cancer Therapeutic Targets; Springer: New York, NY, USA, 2017; Volume 1–2, pp. 183–198. ISBN 9781441907172. [Google Scholar]
- Pardi, N.; Weissman, D. Measuring the Adjuvant Activity of RNA Vaccines. In Methods in Molecular Biology; Humana Press Inc.: Totowa, NJ, USA, 2017; Volume 1499, pp. 143–153. [Google Scholar]
- Wang, Y.; Zhang, Z.; Luo, J.; Han, X.; Wei, Y.; Wei, X. mRNA vaccine: A potential therapeutic strategy. Mol. Cancer 2021, 20, 33. [Google Scholar] [CrossRef]
- Iavarone, C.; O’Hagan, D.T.; Yu, D.; Delahaye, N.F.; Ulmer, J.B. Mechanism of action of mRNA-based vaccines. Expert Rev. Vaccines 2017, 16, 871–881. [Google Scholar] [CrossRef]
- Ho, W.; Gao, M.; Li, F.; Li, Z.; Zhang, X.; Xu, X. Next-Generation Vaccines: Nanoparticle-Mediated DNA and mRNA Delivery. Adv. Healthc. Mater. 2021, 10, e2001812. [Google Scholar] [CrossRef]
- Pardi, N.; Hogan, M.J.; Weissman, D. Recent advances in mRNA vaccine technology. Curr. Opin. Immunol. 2020, 65, 14–20. [Google Scholar] [CrossRef]
- Jahanafrooz, Z.; Baradaran, B.; Mosafer, J.; Hashemzaei, M.; Rezaei, T.; Mokhtarzadeh, A.; Hamblin, M.R. Comparison of DNA and mRNA vaccines against cancer. Drug Discov. Today 2019, 25, 552–560. [Google Scholar] [CrossRef]
- Fuller, D.H.; Berglund, P. Amplifying RNA Vaccine Development. N. Engl. J. Med. 2020, 382, 2469–2471. [Google Scholar] [CrossRef]
- Brenner, S.; Jacob, F.; Meselson, M. An Unstable Intermediate Carrying Information from Genes to Ribosomes for Protein Synthesis. Nature 1961, 190, 576–581. [Google Scholar] [CrossRef]
- Malone, R.W.; Felgner, P.L.; Verma, I.M. Cationic liposome-mediated RNA transfection. Proc. Natl. Acad. Sci. USA 1989, 86, 6077–6081. [Google Scholar] [CrossRef]
- Wolff, J.A.; Malone, R.W.; Williams, P.; Chong, W.; Acsadi, G.; Jani, A.; Felgner, P.L. Direct gene transfer into mouse muscle in vivo. Science 1990, 247, 1465–1468. [Google Scholar] [CrossRef] [PubMed]
- Martinon, F.; Krishnan, S.; Lenzen, G.; Magné, R.; Gomard, E.; Guillet, J.-G.; Lévy, J.-P.; Meulien, P. Induction of virus-specific cytotoxic T lymphocytesin vivo by liposome-entrapped mRNA. Eur. J. Immunol. 1993, 23, 1719–1722. [Google Scholar] [CrossRef] [PubMed]
- Zhou, X.; Berglund, P.; Rhodes, G.; Parker, S.; Jondal, M.; Liljeström, P. Self-replicating Semliki Forest virus RNA as recombinant vaccine. Vaccine 1994, 12, 1510–1514. [Google Scholar] [CrossRef]
- Conry, R.M.; LoBuglio, A.F.; Wright, M.; Sumerel, L.; Pike, M.J.; Johanning, F.; Benjamin, R.; Lu, D.; Curiel, D.T. Characterization of a messenger RNA polynucleotide vaccine vector. Cancer Res. 1995, 55, 1397–1400. [Google Scholar] [PubMed]
- Boczkowski, D.; Nair, S.K.; Snyder, D.; Gilboa, E. Dendritic cells pulsed with RNA are potent antigen-presenting cells in vitro and in vivo. J. Exp. Med. 1996, 184, 465–472. [Google Scholar] [CrossRef] [PubMed]
- Hoerr, I.; Obst, R.; Rammensee, H.G.; Jung, G. In vivo application of RNA leads to induction of specific cytotoxic T lymphocytes and antibodies. Eur. J. Immunol. 2000, 30, 1–7. [Google Scholar] [CrossRef]
- Bloom, K.; Berg, F.V.D.; Arbuthnot, P. Self-amplifying RNA vaccines for infectious diseases. Gene Ther. 2020, 28, 117–129. [Google Scholar] [CrossRef]
- Rosa, S.S.; Prazeres, D.M.; Azevedo, A.M.; Marques, M.P. mRNA vaccines manufacturing: Challenges and bottlenecks. Vaccine 2021, 39, 2190–2200. [Google Scholar] [CrossRef] [PubMed]
- Deering, R.P.; Kommareddy, S.; Ulmer, J.B.; Brito, L.A.; Geall, A.J. Nucleic acid vaccines: Prospects for non-viral delivery of mRNA vaccines. Expert Opin. Drug Deliv. 2014, 11, 885–899. [Google Scholar] [CrossRef]
- Liu, M.A. Immunologic Basis of Vaccine Vectors. Immunity 2010, 33, 504–515. [Google Scholar] [CrossRef]
- Beelman, C.A.; Parker, R. Degradation of mRNA in eukaryotes. Cell 1995, 81, 179–183. [Google Scholar] [CrossRef]
- Pogocki, D.; Schöneich, C. Chemical stability of nucleic acid–derived drugs. J. Pharm. Sci. 2000, 89, 443–456. [Google Scholar] [CrossRef]
- Mintzer, M.A.; Simanek, E.E. Nonviral Vectors for Gene Delivery. Chem. Rev. 2008, 109, 259–302. [Google Scholar] [CrossRef]
- Martin, M.E.; Rice, K.G. Peptide-guided gene delivery. AAPS J. 2007, 9, E18–E29. [Google Scholar] [CrossRef]
- Pack, D.W.; Hoffman, A.S.; Pun, S.; Stayton, P. Design and development of polymers for gene delivery. Nat. Rev. Drug Discov. 2005, 4, 581–593. [Google Scholar] [CrossRef]
- Lee, H.; Lytton-Jean, A.; Chen, Y.; Love, K.T.; Park, A.I.; Karagiannis, E.; Sehgal, A.; Querbes, W.; Zurenko, C.S.; Jayaraman, M.; et al. Molecularly self-assembled nucleic acid nanoparticles for targeted in vivo siRNA delivery. Nat. Nanotechnol. 2012, 7, 389–393. [Google Scholar] [CrossRef] [PubMed]
- Perche, F.; Benvegnu, T.; Berchel, M.; Lebegue, L.; Pichon, C.; Jaffrès, P.-A.; Midoux, P. Enhancement of dendritic cells transfection in vivo and of vaccination against B16F10 melanoma with mannosylated histidylated lipopolyplexes loaded with tumor antigen messenger RNA. Nanomed. Nanotechnol. Biol. Med. 2011, 7, 445–453. [Google Scholar] [CrossRef] [PubMed]
- Thalhamer, J.; Weiss, R.; Scheiblhofer, S. (Eds.) Gene Vaccines; Springer: Vienna, Austria, 2012; ISBN 978-3-7091-0438-5. [Google Scholar]
- Reichmuth, A.M.; Oberli, M.A.; Jaklenec, A.; Langer, R.; Blankschtein, D. mRNA vaccine delivery using lipid nanoparticles. Ther. Deliv. 2016, 7, 319–334. [Google Scholar] [CrossRef] [PubMed]
- De Jong, S.; Chikh, G.; Sekirov, L.; Raney, S.; Semple, S.; Klimuk, S.; Yuan, N.; Hope, M.; Cullis, P.; Tam, Y. Encapsulation in liposomal nanoparticles enhances the immunostimulatory, adjuvant and anti-tumor activity of subcutaneously administered CpG ODN. Cancer Immunol. Immunother. 2007, 56, 1251–1264. [Google Scholar] [CrossRef] [PubMed]
- Pascolo, S. The messenger’s great message for vaccination. Expert Rev. Vaccines 2015, 14, 153–156. [Google Scholar] [CrossRef]
- WHO Evaluation of the Quality, Safety and Efficacy of RNA-Based. Available online: https://www.who.int/docs/default-source/biologicals/ecbs/reg-considerations-on-rna-vaccines_1st-draft_pc_tz_22122020.pdf?sfvrsn=c13e1e20_3 (accessed on 28 January 2021).
- Pardi, N.; Hogan, M.J.; Porter, F.W.; Weissman, D. mRNA vaccines—A new era in vaccinology. Nat. Rev. Drug Discov. 2018, 17, 261–279. [Google Scholar] [CrossRef]
- Coulie, P.G.; Van den Eynde, B.J.; Van Der Bruggen, P.; Boon-Falleur, T. Tumour antigens recognized by T lymphocytes: At the core of cancer immunotherapy. Nat. Rev. Cancer 2014, 14, 135–146. [Google Scholar] [CrossRef]
- Xu, S.; Yang, K.; Li, R.; Zhang, L. mRNA Vaccine Era—Mechanisms, Drug Platform and Clinical Prospection. Int. J. Mol. Sci. 2020, 21, 6582. [Google Scholar] [CrossRef]
- Zhang, R.; Billingsley, M.; Mitchell, M.J. Biomaterials for vaccine-based cancer immunotherapy. J. Control. Release 2018, 292, 256–276. [Google Scholar] [CrossRef] [PubMed]
- McNamara, M.A.; Nair, S.K.; Holl, E.K. RNA-Based Vaccines in Cancer Immunotherapy. J. Immunol. Res. 2015, 2015, 1–9. [Google Scholar] [CrossRef]
- Karikó, K.; Muramatsu, H.; Ludwig, J.; Weissman, D. Generating the optimal mRNA for therapy: HPLC purification eliminates immune activation and improves translation of nucleoside-modified, protein-encoding mRNA. Nucleic Acids Res. 2011, 39, e142. [Google Scholar] [CrossRef]
- Kranz, L.M.; Diken, M.; Haas, H.; Kreiter, S.; Loquai, C.; Reuter, K.C.; Meng, M.; Fritz, D.; Vascotto, F.; Hefesha, H.; et al. Systemic RNA delivery to dendritic cells exploits antiviral defence for cancer immunotherapy. Nature 2016, 534, 396–401. [Google Scholar] [CrossRef] [PubMed]
- Weide, B.; Carralot, J.-P.; Reese, A.; Scheel, B.; Eigentler, T.K.; Hoerr, I.; Rammensee, H.-G.; Garbe, C.; Pascolo, S. Results of the First Phase I/II Clinical Vaccination Trial With Direct Injection of mRNA. J. Immunother. 2008, 31, 180–188. [Google Scholar] [CrossRef] [PubMed]
- Weide, B.; Pascolo, S.; Scheel, B.; Derhovanessian, E.; Pflugfelder, A.; Eigentler, T.; Pawelec, G.; Hoerr, I.; Rammensee, H.-G.; Garbe, C. Direct Injection of Protamine-protected mRNA: Results of a Phase 1/2 Vaccination Trial in Metastatic Melanoma Patients. J. Immunother. 2009, 32, 498–507. [Google Scholar] [CrossRef]
- Kowalski, P.; Rudra, A.; Miao, L.; Anderson, D.G. Delivering the Messenger: Advances in Technologies for Therapeutic mRNA Delivery. Mol. Ther. 2019, 27, 710–728. [Google Scholar] [CrossRef]
- Gilboa, E.; Vieweg, J. Cancer immunotherapy with mRNA-transfected dendritic cells. Immunol. Rev. 2004, 199, 251–263. [Google Scholar] [CrossRef]
- Bol, K.F.; Figdor, C.; Aarntzen, E.H.; Welzen, M.E.; Van Rossum, M.M.; Blokx, W.; Van De Rakt, M.W.; Scharenborg, N.M.; De Boer, A.J.; Pots, J.M.; et al. Intranodal vaccination with mRNA-optimized dendritic cells in metastatic melanoma patients. OncoImmunology 2015, 4, e1019197. [Google Scholar] [CrossRef] [PubMed]
- Timmerman, J.M.; Czerwinski, D.K.; Davis, T.A.; Hsu, F.J.; Benike, C.; Hao, Z.M.; Taidi, B.; Rajapaksa, R.; Caspar, C.B.; Okada, C.Y.; et al. Idiotype-pulsed dendritic cell vaccination for B-cell lymphoma: Clinical and immune responses in 35 patients. Blood 2002, 99, 1517–1526. [Google Scholar] [CrossRef]
- Loveland, B.E.; Zhao, A.; White, S.; Gan, H.; Hamilton, K.; Xing, P.-X.; Pietersz, G.A.; Apostolopoulos, V.; Vaughan, H.; Karanikas, V.; et al. Mannan-MUC1–Pulsed Dendritic Cell Immunotherapy: A Phase I Trial in Patients with Adenocarcinoma. Clin. Cancer Res. 2006, 12, 869–877. [Google Scholar] [CrossRef]
- Yanagisawa, R.; Koizumi, T.; Koya, T.; Sano, K.; Koido, S.; Nagai, K.; Kobayashi, M.; Okamoto, M.; Sugiyama, H.; Shimodaira, S. WT1-pulsed Dendritic Cell Vaccine Combined with Chemotherapy for Resected Pancreatic Cancer in a Phase I Study. Anticancer Res. 2018, 38, 2217–2225. [Google Scholar] [CrossRef]
- Kallen, K.-J.; Heidenreich, R.; Schnee, M.; Petsch, B.; Schlake, T.; Thess, A.; Baumhof, P.; Scheel, B.; Koch, S.D.; Fotin-Mleczek, M. A novel, disruptive vaccination technology. Hum. Vaccines Immunother. 2013, 9, 2263–2276. [Google Scholar] [CrossRef]
- Rauch, S.; Lutz, J.; Kowalczyk, A.; Schlake, T.; Heidenreich, R. RNA Vaccines; Methods in Molecular Biology; Kramps, T., Elbers, K., Eds.; Springer: New York, NY, USA, 2017; Volume 1499, ISBN 978-1-4939-6479-6. [Google Scholar]
- Kübler, H.; Scheel, B.; Gnad-Vogt, U.; Miller, K.; Schultze-Seemann, W.; Dorp, F.V.; Parmiani, G.; Hampel, C.; Wedel, S.; Trojan, L.; et al. Self-adjuvanted mRNA vaccination in advanced prostate cancer patients: A first-in-man phase I/IIa study. J. Immunother. Cancer 2015, 3, 26. [Google Scholar] [CrossRef] [PubMed]
- Sebastian, M.; Schröder, A.; Scheel, B.; Hong, H.S.; Muth, A.; von Boehmer, L.; Zippelius, A.; Mayer, F.; Reck, M.; Atanackovic, D.; et al. A phase I/IIa study of the mRNA-based cancer immunotherapy CV9201 in patients with stage IIIB/IV non-small cell lung cancer. Cancer Immunol. Immunother. 2019, 68, 799–812. [Google Scholar] [CrossRef]
- Lesterhuis, W.J.; De Vries, I.J.M.; Schreibelt, G.; Schuurhuis, D.H.; Aarntzen, E.H.; De Boer, A.; Scharenborg, N.M.; Van De Rakt, M.; Hesselink, E.J.; Figdor, C.; et al. Immunogenicity of dendritic cells pulsed with CEA peptide or transfected with CEA mRNA for vaccination of colorectal cancer patients. Anticancer. Res. 2010, 30, 5091–5098. [Google Scholar]
- Eckl, J.; Raffegerst, S.; Schnorfeil, F.; Prinz-Schulz, P.; Fingerhut, C.; Floisand, Y.; Josefsen, D.; Bigalke, I.; Pinkernell, K.; Kvalheim, G.; et al. DC Vaccination Induces Antigen Specific Immune Responses in AML Patients: A 1-Year Interim Assessment. Blood 2019, 134, 3923. [Google Scholar] [CrossRef]
- Sahin, U.; Derhovanessian, E.; Miller, M.; Kloke, B.-P.; Simon, P.; Löwer, M.; Bukur, V.; Tadmor, A.D.; Luxemburger, U.; Schrörs, B.; et al. Personalized RNA mutanome vaccines mobilize poly-specific therapeutic immunity against cancer. Nature 2017, 547, 222–226. [Google Scholar] [CrossRef] [PubMed]
- Wilgenhof, S.; Van Nuffel, A.; Benteyn, D.; Corthals, J.; Aerts, C.; Heirman, C.; Van Riet, I.; Bonehill, A.; Thielemans, K.; Neyns, B. A phase IB study on intravenous synthetic mRNA electroporated dendritic cell immunotherapy in pretreated advanced melanoma patients. Ann. Oncol. 2013, 24, 2686–2693. [Google Scholar] [CrossRef] [PubMed]
- Borch, T.H.; Engell-Noerregaard, L.; Iversen, T.Z.; Ellebaek, E.; Met, Ö.; Hansen, M.; Andersen, M.H.; Straten, P.T.; Svane, I.M. mRNA-transfected dendritic cell vaccine in combination with metronomic cyclophosphamide as treatment for patients with advanced malignant melanoma. OncoImmunology 2016, 5, e1207842. [Google Scholar] [CrossRef]
- Kongsted, P.; Borch, T.H.; Ellebaek, E.; Iversen, T.Z.; Andersen, R.; Met, Ö.; Hansen, M.; Lindberg, H.; Sengeløv, L.; Svane, I.M. Dendritic cell vaccination in combination with docetaxel for patients with metastatic castration-resistant prostate cancer: A randomized phase II study. Cytotherapy 2017, 19, 500–513. [Google Scholar] [CrossRef] [PubMed]
- Morse, M.A.; Hobeika, A.; Gwin, W.; Osada, T.; Gelles, J.; Rushing, C.; Niedzwiecki, D.; Lyerly, H. Phase I study of alphaviral vector (AVX701) in colorectal cancer patients: Comparison of immune responses in stage III and stage IV patients. J. Immunother. Cancer 2015, 3, P444. [Google Scholar] [CrossRef]
- Krienke, C.; Kolb, L.; Diken, E.; Streuber, M.; Kirchhoff, S.; Bukur, T.; Akilli-Öztürk, Ö.; Kranz, L.M.; Berger, H.; Petschenka, J.; et al. A noninflammatory mRNA vaccine for treatment of experimental autoimmune encephalomyelitis. Science 2021, 371, 145–153. [Google Scholar] [CrossRef]
- Scheiblhofer, S.; Thalhamer, J.; Weiss, R. DNA and mRNA vaccination against allergies. Pediatr. Allergy Immunol. 2018, 29, 679–688. [Google Scholar] [CrossRef]
- Sahin, U.; Karikó, K.; Türeci, Ö. MRNA-based therapeutics-developing a new class of drugs. Nat. Rev. Drug Discov. 2014, 13, 759–780. [Google Scholar] [CrossRef]
- Steinman, L.; Ho, P.P.; Robinson, W.H.; Utz, P.J.; Villoslada, P. Antigen-specific tolerance to self-antigens in protein replacement therapy, gene therapy and autoimmunity. Curr. Opin. Immunol. 2019, 61, 46–53. [Google Scholar] [CrossRef] [PubMed]
- Veiga, N.; Goldsmith, M.; Granot, Y.; Rosenblum, D.; Dammes, N.; Kedmi, R.; Ramishetti, S.; Peer, D. Cell specific delivery of modified mRNA expressing therapeutic proteins to leukocytes. Nat. Commun. 2018, 9, 1–9. [Google Scholar] [CrossRef]
- Wang, L.; Wang, F.-S.; Gershwin, M.E. Human autoimmune diseases: A comprehensive update. J. Intern. Med. 2015, 278, 369–395. [Google Scholar] [CrossRef] [PubMed]
- Steinle, H.; Weber, J.; Stoppelkamp, S.; Große-Berkenbusch, K.; Golombek, S.; Weber, M.; Canak-Ipek, T.; Trenz, S.-M.; Schlensak, C.; Avci-Adali, M. Delivery of synthetic mRNAs for tissue regeneration. Adv. Drug Deliv. Rev. 2021, 179, 114007. [Google Scholar] [CrossRef]
- Curin, M.; Khaitov, M.; Karaulov, A.; Namazova-Baranova, L.; Campana, R.; Garib, V.; Valenta, R. Next-Generation of Allergen-Specific Immunotherapies: Molecular Approaches. Curr. Allergy Asthma Rep. 2018, 18, 39. [Google Scholar] [CrossRef]
- Barnes, P.J. Therapeutic strategies for allergic diseases. Nature 1999, 402, 31–38. [Google Scholar] [CrossRef] [PubMed]
- Linhart, B.; Valenta, R. Vaccines for allergy. Curr. Opin. Immunol. 2012, 24, 354–360. [Google Scholar] [CrossRef] [PubMed]
- Steveling-Klein, E.H.; Durham, S.R. Immunotherapy for Allergy. In Encyclopedia of Respiratory Medicine; Elsevier: Amsterdam, The Netherlands, 2022; pp. 491–502. [Google Scholar]
- Roesler, E.; Weiss, R.; Weinberger, E.E.; Fruehwirth, A.; Stoecklinger, A.; Mostböck, S.; Ferreira, F.; Thalhamer, J.; Scheiblhofer, S. Immunize and disappear—Safety-optimized mRNA vaccination with a panel of 29 allergens. J. Allergy Clin. Immunol. 2009, 124, 1070–1077.e11. [Google Scholar] [CrossRef]
- Hattinger, E.; Scheiblhofer, S.; Roesler, E.; Thalhamer, T.; Weiss, R. Prophylactic mRNA Vaccination against Allergy Confers Long-Term Memory Responses and Persistent Protection in Mice. J. Immunol. Res. 2015, 2015, 1–12. [Google Scholar] [CrossRef]
- Maruggi, G.; Zhang, C.; Li, J.; Ulmer, J.B.; Yu, D. mRNA as a Transformative Technology for Vaccine Development to Control Infectious Diseases. Mol. Ther. 2019, 27, 757–772. [Google Scholar] [CrossRef]
- Blakney, A.; Ip, S.; Geall, A. An Update on Self-Amplifying mRNA Vaccine Development. Vaccines 2021, 9, 97. [Google Scholar] [CrossRef]
- Ljungberg, K.; Liljestrom, P. Self-replicating alphavirus RNA vaccines. Expert Rev. Vaccines 2014, 14, 177–194. [Google Scholar] [CrossRef]
- Ballesteros-Briones, M.C.; Silva-Pilipich, N.; Herrador-Cañete, G.; Vanrell, L.; Smerdou, C. A new generation of vaccines based on alphavirus self-amplifying RNA. Curr. Opin. Virol. 2020, 44, 145–153. [Google Scholar] [CrossRef]
- Lundstrom, K. Self-Amplifying RNA Viruses as RNA Vaccines. Int. J. Mol. Sci. 2020, 21, 5130. [Google Scholar] [CrossRef] [PubMed]
- Vogel, A.B.; Lambert, L.; Kinnear, E.; Busse, D.; Erbar, S.; Reuter, K.C.; Wicke, L.; Perkovic, M.; Beissert, T.; Haas, H.; et al. Self-Amplifying RNA Vaccines Give Equivalent Protection against Influenza to mRNA Vaccines but at Much Lower Doses. Mol. Ther. 2017, 26, 446–455. [Google Scholar] [CrossRef]
- Blakney, A.K.; McKay, P.F.; Christensen, D.; Yus, B.I.; Aldon, Y.; Follmann, F.; Shattock, R.J. Effects of cationic adjuvant formulation particle type, fluidity and immunomodulators on delivery and immunogenicity of saRNA. J. Control. Release 2019, 304, 65–74. [Google Scholar] [CrossRef]
- Luo, F.; Zheng, L.; Hu, Y.; Liu, S.; Wang, Y.; Xiong, Z.; Hu, X.; Tan, F. Induction of Protective Immunity against Toxoplasma gondii in Mice by Nucleoside Triphosphate Hydrolase-II (NTPase-II) Self-amplifying RNA Vaccine Encapsulated in Lipid Nanoparticle (LNP). Front. Microbiol. 2017, 8, 605. [Google Scholar] [CrossRef]
- Garcia, A.B.; Siu, E.; Sun, T.; Exler, V.; Brito, L.; Hekele, A.; Otten, G.; Augustijn, K.; Janse, C.J.; Ulmer, J.B.; et al. Neutralization of the Plasmodium-encoded MIF ortholog confers protective immunity against malaria infection. Nat. Commun. 2018, 9, 2714. [Google Scholar] [CrossRef] [PubMed]
- Duthie, M.; Van Hoeven, N.; Macmillen, Z.; Picone, A.; Mohamath, R.; Erasmus, J.; Hsu, F.-C.; Stinchcomb, D.T.; Reed, S.G. Heterologous Immunization With Defined RNA and Subunit Vaccines Enhances T Cell Responses That Protect Against Leishmania donovani. Front. Immunol. 2018, 9, 2420. [Google Scholar] [CrossRef]
- Raj, D.K.; Das Mohapatra, A.; Jnawali, A.; Zuromski, J.; Jha, A.; Cham-Kpu, G.; Sherman, B.; Rudlaff, R.M.; Nixon, C.E.; Hilton, N.; et al. Anti-PfGARP activates programmed cell death of parasites and reduces severe malaria. Nature 2020, 582, 104–108. [Google Scholar] [CrossRef]
- Maruggi, G.; Chiarot, E.; Giovani, C.; Buccato, S.; Bonacci, S.; Frigimelica, E.; Margarit, I.; Geall, A.; Bensi, G.; Maione, D. Immunogenicity and protective efficacy induced by self-amplifying mRNA vaccines encoding bacterial antigens. Vaccine 2017, 35, 361–368. [Google Scholar] [CrossRef]
- Corbett, K.S.; Edwards, D.K.; Leist, S.R.; Abiona, O.M.; Boyoglu-Barnum, S.; Gillespie, R.A.; Himansu, S.; Schäfer, A.; Ziwawo, C.T.; DiPiazza, A.T.; et al. SARS-CoV-2 mRNA vaccine design enabled by prototype pathogen preparedness. Nature 2020, 586, 567–571. [Google Scholar] [CrossRef] [PubMed]
- Zhong, Z.; Catani, J.P.P.; Mc Cafferty, S.; Couck, L.; Broeck, W.V.D.; Gorlé, N.; Vandenbroucke, R.E.; Devriendt, B.; Ulbert, S.; Cnops, L.; et al. Immunogenicity and Protection Efficacy of a Naked Self-Replicating mRNA-Based Zika Virus Vaccine. Vaccines 2019, 7, 96. [Google Scholar] [CrossRef]
- Roth, C.; Cantaert, T.; Colas, C.; Prot, M.; Casadémont, I.; Levillayer, L.; Thalmensi, J.; Langlade-Demoyen, P.; Gerke, C.; Bahl, K.; et al. A Modified mRNA Vaccine Targeting Immunodominant NS Epitopes Protects Against Dengue Virus Infection in HLA Class I Transgenic Mice. Front. Immunol. 2019, 10, 1424. [Google Scholar] [CrossRef] [PubMed]
- Szurgot, I.; Ljungberg, K.; Kümmerer, B.M.; Liljeström, P. Infectious RNA vaccine protects mice against chikungunya virus infection. Sci. Rep. 2020, 10, 21076. [Google Scholar] [CrossRef]
- Aldrich, C.; Leroux–Roels, I.; Huang, K.B.; Bica, M.A.; Loeliger, E.; Schoenborn-Kellenberger, O.; Walz, L.; Leroux-Roels, G.; von Sonnenburg, F.; Oostvogels, L. Proof-of-concept of a low-dose unmodified mRNA-based rabies vaccine formulated with lipid nanoparticles in human volunteers: A phase 1 trial. Vaccine 2021, 39, 1310–1318. [Google Scholar] [CrossRef]
- Meyer, M.; Huang, E.; Yuzhakov, O.; Ramanathan, P.; Ciaramella, G.; Bukreyev, A. Modified mRNA-Based Vaccines Elicit Robust Immune Responses and Protect Guinea Pigs From Ebola Virus Disease. J. Infect. Dis. 2017, 217, 451–455. [Google Scholar] [CrossRef]
- Egan, K.P.; Hook, L.M.; Naughton, A.; Pardi, N.; Awasthi, S.; Cohen, G.H.; Weissman, D.; Friedman, H.M. An HSV-2 nucleoside-modified mRNA genital herpes vaccine containing glycoproteins gC, gD, and gE protects mice against HSV-1 genital lesions and latent infection. PLoS Pathog. 2020, 16, e1008795. [Google Scholar] [CrossRef] [PubMed]
- Nelson, C.S.; Jenks, J.A.; Pardi, N.; Goodwin, M.; Roark, H.; Edwards, W.; McLellan, J.S.; Pollara, J.; Weissman, D.; Permar, S.R. Human Cytomegalovirus Glycoprotein B Nucleoside-Modified mRNA Vaccine Elicits Antibody Responses with Greater Durability and Breadth than MF59-Adjuvanted gB Protein Immunization. J. Virol. 2020, 94, e00186-20. [Google Scholar] [CrossRef]
- D’Haese, S.; Lacroix, C.; Garcia, F.; Plana, M.; Ruta, S.; Vanham, G.; Verrier, B.; Aerts, J.L. Off the beaten path: Novel mRNA-nanoformulations for therapeutic vaccination against HIV. J. Control. Release 2020, 330, 1016–1033. [Google Scholar] [CrossRef]
- Fleeton, M.N.; Chen, M.S.; Berglund, P.; Rhodes, G.; Parker, S.E.; Murphy, M.; Atkins, G.J.; Liljestrom, P. Self-Replicative RNA Vaccines Elicit Protection against Influenza A Virus, Respiratory Syncytial Virus, and a Tickborne Encephalitis Virus. J. Infect. Dis. 2001, 183, 1395–1398. [Google Scholar] [CrossRef] [PubMed]
- Petsch, B.; Schnee, M.; Vogel, A.B.; Lange, E.; Hoffmann, B.; Voss, D.; Schlake, T.; Thess, A.; Kallen, K.-J.; Stitz, L.; et al. Protective efficacy of in vitro synthesized, specific mRNA vaccines against influenza A virus infection. Nat. Biotechnol. 2012, 30, 1210–1216. [Google Scholar] [CrossRef]
- Wei, C.-J.; Crank, M.C.; Shiver, J.; Graham, B.S.; Mascola, J.R.; Nabel, G.J. Next-generation influenza vaccines: Opportunities and challenges. Nat. Rev. Drug Discov. 2020, 19, 239–252. [Google Scholar] [CrossRef]
- Jacobson, J.M.; Routy, J.-P.; Welles, S.; DeBenedette, M.; Tcherepanova, I.; Angel, J.B.; Asmuth, D.M.; Stein, D.K.; Baril, J.-G.; McKellar, M.; et al. Dendritic Cell Immunotherapy for HIV-1 Infection Using Autologous HIV-1 RNA. JAIDS J. Acquir. Immune Defic. Syndr. 2016, 72, 31–38. [Google Scholar] [CrossRef] [PubMed]
- Gandhi, R.T.; Kwon, D.S.; Macklin, E.; Shopis, J.R.; McLean, A.P.; McBrine, N.; Flynn, T.; Peter, L.; Sbrolla, A.; Kaufmann, D.E.; et al. Immunization of HIV-1-Infected Persons With Autologous Dendritic Cells Transfected With mRNA Encoding HIV-1 Gag and Nef. JAIDS J. Acquir. Immune Defic. Syndr. 2016, 71, 246–253. [Google Scholar] [CrossRef]
- Alberer, M.; Gnad-Vogt, U.; Hong, H.S.; Mehr, K.T.; Backert, L.; Finak, G.; Gottardo, R.; Bica, M.A.; Garofano, A.; Koch, S.D.; et al. Safety and immunogenicity of a mRNA rabies vaccine in healthy adults: An open-label, non-randomised, prospective, first-in-human phase 1 clinical trial. Lancet 2017, 390, 1511–1520. [Google Scholar] [CrossRef]
- Shaw, C.; Lee, H.; Knightly, C.; Kalidindi, S.; Zaks, T.; Smolenov, I.; Panther, L. 2754. Phase 1 Trial of an mRNA-Based Combination Vaccine Against hMPV and PIV3. Open Forum Infect. Dis. 2019, 6, S970. [Google Scholar] [CrossRef]
- Shaw, C.; Panther, L.; August, A.; Zaks, T.; Smolenov, I.; Bart, S.; Watson, M. Safety and immunogenicity of a mRNA-based chikungunya vaccine in a phase 1 dose-ranging trial. Int. J. Infect. Dis. 2019, 79, 17. [Google Scholar] [CrossRef]
- Feldman, R.A.; Fuhr, R.; Smolenov, I.; Ribeiro, A.; Panther, L.; Watson, M.; Senn, J.J.; Smith, M.; Almarsson, Ö.; Pujar, H.S.; et al. mRNA vaccines against H10N8 and H7N9 influenza viruses of pandemic potential are immunogenic and well tolerated in healthy adults in phase 1 randomized clinical trials. Vaccine 2019, 37, 3326–3334. [Google Scholar] [CrossRef] [PubMed]
- Leal, L.; Guardo, A.C.; Morón-López, S.; Salgado, M.; Mothe, B.; Heirman, C.; Pannus, P.; Vanham, G.; Ham, H.J.V.D.; Gruters, R.; et al. Phase I clinical trial of an intranodally administered mRNA-based therapeutic vaccine against HIV-1 infection. AIDS 2018, 32, 2533–2545. [Google Scholar] [CrossRef]
- Bernstein, D.I.; Reap, E.A.; Katen, K.; Watson, A.; Smith, K.; Norberg, P.; Olmsted, R.A.; Hoeper, A.; Morris, J.; Negri, S.; et al. Randomized, double-blind, Phase 1 trial of an alphavirus replicon vaccine for cytomegalovirus in CMV seronegative adult volunteers. Vaccine 2009, 28, 484–493. [Google Scholar] [CrossRef] [PubMed]
- Wecker, M.; Gilbert, P.; Russell, N.; Hural, J.; Allen, M.; Pensiero, M.; Chulay, J.; Chiu, Y.-L.; Karim, S.A.; Burke, D.S. Phase I Safety and Immunogenicity Evaluations of an Alphavirus Replicon HIV-1 Subtype CgagVaccine in Healthy HIV-1-Uninfected Adults. Clin. Vaccine Immunol. 2012, 19, 1651–1660. [Google Scholar] [CrossRef] [PubMed]
- Padron-Regalado, E. Vaccines for SARS-CoV-2: Lessons from Other Coronavirus Strains. Infect. Dis. Ther. 2020, 9, 255–274. [Google Scholar] [CrossRef]
- The Lancet Respiratory Medicine Realising the potential of SARS-CoV-2 vaccines—A long shot? Lancet Respir. Med. 2021, 9, 117. [CrossRef]
- Forni, G.; Mantovani, A. COVID-19 vaccines: Where we stand and challenges ahead. Cell Death Differ. 2021, 28, 626–639. [Google Scholar] [CrossRef]
- Ledford, H. Moderna COVID vaccine becomes second to get US authorization. Nature 2020. [Google Scholar] [CrossRef]
- WHO. Draft Landscape and Tracker of COVID-19 Candidate Vaccines. Available online: https://www.who.int/publications/m/item/draft-landscape-of-covid-19-candidate-vaccines (accessed on 1 October 2021).
- Pallesen, J.; Wang, N.; Corbett, K.S.; Wrapp, D.; Kirchdoerfer, R.N.; Turner, H.L.; Cottrell, C.A.; Becker, M.M.; Wang, L.; Shi, W.; et al. Immunogenicity and structures of a rationally designed prefusion MERS-CoV spike antigen. Proc. Natl. Acad. Sci. USA 2017, 114, E7348–E7357. [Google Scholar] [CrossRef] [PubMed]
- McKay, P.F.; Hu, K.; Blakney, A.K.; Samnuan, K.; Brown, J.C.; Penn, R.; Zhou, J.; Bouton, C.R.; Rogers, P.; Polra, K.; et al. Self-amplifying RNA SARS-CoV-2 lipid nanoparticle vaccine candidate induces high neutralizing antibody titers in mice. Nat. Commun. 2020, 11, 3–9. [Google Scholar] [CrossRef]
- Vogel, A.B.; Kanevsky, I.; Che, Y.; Swanson, K.A.; Muik, A.; Vormehr, M.; Kranz, L.M.; Walzer, K.C.; Hein, S.; Güler, A.; et al. BNT162b vaccines protect rhesus macaques from SARS-CoV-2. Nature 2021, 592, 283–289. [Google Scholar] [CrossRef] [PubMed]
- Erasmus, J.H.; Khandhar, A.P.; O’Connor, M.A.; Walls, A.C.; Hemann, E.A.; Murapa, P.; Archer, J.; Leventhal, S.; Fuller, J.T.; Lewis, T.B.; et al. An Alphavirus -derived replicon RNA vaccine induces SARS-CoV-2 neutralizing antibody and T cell responses in mice and nonhuman primates. Sci. Transl. Med. 2020, 12, eabc9396. [Google Scholar] [CrossRef]
- Rauch, S.; Roth, N.; Schwendt, K.; Fotin-Mleczek, M.; Mueller, S.O.; Petsch, B. mRNA-based SARS-CoV-2 vaccine candidate CVnCoV induces high levels of virus-neutralising antibodies and mediates protection in rodents. npj Vaccines 2021, 6, 1–9. [Google Scholar] [CrossRef]
- Zhang, N.-N.; Li, X.-F.; Deng, Y.-Q.; Zhao, H.; Huang, Y.-J.; Yang, G.; Huang, W.-J.; Gao, P.; Zhou, C.; Zhang, R.-R.; et al. A Thermostable mRNA Vaccine against COVID-19. Cell 2020, 182, 1271–1283.e16. [Google Scholar] [CrossRef] [PubMed]
- Rauch, S.; Gooch, K.; Hall, Y.; Salguero, F.J.; Dennis, M.J.; Gleeson, F.V.; Harris, D.; Ho, C.; Humphries, H.E.; Longet, S.; et al. mRNA vaccine CVnCoV protects non-human primates from SARS-CoV-2 challenge infection. bioRxiv 2020. [Google Scholar] [CrossRef]
- De Alwis, R.; Gan, E.S.; Chen, S.; Leong, Y.S.; Tan, H.C.; Zhang, S.L.; Yau, C.; Low, J.G.; Kalimuddin, S.; Matsuda, D.; et al. A single dose of self-transcribing and replicating RNA-based SARS-CoV-2 vaccine produces protective adaptive immunity in mice. Mol. Ther. 2021, 29, 1970–1983. [Google Scholar] [CrossRef] [PubMed]
- Corbett, K.S.; Flynn, B.; Foulds, K.E.; Francica, J.R.; Boyoglu-Barnum, S.; Werner, A.P.; Flach, B.; O’Connell, S.; Bock, K.W.; Minai, M.; et al. Evaluation of the mRNA-1273 Vaccine against SARS-CoV-2 in Nonhuman Primates. N. Engl. J. Med. 2020, 383, 1544–1555. [Google Scholar] [CrossRef]
- Neumann, P.J.; Cohen, J.T.; Kim, D.D.; Ollendorf, D.A. Consideration Of Value-Based Pricing For Treatments And Vaccines Is Important, Even In The COVID-19 Pandemic. Health Aff. 2021, 40, 53–61. [Google Scholar] [CrossRef]
- Belete, T.M. Review on Up-to-Date Status of Candidate Vaccines for COVID-19 Disease. Infect. Drug Resist. 2021, 14, 151–161. [Google Scholar] [CrossRef] [PubMed]
- Gao, Q.; Bao, L.; Mao, H.; Wang, L.; Xu, K.; Yang, M.; Li, Y.; Zhu, L.; Wang, N.; Lv, Z.; et al. Development of an inactivated vaccine candidate for SARS-CoV-2. Science 2020, 369, 77–81. [Google Scholar] [CrossRef]
- Mullard, A. COVID-19 vaccine development pipeline gears up. Lancet 2020, 395, 1751–1752. [Google Scholar] [CrossRef]
- Anderson, E.J.; Rouphael, N.G.; Widge, A.T.; Jackson, L.A.; Roberts, P.C.; Makhene, M.; Chappell, J.D.; Denison, M.R.; Stevens, L.J.; Pruijssers, A.J.; et al. Safety and Immunogenicity of SARS-CoV-2 mRNA-1273 Vaccine in Older Adults. N. Engl. J. Med. 2020, 383, 2427–2438. [Google Scholar] [CrossRef] [PubMed]
- Widge, A.T.; Rouphael, N.G.; Jackson, L.A.; Anderson, E.J.; Roberts, P.C.; Makhene, M.; Chappell, J.D.; Denison, M.R.; Stevens, L.J.; Pruijssers, A.J.; et al. Durability of Responses after SARS-CoV-2 mRNA-1273 Vaccination. N. Engl. J. Med. 2021, 384, 80–82. [Google Scholar] [CrossRef]
- Shenoy, P. Multi-regional clinical trials and global drug development. Perspect. Clin. Res. 2016, 7, 62–67. [Google Scholar] [CrossRef] [PubMed]
- Baden, L.R.; El Sahly, H.M.; Essink, B.; Kotloff, K.; Frey, S.; Novak, R.; Diemert, D.; Spector, S.A.; Rouphael, N.; Creech, C.B.; et al. Efficacy and Safety of the mRNA-1273 SARS-CoV-2 Vaccine. N. Engl. J. Med. 2021, 384, 403–416. [Google Scholar] [CrossRef] [PubMed]
- Broseta, J.J.; Rodríguez-Espinosa, D.; Soruco, E.; Maduell, F. Weekly seroconversion rate of the mRNA-1273 SARS-CoV-2 vaccine in haemodialysis patients. Nephrol. Dial. Transplant. 2021, 36, 1754–1755. [Google Scholar] [CrossRef]
- Ikizler, T.A.; Coates, P.T.; Rovin, B.H.; Ronco, P. Immune response to SARS-CoV-2 infection and vaccination in patients receiving kidney replacement therapy. Kidney Int. 2021, 99, 1275–1279. [Google Scholar] [CrossRef] [PubMed]
- Caillard, S.; Chavarot, N.; Bertrand, D.; Kamar, N.; Thaunat, O.; Moal, V.; Masset, C.; Hazzan, M.; Gatault, P.; Sicard, A.; et al. Occurrence of severe COVID-19 in vaccinated transplant patients. Kidney Int. 2021, 100, 477–479. [Google Scholar] [CrossRef] [PubMed]
- Boyarsky, B.J.; Werbel, W.A.; Avery, R.K.; Tobian, A.A.R.; Massie, A.B.; Segev, D.L.; Garonzik-Wang, J.M. Immunogenicity of a Single Dose of SARS-CoV-2 Messenger RNA Vaccine in Solid Organ Transplant Recipients. JAMA 2021, 325, 1784. [Google Scholar] [CrossRef] [PubMed]
- Jackson, L.A.; Anderson, E.J.; Rouphael, N.G.; Roberts, P.C.; Makhene, M.; Coler, R.N.; McCullough, M.P.; Chappell, J.D.; Denison, M.R.; Stevens, L.J.; et al. An mRNA Vaccine against SARS-CoV-2—Preliminary Report. N. Engl. J. Med. 2020, 383, 1920–1931. [Google Scholar] [CrossRef]
- Chu, L.; McPhee, R.; Huang, W.; Bennett, H.; Pajon, R.; Nestorova, B.; Leav, B. A preliminary report of a randomized controlled phase 2 trial of the safety and immunogenicity of mRNA-1273 SARS-CoV-2 vaccine. Vaccine 2021, 39, 2791–2799. [Google Scholar] [CrossRef]
- Walsh, E.E.; Frenck, R.W., Jr.; Falsey, A.R.; Kitchin, N.; Absalon, J.; Gurtman, A.; Lockhart, S.; Neuzil, K.; Mulligan, M.J.; Bailey, R.; et al. Safety and Immunogenicity of Two RNA-Based Covid-19 Vaccine Candidates. N. Engl. J. Med. 2020, 383, 2439–2450. [Google Scholar] [CrossRef] [PubMed]
- Polack, F.P.; Thomas, S.J.; Kitchin, N.; Absalon, J.; Gurtman, A.; Lockhart, S.; Perez, J.L.; Pérez Marc, G.; Moreira, E.D.; Zerbini, C.; et al. Safety and efficacy of the BNT162b2 mRNA Covid-19 vaccine. N. Engl. J. Med. 2020, 383, 2603–2615. [Google Scholar] [CrossRef]
- Kremsner, P.G.; Mann, P.; Kroidl, A.; Leroux-Roels, I.; Schindler, C.; Gabor, J.J.; Schunk, M.; Leroux-Roels, G.; Bosch, J.J.; Fendel, R.; et al. Safety and immunogenicity of an mRNA-lipid nanoparticle vaccine candidate against SARS-CoV-2. Wien. Klin. Wochenschr. 2021, 133, 931–941. [Google Scholar] [CrossRef] [PubMed]
- Dolgin, E. CureVac COVID vaccine let-down spotlights mRNA design challenges. Nature 2021, 594, 483. [Google Scholar] [CrossRef]
- Low, J.G.; de Alwis, R.; Chen, S.; Kalimuddin, S.; Leong, Y.S.; Mah, T.K.L.; Yuen, N.; Tan, H.C.; Zhang, S.L.; Sim, J.X.Y.; et al. A phase 1/2 randomized, double-blinded, placebo controlled ascending dose trial to assess the safety, tolerability and immunogenicity of ARCT-021 in healthy adults. medRxiv 2021. [Google Scholar] [CrossRef]
- Liu, K.; Gu, Z.; Islam, S.; Scherngell, T.; Kong, X.; Zhao, J.; Chen, X.; Hu, Y. Global landscape of patents related to human coronaviruses. Int. J. Biol. Sci. 2021, 17, 1588–1599. [Google Scholar] [CrossRef]
- Knezevic, I.; Liu, M.A.; Peden, K.; Zhou, T.; Kang, H.-N. Development of mRNA Vaccines: Scientific and Regulatory Issues. Vaccines 2021, 9, 81. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.-D.; Chi, W.-Y.; Su, J.-H.; Ferrall, L.; Hung, C.-F.; Wu, T.-C. Coronavirus vaccine development: From SARS and MERS to COVID-19. J. Biomed. Sci. 2020, 27, 104. [Google Scholar] [CrossRef] [PubMed]
- Sahin, U.; Muik, A.; Derhovanessian, E.; Vogler, I.; Kranz, L.M.; Vormehr, M.; Baum, A.; Pascal, K.; Quandt, J.; Maurus, D.; et al. COVID-19 vaccine BNT162b1 elicits human antibody and TH1 T-cell responses. Nature 2020, 586, 594–599. [Google Scholar] [CrossRef] [PubMed]
- Mulligan, M.J.; Lyke, K.E.; Kitchin, N.; Absalon, J.; Gurtman, A.; Lockhart, S.; Neuzil, K.; Raabe, V.; Bailey, R.; Swanson, K.A.; et al. Phase I/II study of COVID-19 RNA vaccine BNT162b1 in adults. Nature 2020, 586, 589–593. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Hui, A.; Zhang, X.; Yang, Y.; Tang, R.; Ye, H.; Ji, R.; Lin, M.; Zhu, Z.; Türeci, Ö.; et al. Safety and immunogenicity of the SARS-CoV-2 BNT162b1 mRNA vaccine in younger and older Chinese adults: A randomized, placebo-controlled, double-blind phase 1 study. Nat. Med. 2021, 27, 1062–1070. [Google Scholar] [CrossRef]
- NIH A Trial Investigating the Safety and Effects of Four BNT162 Vaccines Against COVID-2019 in Healthy and Immunocompromised. Adults. Available online: https://clinicaltrials.gov/ct2/show/NCT04380701 (accessed on 13 July 2021).
- Subbaraman, N. Pregnancy and COVID: What the data say. Nature 2021, 591, 193–195. [Google Scholar] [CrossRef]
- Goldshtein, I.; Nevo, D.; Steinberg, D.M.; Rotem, R.S.; Gorfine, M.; Chodick, G.; Segal, Y. Association Between BNT162b2 Vaccination and Incidence of SARS-CoV-2 Infection in Pregnant Women. JAMA 2021, 326, 728. [Google Scholar] [CrossRef]
- Peretz, S.B.; Regev, N.; Novick, L.; Nachshol, M.; Goffer, E.; Ben-David, A.; Asraf, K.; Doolman, R.; Levin, E.G.; Yochay, G.R.; et al. Short-term outcome of pregnant women vaccinated with BNT162b2 mRNA COVID-19 vaccine. Ultrasound Obstet. Gynecol. 2021, 58, 450–456. [Google Scholar] [CrossRef]
- Ledford, H.; Cyranoski, D.; Van Noorden, R. The UK has approved a COVID vaccine—Here’s what scientists now want to know. Nature 2020, 588, 205–206. [Google Scholar] [CrossRef]
- Ledford, H. US authorization of first COVID vaccine marks new phase in safety monitoring. Nature 2020, 588, 377–378. [Google Scholar] [CrossRef]
- Kim, J.H.; Marks, F.; Clemens, J.D. Looking beyond COVID-19 vaccine phase 3 trials. Nat. Med. 2021, 27, 205–211. [Google Scholar] [CrossRef] [PubMed]
- Clemens, J.; Brenner, R.; Rao, M.; Tafari, N.; Lowe, C. Evaluating New Vaccines for Developing Countries. JAMA 1996, 275, 390–397. [Google Scholar] [CrossRef] [PubMed]
- Victora, C.G.; Habicht, J.-P.; Bryce, J. Evidence-Based Public Health: Moving Beyond Randomized Trials. Am. J. Public Health 2004, 94, 400–405. [Google Scholar] [CrossRef] [PubMed]
- Chodick, G.; Tene, L.; Patalon, T.; Gazit, S.; Ben Tov, A.; Cohen, D.; Muhsen, K. Assessment of Effectiveness of 1 Dose of BNT162b2 Vaccine for SARS-CoV-2 Infection 13 to 24 Days After Immunization. JAMA Netw. Open 2021, 4, e2115985. [Google Scholar] [CrossRef] [PubMed]
- Murillo-Zamora, E.; Trujillo, X.; Huerta, M.; Riós-Silva, M.; Mendoza-Cano, O. Effectiveness of BNT162b2 COVID-19 Vaccine in Preventing Severe Symptomatic Infection among Healthcare Workers. Medicina 2021, 57, 746. [Google Scholar] [CrossRef] [PubMed]
- Bernal, J.L.; Andrews, N.; Gower, C.; Robertson, C.; Stowe, J.; Tessier, E.; Simmons, R.; Cottrell, S.; Roberts, R.; O’Doherty, M.; et al. Effectiveness of the Pfizer-BioNTech and Oxford-AstraZeneca vaccines on covid-19 related symptoms, hospital admissions, and mortality in older adults in England: Test negative case-control study. BMJ 2021, 373, n1088. [Google Scholar] [CrossRef]
- Vasileiou, E.; Simpson, C.R.; Shi, T.; Kerr, S.; Agrawal, U.; Akbari, A.; Bedston, S.; Beggs, J.; Bradley, D.; Chuter, A.; et al. Interim findings from first-dose mass COVID-19 vaccination roll-out and COVID-19 hospital admissions in Scotland: A national prospective cohort study. Lancet 2021, 397, 1646–1657. [Google Scholar] [CrossRef]
- Tenforde, M.W.; Patel, M.M.; Ginde, A.A.; Douin, D.J.; Talbot, H.K.; Casey, J.D.; Mohr, N.M.; Zepeski, A.; Gaglani, M.; McNeal, T.; et al. Effectiveness of Severe Acute Respiratory Syndrome Coronavirus 2 Messenger RNA Vaccines for Preventing Coronavirus Disease 2019 Hospitalizations in the United States. Clin. Infect. Dis. 2021, ciab687. [Google Scholar] [CrossRef]
- Zacay, G.; Shasha, D.; Bareket, R.; Kadim, I.; Sikron, F.H.; Tsamir, J.; Mossinson, D.; Heymann, A.D. BNT162b2 Vaccine Effectiveness in Preventing Asymptomatic Infection With SARS-CoV-2 Virus: A Nationwide Historical Cohort Study. Open Forum Infect. Dis. 2021, 8, ofab262. [Google Scholar] [CrossRef]
- Dagan, N.; Barda, N.; Kepten, E.; Miron, O.; Perchik, S.; Katz, M.A.; Hernán, M.A.; Lipsitch, M.; Reis, B.; Balicer, R.D. BNT162b2 mRNA Covid-19 Vaccine in a Nationwide Mass Vaccination Setting. N. Engl. J. Med. 2021, 384, 1412–1423. [Google Scholar] [CrossRef]
- FDA Coronavirus (COVID-19) Update: FDA Authorizes Pfizer-BioNTech COVID-19 Vaccine for Emergency Use in Adolescents in Another Important Action in Fight Against Pandemic. Available online: https://www.fda.gov/news-events/press-announcements/coronavirus-covid-19-update-fda-authorizes-pfizer-biontech-covid-19-vaccine-emergency-use (accessed on 9 August 2021).
- Health Canada Health Canada Authorizes Use of the Pfizer-BioNTech COVID-19 Vaccine in Children 12 to 15 Years of Age. Available online: https://www.canada.ca/en/health-canada/news/2021/05/health-canada-authorizes-use-of-the-pfizer-biontech-covid-19-vaccine-in-children-12-to-15-years-of-age.html (accessed on 9 August 2021).
- Agência Brasil Brazil Authorizes Pfizer Vaccine for Children Aged 12 and Older. Available online: https://agenciabrasil.ebc.com.br/en/saude/noticia/2021-06/brazil-authorizes-pfizer-vaccine-children-aged-12-and-older (accessed on 9 August 2021).
- EMA First COVID-19 Vaccine Approved for Children Aged 12 to 15 in EU. Available online: https://www.ema.europa.eu/en/news/first-covid-19-vaccine-approved-children-aged-12-15-eu (accessed on 9 August 2021).
- Frenck, R.W.; Klein, N.P.; Kitchin, N.; Gurtman, A.; Absalon, J.; Lockhart, S.; Perez, J.L.; Walter, E.B.; Senders, S.; Bailey, R.; et al. Safety, Immunogenicity, and Efficacy of the BNT162b2 Covid-19 Vaccine in Adolescents. N. Engl. J. Med. 2021, 385, 239–250. [Google Scholar] [CrossRef]
- Kim, J.; Eygeris, Y.; Gupta, M.; Sahay, G. Self-assembled mRNA vaccines. Adv. Drug Deliv. Rev. 2021, 170, 83–112. [Google Scholar] [CrossRef]
- Batty, C.J.; Heise, M.T.; Bachelder, E.M.; Ainslie, K.M. Vaccine formulations in clinical development for the prevention of severe acute respiratory syndrome coronavirus 2 infection. Adv. Drug Deliv. Rev. 2020, 169, 168–189. [Google Scholar] [CrossRef]
- Lutz, J.; Lazzaro, S.; Habbeddine, M.; Schmidt, K.E.; Baumhof, P.; Mui, B.L.; Tam, Y.K.; Madden, T.D.; Hope, M.J.; Heidenreich, R.; et al. Unmodified mRNA in LNPs constitutes a competitive technology for prophylactic vaccines. npj Vaccines 2017, 2, 29. [Google Scholar] [CrossRef] [PubMed]
- Baker, N.; Dolgin, E. Coronapod: CureVac disappoints in COVID vaccine trial. Nature 2021. [Google Scholar] [CrossRef] [PubMed]
- Slavov, S.N.; Patané, J.S.L.; Bezerra, R.D.S.; Giovanetti, M.; Fonseca, V.; Martins, A.J.; Viala, V.L.; Rodrigues, E.S.; Santos, E.V.; Barros, C.R.S.; et al. Genomic monitoring unveil the early detection of the SARS-CoV-2 B.1.351 (beta) variant (20H/501Y.V2) in Brazil. J. Med. Virol. 2021, 93, 6782–6787. [Google Scholar] [CrossRef] [PubMed]
- Romero, P.E.; Dávila-Barclay, A.; Salvatierra, G.; González, L.; Cuicapuza, D.; Solis, L.; Marcos-Carbajal, P.; Huancachoque, J.; Maturrano, L.; Tsukayama, P. The Emergence of SARS-CoV-2 Variant Lambda (C.37) in South America. medRxiv 2021. [Google Scholar] [CrossRef]
- Woldemeskel, B.A.; Garliss, C.C.; Blankson, J.N. SARS-CoV-2 mRNA vaccines induce broad CD4+ T cell responses that recognize SARS-CoV-2 variants and HCoV-NL63. J. Clin. Investig. 2021, 131, e149335. [Google Scholar] [CrossRef]
- Zani, A.; Caccuri, F.; Messali, S.; Bonfanti, C.; Caruso, A. Serosurvey in BNT162b2 vaccine-elicited neutralizing antibodies against authentic B.1, B.1.1.7, B.1.351, B.1.525 and P.1 SARS-CoV-2 variants. Emerg. Microbes Infect. 2021, 10, 1241–1243. [Google Scholar] [CrossRef] [PubMed]
- Jalkanen, P.; Kolehmainen, P.; Häkkinen, H.K.; Huttunen, M.; Tähtinen, P.A.; Lundberg, R.; Maljanen, S.; Reinholm, A.; Tauriainen, S.; Pakkanen, S.H.; et al. COVID-19 mRNA vaccine induced antibody responses against three SARS-CoV-2 variants. Nat. Commun. 2021, 12, 3991. [Google Scholar] [CrossRef]
- Verbeke, R.; Lentacker, I.; De Smedt, S.C.; Dewitte, H. The dawn of mRNA vaccines: The COVID-19 case. J. Control. Release 2021, 333, 511–520. [Google Scholar] [CrossRef] [PubMed]
- Karikó, K.; Muramatsu, H.; Welsh, F.A.; Ludwig, J.; Kato, H.; Akira, S.; Weissman, D. Incorporation of Pseudouridine Into mRNA Yields Superior Nonimmunogenic Vector With Increased Translational Capacity and Biological Stability. Mol. Ther. 2008, 16, 1833–1840. [Google Scholar] [CrossRef] [PubMed]
- Hoffmann, D.; Corleis, B.; Rauch, S.; Roth, N.; Mühe, J.; Halwe, N.J.; Ulrich, L.; Fricke, C.; Schön, J.; Kraft, A.; et al. CVnCoV and CV2CoV protect human ACE2 transgenic mice from ancestral B BavPat1 and emerging B.1.351 SARS-CoV-2. Nat. Commun. 2021, 12, 4048. [Google Scholar] [CrossRef] [PubMed]
- Li, D.-D.; Li, Q.-H. SARS-CoV-2: Vaccines in the pandemic era. Mil. Med. Res. 2021, 8, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Pinghui, Z. Coronavirus: Late-Stage Trial of Chinese mRNA Vaccine Candidate to Begin in Mexico. Available online: https://www.scmp.com/news/china/science/article/3133200/coronavirus-late-stage-trial-chinese-mrna-vaccine-candidate (accessed on 7 August 2021).
- Therapeutics, A. Arcturus—A clinical-stage mRNA therapeutics and vaccines company. Biopharma Deal 2021, 2682, B9. Available online: https://www.nature.com/articles/d43747-021-00073-3 (accessed on 30 September 2021).
- Zhao, J.; Zhao, S.; Ou, J.; Zhang, J.; Lan, W.; Guan, W.; Wu, X.; Yan, Y.; Zhao, W.; Wu, J.; et al. COVID-19: Coronavirus Vaccine Development Updates. Front. Immunol. 2020, 11, 3435. [Google Scholar] [CrossRef]
- Scheuber, A. COVID-19 Vaccine Secures New Government Investment. Available online: https://www.imperial.ac.uk/news/197573/covid-19-vaccine-secures-government-investment/ (accessed on 10 August 2021).
- Scheuber, A. Imperial Social Enterprise to Accelerate Low-Cost COVID-19 Vaccine. Available online: https://www.imperial.ac.uk/news/198053/imperial-social-enterprise-accelerate-low-cost-covid-19/ (accessed on 10 August 2021).
- Cagigi, A.; Loré, K. Immune Responses Induced by mRNA Vaccination in Mice, Monkeys and Humans. Vaccines 2021, 9, 61. [Google Scholar] [CrossRef]
- Abu-Raddad, L.J.; Chemaitelly, H.; Butt, A.A. Effectiveness of the BNT162b2 Covid-19 Vaccine against the B.1.1.7 and B.1.351 Variants. N. Engl. J. Med. 2021, 385, 187–189. [Google Scholar] [CrossRef]
- Emary, K.R.W.; Golubchik, T.; Aley, P.K.; Ariani, C.V.; Angus, B.; Bibi, S.; Blane, B.; Bonsall, D.; Cicconi, P.; Charlton, S.; et al. Efficacy of ChAdOx1 nCoV-19 (AZD1222) vaccine against SARS-CoV-2 variant of concern 202012/01 (B.1.1.7): An exploratory analysis of a randomised controlled trial. Lancet 2021, 397, 1351–1362. [Google Scholar] [CrossRef]
- Hitchings, M.D.; Ranzani, O.T.; Torres, M.S.S.; de Oliveira, S.B.; Almiron, M.; Said, R.; Borg, R.; Schulz, W.L.; de Oliveira, R.D.; da Silva, P.V.; et al. Effectiveness of CoronaVac among healthcare workers in the setting of high SARS-CoV-2 Gamma variant transmission in Manaus, Brazil: A test-negative case-control study. Lancet Reg. Health—Am. 2021, 1, 100025. [Google Scholar] [CrossRef]
- Wang, R.; Chen, J.; Gao, K.; Hozumi, Y.; Yin, C.; Wei, G.-W. Analysis of SARS-CoV-2 mutations in the United States suggests presence of four substrains and novel variants. Commun. Biol. 2021, 4, 228. [Google Scholar] [CrossRef]
- Cai, C.; Peng, Y.; Shen, E.; Huang, Q.; Chen, Y.; Liu, P.; Guo, C.; Feng, Z.; Gao, L.; Zhang, X.; et al. A comprehensive analysis of the efficacy and safety of COVID-19 vaccines. Mol. Ther. 2021, 29, 2794–2805. [Google Scholar] [CrossRef]
- Blasi, F.; Gramegna, A.; Sotgiu, G.; Saderi, L.; Voza, A.; Aliberti, S.; Amati, F. SARS-CoV-2 vaccines: A critical perspective through efficacy data and barriers to herd immunity. Respir. Med. 2021, 180, 106355. [Google Scholar] [CrossRef]
- Liu, Y.; Liu, J.; Xia, H.; Zhang, X.; Fontes-Garfias, C.R.; Swanson, K.A.; Cai, H.; Sarkar, R.; Chen, W.; Cutler, M.; et al. Neutralizing Activity of BNT162b2-Elicited Serum. N. Engl. J. Med. 2021, 384, 1466–1468. [Google Scholar] [CrossRef]
- Wu, K.; Werner, A.P.; Koch, M.; Choi, A.; Narayanan, E.; Stewart-Jones, G.B.; Colpitts, T.; Bennett, H.; Boyoglu-Barnum, S.; Shi, W.; et al. Serum Neutralizing Activity Elicited by mRNA-1273 Vaccine. N. Engl. J. Med. 2021, 384, 1468–1470. [Google Scholar] [CrossRef]
- Xie, X.; Liu, Y.; Liu, J.; Zhang, X.; Zou, J.; Fontes-Garfias, C.R.; Xia, H.; Swanson, K.A.; Cutler, M.; Cooper, D.; et al. Neutralization of SARS-CoV-2 spike 69/70 deletion, E484K and N501Y variants by BNT162b2 vaccine-elicited sera. Nat. Med. 2021, 27, 620–621. [Google Scholar] [CrossRef]
- Supasa, P.; Zhou, D.; Dejnirattisai, W.; Liu, C.; Mentzer, A.J.; Ginn, H.M.E.; Zhao, Y.; Duyvesteyn, H.M.; Nutalai, R.; Tuekprakhon, A.; et al. Reduced neutralization of SARS-CoV-2 B.1.1.7 variant by convalescent and vaccine sera. Cell 2021, 184, 2201–2211.e7. [Google Scholar] [CrossRef] [PubMed]
- Shen, X.; Tang, H.; McDanal, C.; Wagh, K.; Fischer, W.; Theiler, J.; Yoon, H.; Li, D.; Haynes, B.F.; Sanders, K.O.; et al. SARS-CoV-2 variant B.1.1.7 is susceptible to neutralizing antibodies elicited by ancestral spike vaccines. Cell Host Microbe 2021, 29, 529–539.e3. [Google Scholar] [CrossRef] [PubMed]
- Garcia-Beltran, W.F.; Lam, E.C.; Denis, K.S.; Nitido, A.D.; Garcia, Z.H.; Hauser, B.M.; Feldman, J.; Pavlovic, M.N.; Gregory, D.J.; Poznansky, M.C.; et al. Multiple SARS-CoV-2 variants escape neutralization by vaccine-induced humoral immunity. Cell 2021, 184, 2372–2383.e9. [Google Scholar] [CrossRef] [PubMed]
- Wang, P.; Nair, M.S.; Liu, L.; Iketani, S.; Luo, Y.; Guo, Y.; Wang, M.; Yu, J.; Zhang, B.; Kwong, P.D.; et al. Antibody resistance of SARS-CoV-2 variants B.1.351 and B.1.1.7. Nature 2021, 593, 130–135. [Google Scholar] [CrossRef]
- Faria, N.R.; Mellan, T.A.; Whittaker, C.; Claro, I.M.; Candido, D.D.S.; Mishra, S.; Crispim, M.A.E.; Sales, F.C.S.; Hawryluk, I.; McCrone, J.T.; et al. Genomics and epidemiology of the P.1 SARS-CoV-2 lineage in Manaus, Brazil. Science 2021, 372, 815–821. [Google Scholar] [CrossRef]
- Moore, J.P.; Offit, P.A. SARS-CoV-2 Vaccines and the Growing Threat of Viral Variants. JAMA 2021, 325, 821. [Google Scholar] [CrossRef] [PubMed]
- Guevara, M.L.; Persano, F.; Persano, S. Advances in Lipid Nanoparticles for mRNA-Based Cancer Immunotherapy. Front. Chem. 2020, 8, 589959. [Google Scholar] [CrossRef] [PubMed]
- Blakney, A.K.; McKay, P.F.; Yus, B.I.; Aldon, Y.; Shattock, R.J. Inside out: Optimization of lipid nanoparticle formulations for exterior complexation and in vivo delivery of saRNA. Gene Ther. 2019, 26, 363–372. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.; Tam, Y.Y.C.; Lin, P.J.; Sung, M.M.H.; Tam, Y.K.; Cullis, P.R. Influence of particle size on the in vivo potency of lipid nanoparticle formulations of siRNA. J. Control. Release 2016, 235, 236–244. [Google Scholar] [CrossRef]
- Park, K.S.; Sun, X.; Aikins, M.E.; Moon, J.J. Non-viral COVID-19 vaccine delivery systems. Adv. Drug Deliv. Rev. 2020, 169, 137–151. [Google Scholar] [CrossRef] [PubMed]
- Habrant, D.; Peuziat, P.; Colombani, T.; Dallet, L.; Gehin, J.; Goudeau, E.; Evrard, B.; Lambert, O.; Haudebourg, T.; Pitard, B. Design of Ionizable Lipids To Overcome the Limiting Step of Endosomal Escape: Application in the Intracellular Delivery of mRNA, DNA, and siRNA. J. Med. Chem. 2016, 59, 3046–3062. [Google Scholar] [CrossRef]
- Witzigmann, D.; Kulkarni, J.A.; Leung, J.; Chen, S.; Cullis, P.R.; van der Meel, R. Lipid nanoparticle technology for therapeutic gene regulation in the liver. Adv. Drug Deliv. Rev. 2020, 159, 344–363. [Google Scholar] [CrossRef] [PubMed]
- Messengers of hope. Nat. Biotechnol. 2020, 39, 1. [CrossRef]
- Buschmann, M.; Carrasco, M.; Alishetty, S.; Paige, M.; Alameh, M.; Weissman, D. Nanomaterial Delivery Systems for mRNA Vaccines. Vaccines 2021, 9, 65. [Google Scholar] [CrossRef] [PubMed]
- Arcturus Therapeutics Addressing Key Challenges of Lipid-Mediated Delivery Systems for mRNA through Innovation. Available online: https://endpts.com/sp/addressing-key-challenges-of-lipid-mediated-delivery-systems-for-mrna-through-innovation/ (accessed on 12 August 2021).
- Thi, T.; Suys, E.; Lee, J.; Nguyen, D.; Park, K.; Truong, N. Lipid-Based Nanoparticles in the Clinic and Clinical Trials: From Cancer Nanomedicine to COVID-19 Vaccines. Vaccines 2021, 9, 359. [Google Scholar] [CrossRef]
- Brito, L.A.; Chan, M.; Shaw, C.A.; Hekele, A.; Carsillo, T.; Schaefer, M.; Archer, J.; Seubert, A.; Otten, G.R.; Beard, C.W.; et al. A Cationic Nanoemulsion for the Delivery of Next-generation RNA Vaccines. Mol. Ther. 2014, 22, 2118–2129. [Google Scholar] [CrossRef]
- Decroly, E.; Ferron, F.; Lescar, J.; Canard, B. Conventional and unconventional mechanisms for capping viral mRNA. Nat. Rev. Microbiol. 2011, 10, 51–65. [Google Scholar] [CrossRef]
- Linares-Fernández, S.; Lacroix, C.; Exposito, J.-Y.; Verrier, B. Tailoring mRNA Vaccine to Balance Innate/Adaptive Immune Response. Trends Mol. Med. 2019, 26, 311–323. [Google Scholar] [CrossRef]
- Granados-Riveron, J.T.; Aquino-Jarquin, G. Engineering of the current nucleoside-modified mRNA-LNP vaccines against SARS-CoV-2. Biomed. Pharmacother. 2021, 142, 111953. [Google Scholar] [CrossRef] [PubMed]
- Von Niessen, A.G.O.; Poleganov, M.A.; Rechner, C.; Plaschke, A.; Kranz, L.; Fesser, S.; Diken, M.; Löwer, M.; Vallazza, B.; Beissert, T.; et al. Improving mRNA-Based Therapeutic Gene Delivery by Expression-Augmenting 3′ UTRs Identified by Cellular Library Screening. Mol. Ther. 2018, 27, 824–836. [Google Scholar] [CrossRef] [PubMed]
- Mauger, D.M.; Cabral, B.J.; Presnyak, V.; Su, S.V.; Reid, D.W.; Goodman, B.; Link, K.; Khatwani, N.; Reynders, J.; Moore, M.J.; et al. mRNA structure regulates protein expression through changes in functional half-life. Proc. Natl. Acad. Sci. USA 2019, 116, 24075–24083. [Google Scholar] [CrossRef] [PubMed]
- Kierzek, E.; Malgowska, M.; Lisowiec, J.; Turner, D.H.; Gdaniec, Z.; Kierzek, R. The contribution of pseudouridine to stabilities and structure of RNAs. Nucleic Acids Res. 2013, 42, 3492–3501. [Google Scholar] [CrossRef]
- Nance, K.D.; Meier, J.L. Modifications in an Emergency: The Role of N1-Methylpseudouridine in COVID-19 Vaccines. ACS Central Sci. 2021, 7, 748–756. [Google Scholar] [CrossRef]
- Schoenmaker, L.; Witzigmann, D.; Kulkarni, J.A.; Verbeke, R.; Kersten, G.; Jiskoot, W.; Crommelin, D.J. mRNA-lipid nanoparticle COVID-19 vaccines: Structure and stability. Int. J. Pharm. 2021, 601, 120586. [Google Scholar] [CrossRef] [PubMed]
- He, Y.; Li, J.; Heck, S.; Lustigman, S.; Jiang, S. Antigenic and Immunogenic Characterization of Recombinant Baculovirus-Expressed Severe Acute Respiratory Syndrome Coronavirus Spike Protein: Implication for Vaccine Design. J. Virol. 2006, 80, 5757–5767. [Google Scholar] [CrossRef]
- Volz, A.; Kupke, A.; Song, F.; Jany, S.; Fux, R.; Shams-Eldin, H.; Schmidt, J.C.; Becker, C.; Eickmann, M.; Becker, S.; et al. Protective Efficacy of Recombinant Modified Vaccinia Virus Ankara Delivering Middle East Respiratory Syndrome Coronavirus Spike Glycoprotein. J. Virol. 2015, 89, 8651–8656. [Google Scholar] [CrossRef]
- Bisht, H.; Roberts, A.; Vogel, L.; Bukreyev, A.; Collins, P.L.; Murphy, B.R.; Subbarao, K.; Moss, B. Severe acute respiratory syndrome coronavirus spike protein expressed by attenuated vaccinia virus protectively immunizes mice. Proc. Natl. Acad. Sci. USA 2004, 101, 6641–6646. [Google Scholar] [CrossRef]
- Dai, L.; Gao, G.F. Viral targets for vaccines against COVID-19. Nat. Rev. Immunol. 2020, 21, 73–82. [Google Scholar] [CrossRef]
- Wang, S.-F.; Tseng, S.-P.; Yen, C.-H.; Yang, J.-Y.; Tsao, C.-H.; Shen, C.-W.; Chen, K.-H.; Liu, F.-T.; Liu, W.-T.; Chen, Y.-M.A.; et al. Antibody-dependent SARS coronavirus infection is mediated by antibodies against spike proteins. Biochem. Biophys. Res. Commun. 2014, 451, 208–214. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.; Zhang, L.; Kuwahara, K.; Li, L.; Liu, Z.; Li, T.; Zhu, H.; Liu, J.; Xu, Y.; Xie, J.; et al. Immunodominant SARS Coronavirus Epitopes in Humans Elicited both Enhancing and Neutralizing Effects on Infection in Non-human Primates. ACS Infect. Dis. 2016, 2, 361–376. [Google Scholar] [CrossRef]
- Tseng, C.-T.; Sbrana, E.; Iwata-Yoshikawa, N.; Newman, P.C.; Garron, T.; Atmar, R.L.; Peters, C.J.; Couch, R.B. Immunization with SARS Coronavirus Vaccines Leads to Pulmonary Immunopathology on Challenge with the SARS Virus. PLoS ONE 2012, 7, e35421. [Google Scholar] [CrossRef]
- He, Y.; Zhou, Y.; Liu, S.; Kou, Z.; Li, W.; Farzan, M.; Jiang, S. Receptor-binding domain of SARS-CoV spike protein induces highly potent neutralizing antibodies: Implication for developing subunit vaccine. Biochem. Biophys. Res. Commun. 2004, 324, 773–781. [Google Scholar] [CrossRef] [PubMed]
- He, Y.; Lu, H.; Siddiqui, P.; Zhou, Y.; Jiang, S. Receptor-Binding Domain of Severe Acute Respiratory Syndrome Coronavirus Spike Protein Contains Multiple Conformation-Dependent Epitopes that Induce Highly Potent Neutralizing Antibodies. J. Immunol. 2005, 174, 4908–4915. [Google Scholar] [CrossRef]
- Kis, Z.; Kontoravdi, C.; Shattock, R.; Shah, N. Resources, Production Scales and Time Required for Producing RNA Vaccines for the Global Pandemic Demand. Vaccines 2020, 9, 3. [Google Scholar] [CrossRef] [PubMed]
- Irwin, A. What it will take to vaccinate the world against COVID-19. Nature 2021, 592, 176–178. [Google Scholar] [CrossRef]
- WHO COVID19 Vaccine Tracker. Available online: https://covid19.trackvaccines.org/ (accessed on 9 October 2021).
- Levin, E.G.; Lustig, Y.; Cohen, C.; Fluss, R.; Indenbaum, V.; Amit, S.; Doolman, R.; Asraf, K.; Mendelson, E.; Ziv, A.; et al. Waning Immune Humoral Response to BNT162b2 Covid-19 Vaccine over 6 Months. N. Engl. J. Med. 2021. [Google Scholar] [CrossRef]
- Favresse, J.; Bayart, J.-L.; Mullier, F.; Elsen, M.; Eucher, C.; Van Eeckhoudt, S.; Roy, T.; Wieers, G.; Laurent, C.; Dogné, J.-M.; et al. Antibody titres decline 3-month post-vaccination with BNT162b2. Emerg. Microbes Infect. 2021, 10, 1495–1498. [Google Scholar] [CrossRef]
- Khoury, D.S.; Cromer, D.; Reynaldi, A.; Schlub, T.E.; Wheatley, A.K.; Juno, J.A.; Subbarao, K.; Kent, S.J.; Triccas, J.A.; Davenport, M.P. Neutralizing antibody levels are highly predictive of immune protection from symptomatic SARS-CoV-2 infection. Nat. Med. 2021, 27, 1205–1211. [Google Scholar] [CrossRef] [PubMed]
- Thomas, S.J.; Moreira, E.D.; Kitchin, N.; Absalon, J.; Gurtman, A.; Lockhart, S.; Perez, J.L.; Pérez Marc, G.; Polack, F.P.; Zerbini, C.; et al. Six Month Safety and Efficacy of the BNT162b2 mRNA COVID-19 Vaccine. medRxiv 2021. [Google Scholar] [CrossRef]
- Benotmane, I.; Gautier, G.; Perrin, P.; Olagne, J.; Cognard, N.; Fafi-Kremer, S.; Caillard, S. Antibody Response After a Third Dose of the mRNA-1273 SARS-CoV-2 Vaccine in Kidney Transplant Recipients With Minimal Serologic Response to 2 Doses. JAMA 2021, 326, 1063. [Google Scholar] [CrossRef] [PubMed]
- Hall, V.G.; Ferreira, V.H.; Ku, T.; Ierullo, M.; Majchrzak-Kita, B.; Chaparro, C.; Selzner, N.; Schiff, J.; McDonald, M.; Tomlinson, G.; et al. Randomized Trial of a Third Dose of mRNA-1273 Vaccine in Transplant Recipients. N. Engl. J. Med. 2021, 385, 1244–1246. [Google Scholar] [CrossRef]
- Del Bello, A.; Abravanel, F.; Marion, O.; Couat, C.; Esposito, L.; Lavayssière, L.; Izopet, J.; Kamar, N. Efficiency of a boost with a third dose of anti-SARS-CoV-2 messenger RNA-based vaccines in solid organ transplant recipients. Am. J. Transplant. 2021. [Google Scholar] [CrossRef]
- Burki, T. Booster shots for COVID-19—The debate continues. Lancet Infect. Dis. 2021, 21, 1359–1360. [Google Scholar] [CrossRef]
- Callaway, E.; Ledford, H. How to redesign COVID vaccines so they protect against variants. Nature 2021, 590, 15–16. [Google Scholar] [CrossRef] [PubMed]
- Engelbrecht, F.A.; Scholes, R.J. Test for Covid-19 seasonality and the risk of second waves. One Health 2020, 12, 100202. [Google Scholar] [CrossRef] [PubMed]
- Abdulla, Z.; Al-Bashir, S.; Al-Salih, N.; Aldamen, A.; Abdulazeez, M. A Summary of the SARS-CoV-2 Vaccines and Technologies Available or under Development. Pathogens 2021, 10, 788. [Google Scholar] [CrossRef]
- Sandbrink, J.B.; Shattock, R.J. RNA Vaccines: A Suitable Platform for Tackling Emerging Pandemics? Front. Immunol. 2020, 11, 3329. [Google Scholar] [CrossRef]
- Crommelin, D.J.; Anchordoquy, T.J.; Volkin, D.B.; Jiskoot, W.; Mastrobattista, E. Addressing the Cold Reality of mRNA Vaccine Stability. J. Pharm. Sci. 2020, 110, 997–1001. [Google Scholar] [CrossRef]
- The Lancet Infectious Diseases. COVID-19 vaccine equity and booster doses. Lancet Infect. Dis. 2021, 21, 1193. [Google Scholar] [CrossRef]
- Choi, E.M. COVID-19 vaccines for low- and middle-income countries. Trans. R. Soc. Trop. Med. Hyg. 2021, 115, 447–456. [Google Scholar] [CrossRef]
- Santos, A.; Gaspar, P.; de Souza, H. Refrigeration of COVID-19 Vaccines: Ideal Storage Characteristics, Energy Efficiency and Environmental Impacts of Various Vaccine Options. Energies 2021, 14, 1849. [Google Scholar] [CrossRef]
- Ball, R.L.; Bajaj, P.; Whitehead, K.A. Achieving long-term stability of lipid nanoparticles: Examining the effect of pH, temperature, and lyophilization. Int. J. Nanomed. 2016, 12, 305–315. [Google Scholar] [CrossRef]
- Maxmen, A. The fight to manufacture COVID vaccines in lower-income countries. Nature 2021, 597, 455–457. [Google Scholar] [CrossRef]
- Hotez, P.J.; Narayan, K.M.V. Restoring Vaccine Diplomacy. JAMA 2021, 325, 2337. [Google Scholar] [CrossRef] [PubMed]
- Kavanagh, M.M.; Gostin, L.O.; Sunder, M. Sharing Technology and Vaccine Doses to Address Global Vaccine Inequity and End the COVID-19 Pandemic. JAMA 2021, 326, 219. [Google Scholar] [CrossRef]
- Machingaidze, S.; Wiysonge, C.S. Understanding COVID-19 vaccine hesitancy. Nat. Med. 2021, 27, 1338–1339. [Google Scholar] [CrossRef]
- Ball, P. Anti-vaccine movement could undermine efforts to end coronavirus pandemic, researchers warn. Nature 2020, 581, 251. [Google Scholar] [CrossRef]
- Arce, J.S.S.; Warren, S.S.; Meriggi, N.F.; Scacco, A.; McMurry, N.; Voors, M.; Syunyaev, G.; Malik, A.A.; Aboutajdine, S.; Adeojo, O.; et al. COVID-19 vaccine acceptance and hesitancy in low- and middle-income countries. Nat. Med. 2021, 27, 1385–1394. [Google Scholar] [CrossRef] [PubMed]
- Mathioudakis, A.G.; Ghrew, M.; Ustianowski, A.; Ahmad, S.; Borrow, R.; Papavasileiou, L.P.; Petrakis, D.; Bakerly, N.D. Self-Reported Real-World Safety and Reactogenicity of COVID-19 Vaccines: A Vaccine Recipient Survey. Life 2021, 11, 249. [Google Scholar] [CrossRef] [PubMed]
- Hacknen, D. Por que os Brasileiros Estão Preferindo Tomar a Vacina da Pfizer? Available online: https://jc.ne10.uol.com.br/brasil/2021/06/12133656-por-que-os-brasileiros-estao-preferindo-tomar-a-vacina-da-pfizer.html (accessed on 14 August 2021).
- Pardi, N.; Hogan, M.; Pelc, R.; Muramatsu, H.; Andersen, H.; DeMaso, C.R.; Dowd, K.A.; Sutherland, L.L.; Scearce, R.M.; Parks, R.; et al. Zika virus protection by a single low-dose nucleoside-modified mRNA vaccination. Nature 2017, 543, 248–251. [Google Scholar] [CrossRef] [PubMed]
| Sponsor | NCT-Number | Country | Vaccine Type | Targets | Trial Phase | Status | Reference |
|---|---|---|---|---|---|---|---|
| Argos Therapeutics | NCT00087984 | USA and Canada | DCs electroporated with autologous tumor mRNA | Metastatic renal cell carcinoma | 1/2 | Completed | NA |
| NCT00664482 | USA | Pancreatic cancer | Not Applicable | ||||
| University Hospital Tuebingen | NCT00204516 | Germany | Naked mRNA | Melanoma | 1/2 | Completed | NA |
| Radboud University | NCT00228189 | Netherlands | DCs electroporated with tumor-associated antigen mRNA | Colorectal cancer | 1/2 | Completed | [84] |
| Medigene AG | NCT02405338 | Norway | Autologous DCs with tumor-associated antigen mRNA | Acute myeloid leukemia | 1/2 | Completed | [85] |
| BioNTech RNA Pharmaceuticals | NCT02035956 | Austria and Germany | Naked RNA | Melanoma | 1 | Completed | [86] |
| Universitair Ziekenhuis Brussel | NCT01302496 | Belgium | DCs electroporated with tumor-associated antigen and TriMix (CD70, CD40L, TLR4) mRNA | Melanoma | 2 | Completed | [87] |
| Herlev Hospital | NCT00978913 | Denmark | DCs loaded with tumor-associated antigen mRNA | Breast cancer, melanoma | 1 | Completed | [88] |
| NCT01446731 | Prostate cancer | 2 | [89] | ||||
| Duke Cancer Institute | NCT01890213 | USA | Alphavirus replicon encoding the tumor-associated antigen | Colon cancer | 1 | Completed | [90] |
| NCT00003433 | Antigen-RNA-pulsed DCs | IV colon cancer, liver metastases | 1/2 | NA | |||
| NCT00003432 | Breast cancer | Terminated (low accrual) | |||||
| University Hospital Erlangen | NCT01983748 | Germany | DCs loaded with autologous tumor mRNA | Melanoma | 3 | Recruiting | NA |
| University Hospital, Antwerp | NCT02649582 | Belgium | DCs electroporated with tumor-associated antigen mRNA | Glioblastoma | 1/2 | Recruiting | NA |
| Changhai Hospital | NCT03468244 | China | Naked mRNA | Advanced esophageal squamous carcinoma, gastric, colorectal, and pancreatic adenocarcinomas | Not Applicable | Recruiting | NA |
| University of Campinas | NCT03083054 | Brazil | DCs electroporated with tumor-associated antigen mRNA | Acute myeloid leukemia | 1/2 | Active, not recruiting | NA |
| Guangdong 999 Brain Hospital | NCT02808416 | China | DCs pulsed with tumor-associated antigen mRNA | Brain metastases | 2 | Active, not recruiting | NA |
| NCT02709616 | Glioblastoma | 1 | |||||
| Life Research Technologies | NCT01456065 | Hungary and Austria | DCs loaded with tumor-associated antigen mRNA | Ovarian cancer | 1 | Unknown | NA |
| Sponsor | NCT-Number | Country | Vaccine Type | Targets | Trial Phase | Status | Reference |
|---|---|---|---|---|---|---|---|
| Argos Therapeutics | NCT00672191 | United States and Canada | DCs loaded with autologous viral antigen and CD40L mRNAs | HIV | 2 | Completed | [128] |
| Massachusetts General Hospital | NCT00833781 | United States | Autologous DCs loaded with viral antigen mRNA | HIV | 2 | Completed | [129] |
| CureVac AG | NCT02241135 | Germany | RNActive viral Ag mRNA | Rabies | 1 | Completed | [130] |
| Moderna Therapeutics | NCT03392389 | United States | Lipid-nanoparticle-encapsulated, chemically modified viral antigen mRNA | Human Metapneumovirus and Human Parainfluenza | 1 | Completed | [131] |
| NCT03014089 | Zika | NA | |||||
| NCT03325075 | Chikungunya | [132] | |||||
| NCT03382405 | Cytomegalovirus | NA | |||||
| NCT03076385 | Germany | Influenza | [133] | ||||
| Fundacion Clinic per a la Recerca Biomédica | NCT02413645 | Spain | Naked Viral antigen and TriMix (CD40, CD70 and IL2) mRNA | HIV | 1 | Completed | [134] |
| AlphaVax | NCT00440362 | United States | Alphavirus replicon vaccine expressing a viral hemagglutinin protein | Influenza | 1/2 | Completed | NA |
| NCT00439803 | Alphavirus replicon vaccine expressing viral genes | Cytomegalovirus | 1 | [135] | |||
| NCT00097838 | United States, Botswana, and South Africa | Alphavirus replicon vaccine expressing viral protein | HIV | [136] |
| Evaluated Vaccine | Doses | Country | Participants | COVID-19 Outcomes | Effectiveness (%) | Reference |
|---|---|---|---|---|---|---|
| BNT162b2 | 1 | England | 156,930 | Symptomatic infection | 61 | [187] |
| Hospitalization | 80 | |||||
| Deaths | 85 | |||||
| BNT162b2 | 1 | Scotland | 1,331,993 | Hospitalization | 91 | [188] |
| BNT162b2 and mRNA-1273 (Spikevax) | 2 | USA | 1212 | Hospitalization | 87.1 | [189] |
| BNT162b2 | 2 | Israel | 6286 | Symptomatic infection | 61 (first dose) to 89 (second dose) | [190] |
| BNT162b2 | 2 | Israel | 596,618 | Documented infection | 46 (first dose) and 92 (second dose) | [191] |
| Symptomatic infection | 57 (first dose) and 94 (second dose) | |||||
| Hospitalization | 74 (first dose) and 87 (second dose) | |||||
| Severe disease | 62 (first dose) and 92 (second dose) |
| Vaccine Name | mRNA or saRNA Construct | Antigen | Delivery Platform Composition | Diluent |
|---|---|---|---|---|
| mRNA-1273 (Spikevax) | N1-methylpseudouridine Modified 5′ CAP1 structure (m7GpppNmN) Codon optimization (GC-enriched sequence) | Full-length S protein with two proline substitutions (K986P and V987P) Wuhan-Hu-1 (GenBank: MN908947.3) | SM-102 Cholesterol DSPC PEG2000-DMG | Sodium chloride |
| BNT162b2 | N1-methylpseudouridine Modified 5′ CAP1 structure Codon optimization | Full-length S protein with two proline substitutions (K986P and V987P) Wuhan-Hu-1 (GenBank: MN908947) | ALC-0315 (proprietary to Acuitas) ALC-0159 DSPC Cholesterol | Sodium chloride |
| CVnCoV | Modified 5′ CAP1 structure (m7GpppNmN) Codon optimization (GC-enriched sequence | Full-length S protein with two proline substitutions (K986P and V987P) Wuhan-Hu-1 (GenBank: YP_009724390.1) | Cholesterol DSPC PEG-ylated lipid Ionizable lipid (undisclosed/proprietary to Acuitas) | Sodium chloride |
| ARCT-021 (LUNAR-COV19) | N1-methylpseudouridine Modified 5′ CAP1 structure (m7GpppNmN) Codon optimization Self-replicating replicon of VEEV | Full-length, unmodified S protein Wuhan-Hu-1 (GenBank: YP_009724390.1) | Ionizable lipid (undisclosed) DSPC Cholesterol PEG2000-DMG | Sodium chloride |
| COVAC1 (LNP-nCoVsaRNA) | Modified 5′ CAP1 structure (m7GpppNmN) Self-replicating replicon of VEEV F318V amino acid substitution | Full-length S protein with two proline substitutions (K986P and V987P) and GGGGSGGGGS linker Wuhan-Hu-1 (GenBank: QHD43416.1) | Ionizable cationic lipid (undisclosed) Phosphatidylcholine (undisclosed) Cholesterol PEG-lipid (undisclosed) | Phosphate-buffered saline |
| ARCoV | Modified 5′ CAP1 structure (m7GpppNmN) Codon optimization | RBD region of S protein Wuhan-Hu-1 (GenBank: MN908947.3) | Ionizable lipid (undisclosed/proprietary to Acuitas) DSPC Cholesterol PEG-lipid (undisclosed) | Not reported |
| HDT-301 (LION/repRNA-CoV2S) | Modified 5′ CAP1 structure Self-replicating replicon of VEEV | Full-length, unmodified S protein Wuhan-Hu-1 (GenBank: MN908947.3) | DOTAP Squalene Span 60 Polysorbate 80 oleic-acid-coated iron oxide nanoparticles | Sodium citrate buffer |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Machado, B.A.S.; Hodel, K.V.S.; Fonseca, L.M.d.S.; Mascarenhas, L.A.B.; Andrade, L.P.C.d.S.; Rocha, V.P.C.; Soares, M.B.P.; Berglund, P.; Duthie, M.S.; Reed, S.G.; et al. The Importance of RNA-Based Vaccines in the Fight against COVID-19: An Overview. Vaccines 2021, 9, 1345. https://doi.org/10.3390/vaccines9111345
Machado BAS, Hodel KVS, Fonseca LMdS, Mascarenhas LAB, Andrade LPCdS, Rocha VPC, Soares MBP, Berglund P, Duthie MS, Reed SG, et al. The Importance of RNA-Based Vaccines in the Fight against COVID-19: An Overview. Vaccines. 2021; 9(11):1345. https://doi.org/10.3390/vaccines9111345
Chicago/Turabian StyleMachado, Bruna Aparecida Souza, Katharine Valéria Saraiva Hodel, Larissa Moraes dos Santos Fonseca, Luís Alberto Brêda Mascarenhas, Leone Peter Correia da Silva Andrade, Vinícius Pinto Costa Rocha, Milena Botelho Pereira Soares, Peter Berglund, Malcolm S. Duthie, Steven G. Reed, and et al. 2021. "The Importance of RNA-Based Vaccines in the Fight against COVID-19: An Overview" Vaccines 9, no. 11: 1345. https://doi.org/10.3390/vaccines9111345
APA StyleMachado, B. A. S., Hodel, K. V. S., Fonseca, L. M. d. S., Mascarenhas, L. A. B., Andrade, L. P. C. d. S., Rocha, V. P. C., Soares, M. B. P., Berglund, P., Duthie, M. S., Reed, S. G., & Badaró, R. (2021). The Importance of RNA-Based Vaccines in the Fight against COVID-19: An Overview. Vaccines, 9(11), 1345. https://doi.org/10.3390/vaccines9111345