Immunogenetic Association Underlying Severe COVID-19
Abstract
1. The Broad Spectrum and Critical Illness in COVID-19 Progression
2. Genetic Association: Interferon and Chemokine Response Representing the Centric Immune Determinants Underlying Severe COVID-19
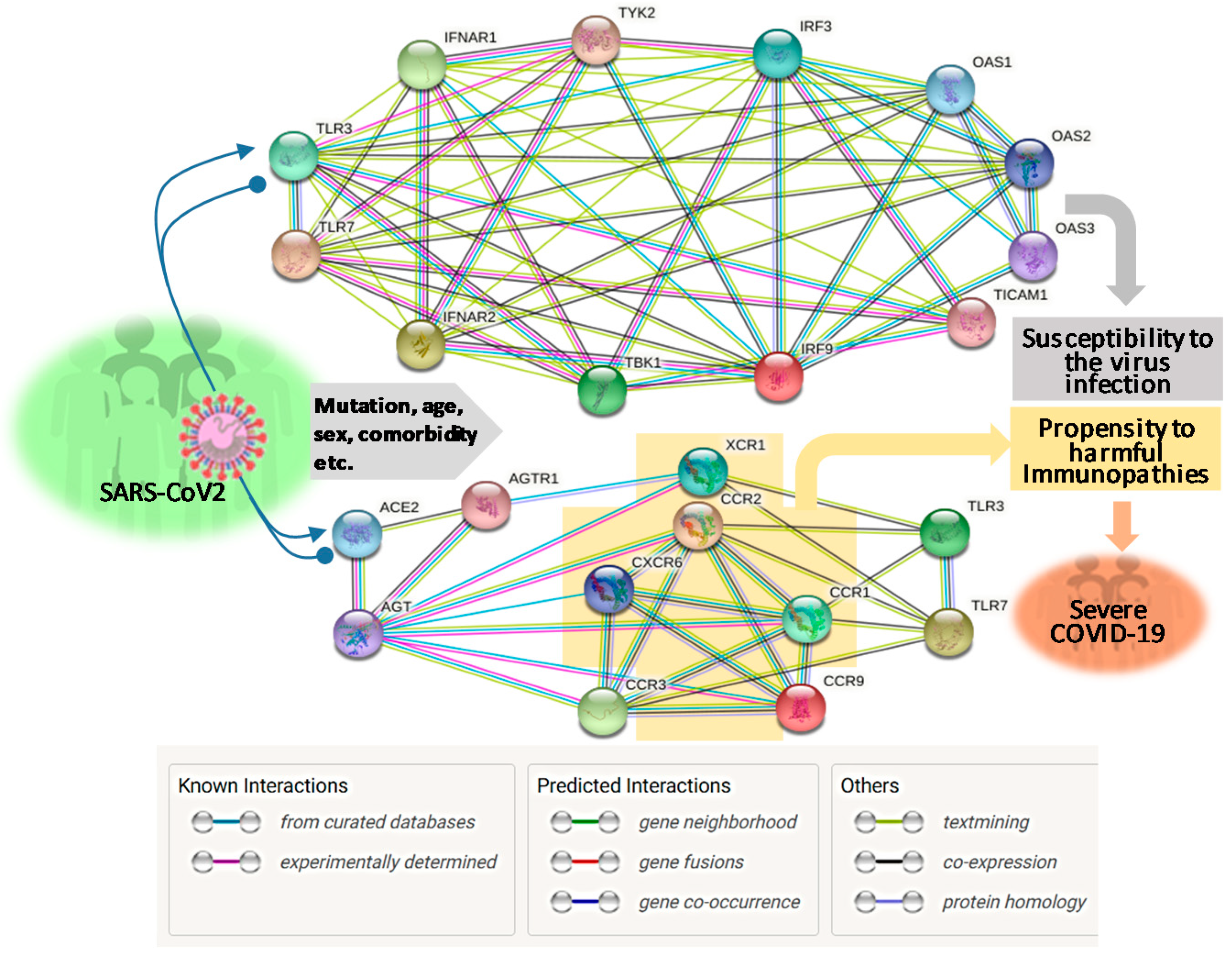
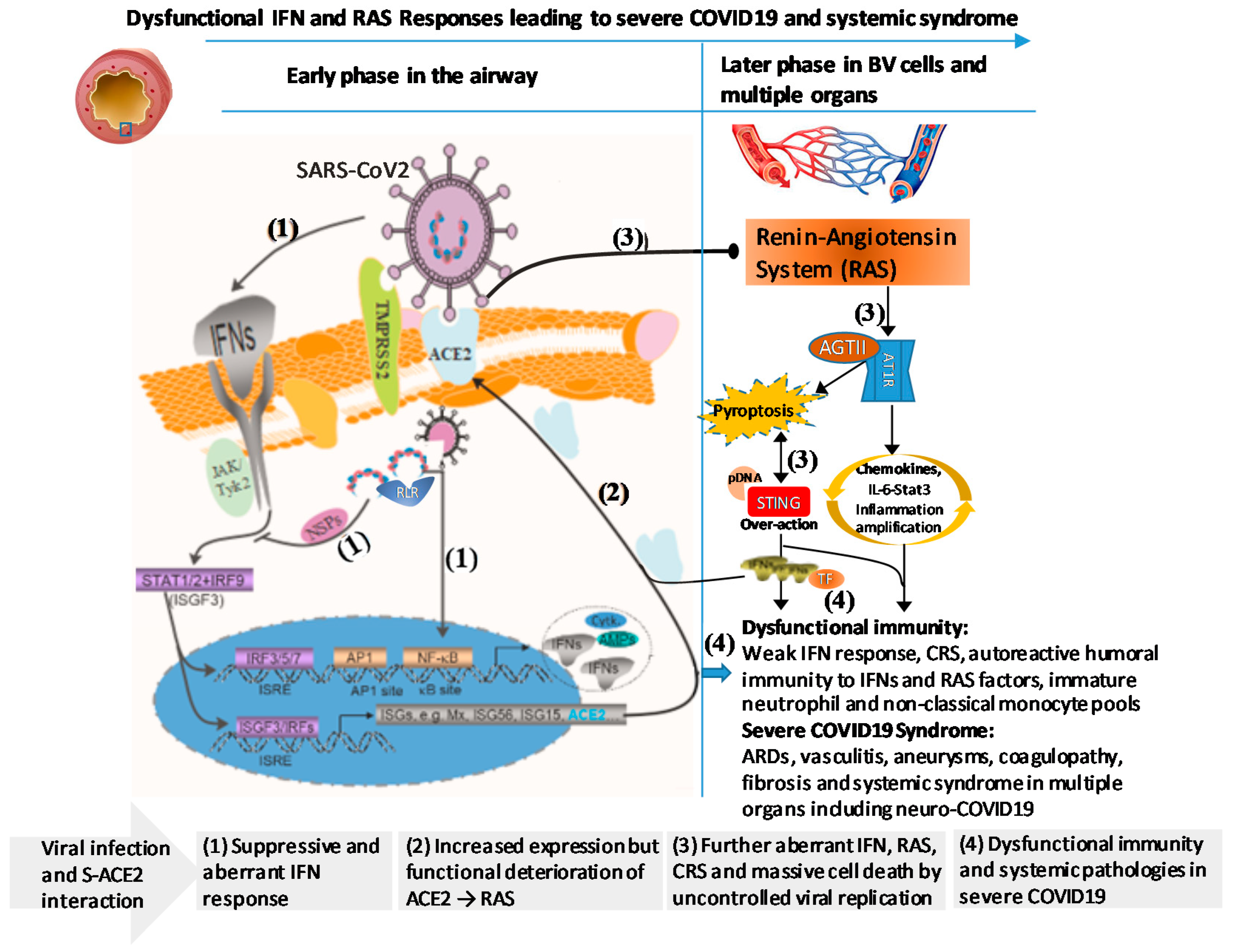
3. Epigenetic Association: Undermined Interferon and RAAS Responses Leading to Antiviral Dysfunction, Hyperinflammation, and Autoimmunity
| Chr. Location (Key Genes Covered, or Epigenetic Effect) | Association (Appr./OR: Freq.) | Major Immune Pathway Involved | References & Notes |
|---|---|---|---|
| 3p21.31 (SLC6A20, LZTFL1, FYCO1, CXCR6, XCR1, and CCR9; Neanderthal-originated allelic region) | GWAS 95% CI/ (1.95–2.79: 1610 vs. 2205) [22] (2.14: 2244 vs. ~5X 2244) [23] | ACE2 mediated amino acid transport (SLC6A20); Chemokine and Inflammation signaling, chemotaxis, immunopathies for lung injury (others) | Associated at [22,23,24] |
| 6p22.1–33 (HLA-G, CCHCR1, NOTCH4) | GWAS 95% CI/ (1.30–1.85: 2244 vs. ~5X 2244) | Antigen processing and presentation (HLA); P-body component for RNA metabolism, associated with psoriasis (CCHCR1); lymphocyte development (NOTCH4) | Associated by [23] |
| 9q34.2 (ABO blood type locus) | GWAS 95% CI/ (1.37–1.45: 1610 patient vs. 2205 control) | Blood type-dependent pathological reaction, such as coagulation and thrombolysis | Associated by [22] |
| 12q24.13 (OAS1, OAS2, OAS3) | GWAS 95% CI/ (1.29: 2244 vs. ~5X 2244) [23] | IFN-mediated antiviral signaling | Associated by [23] |
| 19p13.3 (DPP9, TYK2) | GWAS 95% CI/ (1.36–1.59: 2244 vs. ~5X 2244) [23] | Innate antiviral defense (TYK2), and antigen presentation, CXCL10 signaling, and associated to obesity, diabetes, and cancer (DPP9) | Associated by [23] |
| 21q22.1 (IFNAR2) | GWAS 95% CI/ (1.28: 2244 vs. ~5X 2244) [23] | IFN-mediated immune signaling | Associated by [23] |
| Several Chr. (TLR3, UNC93B1, TICAM1, TRAF3, TBK1, IRF3/7/9, IFNAR1/2, STAT1/2) | NGS and variant calling, wet-bench validation (3.5% of 659 severe COVID-19 vs few in 534 control) | IFN mediated immune signaling | Detected by [31] |
| Epigenetic obtaining (Autoantibody against IFNs, 94% in male) | Wet-bench detection (13.7% of 987 severe COVID-19 vs. 0.33% in 1227 control) | IFN mediated immune signaling | Detected by [39] |
| Epigenetic obtaining (Higher incidence of severe COVID-19 in aged, male, and comorbid patients) | Inclusive studies and evidence (Higher incidence of severe COVID-19 in aged, male, and comorbid patients) | Dysregulated IFN and chemokine responses, chronic/systemic inflammation, impaired other immune responses | Exemplified by [18,19,20,39,44,45] |
4. Conclusions Remarks: Precise Early-Risk Diagnosis and Drug Repurposing for Severe COVID-19 Based on Immunogenetic Association
Author Contributions
Funding
Conflicts of Interest
References
- COVID-19 Dashboard by the Center for Systems Science and Engineering (CSSE) at Johns Hopkins University (JHU). Available online: https://coronavirus.jhu.edu/map.html (accessed on 25 October 2020).
- Epidemiology Working Group for NCIP Epidemic Response, Chinese Center for Disease Control and Prevention. The Epidemiological Characteristics of an Outbreak of 2019 Novel Coronavirus Diseases (COVID-19). Zhonghua Liu Xing Bing Xue Za Zhi 2020, 41, 145–151. [Google Scholar]
- World Health Organization. Report of the WHO-China Joint Mission on Coronavirus Disease 2019 (COVID-19) [Pdf]. Available online: https://www.who.int/docs/default-source/coronaviruse/who-china-joint-mission-on-covid-19-final-report.pdf (accessed on 25 October 2020).
- COVID-19 Pandemic Planning Scenarios. Available online: https://www.cdc.gov/coronavirus/2019-ncov/hcp/planning-scenarios.html (accessed on 25 October 2020).
- Coronavirus disease 2019 (COVID-19). Available online: https://bestpractice.bmj.com/topics/en-gb/3000201 (accessed on 25 October 2020).
- Sanche, S.; Lin, Y.T.; Xu, C.; Romero-Severson, E.; Hengartner, N.; Ke, R. High Contagiousness and Rapid Spread of Severe Acute Respiratory Syndrome Coronavirus 2. Emerg. Infect. Dis. 2020, 26, 1470–1477. [Google Scholar] [CrossRef] [PubMed]
- Courtney, E.P.; Goldenberg, J.L.; Boyd, P. The contagion of mortality: A terror management health model for pandemics. Br. J. Soc. Psychol. 2020, 59, 607–617. [Google Scholar] [CrossRef] [PubMed]
- Age, Sex, Existing Conditions of COVID-19 Cases and Deaths. Available online: https://www.worldometers.info/coronavirus/coronavirus-age-sex-demographics/ (accessed on 25 October 2020).
- Scully, E.P.; Haverfield, J.; Ursin, R.L.; Tannenbaum, C.; Klein, S.L. Considering how biological sex impacts immune responses and COVID-19 outcomes. Nat. Rev. Immunol. 2020, 20, 442–447. [Google Scholar] [CrossRef]
- Gebhard, C.; Regitz-Zagrosek, V.; Neuhauser, H.K.; Morgan, R.; Klein, S.L. Impact of sex and gender on COVID-19 outcomes in Europe. Biol. Sex. Differ. 2020, 11, 29. [Google Scholar] [CrossRef]
- Jutzeler, C.R.; Bourguignon, L.; Weis, C.V.; Tong, B.; Wong, C.; Rieck, B.; Pargger, H.; Tschudin-Sutter, S.; Egli, A.; Borgwardt, K.; et al. Comorbidities, clinical signs and symptoms, laboratory findings, imaging features, treatment strategies, and outcomes in adult and pediatric patients with COVID-19: A systematic review and meta-analysis. Travel Med. Infect. Dis. 2020, 37, 101825. [Google Scholar] [CrossRef]
- Hadjadj, J.; Yatim, N.; Barnabei, L.; Corneau, A.; Boussier, J.; Pere, H.; Charbit, B.; Bondet, V.; Chenevier-Gobeaux, C.; Breillat, P.; et al. Impaired type I interferon activity and inflammatory responses in severe COVID-19 patients. Science 2020, 369, 718–724. [Google Scholar] [CrossRef]
- Blanco-Melo, D.; Nilsson-Payant, B.E.; Liu, W.C.; Uhl, S.; Hoagland, D.; Møller, R.; Jordan, T.X.; Oishi, K.; Panis, M.; Sachs, D.; et al. Imbalanced Host Response to SARS-CoV-2 Drives Development of COVID-19. Cell 2020, 181, 1036–1045. [Google Scholar] [CrossRef]
- Acharya, D.; Liu, G.; Gack, M.U. Dysregulation of type I interferon responses in COVID-19. Nat. Rev. Immunol. 2020, 20, 397–398. [Google Scholar] [CrossRef] [PubMed]
- Vabret, N.; Britton, G.J.; Gruber, C.; Hegde, S.; Kim, J.; Kuksin, M.; Levantovsky, R.; Malle, L.; Morieira, A.; Park, M.D.; et al. Immunology of COVID-19: Current State of the Science. Immunity 2020, 52, 910–941. [Google Scholar] [CrossRef] [PubMed]
- Rydyznski Moderbacher, C.; Ramirez, S.I.; Dan, J.M.; Grifoni, A.; Hastie, K.M.; Weiskopf, D.; Belanger, S.; Abbott, R.K.; Kim, C.; Choi, J.; et al. Antigen-Specific Adaptive Immunity to SARS-CoV-2 in Acute COVID-19 and Associations with Age and Disease Severity. Cell 2020, 183, 996–1012. [Google Scholar] [CrossRef] [PubMed]
- Schultheiß, C.; Paschold, L.; Simnica, D.; Mohme, M.; Willscher, E.; von Wenserski, L.; Scholz, R.; Wieters, I.; Dahlke, C.; Tolosa, E.; et al. Next-Generation Sequencing of T and B Cell Receptor Repertoires from COVID-19 Patients Showed Signatures Associated with Severity of Disease. Immunity 2020, 53, 442–455. [Google Scholar] [CrossRef] [PubMed]
- Mantovani, A.; Netea, M.G. Trained Innate Immunity, Epigenetics, and Covid-19. N. Engl. J. Med. 2020, 383, 1078–1080. [Google Scholar] [CrossRef] [PubMed]
- Kleen, T.O.; Galdon, A.A.; MacDonald, A.S.; Dalgleish, A.G. Mitigating Coronavirus Induced Dysfunctional Immunity for At-Risk Populations in COVID-19: Trained Immunity, BCG and “New Old Friends”. Front. Immunol. 2020, 11, 2059. [Google Scholar] [CrossRef] [PubMed]
- Sang, E.R.; Tian, Y.; Miller, L.C.; Sang, Y. Epigenetic Evolution of ACE2 and IL-6 Genes as Non-Canonical Interferon-Stimulated Genes Correlate to COVID-19 Susceptibility in Vertebrates. bioRxiv 2020. [Google Scholar] [CrossRef]
- COVID-19 Host Genetics Initiative. The COVID-19 Host Genetics Initiative, a global initiative to elucidate the role of host genetic factors in susceptibility and severity of the SARS-CoV-2 virus pandemic. Eur. J. Hum. Genet. 2020, 28, 715–718. [Google Scholar] [CrossRef]
- Severe Covid-19 GWAS Group; Ellinghaus, D.; Degenhardt, F.; Bujanda, L.; Buti, M.; Albillos, A.; Invernizzi, P.; Fernández, J.; Prati, D.; Baselli, G.; et al. Genomewide Association Study of Severe Covid-19 with Respiratory Failure. N. Engl. J. Med. 2020, 383, 1522–1534. [Google Scholar] [CrossRef]
- Pairo-Castineira, E.; Clohisey, S.; Klaric, L.; Bretherick, A.; Rawlik, K.; Parkinson, N.; Pasko, D.; Walker, S.; Richmond, A.; Fourman, M.H.; et al. Genetic mechanisms of critical illness in Covid-19. medRxiv 2020. [Google Scholar] [CrossRef]
- Zeberg, H.; Pääbo, S. The major genetic risk factor for severe COVID-19 is inherited from Neanderthals. Nature 2020. [Google Scholar] [CrossRef]
- Latz, C.A.; DeCarlo, C.; Boitano, L.; Png, C.Y.M.; Patell, R.; Conrad, M.F.; Eagleton, M.; Dua, A. Blood type and outcomes in patients with COVID-19. Ann. Hematol. 2020, 99, 2113–2118. [Google Scholar] [CrossRef]
- Gérard, C.; Maggipinto, G.; Minon, J.M. COVID-19 and ABO blood group: Another viewpoint. Br. J. Haematol. 2020, 190, e93–e94. [Google Scholar] [CrossRef]
- Wu, Y.; Feng, Z.; Li, P.; Yu, Q. Relationship between ABO blood group distribution and clinical characteristics in patients with COVID-19. Clin. Chim. Acta 2020, 509, 220–223. [Google Scholar] [CrossRef] [PubMed]
- Ehrenfeld, M.; Tincani, A.; Andreoli, L.; Cattalini, M.; Greenbaum, A.; Kanduc, D.; Alijotas-Reig, J.; Zinserling, V.; Semenova, N.; Amital, H.; et al. Covid-19 and autoimmunity. Autoimmun. Rev. 2020, 19, 102597. [Google Scholar] [CrossRef] [PubMed]
- Turner, M.D.; Nedjai, B.; Hurst, T.; Pennington, D.J. Cytokines and chemokines: At the crossroads of cell signalling and inflammatory disease. Biochim. Biophys. Acta 2014, 1843, 2563–2582. [Google Scholar] [CrossRef] [PubMed]
- Griffith, J.W.; Sokol, C.L.; Luster, A.D. Chemokines and chemokine receptors: Positioning cells for host defense and immunity. Annu. Rev. Immunol. 2014, 32, 659–702. [Google Scholar] [CrossRef]
- Zhang, Q.; Bastard, P.; Liu, Z.; Le Pen, J.; Moncada-Velez, M.; Chen, J.; Ogishi, M.; Sabil, I.K.; Hodeib, S.; Korol, C.; et al. Inborn errors of type I IFN immunity in patients with life-threatening COVID-19. Science 2020, 370, eabd4570. [Google Scholar] [CrossRef]
- Cooper, M.D.; Alder, M.N. The evolution of adaptive immune systems. Cell 2006, 124, 815–822. [Google Scholar] [CrossRef]
- Boehm, T.; Swann, J.B. Origin and evolution of adaptive immunity. Annu. Rev. Anim. Biosci. 2014, 2, 259–283. [Google Scholar] [CrossRef]
- Gourbal, B.; Pinaud, S.; Beckers, G.J.M.; Van Der Meer, J.W.M.; Conrath, U.; Netea, M.G. Innate immune memory: An evolutionary perspective. Immunol. Rev. 2018, 283, 21–40. [Google Scholar] [CrossRef]
- Icenogle, T. COVID-19: Infection or Autoimmunity. Front. Immunol. 2020, 11, 2055. [Google Scholar] [CrossRef]
- Viner, R.M.; Whittaker, E. Kawasaki-like disease: Emerging complication during the COVID-19 pandemic. Lancet 2020, 395, 1741–1743. [Google Scholar] [CrossRef]
- Toubiana, J.; Poirault, C.; Corsia, A.; Bajolle, F.; Fourgeaud, J.; Angoulvant, F.; Debray, A.; Basmaci, R.; Salvador, E.; Biscardi, S.; et al. Kawasaki-like multisystem inflammatory syndrome in children during the covid-19 pandemic in Paris, France: Prospective observational study. BMJ 2020, 369, m2094. [Google Scholar] [CrossRef] [PubMed]
- Verdoni, L.; Mazza, A.; Gervasoni, A.; Martelli, L.; Ruggeri, M.; Ciuffreda, M.; Bonanomi, E.; D’Antiga, L. An outbreak of severe Kawasaki-like disease at the Italian epicentre of the SARS-CoV-2 epidemic: An observational cohort study. Lancet 2020, 395, 1771–1778. [Google Scholar] [CrossRef]
- Bastard, P.; Rosen, L.B.; Zhang, Q.; Michailidis, E.; Hoffmann, H.H.; Zhang, Y.; Dorgham, K.; Philippot, Q.; Rosain, J.; Béziat, V.; et al. Auto-antibodies against type I IFNs in patients with life-threatening COVID-19. Science 2020, 370, eabd4585. [Google Scholar] [CrossRef] [PubMed]
- Channappanavar, R.; Fehr, A.R.; Vijay, R.; Mack, M.; Zhao, J.; Meyerholz, D.K.; Perlman, S. Dysregulated Type I Interferon and Inflammatory Monocyte-Macrophage Responses Cause Lethal Pneumonia in SARS-CoV-Infected Mice. Cell Host Microbe. 2016, 19, 181–193. [Google Scholar] [CrossRef]
- Newton, A.H.; Cardani, A.; Braciale, T.J. The host immune response in respiratory virus infection: Balancing virus clearance and immunopathology. Semin. Immunopathol. 2016, 38, 471–482. [Google Scholar] [CrossRef]
- McNab, F.; Mayer-Barber, K.; Sher, A.; Wack, A.; O’Garra, A. Type I interferons in infectious disease. Nat. Rev. Immunol. 2015, 15, 87–103. [Google Scholar] [CrossRef]
- Crow, M.K.; Olferiev, M.; Kirou, K.A. Type I Interferons in Autoimmune Disease. Annu. Rev. Pathol. 2019, 14, 369–393. [Google Scholar] [CrossRef]
- Chlamydas, S.; Papavassiliou, A.G.; Piperi, C. Epigenetic mechanisms regulating COVID-19 infection. Epigenetics 2020, 30, 1–8. [Google Scholar] [CrossRef]
- Sawalha, A.H.; Zhao, M.; Coit, P.; Lu, Q. Epigenetic dysregulation of ACE2 and interferon-regulated genes might suggest increased COVID-19 susceptibility and severity in lupus patients. Clin. Immunol. 2020, 215, 108410. [Google Scholar] [CrossRef]
- Netea, M.G.; Giamarellos-Bourboulis, E.J.; Domínguez-Andrés, J.; Curtis, N.; van Crevel, R.; van de Veerdonk, F.L.; Bonten, M. Trained Immunity: A Tool for Reducing Susceptibility to and the Severity of SARS-CoV-2 Infection. Cell 2020, 181, 969–977. [Google Scholar] [CrossRef] [PubMed]
- Pruimboom, L. Methylation Pathways and SARS-CoV-2 Lung Infiltration and Cell Membrane-Virus Fusion Are Both Subject to Epigenetics. Front. Cell Infect. Microbiol. 2020, 10, 290. [Google Scholar] [CrossRef] [PubMed]
- Agrawal, A. Dendritic Cell-Airway Epithelial Cell Cross-Talk Changes with Age and Contributes to Chronic Lung Inflammatory Diseases in the Elderly. Int. J. Mol. Sci. 2017, 18, 1206. [Google Scholar] [CrossRef] [PubMed]
- Prakash, S.; Agrawal, S.; Vahed, H.; Ngyuen, M.; BenMohamed, L.; Gupta, S.; Agrawal, A. Dendritic cells from aged subjects contribute to chronic airway inflammation by activating bronchial epithelial cells under steady state. Mucosal Immunol. 2014, 7, 1386–1394. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Kelley, W.J.; Goldstein, D.R. Role of Aging and the Immune Response to Respiratory Viral Infections: Potential Implications for COVID-19. J. Immunol. 2020, 205, 313–320. [Google Scholar] [CrossRef] [PubMed]
- Sanyaolu, A.; Okorie, C.; Marinkovic, A.; Patidar, R.; Younis, K.; Desai, P.; Hosein, Z.; Padda, I.; Mangat, J. Comorbidity and its Impact on Patients with COVID-19. SN Compr. Clin. Med. 2020, 2, 1069–1076. [Google Scholar] [CrossRef] [PubMed]
- Hamilton, J.A.; Hsu, H.C.; Mountz, J.D. Autoreactive B cells in SLE, villains or innocent bystanders? Immunol. Rev. 2019, 292, 120–138. [Google Scholar] [CrossRef]
- O’Shea, J.; Ma, A.; Lipsky, P. Cytokines and autoimmunity. Nat. Rev. Immunol. 2002, 2, 37–45. [Google Scholar] [CrossRef]
- McMillan, P.; Uhal, B.D. COVID-19-A theory of autoimmunity to ACE-2. MOJ Immunol. 2020, 7, 17–19. [Google Scholar]
- Berthelot, J.M.; Drouet, L.; Lioté, F. Kawasaki-like diseases and thrombotic coagulopathy in COVID-19: Delayed over-activation of the STING pathway? Emerg. Microbes Infect. 2020, 9, 1514–1522. [Google Scholar] [CrossRef]
- Abe, T.; Marutani, Y.; Shoji, I. Cytosolic DNA-sensing immune response and viral infection. Microbiol. Immunol. 2019, 63, 51–64. [Google Scholar] [CrossRef] [PubMed]
- Gustine, J.N.; Jones, D. Immunopathology of Hyperinflammation in COVID-19. Am. J. Pathol. 2020. [Google Scholar] [CrossRef] [PubMed]
- Zuo, Y.; Yalavarthi, S.; Shi, H.; Gockman, K.; Zuo, M.; Madison, J.A.; Blair, C.; Weber, A.; Barnes, B.J.; Egeblad, M.; et al. Neutrophil extracellular traps in COVID-19. JCI Insight. 2020, 5, e138999. [Google Scholar] [CrossRef] [PubMed]
- Webb, K.; Peckham, H.; Radziszewska, A.; Menon, M.; Oliveri, P.; Simpson, F.; Deakin, C.T.; Lee, S.; Ciurtin, C.; Butler, G.; et al. Sex and Pubertal Differences in the Type 1 Interferon Pathway Associate With Both X Chromosome Number and Serum Sex Hormone Concentration. Front. Immunol. 2019, 9, 3167. [Google Scholar] [CrossRef] [PubMed]
- Capone, I.; Marchetti, P.; Ascierto, P.A.; Malorni, W.; Gabriele, L. Sexual Dimorphism of Immune Responses: A New Perspective in Cancer Immunotherapy. Front. Immunol. 2018, 9, 552. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.S.; Shin, E. The type I interferon response in COVID-19: Implications for treatment. Nat. Rev. Immunol. 2020, 20, 585–586. [Google Scholar] [CrossRef] [PubMed]
- Rudemiller, N.P.; Crowley, S.D. Interactions Between the Immune and the Renin-Angiotensin Systems in Hypertension. Hypertension 2016, 68, 289–296. [Google Scholar] [CrossRef]
- Crowley, S.D.; Rudemiller, N.P. Immunologic Effects of the Renin-Angiotensin System. J. Am. Soc. Nephrol. 2017, 28, 1350–1361. [Google Scholar] [CrossRef]
- Santos, R.A.S.; Oudit, G.Y.; Verano-Braga, T.; Canta, G.; Steckelings, U.M.; Bader, M. The renin-angiotensin system: Going beyond the classical paradigms. Am. J. Physiol. Heart Circ. Physiol. 2019, 316, H958–H970. [Google Scholar] [CrossRef]
- Ziegler, C.G.K.; Allon, S.J.; Nyquist, S.K.; Mbano, I.M.; Miao, V.N.; Tzouanas, C.N.; Cao, Y.; Yousif, A.S.; Bals, J.; Hauser, B.M.; et al. SARS-CoV-2 Receptor ACE2 Is an Interferon-Stimulated Gene in Human Airway Epithelial Cells and Is Detected in Specific Cell Subsets across Tissues. Cell 2020, 181, 1016–1035. [Google Scholar] [CrossRef]
- Datta, P.K.; Liu, F.; Fischer, T.; Rappaport, J.; Qin, X. SARS-CoV-2 pandemic and research gaps: Understanding SARS-CoV-2 interaction with the ACE2 receptor and implications for therapy. Theranostics 2020, 10, 7448–7464. [Google Scholar] [CrossRef] [PubMed]
- de Abajo, F.J.; Rodríguez-Martín, S.; Lerma, V.; Mejía-Abril, G.; Aguilar, M.; García-Luque, A.; Laredo, L.; Laosa, O.; Centeno-Soto, G.A.; Gálvez, M.A.; et al. Use of renin-angiotensin-aldosterone system inhibitors and risk of COVID-19 requiring admission to hospital: A case-population study. Lancet 2020, 395, 1705–1714. [Google Scholar] [CrossRef]
- Viana, S.D.; Nunes, S.; Reis, F. ACE2 imbalance as a key player for the poor outcomes in COVID-19 patients with age-related comorbidities—Role of gut microbiota dysbiosis. Ageing Res. Rev. 2020, 62, 101123. [Google Scholar] [CrossRef] [PubMed]
- Gheblawi, M.; Wang, K.; Viveiros, A.; Nguyen, Q.; Zhong, J.C.; Turner, A.J.; Raizada, M.K.; Grant, M.B.; Oudit, G.Y. Angiotensin-Converting Enzyme 2: SARS-CoV-2 Receptor and Regulator of the Renin-Angiotensin System: Celebrating the 20th Anniversary of the Discovery of ACE2. Circ. Res. 2020, 126, 1456–1474. [Google Scholar] [CrossRef] [PubMed]
- Curran, C.S.; Rivera, D.R.; Kopp, J.B. COVID-19 Usurps Host Regulatory Networks. Front. Pharmacol. 2020, 11, 1278. [Google Scholar] [CrossRef]
- Chen, L.Y.C.; Hoiland, R.L.; Stukas, S.; Wellington, C.L.; Sekhon, M.S. Confronting the controversy: Interleukin-6 and the COVID-19 cytokine storm syndrome. Eur. Respir. J. 2020, 56, 2003006. [Google Scholar] [CrossRef]
- Silvin, A.; Chapuis, N.; Dunsmore, G.; Goubet, A.G.; Dubuisson, A.; Derosa, L.; Almire, C.; Hénon, C.; Kosmider, O.; Droin, N.; et al. Elevated Calprotectin and Abnormal Myeloid Cell Subsets Discriminate Severe from Mild COVID-19. Cell 2020, 182, 1401–1418. [Google Scholar] [CrossRef]
- Herold, T.; Jurinovic, V.; Arnreich, C.; Lipworth, B.J.; Hellmuth, J.C.; von Bergwelt-Baildon, M.; Klein, M.; Weinberger, T. Elevated levels of IL-6 and CRP predict the need for mechanical ventilation in COVID-19. J. Allergy Clin. Immunol 2020, 146, 128–136. [Google Scholar] [CrossRef]
- Laguna-Goya, R.; Utrero-Rico, A.; Talayero, P.; Lasa-Lazaro, M.; Ramirez-Fernandez, A.; Naranjo, L.; Segura-Tudela, A.; Cabrera-Marante, O.; de Frias, E.R.; Garcia-Garcia, R.; et al. IL-6-based mortality risk model for hospitalized patients with COVID-19. J. Allergy Clin. Immunol. 2020, 146, 799–807. [Google Scholar] [CrossRef]
- Price, C.C.; Altice, F.L.; Shyr, Y.; Koff, A.; Pischel, L.; Goshua, G.; Azar, M.M.; Mcmanus, D.; Chen, S.C.; Gleeson, S.E.; et al. Tocilizumab Treatment for Cytokine Release Syndrome in Hospitalized Patients With Coronavirus Disease 2019: Survival and Clinical Outcomes. Chest 2020, 158, 1397–1408. [Google Scholar] [CrossRef]
- El Baba, R.; Herbein, G. Management of epigenomic networks entailed in coronavirus infections and COVID-19. Clin. Epigenet. 2020, 12, 118. [Google Scholar] [CrossRef] [PubMed]
- Park, A.; Iwasaki, A. Type I and Type III Interferons—Induction, Signaling, Evasion, and Application to Combat COVID-19. Cell Host Microbe. 2020, 27, 870–878. [Google Scholar] [CrossRef] [PubMed]
- Ingraham, N.E.; Barakat, A.G.; Reilkoff, R.; Bezdicek, T.; Schacker, T.; Chipman, J.G.; Tignanelli, C.J.; Puskarich, M.A. Understanding the renin-angiotensin-aldosterone-SARS-CoV axis: A comprehensive review. Eur. Respir. J. 2020, 56, 2000912. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.; Wang, F.; Tang, J.; Nussinov, R.; Cheng, F. Artificial intelligence in COVID-19 drug repurposing. Lancet Digit. Health. 2020. [Google Scholar] [CrossRef]
- El Bairi, K.; Trapani, D.; Petrillo, A.; Le Page, C.; Zbakh, H.; Daniele, B.; Belbaraka, R.; Curigliano, G.; Afqir, S. Repurposing anticancer drugs for the management of COVID-19. Eur. J. Cancer. 2020. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.; Hou, Y.; Shen, J.; Huang, Y.; Martin, W.; Cheng, F. Network-based drug repurposing for novel coronavirus 2019-nCoV/SARS-CoV-2. Cell Discov. 2020, 6, 14. [Google Scholar] [CrossRef]
- Zoufaly, A.; Poglitsch, M.; Aberle, J.H.; Hoepler, W.; Seitz, T.; Traugott, M.; Grieb, A.; Pawelka, E.; Laferl, H.; Wenisch, C.; et al. Human recombinant soluble ACE2 in severe COVID-19. Lancet Respir. Med. 2020. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
McCoy, K.; Peterson, A.; Tian, Y.; Sang, Y. Immunogenetic Association Underlying Severe COVID-19. Vaccines 2020, 8, 700. https://doi.org/10.3390/vaccines8040700
McCoy K, Peterson A, Tian Y, Sang Y. Immunogenetic Association Underlying Severe COVID-19. Vaccines. 2020; 8(4):700. https://doi.org/10.3390/vaccines8040700
Chicago/Turabian StyleMcCoy, Kendall, Autumn Peterson, Yun Tian, and Yongming Sang. 2020. "Immunogenetic Association Underlying Severe COVID-19" Vaccines 8, no. 4: 700. https://doi.org/10.3390/vaccines8040700
APA StyleMcCoy, K., Peterson, A., Tian, Y., & Sang, Y. (2020). Immunogenetic Association Underlying Severe COVID-19. Vaccines, 8(4), 700. https://doi.org/10.3390/vaccines8040700






