Mass Spectrometric Characterization of Narcolepsy-Associated Pandemic 2009 Influenza Vaccines
Abstract
1. Introduction
2. Methods
2.1. Vaccines
2.2. Mass Spectrometry (MS)
2.3. Statistical Analysis
2.4. DQ0602 Binding
2.5. Tetramer Analysis
3. Results
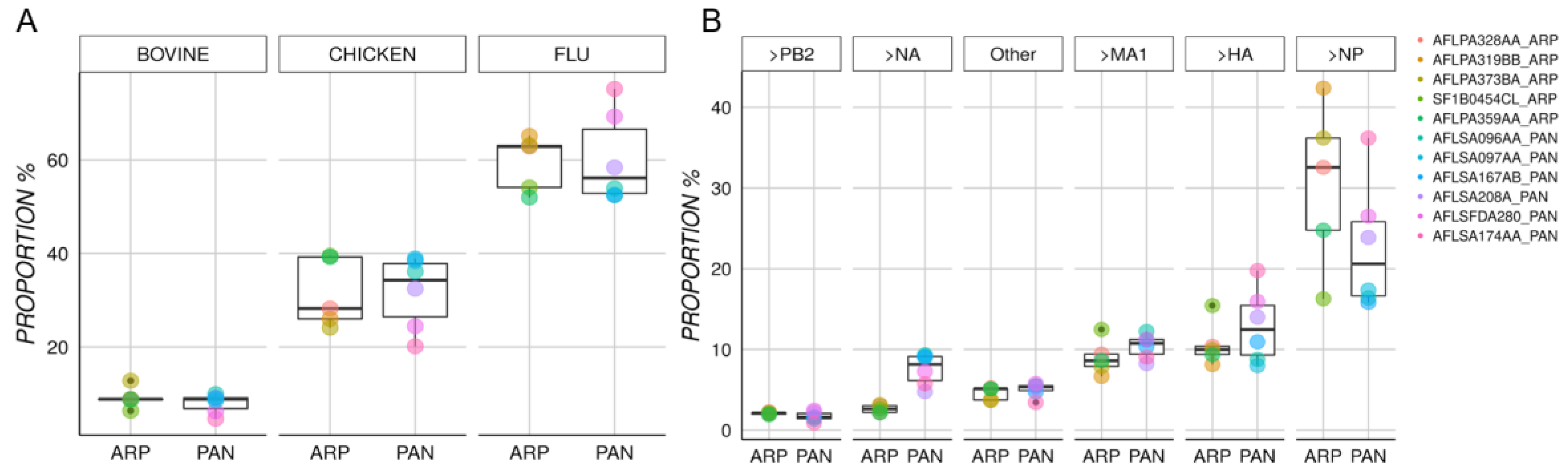
3.1. Characterization of Protein Content in Arepanrix and Pandemrix
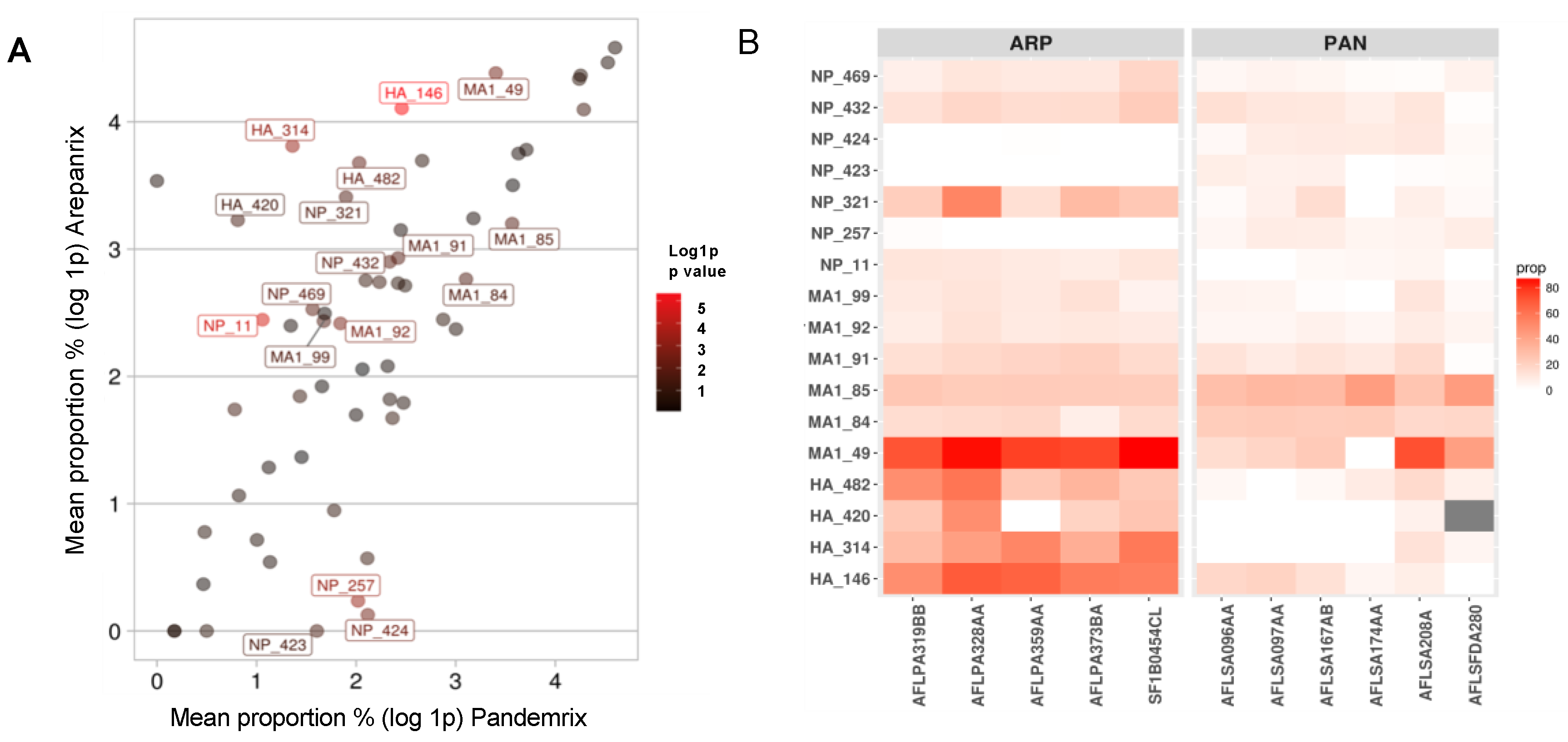
3.2. Differential Mutation Proportions among Arepanrix and Pandemrix
3.3. DQ0602 Binding of Mutated Motifs in Arepanrix and Pandemrix
3.4. Tetramer Studies of Four Mutated Motifs that Could Have Impacted Narcolepsy Risk
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Mahoney, C.E.; Cogswell, A.; Koralnik, I.J.; Scammell, T.E. The neurobiological basis of narcolepsy. Nat. Rev. Neurosci. 2019, 20, 83–93. [Google Scholar] [CrossRef] [PubMed]
- Thannickal, T.C.; Moore, R.Y.; Nienhuis, R.; Ramanathan, L.; Gulyani, S.; Aldrich, M.; Cornford, M.; Siegel, J.M. Reduced number of hypocretin neurons in human narcolepsy. Neuron 2000, 27, 469–474. [Google Scholar] [CrossRef]
- Peyron, C.; Faraco, J.; Rogers, W.; Ripley, B.; Overeem, S.; Charnay, Y.; Nevsimalova, S.; Aldrich, M.; Reynolds, D.; Albin, R.; et al. A mutation in a case of early onset narcolepsy and a generalized absence of hypocretin peptides in human narcoleptic brains. Nat. Med. 2000, 6, 991–997. [Google Scholar] [CrossRef] [PubMed]
- Hagan, J.J.; Leslie, R.A.; Patel, S.; Evans, M.L.; Wattam, T.A.; Holmes, S.; Benham, C.D.; Taylor, S.G.; Routledge, C.; Hemmati, P.; et al. Orexin A activates locus coeruleus cell firing and increases arousal in the rat. Proc. Natl. Acad. Sci. USA 1999, 96, 10911–10916. [Google Scholar] [CrossRef]
- Piper, D.C.; Upton, N.; Smith, M.I.; Hunter, A.J. The novel brain neuropeptide, orexin-A, modulates the sleep–wake cycle of rats. Eur. J. Neurosci. 2000, 12, 726–730. [Google Scholar] [CrossRef] [PubMed]
- Mignot, E.; Hayduk, R.; Black, J.; Grumet, F.C.; Guilleminault, C. HLA DQB1*0602 is associated with cataplexy in 509 narcoleptic patients. Sleep 1997, 20, 1012–1020. [Google Scholar] [PubMed]
- Mignot, E.; Lin, L.; Rogers, W.; Honda, Y.; Qiu, X.; Lin, X.; Okun, M.; Hohjoh, H.; Miki, T.; Hsu, S.; et al. Complex HLA-DR and -DQ interactions confer risk of narcolepsy-cataplexy in three ethnic groups. Am. J. Hum. Genet. 2001, 68, 686–699. [Google Scholar] [CrossRef]
- Chen, Y.H.; Huang, Y.S.; Chien, W.H.; Chen, C.H. Association analysis of the major histocompatibility complex, class II, DQ beta1 gene, HLA-DQB1, with narcolepsy in Han Chinese patients from Taiwan. Sleep Med. 2013, 14, 1393–1397. [Google Scholar] [CrossRef]
- Ollila, H.M.; Ravel, J.M.; Han, F.; Faraco, J.; Lin, L.; Zheng, X.; Plazzi, G.; Dauvilliers, Y.; Pizza, F.; Hong, S.C.; et al. HLA-DPB1 and HLA class I confer risk of and protection from narcolepsy. Am. J. Hum. Genet. 2015, 96, 136–146. [Google Scholar] [CrossRef]
- Han, F.; Faraco, J.; Dong, X.S.; Ollila, H.M.; Lin, L.; Li, J.; An, P.; Wang, S.; Jiang, K.W.; Gao, Z.C.; et al. Genome wide analysis of narcolepsy in China implicates novel immune loci and reveals changes in association prior to versus after the 2009 H1N1 influenza pandemic. PLoS Genet. 2013, 9, e1003880. [Google Scholar] [CrossRef]
- Hallmayer, J.; Faraco, J.; Lin, L.; Hesselson, S.; Winkelmann, J.; Kawashima, M.; Mayer, G.; Plazzi, G.; Nevsimalova, S.; Bourgin, P.; et al. Narcolepsy is strongly associated with the T-cell receptor alpha locus. Nat. Genet. 2009, 41, 708–711. [Google Scholar] [CrossRef]
- Faraco, J.; Lin, L.; Kornum, B.R.; Kenny, E.E.; Trynka, G.; Einen, M.; Rico, T.J.; Lichtner, P.; Dauvilliers, Y.; Arnulf, I.; et al. ImmunoChip study implicates antigen presentation to T cells in narcolepsy. PLoS Genet. 2013, 9, e1003270. [Google Scholar] [CrossRef] [PubMed]
- Aran, A.; Lin, L.; Nevsimalova, S.; Plazzi, G.; Hong, S.C.; Weiner, K.; Zeitzer, J.; Mignot, E. Elevated anti-streptococcal antibodies in patients with recent narcolepsy onset. Sleep 2009, 32, 979–983. [Google Scholar] [CrossRef] [PubMed]
- Ambati, A.; Poiret, T.; Svahn, B.M.; Valentini, D.; Khademi, M.; Kockum, I.; Lima, I.; Arnheim-Dahlstrom, L.; Lamb, F.; Fink, K.; et al. Increased beta-haemolytic group A streptococcal M6 serotype and streptodornase B-specific cellular immune responses in Swedish narcolepsy cases. J. Intern. Med. 2015, 278, 264–276. [Google Scholar] [CrossRef] [PubMed]
- Ding, Q.; Li, J.; Xiao, F.; Zhang, C.; Dong, X.; Han, F. Anti-streptococcal antibodies in Chinese patients with type -1 narcolepsy. Sleep Med. 2020, 72, 37–40. [Google Scholar] [CrossRef]
- Longstreth, W.T., Jr.; Ton, T.G.; Koepsell, T.; Gersuk, V.H.; Hendrickson, A.; Velde, S. Prevalence of narcolepsy in King County, Washington, USA. Sleep Med. 2009, 10, 422–426. [Google Scholar] [CrossRef]
- Sarkanen, T.; Alakuijala, A.; Julkunen, I.; Partinen, M. Narcolepsy Associated with Pandemrix Vaccine. Curr. Neurol. Neurosci. Rep. 2018, 18, 43. [Google Scholar] [CrossRef]
- Dauvilliers, Y.; Arnulf, I.; Lecendreux, M.; Monaca Charley, C.; Franco, P.; Drouot, X.; d’Ortho, M.P.; Launois, S.; Lignot, S.; Bourgin, P.; et al. Increased risk of narcolepsy in children and adults after pandemic H1N1 vaccination in France. Brain 2013, 136, 2486–2496. [Google Scholar] [CrossRef]
- Nohynek, H.; Jokinen, J.; Partinen, M.; Vaarala, O.; Kirjavainen, T.; Sundman, J.; Himanen, S.L.; Hublin, C.; Julkunen, I.; Olsen, P.; et al. AS03 adjuvanted AH1N1 vaccine associated with an abrupt increase in the incidence of childhood narcolepsy in Finland. PLoS ONE 2012, 7, e33536. [Google Scholar] [CrossRef]
- Partinen, M.; Saarenpää-Heikkilä, O.; Ilveskoski, I.; Hublin, C.; Linna, M.; Olsén, P.; Nokelainen, P.; Alén, R.; Wallden, T.; Espo, M.; et al. Increased incidence and clinical picture of childhood narcolepsy following the 2009 H1N1 pandemic vaccination campaign in Finland. PLoS ONE 2012, 7, e33723. [Google Scholar] [CrossRef]
- Miller, E.; Andrews, N.; Stellitano, L.; Stowe, J.; Winstone, A.M.; Shneerson, J.; Verity, C. Risk of narcolepsy in children and young people receiving AS03 adjuvanted pandemic A/H1N1 2009 influenza vaccine: Retrospective analysis. BMJ Br. Med. J. 2013, 346, f794. [Google Scholar] [CrossRef] [PubMed]
- Heier, M.S.; Gautvik, K.M.; Wannag, E.; Bronder, K.H.; Midtlyng, E.; Kamaleri, Y.; Storsaeter, J. Incidence of narcolepsy in Norwegian children and adolescents after vaccination against H1N1 influenza A. Sleep Med. 2013, 14, 867–871. [Google Scholar] [CrossRef] [PubMed]
- Persson, I.; Granath, F.; Askling, J.; Ludvigsson, J.F.; Olsson, T.; Feltelius, N. Risks of neurological and immune-related diseases, including narcolepsy, after vaccination with Pandemrix: A population- and registry-based cohort study with over 2 years of follow-up. J. Intern. Med. 2014, 275, 172–190. [Google Scholar] [CrossRef]
- O’Flanagan, D.; Barret, A.S.; Foley, M.; Cotter, S.; Bonner, C.; Crowe, C.; Lynch, B.; Sweeney, B.; Johnson, H.; McCoy, B.; et al. Investigation of an association between onset of narcolepsy and vaccination with pandemic influenza vaccine, Ireland April 2009-December 2010. Eurosurveillance 2014, 19, 20789. [Google Scholar] [CrossRef] [PubMed]
- Montplaisir, J.; Petit, D.; Quinn, M.-J.; Ouakki, M.; Deceuninck, G.; Desautels, A.; Mignot, E.; De Wals, P. Risk of narcolepsy associated with inactivated adjuvanted (AS03) A/H1N1 (2009) pandemic influenza vaccine in Quebec. PLoS ONE 2014, 9, e108489. [Google Scholar] [CrossRef] [PubMed]
- Robertson, J.S.; Nicolson, C.; Harvey, R.; Johnson, R.; Major, D.; Guilfoyle, K.; Roseby, S.; Newman, R.; Collin, R.; Wallis, C.; et al. The development of vaccine viruses against pandemic A(H1N1) influenza. Vaccine 2011, 29, 1836–1843. [Google Scholar] [CrossRef]
- Roman, F.; Vaman, T.; Kafeja, F.; Hanon, E.; Van Damme, P. AS03(A)-Adjuvanted influenza A (H1N1) 2009 vaccine for adults up to 85 years of age. Clin. Infect. Dis. Off. Publ. Infect. Dis. Soc. Am. 2010, 51, 668–677. [Google Scholar] [CrossRef]
- Leroux-Roels, I.; Borkowski, A.; Vanwolleghem, T.; Drame, M.; Clement, F.; Hons, E.; Devaster, J.M.; Leroux-Roels, G. Antigen sparing and cross-reactive immunity with an adjuvanted rH5N1 prototype pandemic influenza vaccine: A randomised controlled trial. Lancet 2007, 370, 580–589. [Google Scholar] [CrossRef]
- Barker, C.I.; Snape, M.D. Pandemic influenza A H1N1 vaccines and narcolepsy: Vaccine safety surveillance in action. Lancet. Infect. Dis. 2014, 14, 227–238. [Google Scholar] [CrossRef]
- Jacob, L.; Leib, R.; Ollila, H.M.; Bonvalet, M.; Adams, C.M.; Mignot, E. Comparison of Pandemrix and Arepanrix, two pH1N1 AS03-adjuvanted vaccines differentially associated with narcolepsy development. Brain Behav. Immun. 2015, 47, 44–57. [Google Scholar] [CrossRef]
- Latorre, D.; Kallweit, U.; Armentani, E.; Foglierini, M.; Mele, F.; Cassotta, A.; Jovic, S.; Jarrossay, D.; Mathis, J.; Zellini, F.; et al. T cells in patients with narcolepsy target self-antigens of hypocretin neurons. Nature 2018, 562, 63–68. [Google Scholar] [CrossRef] [PubMed]
- Luo, G.; Ambati, A.; Lin, L.; Bonvalet, M.; Partinen, M.; Ji, X.; Maecker, H.T.; Mignot, E.J.-M. Autoimmunity to hypocretin and molecular mimicry to flu in type 1 narcolepsy. Proc. Natl. Acad. Sci. USA 2018, 115, E12323–E12332. [Google Scholar] [CrossRef]
- Vaarala, O.; Vuorela, A.; Partinen, M.; Baumann, M.; Freitag, T.L.; Meri, S.; Saavalainen, P.; Jauhiainen, M.; Soliymani, R.; Kirjavainen, T.; et al. Antigenic Differences between AS03 Adjuvanted Influenza A (H1N1) Pandemic Vaccines: Implications for Pandemrix-Associated Narcolepsy Risk. PLoS ONE 2014, 9, e114361. [Google Scholar] [CrossRef] [PubMed]
- Ahmed, S.S.; Volkmuth, W.; Duca, J.; Corti, L.; Pallaoro, M.; Pezzicoli, A.; Karle, A.; Rigat, F.; Rappuoli, R.; Narasimhan, V.; et al. Antibodies to influenza nucleoprotein cross-react with human hypocretin receptor 2. Sci. Transl. Med. 2015, 7, 294ra105. [Google Scholar] [CrossRef]
- Luo, G.; Lin, L.; Jacob, L.; Bonvalet, M.; Ambati, A.; Plazzi, G.; Pizza, F.; Leib, R.; Adams, C.M.; Partinen, M.; et al. Absence of anti-hypocretin receptor 2 autoantibodies in post pandemrix narcolepsy cases. PLoS ONE 2017, 12, e0187305. [Google Scholar] [CrossRef]
- Andreatta, M.; Karosiene, E.; Rasmussen, M.; Stryhn, A.; Buus, S.; Nielsen, M. Accurate pan-specific prediction of peptide-MHC class II binding affinity with improved binding core identification. Immunogenetics 2015, 67, 641–650. [Google Scholar] [CrossRef]
- Skowronski, D.M.; Janjua, N.Z.; De Serres, G.; Sabaiduc, S.; Eshaghi, A.; Dickinson, J.A.; Fonseca, K.; Winter, A.-L.; Gubbay, J.B.; Krajden, M.; et al. Low 2012–13 Influenza Vaccine Effectiveness Associated with Mutation in the Egg-Adapted H3N2 Vaccine Strain Not Antigenic Drift in Circulating Viruses. PLoS ONE 2014, 9, e92153. [Google Scholar] [CrossRef] [PubMed]
- Rudd, P.M.; Elliott, T.; Cresswell, P.; Wilson, I.A.; Dwek, R.A. Glycosylation and the immune system. Science 2001, 291, 2370–2376. [Google Scholar] [CrossRef] [PubMed]
- Bonvalet, M.; Ollila, H.M.; Ambati, A.; Mignot, E. Autoimmunity in narcolepsy. Curr. Opin. Pulm. Med. 2017, 23, 522–529. [Google Scholar] [CrossRef] [PubMed]
- De la Herran-Arita, A.K.; Kornum, B.R.; Mahlios, J.; Jiang, W.; Lin, L.; Hou, T.; Macaubas, C.; Einen, M.; Plazzi, G.; Crowe, C.; et al. CD4+ T cell autoimmunity to hypocretin/orexin and cross-reactivity to a 2009 H1N1 influenza A epitope in narcolepsy. Sci. Transl. Med. 2013, 5, 216ra176. [Google Scholar] [CrossRef]
- De la Herran-Arita, A.K.; Kornum, B.R.; Mahlios, J.; Jiang, W.; Lin, L.; Hou, T.; Macaubas, C.; Einen, M.; Plazzi, G.; Crowe, C.; et al. Retraction of the research article: “CD4(+) T cell autoimmunity to hypocretin/orexin and cross-reactivity to a 2009 H1N1 influenza A epitope in narcolepsy”. Sci. Transl. Med. 2014, 6, 247rt1. [Google Scholar] [CrossRef] [PubMed]
| Vaccine | Batch | HA (ug/mL) | Type | Origin | Viral Strain |
|---|---|---|---|---|---|
| Arepanrix | AFLPA328AA | 15 | Vaccine | Canada | X179A |
| Arepanrix | AFLPA359AA | 15 | Vaccine | Canada | X179A |
| Arepanrix | AFLPA373BA | 15 | Vaccine | Canada | X179A |
| Arepanrix | AFLPA319BB | 15 | Vaccine | Canada | X179A |
| Arepanrix | SF1B0454CL | 457 | Bulk | GSK | X179A |
| Pandemrix | AFLSA208A | 15 | Vaccine | GSK | X179A |
| Pandemrix | AFLSA174AA | 15 | Vaccine | France | X179A |
| Pandemrix | AFLSFDA280 | 139 | Bulk | GSK | X179A |
| Pandemrix | AFLSA167AB | 15 | Vaccine | Sweden | X179A |
| Pandemrix | AFLSA097AA | 15 | Vaccine | Sweden | X179A |
| Pandemrix | AFLSA096AA | 15 | Vaccine | Sweden | X179A |
| Protein | Organism | Pandemrix | Arepanrix | CI95 U ± L | Statistic | p-Value |
|---|---|---|---|---|---|---|
| Nucleoprotein NP | PR8 | 22.69% (1600) | 30.44% (1092) | 20.8 ± −5.3 | 1.389 | 0.2044 |
| Hemagglutinin HA | pH1N1 | 12.9% (894) | 10.66% (420) | 2.9 ± −7.3 | −1.003 | 0.3436 |
| Matrix Protein 1 MA1 | PR8 | 10.39% (724) | 9.01% (357) | 1.4 ± −4.1 | −1.197 | 0.2709 |
| Neuraminidase NA | pH1N1 | 7.56% (533) | 2.64% (99) | −2.9 ± −7 | −5.997 | 0.0012 |
| Polymerase PB2 | PR8 | 1.7% (109) | 2.09% (80) | 1 ± −0.2 | 1.655 | 0.1528 |
| Polymerase PA | pH1N1 | 1.81% (124) | 1.34% (56) | 0.5 ± −1.5 | −1.12 | 0.3003 |
| Polymerase PB1 | pH1N1 | 1.4% (91) | 1.33% (52) | 0.6 ± −0.7 | −0.266 | 0.7985 |
| Nonstructural protein 1 | PR8 | 0.52% (34) | 0.45% (18) | 0.1 ± −0.3 | −0.821 | 0.4404 |
| Nuclear export protein | PR8 | 0.11% (10) | 0.26% (11) | 0.4 ± −0.1 | 1.605 | 0.1496 |
| Apolipoprotein B | Gallus gallus | 0.16% (11) | 2.07% (87) | 3.2 ± 0.6 | 3.965 | 0.016 |
| Vitellogenin−2 | Gallus gallus | 0.12% (8) | 1.15% (47) | 1.5 ± 0.6 | 5.772 | 0.0024 |
| Glucose−6−phosphate isomerase | Gallus gallus | 0.49% (35) | 0.87% (31) | 0.7 ± 0 | 2.684 | 0.0442 |
| Ovalbumin | Gallus gallus | 0.39% (28) | 0.85% (33) | 0.8 ± 0.1 | 2.872 | 0.02 |
| Annexin A2 | Gallus gallus | 0.55% (37) | 0.73% (28) | 0.4 ± 0 | 2.382 | 0.0434 |
| Ezrin | Gallus gallus | 0.58% (40) | 0.73% (28) | 0.3 ± 0 | 2.754 | 0.0246 |
| Junction plakoglobin | Bos taurus | 0.15% (9) | 0.69% (23) | 1.1 ± 0 | 2.586 | 0.0496 |
| Tubulin alpha−1B chain | Bos taurus | 0.88% (60) | 0.54% (21) | −0.1 ± −0.6 | −3.212 | 0.0147 |
| Protein Pos (Intial > Mut) | p-Value | T−Stat (CI U ± L) | Mean Proportion% | |
|---|---|---|---|---|
| Arepanrix | Pandemrix | |||
| HA 146 (N > D) | 4.40 × 10−6 | 10.2 (60 ± 38) | 59.7 | 10.7 |
| HA 314 (P > Q) | 5.70 × 10−4 | 7.3 (55.6 ± 27) | 44.2 | 2.9 |
| HA 420 (R > I) | 4.42 × 10−2 | 2.8 (45 ± 1) | 24.2 | 1.3 |
| HA 482 (F > Y) | 8.24 × 10−3 | 4.3 (51.2 ± 12.6) | 38.5 | 6.6 |
| M1 49 (R > I) | 3.90 × 10−3 | 4.6 (76.8 ± 23.4) | 79.1 | 29 |
| M1 84 (L > P) | 2.24 × 10−2 | −2.9 (−1.2 ± −11.6) | 14.9 | 21.3 |
| M1 85 (N > D) | 1.78 × 10−2 | −3.4 (−2.8 ± −19) | 23.5 | 34.4 |
| M1 91 (N > D) | 1.66 × 10−2 | 3.1 (13.2 ± 1.8) | 17.7 | 10.3 |
| M1 92 (N > D) | 2.61 × 10−3 | 4.2 (7.6 ± 2.2) | 10.2 | 5.3 |
| M1 99 (L > M) | 2.12 × 10−2 | 2.8 (11 ± 1.1) | 10.4 | 4.3 |
| NP 11 (E > Q) | 2.80 × 10−5 | 7.9 (11.1 ± 6.2) | 10.5 | 1.9 |
| NP 257 (T > A) | 5.20 × 10−4 | −6.8 (−4 ± −8.5) | 0.3 | 6.5 |
| NP 321 (N > D) | 2.14 × 10−2 | 3.3 (41.8 ± 5.3) | 29.2 | 5.7 |
| NP 423 (T > R) | 3.19 × 10−2 | −2.9 (−0.5 ± −7.4) | 0 | 4 |
| NP 424 (T > I) | 3.65 × 10−3 | −5.1 (−3.6 ± −10.8) | 0.1 | 7.3 |
| NP 432 (N > D) | 1.28 × 10−2 | 3.1 (13.6 ± 2.1) | 17.2 | 9.3 |
| NP 469 (E > D) | 1.13 × 10−2 | 3.9 (12.9 ± 2.6) | 11.5 | 3.8 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ambati, A.; Luo, G.; Pradhan, E.; Louis, J.; Lin, L.; Leib, R.D.; Ollila, H.M.; Poiret, T.; Adams, C.; Mignot, E. Mass Spectrometric Characterization of Narcolepsy-Associated Pandemic 2009 Influenza Vaccines. Vaccines 2020, 8, 630. https://doi.org/10.3390/vaccines8040630
Ambati A, Luo G, Pradhan E, Louis J, Lin L, Leib RD, Ollila HM, Poiret T, Adams C, Mignot E. Mass Spectrometric Characterization of Narcolepsy-Associated Pandemic 2009 Influenza Vaccines. Vaccines. 2020; 8(4):630. https://doi.org/10.3390/vaccines8040630
Chicago/Turabian StyleAmbati, Aditya, Guo Luo, Elora Pradhan, Jacob Louis, Ling Lin, Ryan D. Leib, Hanna Maria Ollila, Thomas Poiret, Christopher Adams, and Emmanuel Mignot. 2020. "Mass Spectrometric Characterization of Narcolepsy-Associated Pandemic 2009 Influenza Vaccines" Vaccines 8, no. 4: 630. https://doi.org/10.3390/vaccines8040630
APA StyleAmbati, A., Luo, G., Pradhan, E., Louis, J., Lin, L., Leib, R. D., Ollila, H. M., Poiret, T., Adams, C., & Mignot, E. (2020). Mass Spectrometric Characterization of Narcolepsy-Associated Pandemic 2009 Influenza Vaccines. Vaccines, 8(4), 630. https://doi.org/10.3390/vaccines8040630






