A Phase 1 Randomized Placebo-Controlled Study to Assess the Safety, Immunogenicity and Genetic Stability of a New Potential Pandemic H7N9 Live Attenuated Influenza Vaccine in Healthy Adults
Abstract
1. Introduction
2. Materials and Methods
2.1. Vaccine
2.2. Study Design
2.3. Study Population
2.4. Intervention
2.5. Outcomes
2.6. Trial Registration
2.7. Clinical Observation
2.8. Ethical Approval
2.9. Vaccine Virus Isolation in Chicken Eggs
2.10. PCR-Based Vaccine Virus Detection
2.11. Determining ts/ca Phenotype Stability of Clinical Isolates
2.12. Determining Genotype Stability of Clinical Isolates
2.13. Hemagglutination Inhibition (HI) Test
2.14. Microneutralization (MN) Assay
2.15. Measurement of Virus-Specific Antibodies
2.16. T-Cell Immune Responses
2.17. Statistical Analysis
3. Results
3.1. Study Population Characteristics
3.2. Reactogenicity
3.3. Vaccine Virus Shedding
3.3.1. Vaccine Virus Isolation in Embryonated Chicken Eggs
3.3.2. Vaccine Virus Detection by RT-PCR
3.4. Vaccine Virus trAnsmission to Placebo Recipients
3.5. Vaccine Virus Stability after Replication in Humans
3.5.1. Confirmation of 6:2 Genotype and Genetic Stability
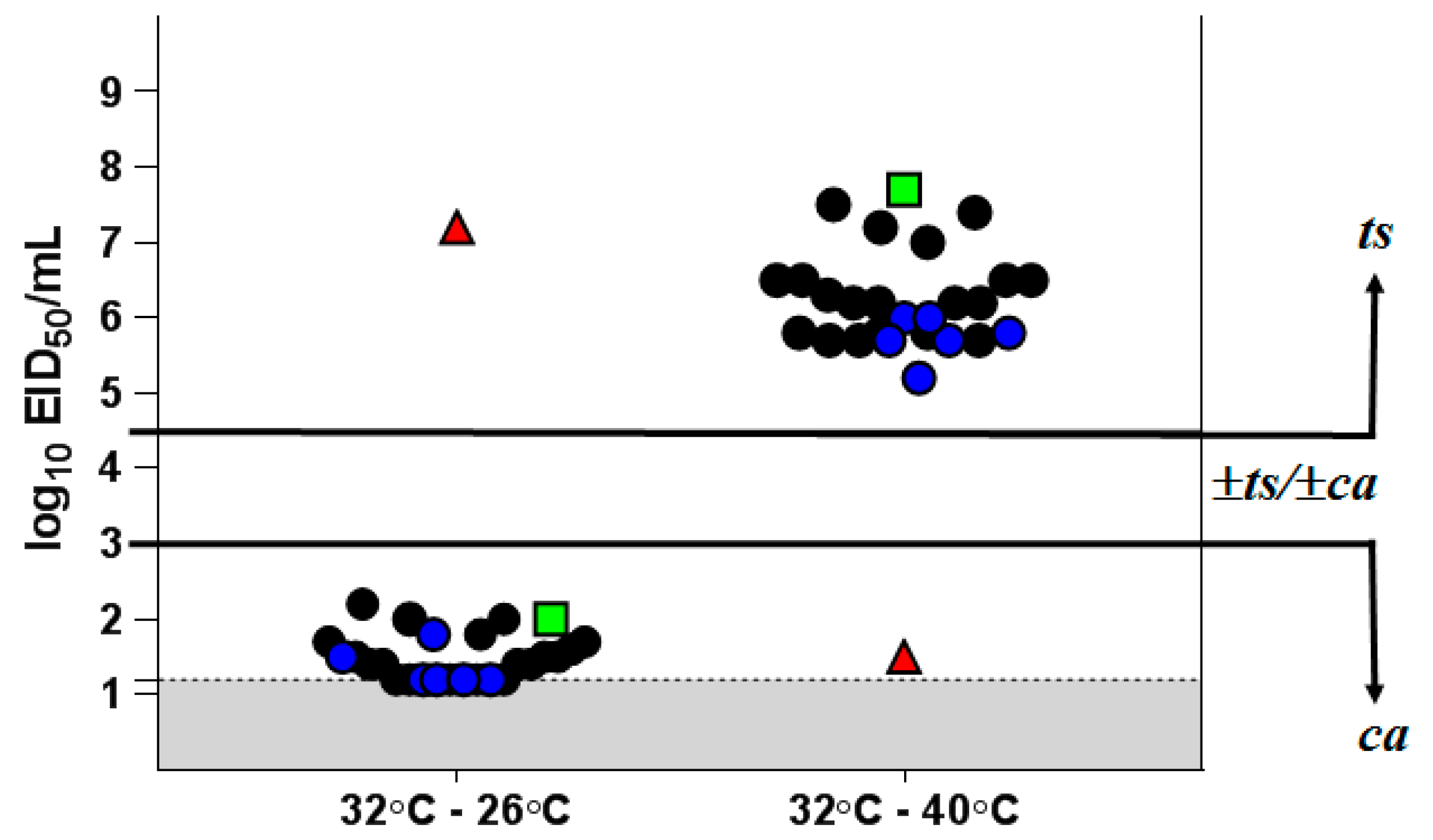
3.5.2. Phenotypic Stability of Vaccine Virus Clinical Isolates
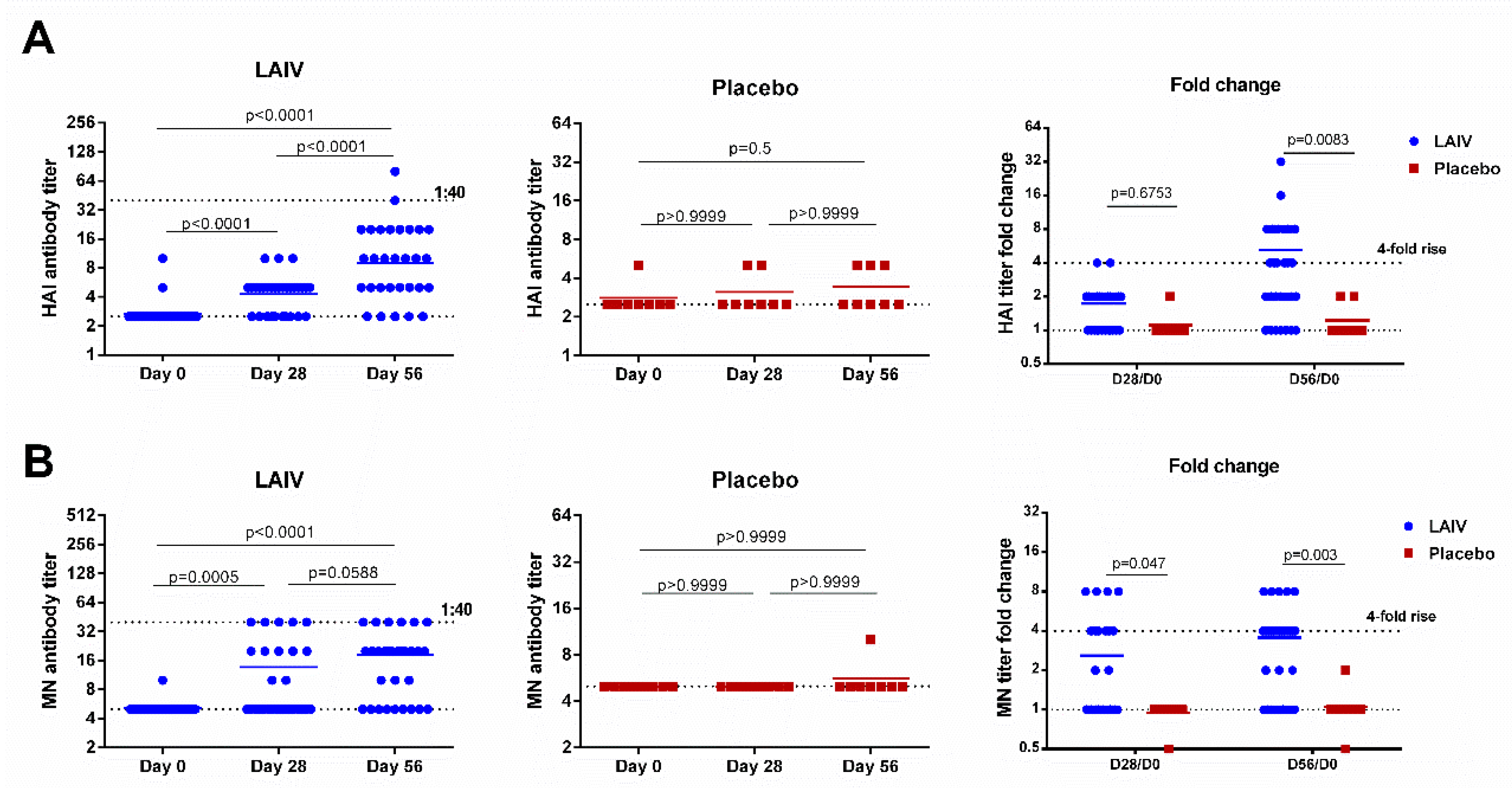
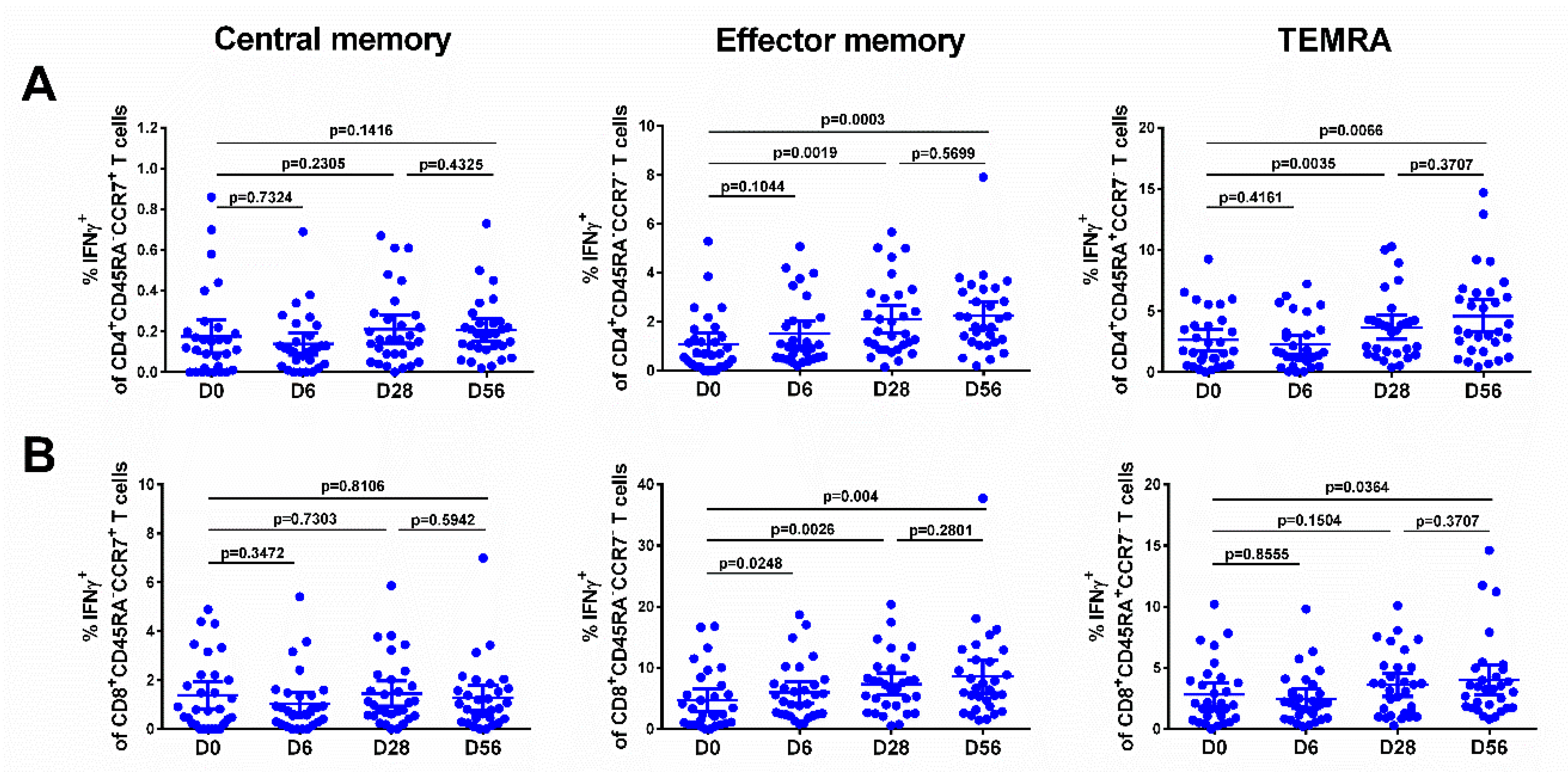
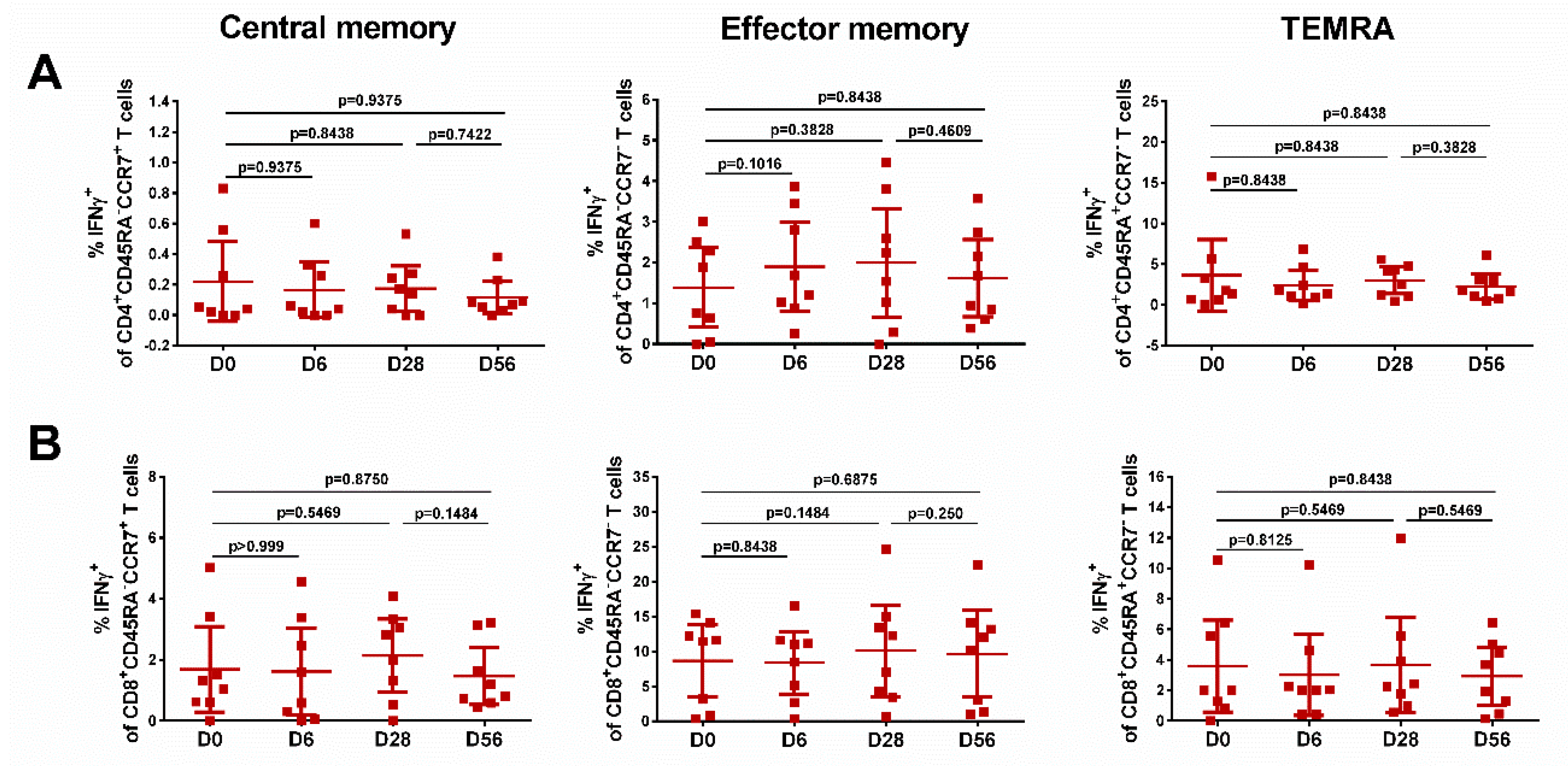
3.6. Immune Responses
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Disclaimer
References
- World Health Organization. Coronavirus. Available online: https://www.who.int/health-topics/coronavirus (accessed on 14 May 2020).
- Mossad, S.B. The resurgence of swine-origin influenza A (H1N1). Cleve. Clin. J. Med. 2009, 76, 337–343. [Google Scholar] [CrossRef] [PubMed]
- The Centers for Disease Control. Information on Avian Influenza. Available online: https://www.cdc.gov/flu/avianflu/index.htm (accessed on 14 May 2020).
- World Health Organization. Human Infection with Influenza A (H7N9) Virus in China. Available online: http://www.who.int/csr/don/2013_04_01/en/ (accessed on 14 May 2020).
- Gao, R.; Cao, B.; Hu, Y.; Feng, Z.; Wang, D.-Y.; Hu, W.; Chen, J.; Jie, Z.; Qiu, H.; Xu, K.; et al. Human Infection with a Novel Avian-Origin Influenza A (H7N9) Virus. N. Engl. J. Med. 2013, 368, 1888–1897. [Google Scholar] [CrossRef] [PubMed]
- Kageyama, T.; Fujisaki, S.; Takashita, E.; Xu, H.; Yamada, S.; Uchida, Y.; Neumann, G.; Saito, T.; Kawaoka, Y.; Tashiro, M. Genetic analysis of novel avian A(H7N9) influenza viruses isolated from patients in China, February to April 2013. Euro Surveill. 2013, 18, 20453. [Google Scholar] [PubMed]
- World Health Organization. Antigenic and Genetic Characteristics of Zoonotic Influenza A Viruses and Development of Candidate Vaccine Viruses for Pandemic Preparedness. 30 September 2019. Available online: https://www.who.int/influenza/vaccines/virus/201909_zoonotic_vaccinevirusupdate.pdf?ua=1 (accessed on 14 May 2020).
- Chung, K.Y.; Coyle, E.M.; Jani, D.; King, L.; Bhardwaj, R.; Fries, L.; Smith, G.; Glenn, G.M.; Golding, H.; Khurana, S. ISCOMATRIX™ adjuvant promotes epitope spreading and antibody affinity maturation of influenza A H7N9 virus like particle vaccine that correlate with virus neutralization in humans. Vaccine 2015, 33, 3953–3962. [Google Scholar] [CrossRef] [PubMed]
- Fries, L.F.; Smith, G.E.; Glenn, G.M. A Recombinant Viruslike Particle Influenza A (H7N9) Vaccine. N. Engl. J. Med. 2013, 369, 2564–2566. [Google Scholar] [CrossRef] [PubMed]
- Nakamura, K.; Shirakura, M.; Suzuki, Y.; Naito, T.; Fujisaki, S.; Tashiro, M.; Nobusawa, E. Development of a high-yield reassortant influenza vaccine virus derived from the A/Anhui/1/2013 (H7N9) strain. Vaccine 2016, 34, 328–333. [Google Scholar] [CrossRef] [PubMed]
- Rudenko, L.; Isakova-Sivak, I.; Naykhin, A.; Kiseleva, I.; Stukova, M.; Erofeeva, M.; Korenkov, D.; Matyushenko, V.; Sparrow, E.; Kieny, M.-P. H7N9 live attenuated influenza vaccine in healthy adults: a randomised, double-blind, placebo-controlled, phase 1 trial. Lancet Infect. Dis. 2015, 16, 303–310. [Google Scholar] [CrossRef]
- Shcherbik, S.; Pearce, N.; Balish, A.; Jones, J.; Thor, S.; Davis, C.T.; Pearce, M.; Tumpey, T.; Cureton, D.; Chen, L.-M.; et al. Generation and Characterization of Live Attenuated Influenza A(H7N9) Candidate Vaccine Virus Based on Russian Donor of Attenuation. PLoS ONE 2015, 10, e0138951. [Google Scholar] [CrossRef] [PubMed]
- Sobhanie, M.; Matsuoka, Y.; Jegaskanda, S.; Fitzgerald, T.; Mallory, R.; Chen, Z.; Luke, C.; Treanor, J.; Subbarao, K. Evaluation of the Safety and Immunogenicity of a Candidate Pandemic Live Attenuated Influenza Vaccine (pLAIV) Against Influenza A(H7N9). J. Infect. Dis. 2016, 213, 922–929. [Google Scholar] [CrossRef] [PubMed]
- Suzuki, Y.; Odagiri, T.; Tashiro, M.; Nobusawa, E. Development of an Influenza A Master Virus for Generating High-Growth Reassortants for A/Anhui/1/2013(H7N9) Vaccine Production in Qualified MDCK Cells. PLoS ONE 2016, 11, e0160040. [Google Scholar] [CrossRef] [PubMed]
- Wu, U.-I.; Hsieh, S.-M.; Lee, W.-S.; Wang, N.-C.; Kung, H.-C.; Ou, T.-Y.; Chen, F.-L.; Lin, T.-Y.; Chen, Y.-C.; Chang, S.-Y. Safety and immunogenicity of an inactivated cell culture-derived H7N9 influenza vaccine in healthy adults: A phase I/II, prospective, randomized, open-label trial. Vaccine 2017, 35, 4099–4104. [Google Scholar] [CrossRef] [PubMed]
- Zheng, D.; Gao, F.; Zhao, C.; Ding, Y.; Cao, Y.; Yang, T.; Xu, X.; Chen, Z. Comparative effectiveness of H7N9 vaccines in healthy individuals. Hum. Vaccines Immunother. 2018, 15, 80–90. [Google Scholar] [CrossRef] [PubMed]
- Bart, S.A.; Hohenboken, M.; Della Cioppa, G.; Narasimhan, V.; Dormitzer, P.R.; Kanesa-Thasan, N. A Cell Culture-Derived MF59-Adjuvanted Pandemic A/H7N9 Vaccine Is Immunogenic in Adults. Sci. Transl. Med. 2014, 6, 234ra55. [Google Scholar] [CrossRef] [PubMed]
- DeZure, A.D.; Coates, E.E.; Hu, Z.; Yamshchikov, G.V.; Zephir, K.L.; Enama, M.E.; Plummer, S.H.; Gordon, I.J.; Kaltovich, F.; Andrews, S.; et al. An avian influenza H7 DNA priming vaccine is safe and immunogenic in a randomized phase I clinical trial. NPJ Vaccines 2017, 2, 15. [Google Scholar] [CrossRef] [PubMed]
- Jackson, L.A.; Campbell, J.D.; Frey, S.E.; Edwards, K.M.; Keitel, W.A.; Kotloff, K.L.; Berry, A.A.; Graham, I.; Atmar, R.L.; Creech, C.B.; et al. Effect of Varying Doses of a Monovalent H7N9 Influenza Vaccine With and Without AS03 and MF59 Adjuvants on Immune Response. JAMA 2015, 314, 237–246. [Google Scholar] [CrossRef] [PubMed]
- Madan, A.; Segall, N.; Ferguson, M.; Frenette, L.; Kroll, R.; Friel, D.; Soni, J.; Li, P.; Innis, B.L.; Schuind, A. Immunogenicity and Safety of an AS03-Adjuvanted H7N9 Pandemic Influenza Vaccine in a Randomized Trial in Healthy Adults. J. Infect. Dis. 2016, 214, 1717–1727. [Google Scholar] [CrossRef] [PubMed]
- Mulligan, M.J.; I Bernstein, D.; Winokur, P.L.; E Rupp, R.; Anderson, E.; Rouphael, N.; Dickey, M.; Stapleton, J.T.; Edupuganti, S.; Spearman, P.; et al. Serological Responses to an Avian Influenza A/H7N9 Vaccine Mixed at the Point-of-Use With MF59 Adjuvant;A randomized clinical trial. JAMA 2014, 312, 1409–1419. [Google Scholar] [CrossRef] [PubMed]
- Ridenour, C.; Johnson, A.; Winne, E.; Hossain, J.; Mateu-Petit, G.; Balish, A.; Santana, W.; Kim, T.; Davis, C.; Cox, N.J.; et al. Development of influenza A(H7N9) candidate vaccine viruses with improved hemagglutinin antigen yield in eggs. Influenza Other Respir. Viruses 2015, 9, 263–270. [Google Scholar] [CrossRef] [PubMed]
- Isakova-Sivak, I.; Rudenko, L. Tackling a novel lethal virus: A focus on H7N9 vaccine development. Expert Rev. Vaccines 2017, 16, 709–721. [Google Scholar] [CrossRef] [PubMed]
- World Health Organization. Tables on Clinical Evaluation of Influenza Vaccines. Pandemic/Potentially Pandemic Influenza Vaccines. Updated August 2019. Available online: https://www.who.int/immunization/diseases/influenza/clinical_evaluation_tables/en/ (accessed on 14 May 2020).
- World Health Organization. Antigenic and Genetic Characteristics of Zoonotic Influenza Viruses and Development of Candidate Vaccine Viruses for Pandemic Preparedness. March 2017. Available online: https://www.who.int/influenza/vaccines/virus/201703_zoonotic_vaccinevirusupdate.pdf?ua=1 (accessed on 14 May 2020).
- Rudenko, L.; Kiseleva, I.; Krutikova, E.V.; Stepanova, E.; Isakova-Sivak, I.; Donina, S.; Rekstin, A.; Pisareva, M.; Bazhenova, E.; Kotomina, T.; et al. Two Live Attenuated Vaccines against Recent Low–and Highly Pathogenic H7N9 Influenza Viruses Are Safe and Immunogenic in Ferrets. Vaccines 2018, 6, 74. [Google Scholar] [CrossRef]
- Gelfand, A.S.; Bryzgalova, S.I.; Melnikov, S.J. A Method of Producing Live Influenza Vaccine for the Prevention of Influenza. Patent RU 2290205, 13 September 2005. [Google Scholar]
- World Health Organization. Manual on animal influenza diagnosis and surveillance. Available online: http://www.who.int/csr/resources/publications/influenza/whocdscsrncs20025rev.pdf (accessed on 10 June 2020).
- Reed, L.; Muench, H. A simple method of estimating fifty per cent endpoints. Am. J. Epidemiol. 1938, 27, 493–497. [Google Scholar] [CrossRef]
- Hoffmann, E.; Stech, J.; Guan, Y.; Webster, R.G.; Perez, D.R. Universal primer set for the full-length amplification of all influenza A viruses. Arch. Virol. 2001, 146, 2275–2289. [Google Scholar] [CrossRef] [PubMed]
- Matyushenko, V.; Isakova-Sivak, I.; Smolonogina, T.; Dubrovina, I.; Tretiak, T.; Rudenko, L. Genotyping assay for differentiation of wild-type and vaccine viruses in subjects immunized with live attenuated influenza vaccine. PLoS ONE 2017, 12, e0180497. [Google Scholar] [CrossRef] [PubMed]
- World Health Organization. Serological Detection of Avian Influenza A(H7N9) Virus Infections by Modified Horse Red Blood Cells Haemagglutination–Inhibition Assay. Laboratory Procedure. Available online: https://www.who.int/influenza/gisrs_laboratory/cnic_serological_diagnosis_hai_a_h7n9_20131220.pdf?ua=1 (accessed on 21 February 2020).
- Rowe, T.; Abernathy, R.A.; Hu-Primmer, J.; Thompson, W.W.; Lu, X.; Lim, W.; Fukuda, K.; Cox, N.J.; Katz, J.M. Detection of Antibody to Avian Influenza A (H5N1) Virus in Human Serum by Using a Combination of Serologic Assays. J. Clin. Microbiol. 1999, 37, 937–943. [Google Scholar] [CrossRef] [PubMed]
- Fujimoto, C.; Takeda, N.; Matsunaga, A.; Sawada, A.; Tanaka, T.; Kimoto, T.; Shinahara, W.; Sawabuchi, T.; Yamaguchi, M.; Hayama, M.; et al. Induction and maintenance of anti-influenza antigen-specific nasal secretory IgA levels and serum IgG levels after influenza infection in adults. Influenza Other Respir. Viruses 2012, 6, 396–403. [Google Scholar] [CrossRef] [PubMed]
- Sallusto, F.; Geginat, J.; Lanzavecchia, A. CentralMemory andEffectorMemoryT CellSubsets: Function, Generation, and Maintenance. Annu. Rev. Immunol. 2004, 22, 745–763. [Google Scholar] [CrossRef] [PubMed]
- Burel, J.G.; Qian, Y.; Arlehamn, C.S.L.; Weiskopf, D.; Zapardiel-Gonzalo, J.; Taplitz, R.; Gilman, R.H.; Saito, M.; De Silva, A.D.; Vijayanand, P.; et al. An integrated workflow to assess technical and biological variability of cell population frequencies in human peripheral blood by flow cytometry. J. Immunol. 2017, 198, 1748–1758. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.; Zhou, X.; Zheng, J.; Kwoh, C.K. Rules of co-occurring mutations characterize the antigenic evolution of human influenza A/H3N2, A/H1N1 and B viruses. BMC Med. Genom. 2016, 9, 69. [Google Scholar] [CrossRef] [PubMed]
- Yoon, S.-W.; Webby, R.J.; Webster, R.G. Evolution and Ecology of Influenza A Viruses. Curr. Top. Microbiol. Immunol. 2014, 385, 359–375. [Google Scholar] [CrossRef] [PubMed]
- World Health Organization. Recommendations on the Composition of Influenza Virus Vaccines. Available online: https://www.who.int/influenza/vaccines/virus/recommendations/en/ (accessed on 14 May 2020).
- World Health Organization. Global Influenza Preparedness Plan. 2005. Available online: http://www.who.int/csr/resources/publications/influenza/WHO_CDS_CSR_GIP_2005_5.pdf (accessed on 21 February 2020).
- Chen, T.; Zhang, R. Symptoms seem to be mild in children infected with avian influenza A (H5N6) and other subtypes. J. Infect. 2015, 71, 702–703. [Google Scholar] [CrossRef]
- Pan, M.; Gao, R.; Lv, Q.; Huang, S.; Zhou, Z.; Yang, L.; Li, X.; Zhao, X.; Zou, X.; Tong, W.; et al. Human infection with a novel, highly pathogenic avian influenza A (H5N6) virus: Virological and clinical findings. J. Infect. 2016, 72, 52–59. [Google Scholar] [CrossRef] [PubMed]
- Yang, Z.F.; Mok, C.K.P.; Peiris, J.S.M.; Zhong, N.S. Human Infection with a Novel Avian Influenza A(H5N6) Virus. N. Engl. J. Med. 2015, 373, 487–489. [Google Scholar] [CrossRef] [PubMed]
- White, V.C. A review of influenza viruses in seals and the implications for public health. U.S. Army Med. Dep. J. 2013, 45–50. [Google Scholar]
- Subbarao, K.; Klimov, A.; Katz, J.; Regnery, H.; Lim, W.; Hall, H.; Perdue, M.; Swayne, D.; Bender, C.; Huang, J.; et al. Characterization of an avian influenza A (H5N1) virus isolated from a child with a fatal respiratory illness. Science 1998, 279, 393–396. [Google Scholar] [CrossRef] [PubMed]
- Butt, K.M.; Smith, G.; Chen, H.; Zhang, L.J.; Leung, Y.H.C.; Xu, K.M.; Lim, W.; Webster, R.G.; Yuen, K.-Y.; Peiris, J.S.M.; et al. Human Infection with an Avian H9N2 Influenza A Virus in Hong Kong in 2003. J. Clin. Microbiol. 2005, 43, 5760–5767. [Google Scholar] [CrossRef] [PubMed]
- Cheng, V.; Chan, J.F.-W.; Wen, X.; Wu, W.; Que, T.; Chen, H.; Chan, K.; Yuen, K.-Y. Infection of immunocompromised patients by avian H9N2 influenza A virus. J. Infect. 2011, 62, 394–399. [Google Scholar] [CrossRef] [PubMed]
- Allen, G.M. Bats: Biology, Behavior, and Folklore; Dover Publications: Mineola, NY, USA, 2004; 368p. [Google Scholar]
- Yu, H.; Cowling, B.J.; Feng, L.; Lau, E.H.; Liao, Q.; Tsang, T.K.; Peng, Z.; Wu, P.; Liu, F.; Fang, V.J.; et al. Human infection with avian influenza A H7N9 virus: an assessment of clinical severity. Lancet 2013, 382, 138–145. [Google Scholar] [CrossRef]
- De Jong, J.C.; Claas, E.C.J.; Osterhaus, A.D.M.E.; Webster, R.G.; Lim, W.L. A pandemic warning? Nature 1997, 389, 554. [Google Scholar] [CrossRef] [PubMed]
- World Health Organization. Antigenic and Genetic Characteristics of Zoonotic influenza A Viruses and Development of Candidate Vaccine Viruses for Pandemic Preparedness. 28 February 2020. Available online: https://www.who.int/influenza/vaccines/virus/202002_zoonotic_vaccinevirusupdate.pdf?ua=1 (accessed on 14 May 2020).
- World Health Organization. Global Action Plan to Increase Vaccine Supply for Influenza Vaccines. Available online: http://whqlibdoc.who.int/hq/2006/WHO_IVB_06.13_eng.pdf (accessed on 14 May 2020).
- Isakova-Sivak, I.; Stukova, M.; Erofeeva, M.; Naykhin, A.; Donina, S.; Petukhova, G.; Kuznetsova, V.; Kiseleva, I.; Smolonogina, T.; Dubrovina, I.; et al. H2N2 live attenuated influenza vaccine is safe and immunogenic for healthy adult volunteers. Hum. Vaccines Immunother. 2015, 11, 970–982. [Google Scholar] [CrossRef] [PubMed]
- Rudenko, L.; Desheva, J.; Korovkin, S.; Mironov, A.; Rekstin, A.; Grigorieva, E.; Donina, S.; Gambaryan, A.; Katlinsky, A. Safety and immunogenicity of live attenuated influenza reassortant H5 vaccine (phase I-II clinical trials). Influenza Other Respir. Viruses 2008, 2, 203–209. [Google Scholar] [CrossRef] [PubMed]
- Rudenko, L.; Kiseleva, I.; Naykhin, A.N.; Erofeeva, M.; Stukova, M.; Donina, S.; Petukhova, G.; Pisareva, M.; Krivitskaya, V.; Grudinin, M.; et al. Assessment of Human Immune Responses to H7 Avian Influenza Virus of Pandemic Potential: Results from a Placebo–Controlled, Randomized Double–Blind Phase I Study of Live Attenuated H7N3 Influenza Vaccine. PLoS ONE 2014, 9, e87962. [Google Scholar] [CrossRef] [PubMed]
- Rudenko, L.; Kiseleva, I.; Stukova, M.; Erofeeva, M.; Naykhin, A.; Donina, S.; Larionova, N.; Pisareva, M.; Krivitskaya, V.; Flores, J.; et al. Clinical testing of pre-pandemic live attenuated A/H5N2 influenza candidate vaccine in adult volunteers: Results from a placebo-controlled, randomized double-blind phase I study. Vaccine 2015, 33, 5110–5117. [Google Scholar] [CrossRef] [PubMed]
- Isakova-Sivak, I.; Rudenko, L. Safety, immunogenicity and infectivity of new live attenuated influenza vaccines. Expert Rev. Vaccines 2015, 14, 1313–1329. [Google Scholar] [CrossRef] [PubMed]
- Klimov, A.I.; Kiseleva, I.V.; Desheva, J.A.; Alexandrova, G.I.; Cox, N.J.; Rudenko, L.G. Live attenuated reassortant influenza vaccine prepared using A/Leningrad/134/17/57 (H2N2) donor strain is genetically stable after replication in children 3–6 years of age. In Proceedings of the Options for the Control of Influenza IV, Crete, Greece, 23–28 September 2000; pp. 951–954. [Google Scholar]
- Kiseleva, I.; Rudenko, L. Potentially Pandemic Live Influenza Vaccines Based on Russian Master Donor Virus are Genetically Stable after Replication in Humans. J. Vaccines Vaccin. 2016, 7, 1–3. [Google Scholar] [CrossRef]
- Kiseleva, I.; Dubrovina, I.; Fedorova, E.; Larionova, N.; Isakova-Sivak, I.; Bazhenova, E.; Pisareva, M.; Kuznetsova, V.; Flores, J.; Rudenko, L. Genetic stability of live attenuated vaccines against potentially pandemic influenza viruses. Vaccine 2015, 33, 7008–7014. [Google Scholar] [CrossRef] [PubMed]
- Shcherbik, S.; Pearce, N.; Carney, P.; Bazhenova, E.; Larionova, N.; Kiseleva, I.; Rudenko, L.; Kumar, A.; Goldsmith, C.S.; Dugan, V.; et al. Evaluation of A(H1N1)pdm09 LAIV vaccine candidates stability and replication efficiency in primary human nasal epithelial cells. Vaccine X 2019, 2, 100031. [Google Scholar] [CrossRef] [PubMed]
- Davenport, F.M.; Hennessy, A.V.; Maassab, H.F.; Minuse, E.; Clark, L.C.; Abrams, G.D.; Mitchell, J.R. Pilot Studies on Recombinant Cold-Adapted Live Type A and B Influenza Virus Vaccines. J. Infect. Dis. 1977, 136, 17–25. [Google Scholar] [CrossRef] [PubMed]
- Rudenko, L.G.; Lonskaya, N.I.; Klimov, A.I.; Vasilieva, R.I.; Ramirez, A. Clinical and epidemiological evaluation of a live, cold-adapted influenza vaccine for 3–14-year-olds. Bull. World Health Organ. 1996, 74, 77–84. [Google Scholar] [PubMed]
- Vesikari, T.; Karvonen, A.; Korhonen, T.; Edelman, K.; Vainionp, R.; Salmi, A.; Saville, M.K.; Cho, I.; Razmpour, A.; Rappaport, R.; et al. A Randomized, Double-Blind Study of the Safety, Transmissibility and Phenotypic and Genotypic Stability of Cold-Adapted Influenza Virus Vaccine. Pediatr. Infect. Dis. J. 2006, 25, 590–595. [Google Scholar] [CrossRef]
- Krammer, F. The human antibody response to influenza A virus infection and vaccination. Nat. Rev. Immunol. 2019, 19, 383–397. [Google Scholar] [CrossRef] [PubMed]
- Schotsaert, M.; Saelens, X.; Leroux-Roels, G. Influenza vaccines: T-cell responses deserve more attention. Expert Rev. Vaccines 2012, 11, 949–962. [Google Scholar] [CrossRef] [PubMed]
- Jansen, J.M.; Gerlach, T.; Elbahesh, H.; Rimmelzwaan, G.F.; Saletti, G. Influenza virus-specific CD4+ and CD8+ T cell-mediated immunity induced by infection and vaccination. J. Clin. Virol. 2019, 119, 44–52. [Google Scholar] [CrossRef] [PubMed]
- Kedzierska, K.; Valkenburg, S.A.; Doherty, P.C.; Davenport, M.P.; Venturi, V. Use it or lose it: establishment and persistence of T cell memory. Front. Immunol. 2012, 3, 357. [Google Scholar] [CrossRef] [PubMed]


| Characteristics | Group | ||
|---|---|---|---|
| Vaccine, n (%) | Placebo, n (%) | ||
| Gender | Male | 18/30 (60.0) | 6/10 (60.0) |
| Female | 12/30 (40.0) | 4/10 (40.0) | |
| Age (years) | Mean (SE 1) | 32.6 (9.8) | 34.8 (9.3) |
| Median | 33.0 | 31.0 | |
| Range | 18.0–48.0 | 25.0–49.0 | |
| Weight (kg) | Mean (SE) | 68.1 (11.5) | 74.6 (3.8) |
| Median | 65.0 | 74.0 | |
| Range | 50.0–94.0 | 70.0–81.0 | |
| Height (cm) | Mean (SE) | 175.6 (10.1) | 175.1 (7.0) |
| Median | 173.5 | 174.5 | |
| Range | 160.0–210.0 | 166.0–185.0 | |
| Adverse Reaction | Worst Grade 2 | Group | |||
|---|---|---|---|---|---|
| Vaccine (n = 30) | Placebo (n = 8) | ||||
| N 3 (%) | 95% CI | N 3 (%) | 95% CI | ||
| Dose 1 | |||||
| Reactions within 2 h | none | 0/30 (0) | 0/8 (0) | ||
| Any solicited local or systemic reaction 1 | 3/30 (10.0) | 3.5–25.6 | 0/8 (0) | 0.0–32.4 | |
| Local reactions (total) | 1/30 (3.3) | 0.6–16.7 | 0/8 (0) | 0.0–32.4 | |
| Nasal congestion | mild | 1/30 (3.3) | 0.6–16.7 | 0/8 (0) | 0.0–32.4 |
| Systemic reactions (total) | 4/30 (13.3) | 5.3–29.7 | 0/8 (0) | 0.0–32.4 | |
| Fatigue | mild | 1/30 (3.3) | 0.6–16.7 | 0/8 (0) | 0.0–32.4 |
| Malaise | mild | 1/30 (3.3) | 0.6–16.7 | 0/8 (0) | 0.0–32.4 |
| Fever | mild | 2/30 (6.7) | 1.8–21.3 | 0/8 (0) | 0.0–32.4 |
| Worst grade (total) | mild (n = 5) | None | |||
| Any serious adverse event | none | None | |||
| Dose 2 | |||||
| Reactions within 2 h | none | 0/30 (0) | 0/8 (0) | ||
| Any solicited local or systemic reaction 1 | 1/30 (3.3) | 0.6–16.7 | 3/8 (37.5) | 13.7–69.4 | |
| Local reactions (total) | 1/30 (3.3) | 0.6–16.7 | 0/8 (0) | 0.0–32.4 | |
| Nasal congestion | mild | 1/30 (3.3) | 0.6–16.7 | 0/8 (0) | 0.0–32.4 |
| Systemic reactions (total) | 0/30 (0) | 0.0–11.4 | 5/8 (62.5) | 30.6–86.3 | |
| Cough | mild | 0/30 (0) | 0.0–11.4 | 1/8 (12.5) | 2.2–47.1 |
| Sore throat | mild | 0/30 (0) | 0.0–11.4 | 1/8 (12.5) | 2.2–47.1 |
| Fever | mild | 0/30 (0) | 0.0–11.4 | 1/8 (12.5) | 2.2–47.1 |
| Herpes | mild | 0/30 (0) | 0.0–11.4 | 1/8 (12.5) | 2.2–47.1 |
| Hyperemia arches throat | mild | 0/30 (0) | 0.0–11.4 | 1/8 (12.5) | 2.2–47.1 |
| Worst grade (total) | mild (n = 1) | mild (n = 5) | |||
| Any serious adverse event | none | None | |||
| Vaccination | Group 1 | Virus Isolation Confirmed by | Clinical Isolates | Total No. of Positive Subjects | ||||||
|---|---|---|---|---|---|---|---|---|---|---|
| D1 | D2 | D3 | D4 | D5 | D6 | Total No. of Isolates | ||||
| Dose 1 | Vaccine (n = 30) | RT–PCR | 27/30 | 8/30 | 5/30 | 3/30 | 1/30 | 1/30 | 45 | 27/30 (90.0%) |
| Culture | 6/30 | 5/30 | 5/30 | 3/30 | 0/10 | 0/10 | 19 | 11/30 (36.7%) | ||
| Placebo (n = 8) | RT–PCR | 0/8 | 0/8 | 0/8 | 0/8 | 0/8 | 0/8 | 0 | 0/8 (0%) | |
| Culture | 0/8 | 0/8 | 0/8 | 0/8 | 0/8 | 0/8 | 0 | 0/8 (0%) | ||
| Dose 2 | Vaccine (n = 30) | RT–PCR | 9/30 | 4/30 | 1/30 | 2/30 | 0/30 | ND 2 | 16 | 4/30 (13.3%) |
| Culture | 1/30 | 2/30 | 1/30 | 2/30 | ND | ND | 6 | 3/30 (10.0%) | ||
| Placebo (n = 8) | RT–PCR | 0/8 | 0/8 | 0/8 | 0/8 | ND | ND | 0 | 0/8 (0%) | |
| Culture | 0/8 | 0/8 | 0/8 | 0/8 | ND | ND | 0 | 0/8 (0%) | ||
| LAIV Based on Vaccine Candidate [ClinicalTrials.Gov Identifier] | Dose Per 0.5 mL | Vaccine Virus Shedding after 1 and 2 Doses, Total Number | Immunogenicity (Any Test) 3 | Reference | |
|---|---|---|---|---|---|
| Isolates 1 | Subjects 2 | ||||
| A/17/duck/Potsdam/88/92 (H5N2) [No data] | 6.9 log10 EID50 | 25 | No data | 80.0% (16/20) | [57] |
| A/17/mallard/Netherlands/90/95 (H7N3) [NCT01511419] | 7.0 log10 EID50 | 4 | 4 (13.3%) | 86.2% (25/29) | [55] |
| A/17/turkey/Turkey/05/133 (H5N2) [NCT01719783] | 8.4 log10 EID50 | 16 | 14 (46.7%) | 96.6% (28/29) | [56] |
| A/17/California/66/395 H2N2) [NCT01982331] | 7.5 log10 EID50 | 20 | 14 (50.0%) | 92.6% (24/26) | [53] |
| A/17/Anhui/2013/61 (H7N9) [NCT02480101] | 7.5 log10 EID50 | 45 | 22 (73.3%) | 93.1% (27/29) | [11] |
| A/17/Hong Kong/2017/75108 (H7N9) [NCT03739229] | 7.0 log10 EID50 | 25 | 12 (40.0%) | 96.6% (29/30) | Current study |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kiseleva, I.; Isakova-Sivak, I.; Stukova, M.; Erofeeva, M.; Donina, S.; Larionova, N.; Krutikova, E.; Bazhenova, E.; Stepanova, E.; Vasilyev, K.; et al. A Phase 1 Randomized Placebo-Controlled Study to Assess the Safety, Immunogenicity and Genetic Stability of a New Potential Pandemic H7N9 Live Attenuated Influenza Vaccine in Healthy Adults. Vaccines 2020, 8, 296. https://doi.org/10.3390/vaccines8020296
Kiseleva I, Isakova-Sivak I, Stukova M, Erofeeva M, Donina S, Larionova N, Krutikova E, Bazhenova E, Stepanova E, Vasilyev K, et al. A Phase 1 Randomized Placebo-Controlled Study to Assess the Safety, Immunogenicity and Genetic Stability of a New Potential Pandemic H7N9 Live Attenuated Influenza Vaccine in Healthy Adults. Vaccines. 2020; 8(2):296. https://doi.org/10.3390/vaccines8020296
Chicago/Turabian StyleKiseleva, Irina, Irina Isakova-Sivak, Marina Stukova, Marianna Erofeeva, Svetlana Donina, Natalie Larionova, Elena Krutikova, Ekaterina Bazhenova, Ekaterina Stepanova, Kirill Vasilyev, and et al. 2020. "A Phase 1 Randomized Placebo-Controlled Study to Assess the Safety, Immunogenicity and Genetic Stability of a New Potential Pandemic H7N9 Live Attenuated Influenza Vaccine in Healthy Adults" Vaccines 8, no. 2: 296. https://doi.org/10.3390/vaccines8020296
APA StyleKiseleva, I., Isakova-Sivak, I., Stukova, M., Erofeeva, M., Donina, S., Larionova, N., Krutikova, E., Bazhenova, E., Stepanova, E., Vasilyev, K., Matyushenko, V., Krylova, M., Galatonova, J., Ershov, A., Lioznov, D., Sparrow, E. G., Torelli, G., & Rudenko, L. (2020). A Phase 1 Randomized Placebo-Controlled Study to Assess the Safety, Immunogenicity and Genetic Stability of a New Potential Pandemic H7N9 Live Attenuated Influenza Vaccine in Healthy Adults. Vaccines, 8(2), 296. https://doi.org/10.3390/vaccines8020296






