Impact of HIV-1 Diversity on Its Sensitivity to Neutralization
Abstract
1. Introduction
2. HIV Env Diversity and Humoral Response
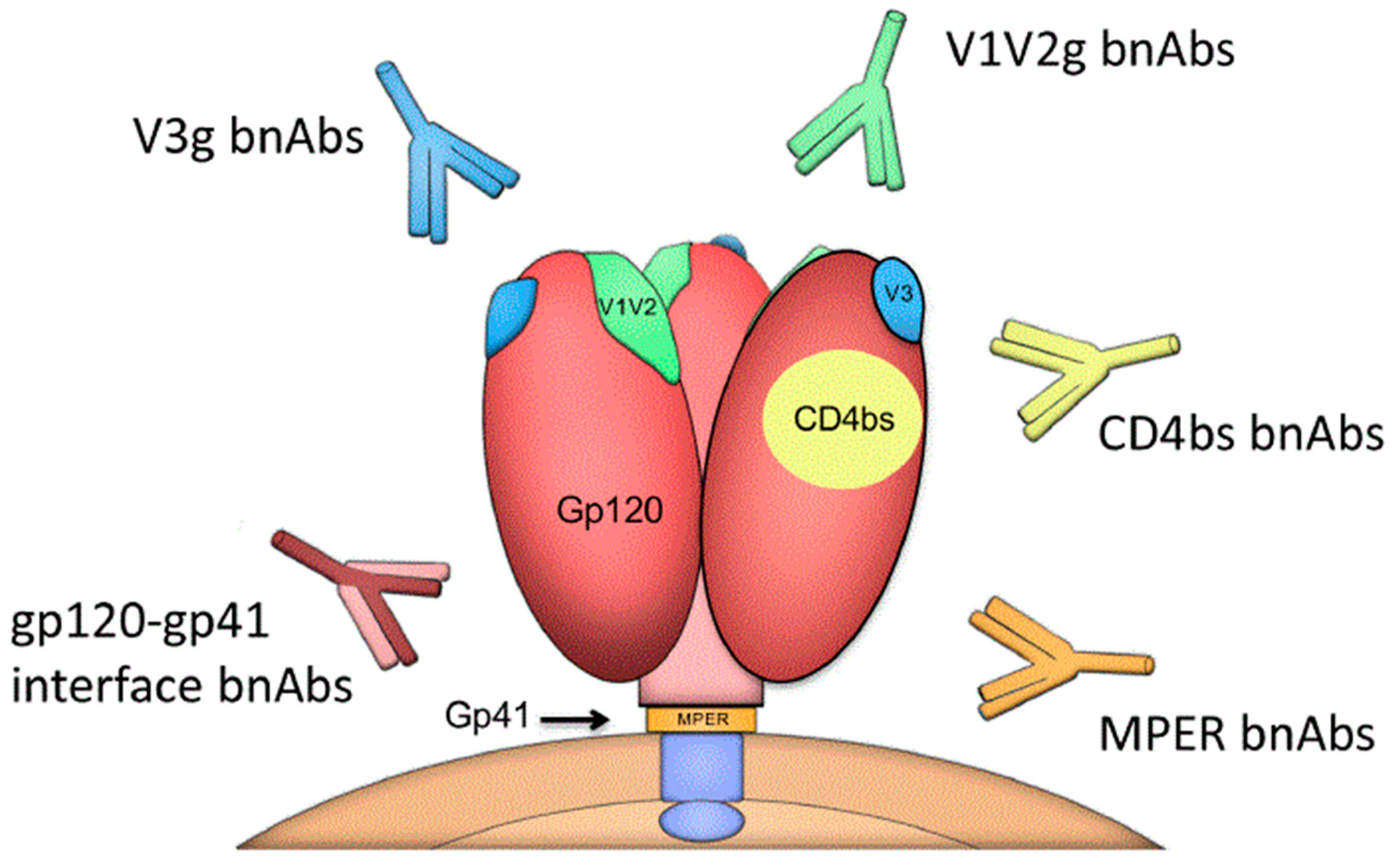
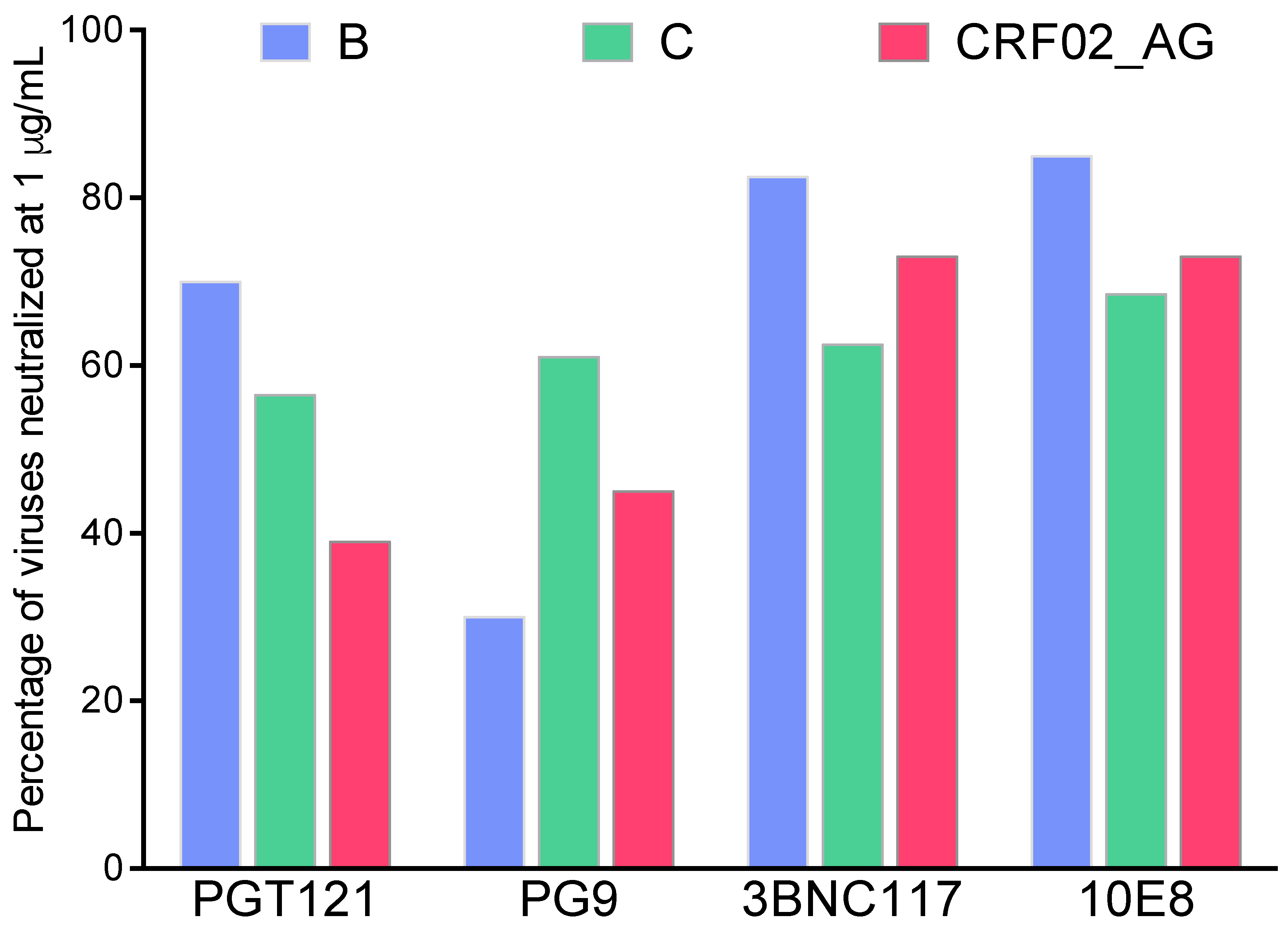
3. BnAbs, Env Targets, and Cross-Subtype Neutralization
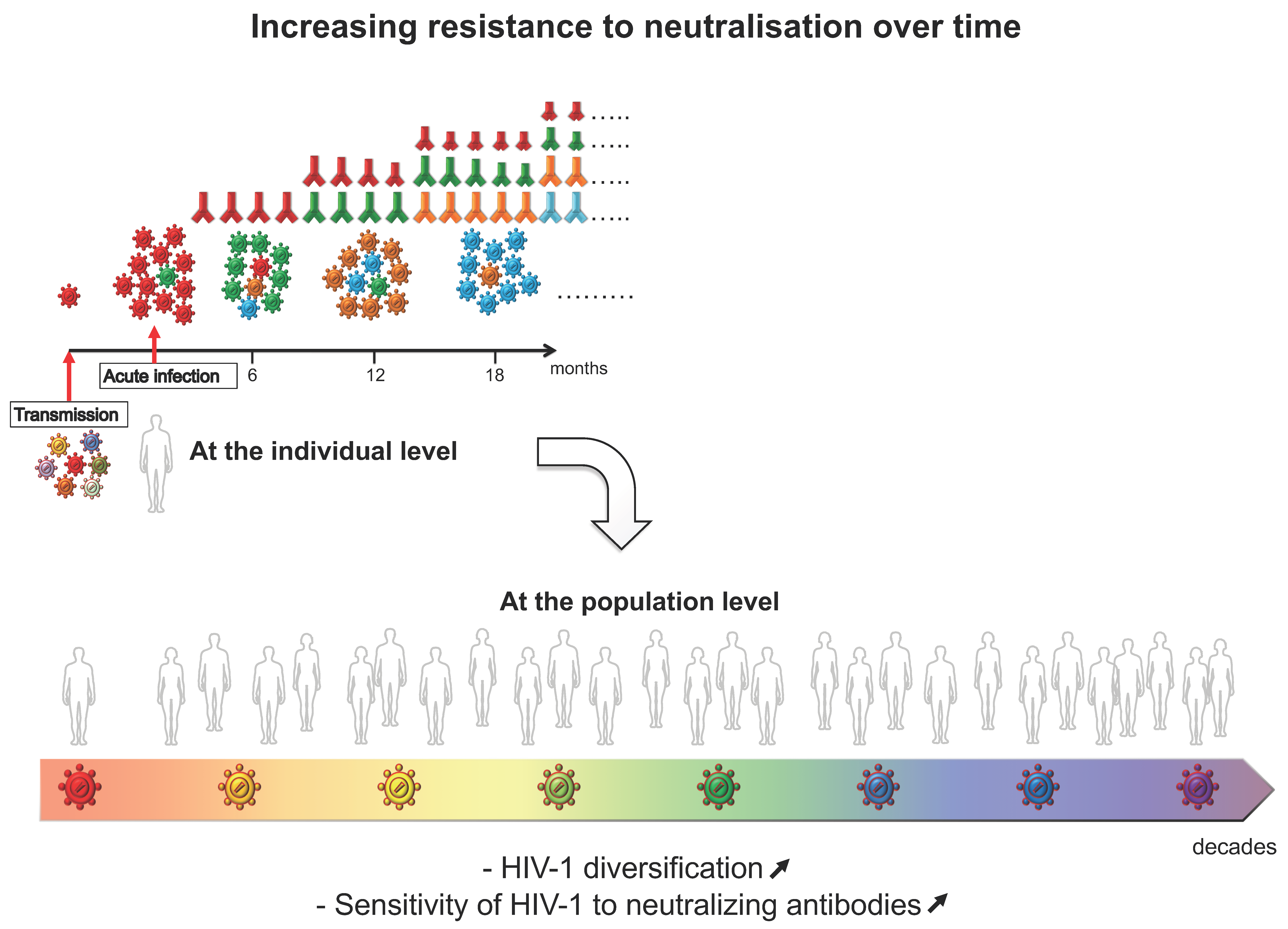
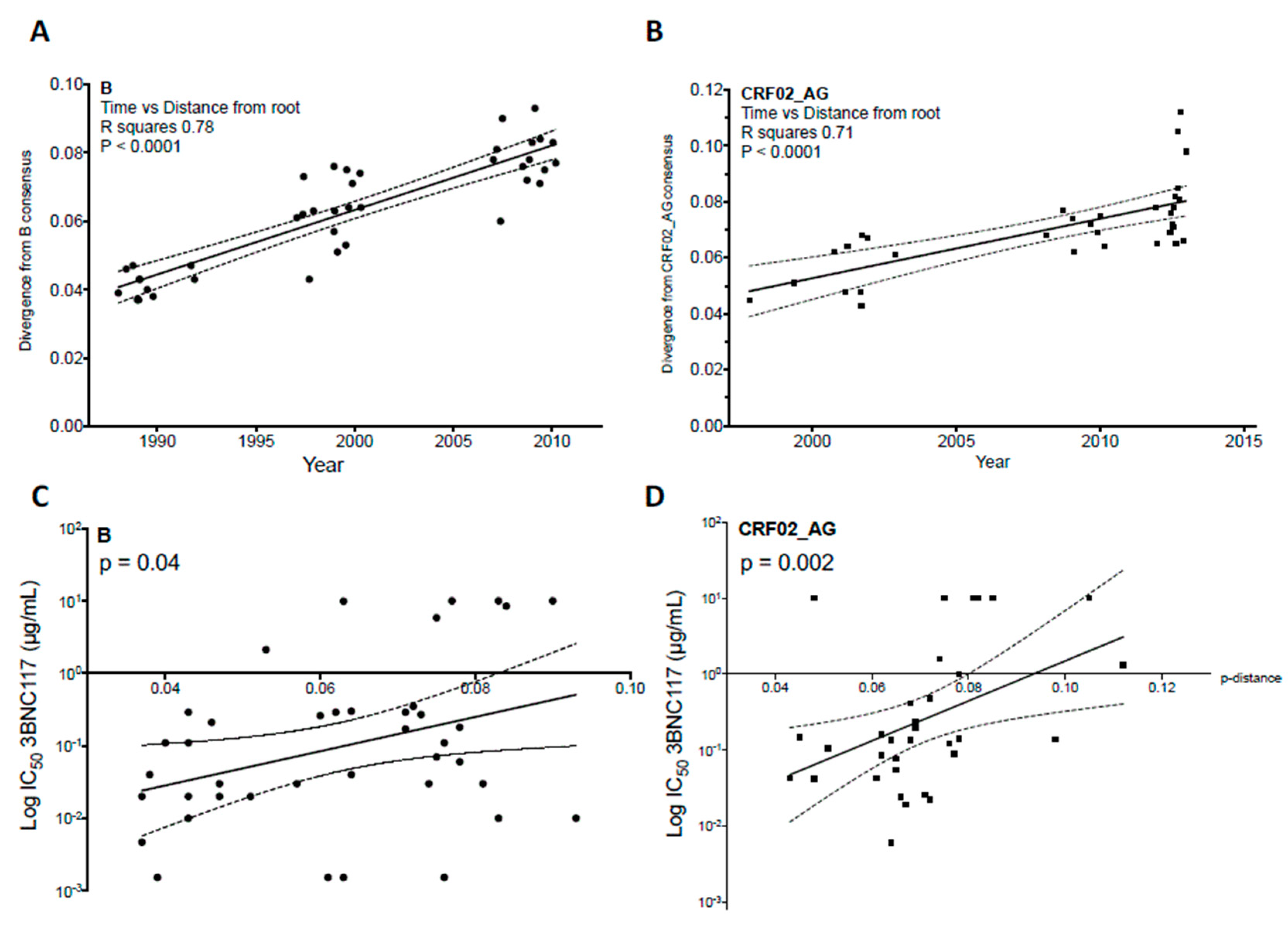
4. Evolution of HIV-1 Env Towards a Greater Resistance to Neutralization over Time
5. A Future Vaccine to Overcome HIV-1 Diversity?
6. Conclusions
Acknowledgments
Conflicts of Interest
References
- Mascola, J.R.; Stiegler, G.; VanCott, T.C.; Katinger, H.; Carpenter, C.B.; Hanson, C.E.; Beary, H.; Hayes, D.; Frankel, S.S.; Birx, D.L.; et al. Protection of macaques against vaginal transmission of a pathogenic HIV-1/SIV chimeric virus by passive infusion of neutralizing antibodies. Nat. Med. 2000, 6, 207–210. [Google Scholar] [CrossRef] [PubMed]
- Shingai, M.; Nishimura, Y.; Klein, F.; Mouquet, H.; Donau, O.K.; Plishka, R.; Buckler-White, A.; Seaman, M.; Piatak, M.; Lifson, J.D.; et al. Antibody-mediated immunotherapy of macaques chronically infected with SHIV suppresses viraemia. Nature 2013, 503, 277–280. [Google Scholar] [CrossRef] [PubMed]
- Julg, B.; Liu, P.-T.; Wagh, K.; Fischer, W.M.; Abbink, P.; Mercado, N.B.; Whitney, J.B.; Nkolola, J.P.; McMahan, K.; Tartaglia, L.J.; et al. Protection against a mixed SHIV challenge by a broadly neutralizing antibody cocktail. Sci. Transl. Med. 2017, 9, eaao4235. [Google Scholar] [CrossRef]
- Nishimura, Y.; Gautam, R.; Chun, T.-W.; Sadjadpour, R.; Foulds, K.E.; Shingai, M.; Klein, F.; Gazumyan, A.; Golijanin, J.; Donaldson, M.; et al. Early antibody therapy can induce long-lasting immunity to SHIV. Nature 2017, 543, 559–563. [Google Scholar] [CrossRef]
- Gilbert, P.B.; Juraska, M.; DeCamp, A.C.; Karuna, S.; Edupuganti, S.; Mgodi, N.; Donnell, D.J.; Bentley, C.; Sista, N.; Andrew, P.; et al. Basis and Statistical Design of the Passive HIV-1 Antibody Mediated Prevention (AMP) Test-of-Concept Efficacy Trials. Stat. Commun. Infect. Dis. 2017, 9, 20160001. [Google Scholar] [CrossRef] [PubMed]
- Gautam, R.; Nishimura, Y.; Gaughan, N.; Gazumyan, A.; Schoofs, T.; Buckler-White, A.; Seaman, M.S.; Swihart, B.J.; Follmann, D.A.; Nussenzweig, M.C.; et al. A single injection of crystallizable fragment domain-modified antibodies elicits durable protection from SHIV infection. Nat. Med. 2018, 24, 610–616. [Google Scholar] [CrossRef] [PubMed]
- Gaschen, B.; Taylor, J.; Yusim, K.; Foley, B.; Gao, F.; Lang, D.; Novitsky, V.; Haynes, B.; Hahn, B.H.; Bhattacharya, T.; et al. Diversity considerations in HIV-1 vaccine selection. Science 2002, 296, 2354–2360. [Google Scholar] [CrossRef]
- Taylor, B.S.; Hammer, S.M. The challenge of HIV-1 subtype diversity. N. Engl. J. Med. 2008, 359, 1965–1966. [Google Scholar] [CrossRef]
- Lynch, R.M.; Shen, T.; Gnanakaran, S.; Derdeyn, C.A. Appreciating HIV type 1 diversity: subtype differences in Env. AIDS Res. Hum. Retroviruses 2009, 25, 237–248. [Google Scholar] [CrossRef]
- Haynes, B.F.; Moody, M.A.; Alam, M.; Bonsignori, M.; Verkoczy, L.; Ferrari, G.; Gao, F.; Tomaras, G.D.; Liao, H.-X.; Kelsoe, G. Progress in HIV-1 vaccine development. J. Allergy Clin. Immunol. 2014, 134, 3–10. [Google Scholar] [CrossRef]
- Hemelaar, J.; Elangovan, R.; Yun, J.; Dickson-Tetteh, L.; Fleminger, I.; Kirtley, S.; Williams, B.; Gouws-Williams, E.; Ghys, P.D.; WHO–UNAIDS Network for HIV Isolation Characterisation. Global and regional molecular epidemiology of HIV-1, 1990-2015: A systematic review, global survey, and trend analysis. Lancet Infect. Dis. 2019, 19, 143–155. [Google Scholar] [CrossRef]
- Richman, D.D.; Wrin, T.; Little, S.J.; Petropoulos, C.J. Rapid evolution of the neutralizing antibody response to HIV type 1 infection. Proc. Natl. Acad. Sci. USA 2003, 100, 4144–4149. [Google Scholar] [CrossRef] [PubMed]
- Li, B.; Decker, J.M.; Johnson, R.W.; Bibollet-Ruche, F.; Wei, X.; Mulenga, J.; Allen, S.; Hunter, E.; Hahn, B.H.; Shaw, G.M.; et al. Evidence for Potent Autologous Neutralizing Antibody Titers and Compact Envelopes in Early Infection with Subtype C Human Immunodeficiency Virus Type 1. J. Virol. 2006, 80, 5211–5218. [Google Scholar] [CrossRef] [PubMed]
- Gray, E.S.; Moore, P.L.; Pantophlet, R.A.; Morris, L. N-Linked Glycan Modifications in gp120 of Human Immunodeficiency Virus Type 1 Subtype C Render Partial Sensitivity to 2G12 Antibody Neutralization. J. Virol. 2007, 81, 10769–10776. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Bunnik, E.M.; Pisas, L.; van Nuenen, A.C.; Schuitemaker, H. Autologous Neutralizing Humoral Immunity and Evolution of the Viral Envelope in the Course of Subtype B Human Immunodeficiency Virus Type 1 Infection. J. Virol. 2008, 82, 7932–7941. [Google Scholar] [CrossRef] [PubMed]
- Wei, X.; Decker, J.M.; Wang, S.; Hui, H.; Kappes, J.C.; Wu, X.; Salazar-Gonzalez, J.F.; Salazar, M.G.; Kilby, J.M.; Saag, M.S.; et al. Antibody neutralization and escape by HIV-1. Nature 2003, 422, 307–312. [Google Scholar] [CrossRef] [PubMed]
- Frost, S.D.W.; Wrin, T.; Smith, D.M.; Kosakovsky Pond, S.L.; Liu, Y.; Paxinos, E.; Chappey, C.; Galovich, J.; Beauchaine, J.; Petropoulos, C.J.; et al. Neutralizing antibody responses drive the evolution of human immunodeficiency virus type 1 envelope during recent HIV infection. Proc. Natl. Acad. Sci. USA 2005, 102, 18514–18519. [Google Scholar] [CrossRef]
- Mahalanabis, M.; Jayaraman, P.; Miura, T.; Pereyra, F.; Chester, E.M.; Richardson, B.; Walker, B.; Haigwood, N.L. Continuous viral escape and selection by autologous neutralizing antibodies in drug-naive human immunodeficiency virus controllers. J. Virol. 2008, 83, 662–672. [Google Scholar] [CrossRef]
- Liao, H.-X.; Lynch, R.; Zhou, T.; Gao, F.; Alam, S.M.; Boyd, S.D.; Fire, A.Z.; Roskin, K.M.; Schramm, C.A.; Zhang, Z.; et al. Co-evolution of a broadly neutralizing HIV-1 antibody and founder virus. Nature 2013, 496, 469–476. [Google Scholar] [CrossRef]
- Moore, P.L.; Ranchobe, N.; Lambson, B.E.; Gray, E.S.; Cave, E.; Abrahams, M.-R.; Bandawe, G.; Mlisana, K.; Abdool Karim, S.S.; Williamson, C.; et al. Limited neutralizing antibody specificities drive neutralization escape in early HIV-1 subtype C infection. PLoS Pathog. 2009, 5, e1000598. [Google Scholar] [CrossRef]
- Rong, R.; Li, B.; Lynch, R.M.; Haaland, R.E.; Murphy, M.K.; Mulenga, J.; Allen, S.A.; Pinter, A.; Shaw, G.M.; Hunter, E.; et al. Escape from autologous neutralizing antibodies in acute/early subtype C HIV-1 infection requires multiple pathways. PLoS Pathog. 2009, 5, e1000594. [Google Scholar] [CrossRef] [PubMed]
- Gray, E.S.; Moody, M.A.; Wibmer, C.K.; Chen, X.; Marshall, D.; Amos, J.; Moore, P.L.; Foulger, A.; Yu, J.-S.; Lambson, B.; et al. Isolation of a monoclonal antibody that targets the alpha-2 helix of gp120 and represents the initial autologous neutralizing-antibody response in an HIV-1 subtype C-infected individual. J. Virol. 2011, 85, 7719–7729. [Google Scholar] [CrossRef] [PubMed]
- van Gils, M.J.; Bunnik, E.M.; Boeser-Nunnink, B.D.; Burger, J.A.; Terlouw-Klein, M.; Verwer, N.; Schuitemaker, H. Longer V1V2 Region with Increased Number of Potential N-Linked Glycosylation Sites in the HIV-1 Envelope Glycoprotein Protects against HIV-Specific Neutralizing Antibodies. J. Virol. 2011, 85, 6986–6995. [Google Scholar] [CrossRef] [PubMed]
- Dacheux, L.; Moreau, A.; Ataman-Onal, Y.; Biron, F.; Verrier, B.; Barin, F. Evolutionary dynamics of the glycan shield of the human immunodeficiency virus envelope during natural infection and implications for exposure of the 2G12 epitope. J. Virol. 2004, 78, 12625–12637. [Google Scholar] [CrossRef] [PubMed]
- Sagar, M.; Wu, X.; Lee, S.; Overbaugh, J. Human Immunodeficiency Virus Type 1 V1-V2 Envelope Loop Sequences Expand and Add Glycosylation Sites over the Course of Infection, and These Modifications Affect Antibody Neutralization Sensitivity. J. Virol. 2006, 80, 9586–9598. [Google Scholar] [CrossRef]
- Mascola, J.R.; Montefiori, D.C. HIV-1: nature’s master of disguise. Nat. Med. 2003, 9, 393–394. [Google Scholar] [CrossRef] [PubMed]
- Simek, M.D.; Rida, W.; Priddy, F.H.; Pung, P.; Carrow, E.; Laufer, D.S.; Lehrman, J.K.; Boaz, M.; Tarragona-Fiol, T.; Miiro, G.; et al. Human Immunodeficiency Virus Type 1 Elite Neutralizers: Individuals with Broad and Potent Neutralizing Activity Identified by Using a High-Throughput Neutralization Assay together with an Analytical Selection Algorithm. J. Virol. 2009, 83, 7337–7348. [Google Scholar] [CrossRef]
- Euler, Z.; van den Kerkhof, T.L.G.M.; van Gils, M.J.; Burger, J.A.; Edo-Matas, D.; Phung, P.; Wrin, T.; Schuitemaker, H. Longitudinal analysis of early HIV-1-specific neutralizing activity in an elite neutralizer and in five patients who developed cross-reactive neutralizing activity. J. Virol. 2012, 86, 2045–2055. [Google Scholar] [CrossRef]
- Rusert, P.; Kouyos, R.D.; Kadelka, C.; Ebner, H.; Schanz, M.; Huber, M.; Braun, D.L.; Hozé, N.; Scherrer, A.; Magnus, C.; et al. Determinants of HIV-1 broadly neutralizing antibody induction. Nat. Med. 2016, 22, 1260–1267. [Google Scholar] [CrossRef]
- Braibant, M.; Agut, H.; Rouzioux, C.; Costagliola, D.; Autran, B.; Barin, F. Characteristics of the env genes of HIV type 1 quasispecies in long-term nonprogressors with broadly neutralizing antibodies. J. Acquir. Immune. Defic. Syndr. 2008, 47, 274–284. [Google Scholar] [CrossRef]
- Mikell, I.; Sather, D.N.; Kalams, S.A.; Altfeld, M.; Alter, G.; Stamatatos, L. Characteristics of the earliest cross-neutralizing antibody response to HIV-1. PLoS Pathog. 2011, 7, e1001251. [Google Scholar] [CrossRef]
- Derdeyn, C.A.; Moore, P.L.; Morris, L. Development of broadly neutralizing antibodies from autologous neutralizing antibody responses in HIV infection. Curr. Opin. HIV AIDS 2014, 9, 210–216. [Google Scholar] [CrossRef] [PubMed]
- Piantadosi, A.; Panteleeff, D.; Blish, C.A.; Baeten, J.M.; Jaoko, W.; McClelland, R.S.; Overbaugh, J. Breadth of neutralizing antibody response to human immunodeficiency virus type 1 is affected by factors early in infection but does not influence disease progression. J. Virol. 2009, 83, 10269–10274. [Google Scholar] [CrossRef] [PubMed]
- Chaillon, A.; Braibant, M.; Hué, S.; Bencharif, S.; Enard, D.; Moreau, A.; Samri, A.; Agut, H.; Barin, F. Human Immunodeficiency Virus Type-1 (HIV-1) Continues to Evolve in Presence of Broadly Neutralizing Antibodies More than Ten Years after Infection. PLoS ONE 2012, 7, e44163. [Google Scholar] [CrossRef]
- Sather, D.N.; Carbonetti, S.; Kehayia, J.; Kraft, Z.; Mikell, I.; Scheid, J.F.; Klein, F.; Stamatatos, L. Broadly neutralizing antibodies developed by an HIV-positive elite neutralizer exact a replication fitness cost on the contemporaneous virus. J. Virol. 2012, 86, 12676–12685. [Google Scholar] [CrossRef]
- Braibant, M.; Xie, J.; Samri, A.; Agut, H.; Autran, B.; Barin, F. Disease progression due to dual infection in an HLA-B57-positive asymptomatic long-term nonprogressor infected with a nef-defective HIV-1 strain. Virology 2010, 405, 81–92. [Google Scholar] [CrossRef]
- Kwong, P.D.; Mascola, J.R.; Nabel, G.J. Mining the B cell repertoire for broadly neutralizing monoclonal antibodies to HIV-1. Cell Host Microbe 2009, 6, 292–294. [Google Scholar] [CrossRef]
- Moir, S.; Malaspina, A.; Fauci, A.S. Prospects for an HIV vaccine: leading B cells down the right path. Nat. Struct. Mol. Biol. 2011, 18, 1317–1321. [Google Scholar] [CrossRef]
- Sok, D.; Burton, D.R. Recent progress in broadly neutralizing antibodies to HIV. Nat. Immunol. 2018, 19, 1179–1188. [Google Scholar] [CrossRef]
- Walker, L.M.; Burton, D.R. Passive immunotherapy of viral infections: “Super-antibodies” enter the fray. Nat. Rev. Immunol. 2018, 18, 297–308. [Google Scholar] [CrossRef]
- Moldt, B.; Rakasz, E.G.; Schultz, N.; Chan-Hui, P.Y.; Swiderek, K.; Weisgrau, K.L.; Piaskowski, S.M.; Bergman, Z.; Watkins, D.I.; Poignard, P.; et al. Highly potent HIV-specific antibody neutralization in vitro translates into effective protection against mucosal SHIV challenge in vivo. Proc. Natl. Acad. Sci. USA 2012, 109, 18921–18925. [Google Scholar] [CrossRef]
- Shingai, M.; Donau, O.K.; Plishka, R.J.; Buckler-White, A.; Mascola, J.R.; Nabel, G.J.; Nason, M.C.; Montefiori, D.; Moldt, B.; Poignard, P.; et al. Passive transfer of modest titers of potent and broadly neutralizing anti-HIV monoclonal antibodies block SHIV infection in macaques. J. Exp. Med. 2014, 211, 2061–2074. [Google Scholar] [CrossRef]
- Hessell, A.J.; Haigwood, N.L. Animal models in HIV-1 protection and therapy. Curr. Opin. HIV AIDS 2015, 10, 170–176. [Google Scholar] [CrossRef]
- Chun, T.-W.; Murray, D.; Justement, J.S.; Blazkova, J.; Hallahan, C.W.; Fankuchen, O.; Gittens, K.; Benko, E.; Kovacs, C.; Moir, S.; et al. Broadly neutralizing antibodies suppress HIV in the persistent viral reservoir. Proc. Natl. Acad. Sci. USA 2014, 111, 13151–13156. [Google Scholar] [CrossRef]
- Barin, F.; Braibant, M. HIV-1 antibodies in prevention of transmission. Curr. Opin. HIV AIDS 2019, 14, 273–278. [Google Scholar] [CrossRef]
- Julien, J.P.; Cupo, A.; Sok, D.; Stanfield, R.L.; Lyumkis, D.; Deller, M.C.; Klasse, P.J.; Burton, D.R.; Sanders, R.W.; Moore, J.P.; et al. Crystal Structure of a Soluble Cleaved HIV-1 Envelope Trimer. Science 2013, 342, 1477–1483. [Google Scholar] [CrossRef]
- Lyumkis, D.; Julien, J.P.; de Val, N.; Cupo, A.; Potter, C.S.; Klasse, P.J.; Burton, D.R.; Sanders, R.W.; Moore, J.P.; Carragher, B.; et al. Cryo-EM Structure of a Fully Glycosylated Soluble Cleaved HIV-1 Envelope Trimer. Science 2013, 342, 1484–1490. [Google Scholar] [CrossRef]
- Pancera, M.; Zhou, T.; Druz, A.; Georgiev, I.S.; Soto, C.; Gorman, J.; Huang, J.; Acharya, P.; Chuang, G.-Y.; Ofek, G.; et al. Structure and immune recognition of trimeric pre-fusion HIV-1 Env. Nature 2014, 514, 455–461. [Google Scholar] [CrossRef]
- Kong, R.; Xu, K.; Zhou, T.; Acharya, P.; Lemmin, T. Fusion peptide of HIV-1 as a site of vulnerability to neutralizing antibody. Science 2016, 352, 828–833. [Google Scholar] [CrossRef]
- Dingens, A.S.; Arenz, D.; Weight, H.; Overbaugh, J.; Bloom, J.D. An Antigenic Atlas of HIV-1 Escape from Broadly Neutralizing Antibodies Distinguishes Functional and Structural Epitopes. Immunity 2019, 50, 520–532. [Google Scholar] [CrossRef]
- Joseph, S.B.; Swanstrom, R.; Kashuba, A.D.M.; Cohen, M.S. Bottlenecks in HIV-1 transmission: insights from the study of founder viruses. Nat. Rev. Microbiol. 2015, 13, 414–425. [Google Scholar] [CrossRef]
- Kariuki, S.M.; Selhorst, P.; Ariën, K.K.; Dorfman, J.R. The HIV-1 transmission bottleneck. Retrovirology 2017, 14, 22. [Google Scholar] [CrossRef]
- Beretta, M.; Moreau, A.; Bouvin-Pley, M.; Essat, A.; Goujard, C.; Chaix, M.-L.; Hué, S.; Meyer, L.; Barin, F.; Braibant, M.; et al. Phenotypic properties of envelope glycoproteins of transmitted HIV-1 variants from patients belonging to transmission chains. AIDS 2018, 32, 1917–1926. [Google Scholar] [CrossRef]
- Hraber, P.; Seaman, M.S.; Bailer, R.T.; Mascola, J.R.; Montefiori, D.C.; Korber, B.T. Prevalence of broadly neutralizing antibody responses during chronic HIV-1 infection. AIDS 2014, 28, 163–169. [Google Scholar] [CrossRef]
- Bricault, C.A.; Yusim, K.; Seaman, M.S.; Yoon, H.; Theiler, J.; Giorgi, E.E.; Wagh, K.; Theiler, M.; Hraber, P.; Macke, J.P.; et al. HIV-1 Neutralizing Antibody Signatures and Application to Epitope-Targeted Vaccine Design. Cell Host Microbe 2019, 25, 59–72. [Google Scholar] [CrossRef]
- Doria-Rose, N.A.; Bhiman, J.N.; Roark, R.S.; Schramm, C.A.; Gorman, J.; Chuang, G.-Y.; Pancera, M.; Cale, E.M.; Ernandes, M.J.; Louder, M.K.; et al. New Member of the V1V2-Directed CAP256-VRC26 Lineage That Shows Increased Breadth and Exceptional Potency. J. Virol. 2016, 90, 76–91. [Google Scholar] [CrossRef]
- Chuang, G.-Y.; Acharya, P.; Schmidt, S.D.; Yang, Y.; Louder, M.K.; Zhou, T.; Kwon, Y.D.; Pancera, M.; Bailer, R.T.; Doria-Rose, N.A.; et al. Residue-level prediction of HIV-1 antibody epitopes based on neutralization of diverse viral strains. J. Virol. 2013, 87, 10047–10058. [Google Scholar] [CrossRef]
- Georgiev, I.S.; Doria-Rose, N.A.; Zhou, T.; Kwon, Y.D.; Staupe, R.P.; Moquin, S.; Chuang, G.-Y.; Louder, M.K.; Schmidt, S.D.; Altae-Tran, H.R.; et al. Delineating antibody recognition in polyclonal sera from patterns of HIV-1 isolate neutralization. Science 2013, 340, 751–756. [Google Scholar] [CrossRef]
- Kong, L.; Lee, J.H.; Doores, K.J.; Murin, C.D.; Julien, J.-P.; McBride, R.; Liu, Y.; Marozsan, A.; Cupo, A.; Klasse, P.J.; et al. Supersite of immune vulnerability on the glycosylated face of HIV-1 envelope glycoprotein gp120. Nat. Struct. Mol. Biol. 2013, 20, 796–803. [Google Scholar] [CrossRef]
- Rademeyer, C.; Korber, B.; Seaman, M.S.; Giorgi, E.E.; Thebus, R.; Robles, A.; Sheward, D.J.; Wagh, K.; Garrity, J.; Carey, B.R.; et al. Features of Recently Transmitted HIV-1 Clade C Viruses that Impact Antibody Recognition: Implications for Active and Passive Immunization. PLoS Pathog. 2016, 12, e1005742. [Google Scholar] [CrossRef]
- Stefic, K.; Bouvin-Pley, M.; Essat, A.; Visdeloup, C.; Moreau, A.; Goujard, C.; Chaix, M.-L.; Braibant, M.; Meyer, L.; Barin, F. Sensitivity to Broadly Neutralizing Antibodies of Recently Transmitted HIV-1 Clade CRF02_AG Viruses with a Focus on Evolution over Time. J. Virol. 2019, 93, e01492-18. [Google Scholar] [CrossRef]
- Bouvin-Pley, M.; Morgand, M.; Meyer, L.; Goujard, C.; Moreau, A.; Mouquet, H.; Nussenzweig, M.; Pace, C.; Ho, D.; Bjorkman, P.J.; et al. Drift of the HIV-1 Envelope Glycoprotein gp120 toward Increased Neutralization Resistance over the Course of the Epidemic: a Comprehensive Study Using the Most Potent and Broadly Neutralizing Monoclonal Antibodies. J. Virol. 2014, 88, 13910–13917. [Google Scholar] [CrossRef]
- Seaman, M.S.; Janes, H.; Hawkins, N.; Grandpre, L.E.; Devoy, C.; Giri, A.; Coffey, R.T.; Harris, L.; Wood, B.; Daniels, M.G.; et al. Tiered Categorization of a Diverse Panel of HIV-1 Env Pseudoviruses for Assessment of Neutralizing Antibodies. J. Virol. 2010, 84, 1439–1452. [Google Scholar] [CrossRef]
- Korber, B.; Gnanakaran, S. The implications of patterns in HIV diversity for neutralizing antibody induction and susceptibility. Curr. Opin. HIV AIDS 2009, 4, 408–417. [Google Scholar] [CrossRef]
- Ferguson, A.L.; Falkowska, E.; Walker, L.M.; Seaman, M.S.; Burton, D.R.; Chakraborty, A.K. Computational prediction of broadly neutralizing HIV-1 antibody epitopes from neutralization activity data. PLoS ONE 2013, 8, e80562. [Google Scholar] [CrossRef]
- Evans, M.C.; Phung, P.; Paquet, A.C.; Parikh, A.; Petropoulos, C.J.; Wrin, T.; Haddad, M. Predicting HIV-1 broadly neutralizing antibody epitope networks using neutralization titers and a novel computational method. BMC Bioinformatics 2014, 15, 77. [Google Scholar] [CrossRef]
- Crispin, M.; Doores, K.J. Targeting host-derived glycans on enveloped viruses for antibody-based vaccine design. Curr. Opin. Virol. 2015, 11, 63–69. [Google Scholar] [CrossRef]
- Bai, H.; Li, Y.; Michael, N.L.; Robb, M.L.; Rolland, M. The breadth of HIV-1 neutralizing antibodies depends on the conservation of key sites in their epitopes. PLoS Comp. Biol. 2019, 15, e1007056. [Google Scholar] [CrossRef]
- Burton, D.R.; Hangartner, L. Broadly Neutralizing Antibodies to HIV and Their Role in Vaccine Design. Annu. Rev. Immunol. 2016, 34, 635–659. [Google Scholar] [CrossRef]
- Dingens, A.S.; Haddox, H.K.; Overbaugh, J.; Bloom, J.D. Comprehensive Mapping of HIV-1 Escape from a Broadly Neutralizing Antibody. Cell Host Microbe 2017, 21, 777–787.e4. [Google Scholar] [CrossRef]
- Doria-Rose, N.A.; Altae-Tran, H.R.; Roark, R.S.; Schmidt, S.D.; Sutton, M.S.; Louder, M.K.; Chuang, G.-Y.; Bailer, R.T.; Cortez, V.; Kong, R.; et al. Mapping Polyclonal HIV-1 Antibody Responses via Next-Generation Neutralization Fingerprinting. PLoS Pathog. 2017, 13, e1006148. [Google Scholar] [CrossRef]
- Bunnik, E.M.; Euler, Z.; Welkers, M.R.A.; Boeser-Nunnink, B.D.M.; Grijsen, M.L.; Prins, J.M.; Schuitemaker, H. Adaptation of HIV-1 envelope gp120 to humoral immunity at a population level. Nat. Med. 2010, 16, 995–997. [Google Scholar] [CrossRef]
- Bouvin-Pley, M.; Morgand, M.; Moreau, A.; Jestin, P.; Simonnet, C.; Tran, L.; Goujard, C.; Meyer, L.; Barin, F.; Braibant, M. Evidence for a Continuous Drift of the HIV-1 Species towards Higher Resistance to Neutralizing Antibodies over the Course of the Epidemic. PLoS Pathog. 2013, 9, e1003477. [Google Scholar] [CrossRef]
- Kawashima, Y.; Pfafferott, K.; Frater, J.; Matthews, P.; Payne, R.; Addo, M.; Gatanaga, H.; Fujiwara, M.; Hachiya, A.; Koizumi, H.; et al. Adaptation of HIV-1 to human leukocyte antigen class I. Nature 2009, 458, 641–645. [Google Scholar] [CrossRef]
- Gali, Y.; Berkhout, B.; Vanham, G.; Bakker, M.; Back, N.K.T.; Ariën, K.K. Survey of the temporal changes in HIV-1 replicative fitness in the Amsterdam Cohort. Virology 2006, 364, 140–146. [Google Scholar] [CrossRef]
- Herbeck, J.T.; Müller, V.; Maust, B.S.; Ledergerber, B.; Torti, C.; Di Giambenedetto, S.; Gras, L.; Günthard, H.F.; Jacobson, L.P.; Mullins, J.I.; et al. Is the virulence of HIV changing? A meta-analysis of trends in prognostic markers of HIV disease progression and transmission. AIDS 2012, 26, 193–205. [Google Scholar] [CrossRef]
- Bouvin-Pley, M.; Beretta, M.; Moreau, A.; Roch, E.; Essat, A.; Goujard, C.; Chaix, M.-L.; Moiré, N.; Martin, L.; Meyer, L.; et al. Evolution of the Envelope Glycoprotein of HIV-1 Clade B toward Higher Infectious Properties over the Course of the Epidemic. J. Virol. 2019, 93, e01171-18. [Google Scholar] [CrossRef]
- Haynes, B.F.; Bradley, T. Broadly Neutralizing Antibodies and the Development of Vaccines. JAMA 2015, 313, 2419–2420. [Google Scholar] [CrossRef]
- Subbaraman, H.; Schanz, M.; Trkola, A. Broadly neutralizing antibodies: What is needed to move from a rare event in HIV-1 infection to vaccine efficacy? Retrovirology 2018, 15, 635. [Google Scholar] [CrossRef]
- Manrique, A.; Rusert, P.; Joos, B.; Fischer, M.; Kuster, H.; Leemann, C.; Niederöst, B.; Weber, R.; Stiegler, G.; Katinger, H.; et al. In vivo and in vitro escape from neutralizing antibodies 2G12, 2F5, and 4E10. J. Virol. 2007, 81, 8793–8808. [Google Scholar] [CrossRef]
- Walker, L.M.; Simek, M.D.; Priddy, F.; Gach, J.S.; Wagner, D.; Zwick, M.B.; Phogat, S.K.; Poignard, P.; Burton, D.R. A Limited Number of Antibody Specificities Mediate Broad and Potent Serum Neutralization in Selected HIV-1 Infected Individuals. PLoS Pathog. 2010, 6, e1001028. [Google Scholar] [CrossRef]
- Kouyos, R.D.; Rusert, P.; Kadelka, C.; Huber, M.; Marzel, A.; Ebner, H.; Schanz, M.; Liechti, T.; Friedrich, N.; Braun, D.L.; et al. Tracing HIV-1 strains that imprint broadly neutralizing antibody responses. Nature 2018, 561, 406–410. [Google Scholar] [CrossRef]
- Walker, L.M.; Huber, M.; Doores, K.J.; Falkowska, E.; Pejchal, R.; Julien, J.-P.; Wang, S.-K.; Ramos, A.; Chan-Hui, P.-Y.; Moyle, M.; et al. Broad neutralization coverage of HIV by multiple highly potent antibodies. Nature 2011, 477, 466–470. [Google Scholar] [CrossRef]
- Doria-Rose, N.A.; Georgiev, I.; O’Dell, S.; Chuang, G.Y.; Staupe, R.P.; McLellan, J.S.; Gorman, J.; Pancera, M.; Bonsignori, M.; Haynes, B.F.; et al. A Short Segment of the HIV-1 gp120 V1/V2 Region Is a Major Determinant of Resistance to V1/V2 Neutralizing Antibodies. J. Virol. 2012, 86, 8319–8323. [Google Scholar] [CrossRef]
- Moore, P.L.; Gray, E.S.; Wibmer, C.K.; Bhiman, J.N.; Nonyane, M.; Sheward, D.J.; Hermanus, T.; Bajimaya, S.; Tumba, N.L.; Abrahams, M.-R.; et al. Evolution of an HIV glycan-dependent broadly neutralizing antibody epitope through immune escape. Nat. Med. 2012, 18, 1688–1692. [Google Scholar] [CrossRef]
- Caskey, M.; Klein, F.; Lorenzi, J.C.C.; Seaman, M.S.; West, A.P.; Buckley, N.; Kremer, G.; Nogueira, L.; Braunschweig, M.; Scheid, J.F.; et al. Viraemia suppressed in HIV-1-infected humans by broadly neutralizing antibody 3BNC117. Nature 2015, 522, 487–491. [Google Scholar] [CrossRef]
- Lynch, R.M.; Boritz, E.; Coates, E.E.; DeZure, A.; Madden, P.; Costner, P.; Enama, M.E.; Plummer, S.; Holman, L.; Hendel, C.S.; et al. Virologic effects of broadly neutralizing antibody VRC01 administration during chronic HIV-1 infection. Sci. Transl. Med. 2015, 7, 319ra206. [Google Scholar] [CrossRef]
- Schoofs, T.; Klein, F.; Braunschweig, M.; Kreider, E.F.; Feldmann, A.; Nogueira, L.; Oliveira, T.; Lorenzi, J.C.C.; Parrish, E.H.; Learn, G.H.; et al. HIV-1 therapy with monoclonal antibody 3BNC117 elicits host immune responses against HIV-1. Science 2016, 352, 997–1001. [Google Scholar] [CrossRef]
- Caskey, M.; Schoofs, T.; Gruell, H.; Settler, A.; Karagounis, T.; Kreider, E.F.; Murrell, B.; Pfeifer, N.; Nogueira, L.; Oliveira, T.Y.; et al. Antibody 10-1074 suppresses viremia in HIV-1-infected individuals. Nat. Med. 2017, 23, 185–191. [Google Scholar] [CrossRef]
- Julg, B.; Sok, D.; Schmidt, S.D.; Abbink, P.; Newman, R.M.; Broge, T.; Linde, C.; Nkolola, J.; Le, K.; Su, D.; et al. Protective Efficacy of Broadly Neutralizing Antibodies with Incomplete Neutralization Activity against Simian-Human Immunodeficiency Virus in Rhesus Monkeys. J. Virol. 2017, 91, e01187-17. [Google Scholar] [CrossRef]
- Cohen, Y.Z.; Caskey, M. Broadly neutralizing antibodies for treatment and prevention of HIV-1 infection. Curr. Opin. HIV AIDS 2018, 13, 366–373. [Google Scholar] [CrossRef]
- Xu, L.; Pegu, A.; Rao, E.; Doria-Rose, N.; Beninga, J.; McKee, K.; Lord, D.M.; Wei, R.R.; Deng, G.; Louder, M.; et al. Trispecific broadly neutralizing HIV antibodies mediate potent SHIV protection in macaques. Science 2017, 358, 85–90. [Google Scholar] [CrossRef]
- Wagh, K.; Seaman, M.S.; Zingg, M.; Fitzsimons, T.; Barouch, D.H.; Burton, D.R.; Connors, M.; Ho, D.D.; Mascola, J.R.; Nussenzweig, M.C.; et al. Potential of conventional & bispecific broadly neutralizing antibodies for prevention of HIV-1 subtype A, C & D infections. PLoS Pathog. 2018, 14, e1006860. [Google Scholar]
- deCamp, A.; Hraber, P.; Bailer, R.T.; Seaman, M.S.; Ochsenbauer, C.; Kappes, J.; Gottardo, R.; Edlefsen, P.; Self, S.; Tang, H.; et al. Global panel of HIV-1 Env reference strains for standardized assessments of vaccine-elicited neutralizing antibodies. J. Virol. 2014, 88, 2489–2507. [Google Scholar] [CrossRef]
- Mouquet, H. Antibody B cell responses in HIV-1 infection. Trends Immunol. 2014, 35, 549–561. [Google Scholar] [CrossRef]
- Moore, P.L. The Neutralizing Antibody Response to the HIV-1 Env Protein. Curr. HIV Res. 2018, 16, 21–28. [Google Scholar] [CrossRef]
- Alter, G.; Barouch, D. Immune Correlate-Guided HIV Vaccine Design. Cell Host Microbe 2018, 24, 25–33. [Google Scholar] [CrossRef]
- Kwong, P.D.; Mascola, J.R. HIV-1 Vaccines Based on Antibody Identification, B Cell Ontogeny, and Epitope Structure. Immunity 2018, 48, 855–871. [Google Scholar] [CrossRef]
- Rudicell, R.S.; Kwon, Y.D.; Ko, S.-Y.; Pegu, A.; Louder, M.K.; Georgiev, I.S.; Wu, X.; Zhu, J.; Boyington, J.C.; Chen, X.; et al. Enhanced potency of a broadly neutralizing HIV-1 antibody in vitro improves protection against lentiviral infection in vivo. J. Virol. 2014, 88, 12669–12682. [Google Scholar] [CrossRef]
- Huang, J.; Kang, B.H.; Ishida, E.; Zhou, T.; Griesman, T.; Sheng, Z.; Wu, F.; Doria-Rose, N.A.; Zhang, B.; McKee, K.; et al. Identification of a CD4-Binding-Site Antibody to HIV that Evolved Near-Pan Neutralization Breadth. Immunity 2016, 45, 1108–1121. [Google Scholar] [CrossRef]
- Bonsignori, M.; Montefiori, D.C.; Wu, X.; Chen, X.; Hwang, K.-K.; Tsao, C.-Y.; Kozink, D.M.; Parks, R.J.; Tomaras, G.D.; Crump, J.A.; et al. Two distinct broadly neutralizing antibody specificities of different clonal lineages in a single HIV-1-infected donor: implications for vaccine design. J. Virol. 2012, 86, 4688–4692. [Google Scholar] [CrossRef] [PubMed]
- Ditse, Z.; Muenchhoff, M.; Adland, E.; Jooste, P.; Goulder, P.; Moore, P.L.; Morris, L. HIV-1 Subtype C-Infected Children with Exceptional Neutralization Breadth Exhibit Polyclonal Responses Targeting Known Epitopes. J. Virol. 2018, 92, e00878-18. [Google Scholar] [CrossRef] [PubMed]
- Fischer, W.; Perkins, S.; Theiler, J.; Bhattacharya, T.; Yusim, K.; Funkhouser, R.; Kuiken, C.; Haynes, B.; Letvin, N.L.; Walker, B.D.; et al. Polyvalent vaccines for optimal coverage of potential T-cell epitopes in global HIV-1 variants. Nat. Med. 2007, 13, 100–106. [Google Scholar] [CrossRef] [PubMed]
- Barouch, D.H.; Stephenson, K.E.; Borducchi, E.N.; Smith, K.; Stanley, K.; McNally, A.G.; Liu, J.; Abbink, P.; Maxfield, L.F.; Seaman, M.S.; et al. Protective Efficacy of a Global HIV-1 Mosaic Vaccine against Heterologous SHIV Challenges in Rhesus Monkeys. CELL 2013, 155, 531–539. [Google Scholar] [CrossRef] [PubMed]
- Bricault, C.A.; Kovacs, J.M.; Badamchi-Zadeh, A.; McKee, K.; Shields, J.L.; Gunn, B.M.; Neubauer, G.H.; Ghantous, F.; Jennings, J.; Gillis, L.; et al. Neutralizing Antibody Responses following Long-Term Vaccination with HIV-1 Env gp140 in Guinea Pigs. J. Virol. 2018, 92, e00369-18. [Google Scholar] [CrossRef] [PubMed]
- Barouch, D.H.; Tomaka, F.L.; Wegmann, F.; Stieh, D.J.; Alter, G.; Robb, M.L.; Michael, N.L.; Peter, L.; Nkolola, J.P.; Borducchi, E.N.; et al. Evaluation of a mosaic HIV-1 vaccine in a multicentre, randomised, double-blind, placebo-controlled, phase 1/2a clinical trial (APPROACH) and in rhesus monkeys (NHP 13-19). Lancet 2018, 392, 232–243. [Google Scholar] [CrossRef]
- Liao, H.-X.; Tsao, C.-Y.; Alam, S.M.; Muldoon, M.; Vandergrift, N.; Ma, B.-J.; Lu, X.; Sutherland, L.L.; Scearce, R.M.; Bowman, C.; et al. Antigenicity and immunogenicity of transmitted/founder, consensus, and chronic envelope glycoproteins of human immunodeficiency virus type 1. J. Virol. 2013, 87, 4185–4201. [Google Scholar] [CrossRef] [PubMed]
- Hulot, S.L.; Korber, B.; Giorgi, E.E.; Vandergrift, N.; Saunders, K.O.; Balachandran, H.; Mach, L.V.; Lifton, M.A.; Pantaleo, G.; Tartaglia, J.; et al. Comparison of Immunogenicity in Rhesus Macaques of Transmitted-Founder, HIV-1 Group M Consensus, and Trivalent Mosaic Envelope Vaccines Formulated as a DNA Prime, NYVAC, and Envelope Protein Boost. J. Virol. 2015, 89, 6462–6480. [Google Scholar] [CrossRef] [PubMed]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Stefic, K.; Bouvin-Pley, M.; Braibant, M.; Barin, F. Impact of HIV-1 Diversity on Its Sensitivity to Neutralization. Vaccines 2019, 7, 74. https://doi.org/10.3390/vaccines7030074
Stefic K, Bouvin-Pley M, Braibant M, Barin F. Impact of HIV-1 Diversity on Its Sensitivity to Neutralization. Vaccines. 2019; 7(3):74. https://doi.org/10.3390/vaccines7030074
Chicago/Turabian StyleStefic, Karl, Mélanie Bouvin-Pley, Martine Braibant, and Francis Barin. 2019. "Impact of HIV-1 Diversity on Its Sensitivity to Neutralization" Vaccines 7, no. 3: 74. https://doi.org/10.3390/vaccines7030074
APA StyleStefic, K., Bouvin-Pley, M., Braibant, M., & Barin, F. (2019). Impact of HIV-1 Diversity on Its Sensitivity to Neutralization. Vaccines, 7(3), 74. https://doi.org/10.3390/vaccines7030074




