Impact of Pneumococcal Vaccination on Nasopharyngeal Carriage of Streptococcus pneumoniae and Microbiota Profiles in Preschool Children in South East Poland
Abstract
1. Introduction
2. Materials and Methods
2.1. Patients
2.2. Laboratory Procedures
2.3. Real-Time PCR Analysis
2.4. Statistical Analysis
3. Results
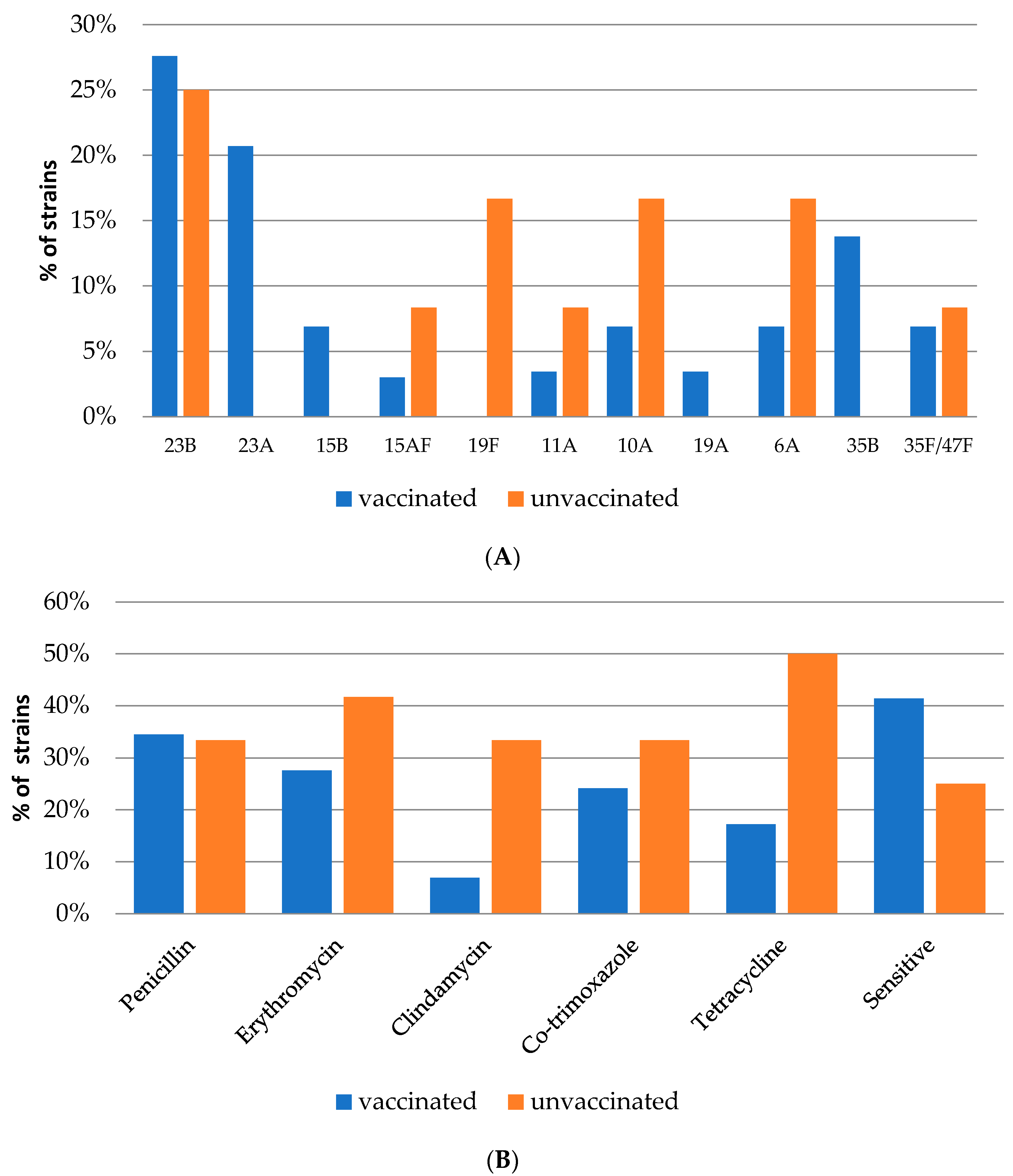
3.1. Serotype Distribution and Vaccine Coverage
3.2. Antimicrobial Susceptibility
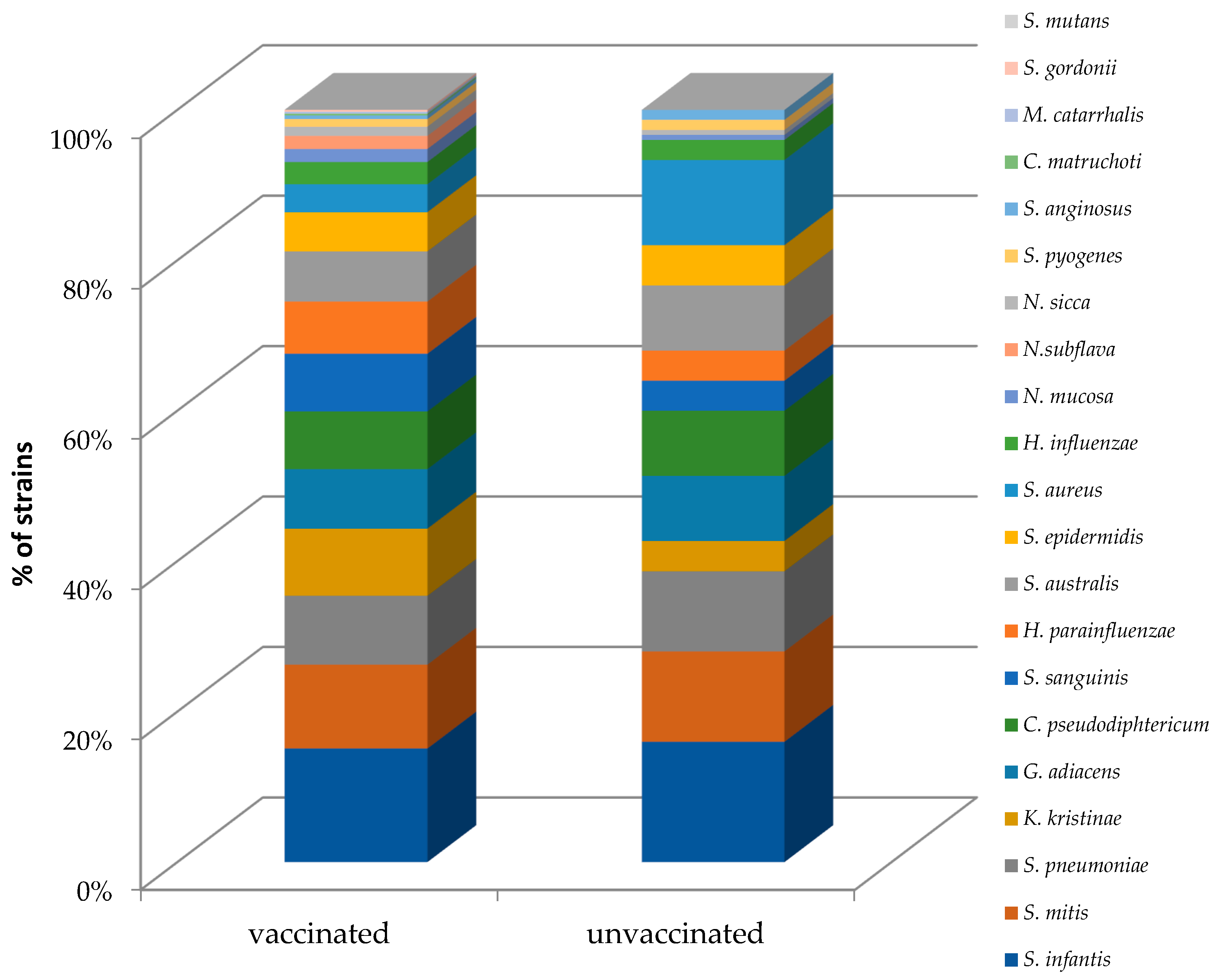
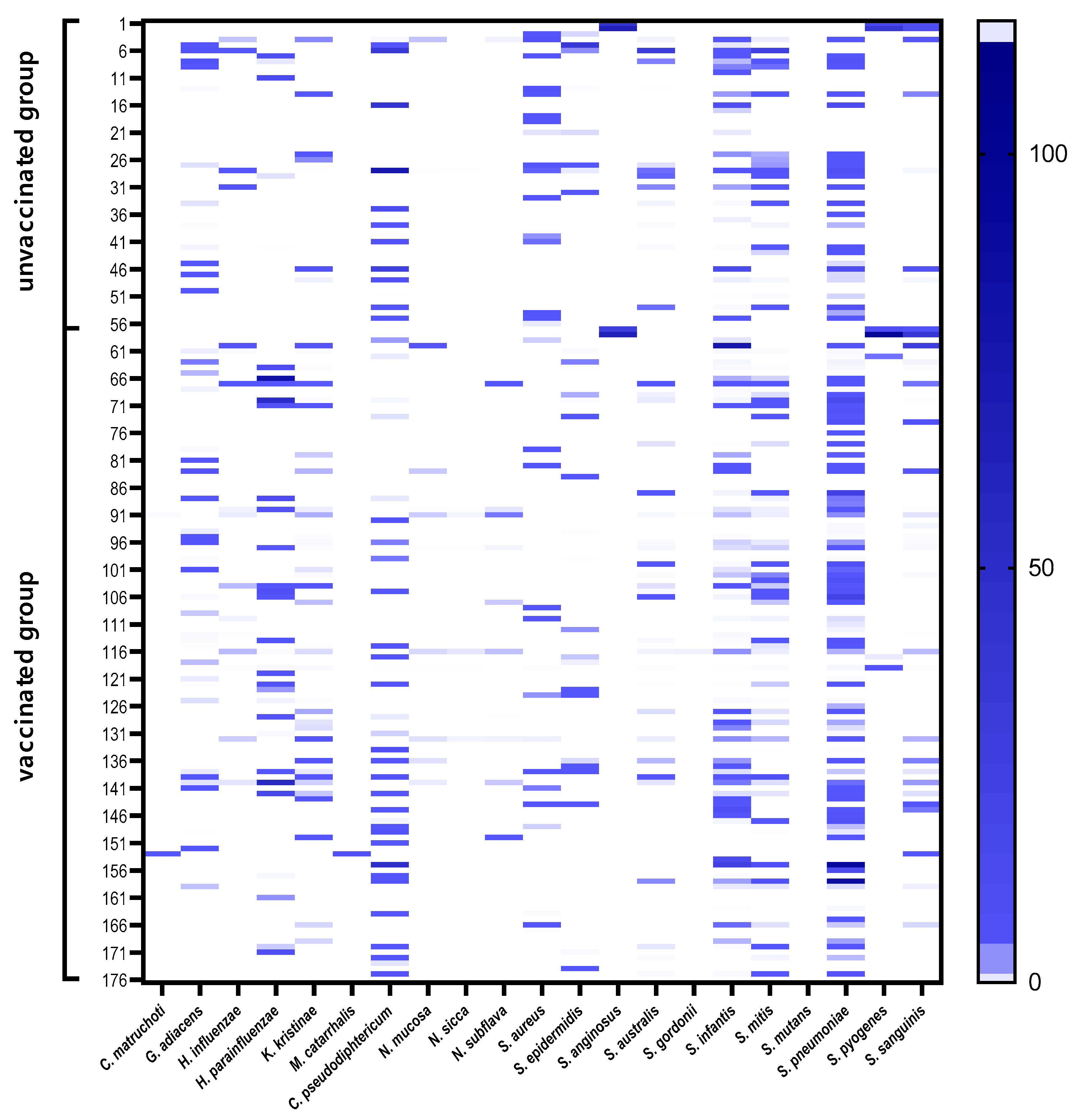
3.3. Nasopharyngeal Microbiota by Real-Time PCR
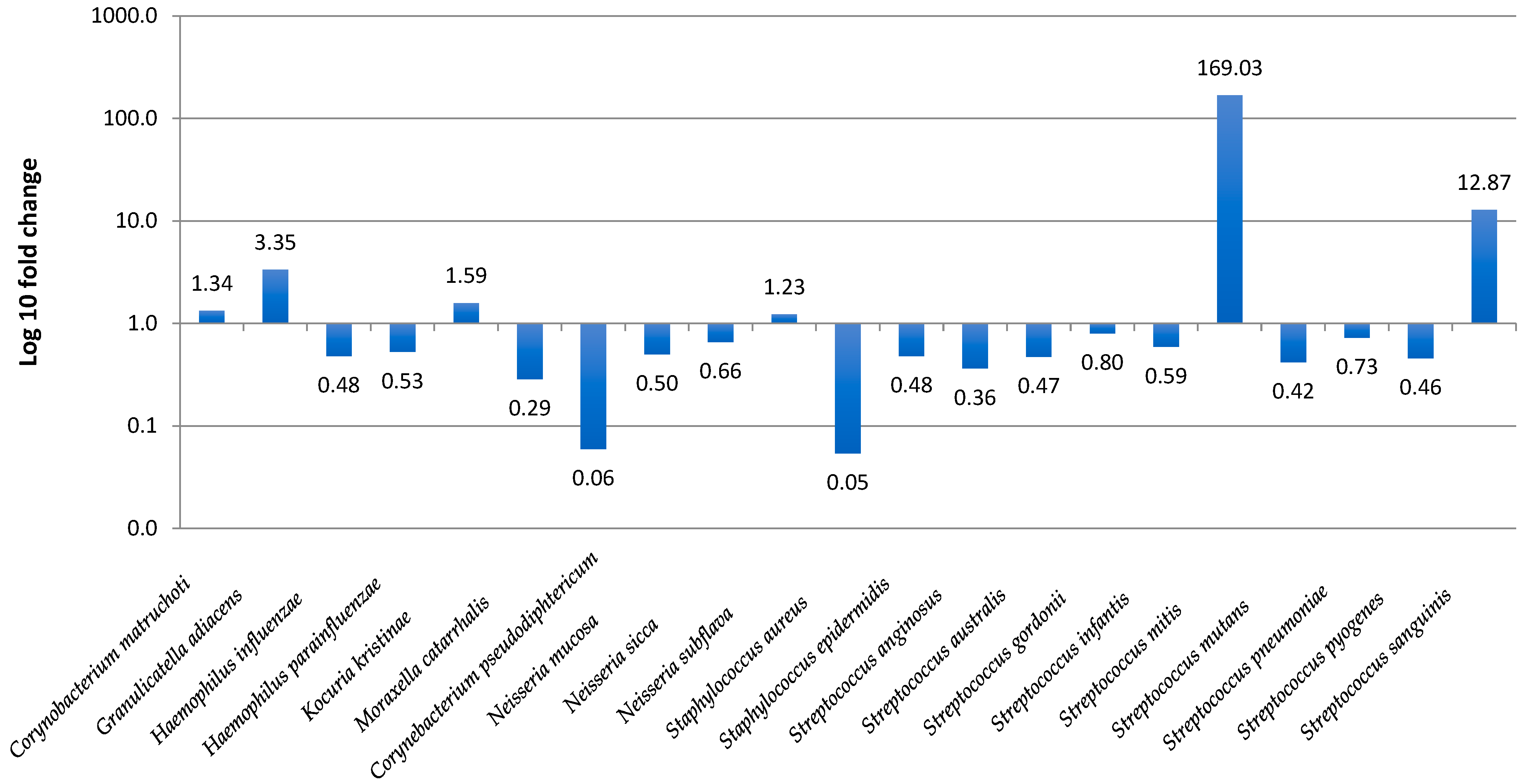
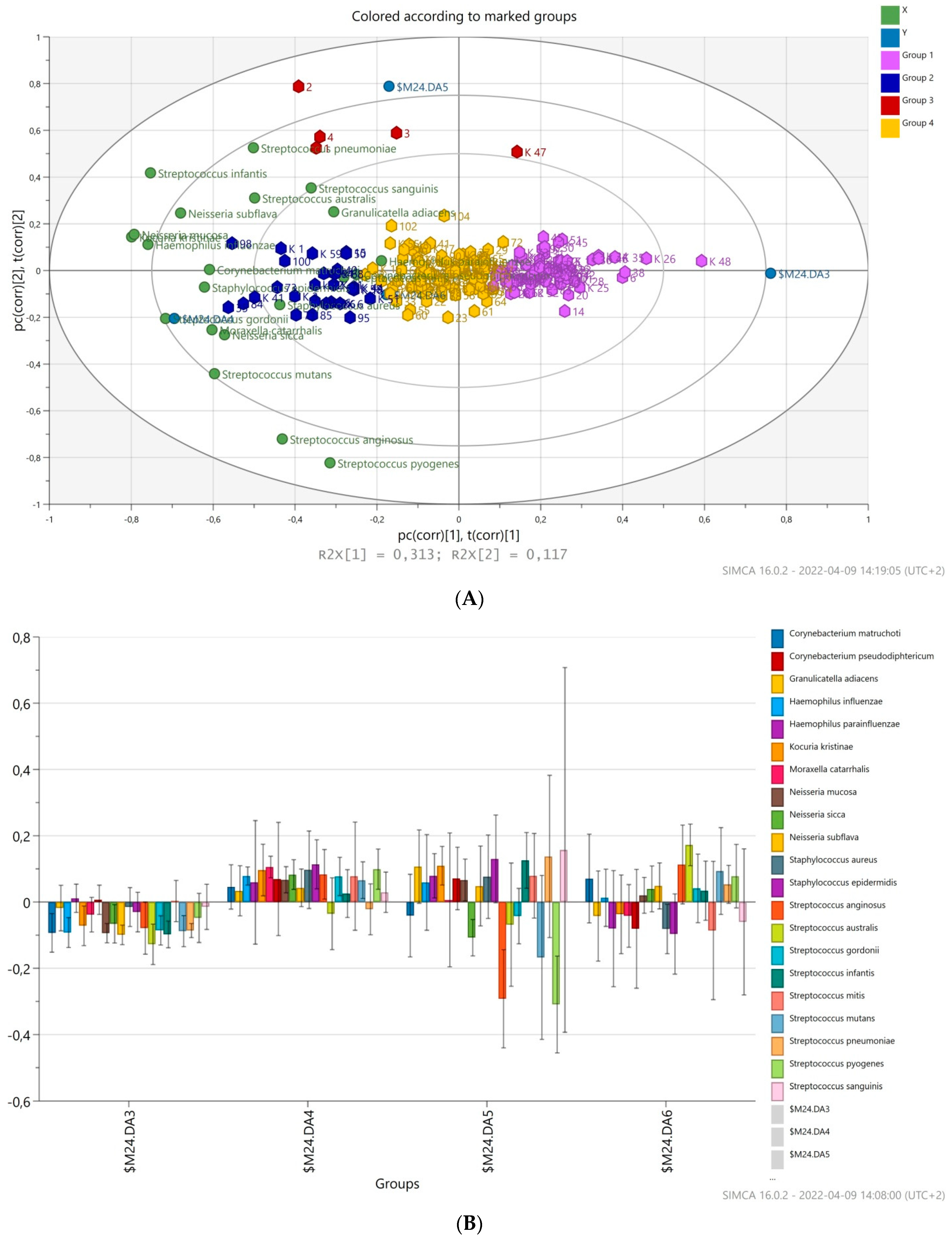
3.4. Prediction of Bacterial Communities Profile
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Givon-Lavi, N.; Fraser, D.; Porat, N.; Dagan, R. Spread of Streptococcus pneumoniae and Antibiotic-Resistant S. Pneumoniae from Day-Care Center Attendees to Their Younger Siblings. J. Infect. Dis. 2002, 186, 1608–1614. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Weiser, J.N.; Ferreira, D.M.; Paton, J.C. Streptococcus pneumoniae: Transmission, colonization and invasion. Nat. Rev. Microbiol. 2018, 16, 355–367. [Google Scholar] [CrossRef]
- Brugger, S.D.; Bomar, L.; Lemon, K.P. Commensal–Pathogen Interactions along the Human Nasal Passages. PLoS Pathog. 2016, 12, e1005633. [Google Scholar] [CrossRef] [PubMed]
- Ghaffar, F.; Friedland, I.R.; McCracken, G.H. Dynamics of nasopharyngeal colonization by Streptococcus pneumoniae. Pediatr. Infect. Dis. J. 1999, 18, 638–646. [Google Scholar] [CrossRef] [PubMed]
- Bogaert, D.; de Groot, R.; Hermans, P.W.M. Streptococcus pneumoniae colonisation: The key to pneumococcal disease. Lancet Infect. Dis. 2004, 4, 144–154. [Google Scholar] [CrossRef]
- de Lencastre, H.; Tomasz, A. From ecological reservoir to disease: The nasopharynx, day-care centres and drug-resistant clones of Streptococcus pneumoniae. J. Antimicrob. Chemother. 2002, 50 (Suppl. S2), 75–81. [Google Scholar] [CrossRef]
- Lynch, J.P.; Zhanel, G.G. Streptococcus pneumoniae: Epidemiology and risk factors, evolution of antimicrobial resistance, and impact of vaccines. Curr. Opin. Pulm. Med. 2010, 16, 217–225. [Google Scholar] [CrossRef]
- Desmet, S.; Wouters, I.; Van Heirstraeten, L.; Beutels, P.; Van Damme, P.; Malhotra-Kumar, S.; Maes, P.; Verhaegen, J.; Peetermans, W.E.; Lagrou, K.; et al. In-depth analysis of pneumococcal serotypes in Belgian children (2015–2018): Diversity, invasive disease potential, and antimicrobial susceptibility in carriage and disease. Vaccine 2021, 39, 372–379. [Google Scholar] [CrossRef]
- Klugman, K.P. Efficacy of pneumococcal conjugate vaccines and their effect on carriage and antimicrobial resistance. Lancet Infect. Dis. 2001, 1, 85–91. [Google Scholar] [CrossRef]
- O’Brien, K.L.; Wolfson, L.J.; Watt, J.P.; Henkle, E.; Deloria-Knoll, M.; McCall, N.; Lee, E.; Mulholland, K.; Levine, O.S.; Cherian, T.; et al. Burden of disease caused by Streptococcus pneumoniae in children younger than 5 years: Global estimates. Lancet 2009, 374, 893–902. [Google Scholar] [CrossRef]
- Moore, M.R.; Link-Gelles, R.; Schaffner, W.; Lynfield, R.; Lexau, C.; Bennett, N.M.; Petit, S.; Zansky, S.M.; Harrison, L.H.; Reingold, A.; et al. Effect of use of 13-valent pneumococcal conjugate vaccine in children on invasive pneumococcal disease in children and adults in the USA: Analysis of multisite, population-based surveillance. Lancet Infect. Dis. 2015, 15, 301–309. [Google Scholar] [CrossRef]
- Alfayate Miguélez, S.; Yague Guirao, G.; Menasalvas Ruíz, A.I.; Sanchez-Solís, M.; Domenech Lucas, M.; González Camacho, F.; Ortíz Romero, M.M.; Espejo García, P.; Guerrero Gómez, C.; Iofrío de Arce, A.; et al. Impact of Pneumococcal Vaccination in the Nasopharyngeal Carriage of Streptococcus pneumoniae in Healthy Children of the Murcia Region in Spain. Vaccines 2020, 9, 14. [Google Scholar] [CrossRef] [PubMed]
- Ceyhan, M.; Karadag-Oncel, E.; Hascelik, G.; Ustundag, G.; Gurbuz, V.; Samlioglu, P.; Yilmaz, N.; Ozsurekci, Y.; Yilmaz, E.; Aykac, K.; et al. Nasopharyngeal carriage of Streptococcus pneumoniae in healthy children aged less than five years. Vaccine 2021, 39, 2041–2047. [Google Scholar] [CrossRef] [PubMed]
- Lindstrand, A.; Galanis, I.; Darenberg, J.; Morfeldt, E.; Naucler, P.; Blennow, M.; Alfven, T.; Henriques-Normark, B.; Örtqvist, Å. Unaltered pneumococcal carriage prevalence due to expansion of non-vaccine types of low invasive potential 8 years after vaccine introduction in Stockholm, Sweden. Vaccine 2016, 34, 4565–4571. [Google Scholar] [CrossRef]
- Spijkerman, J.; Prevaes, S.M.P.J.; Van Gils, E.J.M.; Veenhoven, R.H.; Bruin, J.P.; Bogaert, D.; Wijmenga-Monsuur, A.J.; van den Dobbelsteen, G.P.; Sanders, E.A.M. Long-Term Effects of Pneumococcal Conjugate Vaccine on Nasopharyngeal Carriage of S. pneumoniae, S. aureus, H. influenzae and M. catarrhalis. PLoS ONE 2012, 7, e39730. [Google Scholar] [CrossRef]
- Navne, J.E.; Koch, A.; Slotved, H.-C.; Andersson, M.; Melbye, M.; Ladefoged, K.; Børresen, M. Effect of the 13-valent pneumococcal conjugate vaccine on nasopharyngeal carriage by respiratory pathogens among Greenlandic children. Int. J. Circumpolar Health 2017, 76, 1309504. [Google Scholar] [CrossRef] [PubMed]
- 2017 Poland Introduced 10-Valent Pneumococcal Vaccine in the Immunization Schedule. Szczepienia.Info EN. Available online: http://szczepieniainfo.testa.com.pl/en/stories/introduction_of_pcv_2016/ (accessed on 22 August 2018).
- Richter, S.S.; Heilmann, K.P.; Dohrn, C.L.; Riahi, F.; Diekema, D.J.; Doern, G.V. Evaluation of Pneumococcal Serotyping by Multiplex PCR and Quellung Reactions. J. Clin. Microbiol. 2013, 51, 4193–4195. [Google Scholar] [CrossRef]
- Llull, D.; López, R.; García, E. Characteristic Signatures of the lytA Gene Provide a Basis for Rapid and Reliable Diagnosis of Streptococcus pneumoniae Infections. J. Clin. Microbiol. 2006, 44, 1250–1256. [Google Scholar] [CrossRef]
- Schmittgen, T.D.; Livak, K.J. Analysing real-time PCR data by comparative Ct method. Nat. Protoc. 2008, 3, 1101–1108. [Google Scholar] [CrossRef]
- Patrzalek, M.; Gorynski, P.; Albrecht, P. Indirect population impact of universal PCV7 vaccination of children in a 2 + 1 schedule on the incidence of pneumonia morbidity in Kielce, Poland. Eur. J. Clin. Microbiol. Infect. Dis. 2012, 31, 3023–3028. [Google Scholar] [CrossRef]
- Patrzałek, M.; Kotowska, M.; Goryński, P.; Albrecht, P. Indirect effects of a 7 year PCV7/PCV13 mass vaccination program in children on the incidence of pneumonia among adults: A comparative study based on two Polish cities. Curr. Med. Res. Opin. 2016, 32, 397–403. [Google Scholar] [CrossRef]
- Korona-Glowniak, I.; Malm, A. Characteristics of Streptococcus pneumoniae Strains Colonizing Upper Respiratory Tract of Healthy Preschool Children in Poland. Sci. World J. 2012, 2012, 732901. [Google Scholar] [CrossRef] [PubMed]
- Korona-Glowniak, I.; Niedzielski, A.; Malm, A. Upper respiratory colonization by Streptococcus pneumoniae in healthy pre-school children in south-east Poland. Int. J. Pediatr. Otorhinolaryngol. 2011, 75, 1529–1534. [Google Scholar] [CrossRef]
- Niedzielski, A.; Korona-Glowniak, I.; Malm, A. High prevalence of Streptococcus pneumoniae in adenoids and nasopharynx in preschool children with recurrent upper respiratory tract infections in Poland—Distribution of serotypes and drug resistance patterns. Med. Sci. Monit. 2013, 19, 54–60. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Korona-Glowniak, I.; Niedzielski, A.; Kosikowska, U.; Grzegorczyk, A.; Malm, A. Nasopharyngeal vs. adenoid cultures in children undergoing adenoidectomy: Prevalence of bacterial pathogens, their interactions and risk factors. Epidemiol. Infect. 2014, 143, 821–830. [Google Scholar] [CrossRef] [PubMed]
- van Hoek, A.J.; Sheppard, C.L.; Andrews, N.J.; Waight, P.A.; Slack, M.P.E.; Harrison, T.G.; Ladhani, S.N.; Miller, E. Pneumococcal carriage in children and adults two years after introduction of the thirteen valent pneumococcal conjugate vaccine in England. Vaccine 2014, 32, 4349–4355. [Google Scholar] [CrossRef] [PubMed]
- Løvlie, A.; Vestrheim, D.F.; Aaberge, I.S.; Steens, A. Changes in pneumococcal carriage prevalence and factors associated with carriage in Norwegian children, four years after introduction of PCV13. BMC Infect. Dis. 2020, 20, 29. [Google Scholar] [CrossRef] [PubMed]
- Danino, D.; Givon-Lavi, N.; Ben-Shimol, S.; Greenberg, D.; Dagan, R. Understanding the Evolution of Antibiotic-nonsusceptible Pneumococcal Nasopharyngeal Colonization Following Pneumococcal Conjugate Vaccine Implementation in Young Children. Clin. Infect. Dis. 2019, 69, 648–656. [Google Scholar] [CrossRef]
- Torres, A.; Bonanni, P.; Hryniewicz, W.; Moutschen, M.; Reinert, R.R.; Welte, T. Pneumococcal vaccination: What have we learnt so far and what can we expect in the future? Eur. J. Clin. Microbiol. Infect. Dis. 2015, 34, 19–31. [Google Scholar] [CrossRef]
- Gladstone, R.A.; Jefferies, J.M.; Tocheva, A.S.; Beard, K.R.; Garley, D.; Chong, W.W.; Bentley, S.D.; Faust, S.N.; Clarke, S.C. Five winters of pneumococcal serotype replacement in UK carriage following PCV introduction. Vaccine 2015, 33, 2015–2021. [Google Scholar] [CrossRef]
- Grivea, I.N.; Priftis, K.N.; Giotas, A.; Kotzia, D.; Tsantouli, A.G.; Douros, K.; Michoula, A.N.; Syrogiannopoulos, G.A. Dynamics of pneumococcal carriage among day-care center attendees during the transition from the 7-valent to the higher-valent pneumococcal conjugate vaccines in Greece. Vaccine 2014, 32, 6513–6520. [Google Scholar] [CrossRef] [PubMed]
- Desai, A.P.; Sharma, D.; Crispell, E.K.; Baughman, W.; Thomas, S.; Tunali, A.; Sherwood, L.; Zmitrovich, A.; Jerris, R.; Satola, S.W.; et al. Decline in Pneumococcal Nasopharyngeal Carriage of Vaccine Serotypes After the Introduction of the 13-Valent Pneumococcal Conjugate Vaccine in Children in Atlanta, Georgia. Pediatr. Infect. Dis. J. 2015, 34, 1168–1174. [Google Scholar] [CrossRef] [PubMed]
- Obolski, U.; Lourenço, J.; Thompson, C.; Thompson, R.; Gori, A.; Gupta, S. Vaccination can drive an increase in frequencies of antibiotic resistance among nonvaccine serotypes of Streptococcus pneumoniae. Proc. Natl. Acad. Sci. USA 2018, 115, 3102–3107. [Google Scholar] [CrossRef] [PubMed]
- Yahiaoui, R.Y.; Bootsma, H.J.; den Heijer, C.D.J.; Pluister, G.N.; Paget, W.J.; Spreeuwenberg, P.; Trzcinski, K.; Stobberingh, E.E. Distribution of serotypes and patterns of antimicrobial resistance among commensal Streptococcus pneumoniae in nine European countries. BMC Infect. Dis. 2018, 18, 440. [Google Scholar] [CrossRef]
- Farrell, D.J.; Klugman, K.P.; Pichichero, M. Increased Antimicrobial Resistance Among Nonvaccine Serotypes of Streptococcus pneumoniae in the Pediatric Population After the Introduction of 7-Valent Pneumococcal Vaccine in the United States. Pediatr. Infect. Dis. J. 2007, 26, 123–128. [Google Scholar] [CrossRef]
- Leibovitz, E. The effect of vaccination on Streptococcus pneumoniae resistance. Curr. Infect. Dis. Rep. 2008, 10, 182–191. [Google Scholar] [CrossRef]
- Korona-Glowniak, I.; Maj, M.; Siwiec, R.; Niedzielski, A.; Malm, A. Molecular Epidemiology of Streptococcus pneumoniae Isolates from Children with Recurrent Upper Respiratory Tract Infections. PLoS ONE 2016, 11, e0158909. [Google Scholar] [CrossRef]
- Korona-Glowniak, I.; Zychowski, P.; Siwiec, R.; Mazur, E.; Niedzielska, G.; Malm, A. Resistant Streptococcus pneumoniae strains in children with acute otitis media– high risk of persistent colonization after treatment. BMC Infect. Dis. 2018, 18, 478. [Google Scholar] [CrossRef]
- Wouters, I.; Desmet, S.; Van Heirstraeten, L.; Blaizot, S.; Verhaegen, J.; Van Damme, P.; Malhotra-Kumar, S.; Theeten, H.; NPcarriage Study Group. Follow-up of serotype distribution and antimicrobial susceptibility of Streptococcus pneumoniae in child carriage after a PCV13-to-PCV10 vaccine switch in Belgium. Vaccine 2019, 37, 1080–1086. [Google Scholar] [CrossRef]
- Assefa, A.; Gelaw, B.; Shiferaw, Y.; Tigabu, Z. Nasopharyngeal Carriage and Antimicrobial Susceptibility Pattern of Streptococcus Pneumoniae among Pediatric Outpatients at Gondar University Hospital, North West Ethiopia. Pediatr. Neonatol. 2013, 54, 315–321. [Google Scholar] [CrossRef]
- Harboe, Z.B.; Slotved, H.-C.; Konradsen, H.B.; Kaltoft, M.S. A Pneumococcal Carriage Study in Danish Pre-school Children before the Introduction of Pneumococcal Conjugate Vaccination. Open Microbiol. J. 2012, 6, 40–44. [Google Scholar] [CrossRef] [PubMed]
- Jensen, V.V.S.; Furberg, A.-S.; Slotved, H.-C.; Bazhukova, T.; Haldorsen, B.; Caugant, D.A.; Sundsfjord, A.; Valentiner-Branth, P.; Simonsen, G.S. Epidemiological and molecular characterization of Streptococcus pneumoniae carriage strains in pre-school children in Arkhangelsk, northern European Russia, prior to the introduction of conjugate pneumococcal vaccines. BMC Infect. Dis. 2020, 20, 279. [Google Scholar] [CrossRef]
- Vestrheim, D.F.; Høiby, E.A.; Aaberge, I.S.; Caugant, D.A. Phenotypic and Genotypic Characterization of Streptococcus pneumoniae Strains Colonizing Children Attending Day-Care Centers in Norway. J. Clin. Microbiol. 2008, 46, 2508–2518. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Biesbroek, G.; Wang, X.; Keijser, B.J.F.; Eijkemans, R.M.J.; Trzciński, K.; Rots, N.Y.; Veenhoven, R.H.; Sanders, E.A.M.; Bogaert, D. Seven-Valent Pneumococcal Conjugate Vaccine and Nasopharyngeal Microbiota in Healthy Children. Emerg. Infect. Dis. 2014, 20, 201–210. [Google Scholar] [CrossRef] [PubMed]
- Mika, M.; Maurer, J.; Korten, I.; Allemann, A.; Aebi, S.; Brugger, S.D.; Qi, W.; Frey, U.; Latzin, P.; Hilty, M. Influence of the pneumococcal conjugate vaccines on the temporal variation of pneumococcal carriage and the nasal microbiota in healthy infants: A longitudinal analysis of a case–control study. Microbiome 2017, 5, 85. [Google Scholar] [CrossRef]
- Feazel, L.M.; Santorico, S.A.; Robertson, C.E.; Bashraheil, M.; Scott, J.A.G.; Frank, D.N.; Hammitt, L.L. Effects of Vaccination with 10-Valent Pneumococcal Non-Typeable Haemophilus influenza Protein D Conjugate Vaccine (PHiD-CV) on the Nasopharyngeal Microbiome of Kenyan Toddlers. PLoS ONE 2015, 10, e0128064. [Google Scholar] [CrossRef]
- Hilty, M.; Qi, W.; Brugger, S.D.; Frei, L.; Agyeman, P.; Frey, P.M.; Aebi, S.; Mühlemann, K. Nasopharyngeal Microbiota in Infants With Acute Otitis Media. J. Infect. Dis. 2012, 205, 1048–1055. [Google Scholar] [CrossRef] [PubMed]
- Pettigrew, M.M.; Gent, J.F.; Revai, K.; Patel, J.A.; Chonmaitree, T. Microbial Interactions during Upper Respiratory Tract Infections. Emerg. Infect. Dis. 2008, 14, 1584–1591. [Google Scholar] [CrossRef]
- Van Gils, E.J.M.; Hak, E.; Veenhoven, R.H.; Rodenburg, G.D.; Bogaert, D.; Bruin, J.P.; Van Alphen, L.; Sanders, E.A.M. Effect of Seven-Valent Pneumococcal Conjugate Vaccine on Staphylococcus aureus Colonisation in a Randomised Controlled Trial. PLoS ONE 2011, 6, e20229. [Google Scholar] [CrossRef]
| Parameters | Sp Colonized (n = 41) | Sp Uncolonized (n = 135) | OR (95%CI) | p Value | |
|---|---|---|---|---|---|
| Age (years) | 1–2 | 6 (14.6%) | 19 (14.1%) | 2.0 (0.7–6.5) | 0.22 |
| 3–4 | 25 (61.0%) | 50 (37.0%) | 3.3 (1.4–7.5) | 0.0038 | |
| 5–6 | 10 (24.4%) | 66 (48.9%) | referent | ||
| Sex | Female | 16 (39.0%) | 63 (46.7%) | 0.7 (0.4–1.5) | 0.47 |
| male | 25 (61.0%) | 72 (53.3%) | |||
| Siblings | No | 15 (36.6%) | 45 (33.3%) | Referent | |
| 1 | 15 (36.6%) | 72 (53.3%) | 0.6 (0.3–1.4) | 0.30 | |
| >2 | 11 (26.8%) | 18 (13.3%) | 1.8 (0.7–4.7) | 0.22 | |
| Passive smoking | 3 (7.3%) | 19 (14.1%) | 0.5 (0.1–1.7) | 0.42 | |
| Place of residence | Rural | 12 (29.3%) | 36 (26.7%) | 1.1 (0.5–2.5) | 0.84 |
| urban | 29 (70.7%) | 99 (73.3%) | |||
| DCC/orphanage attendance | 34 (82.9%) | 118 (87.4%) | 0.7 (0.3–1.8) | 0.45 | |
| Antibiotic therapy | AM/AMC | 4 (9.8%) | 28 (20.7%) | Referent | |
| Macrolides | 1 (2.4%) | 9 (6.7%) | 0.3 (0.07–1.4) | 0.14 | |
| Co-trimoxazole | 1 (2.4%) | 2 (1.5%) | 0.2 (0.02–2.5) | 0.35 | |
| Cephalosporins | 5 (12.2%) | 11 (8.1%) | 1.1 (0.08–15.2) | 1.0 | |
| Number of antibiotic therapy | 0 | 28 (68.3%) | 85 (63.0%) | Referent | |
| 1 | 9 (22.0%) | 22 (16.3%) | 1.2 (0.5–3.0) | 0.65 | |
| >2 | 3 (7.3%) | 26 (19.3%) | 0.4 (0.1–1.2) | 0.13 | |
| RTIs | Pharyngitis | 11 (26.8%) | 35 (25.9%) | Referent | |
| Otitis media | 1 (2.4%) | 24 (17.8%) | 0.3 (0.04–2.6) | 0.46 | |
| Sinusitis | 0 (0) | 13 (9.6%) | 0.3 (0.02–5.0) | 0.35 | |
| Laryngitis | 3 (7.3%) | 9 (6.7%) | 2.6 (0.6–11.0) | 0.19 | |
| Number of URTIs | 0 | 12 (29.3%) | 36 (26.7%) | Referent | |
| 1 | 9 (22.0%) | 28 (20.7%) | 0.96 (0.4–2.6) | 1.0 | |
| >2 | 19 (46.3) | 65 (48.1%) | 0.9 (0.4–2.0) | 0.83 | |
| Hospitalization | 10 (24.4%) | 25 (18.5%) | 1.4 (0.6–3.3) | 0.50 | |
| Anti-pneumococcal vaccination | total | 29 (70.7%) | 91 (67.4%) | 1.2 (0.5–2.5) | 0.85 |
| PCV 10 | 10 (24.4%) | 37 (27.4%) | 0.9 (0.4–2.3) | 1.0 | |
| PCV13 | 15 (36.6%) | 51 (37.8%) | |||
| No data | 4 (9.8%) | 3 (2.2%) |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kielbik, K.; Pietras, A.; Jablonska, J.; Bakiera, A.; Borek, A.; Niedzielska, G.; Grzegorczyk, M.; Grywalska, E.; Korona-Glowniak, I. Impact of Pneumococcal Vaccination on Nasopharyngeal Carriage of Streptococcus pneumoniae and Microbiota Profiles in Preschool Children in South East Poland. Vaccines 2022, 10, 791. https://doi.org/10.3390/vaccines10050791
Kielbik K, Pietras A, Jablonska J, Bakiera A, Borek A, Niedzielska G, Grzegorczyk M, Grywalska E, Korona-Glowniak I. Impact of Pneumococcal Vaccination on Nasopharyngeal Carriage of Streptococcus pneumoniae and Microbiota Profiles in Preschool Children in South East Poland. Vaccines. 2022; 10(5):791. https://doi.org/10.3390/vaccines10050791
Chicago/Turabian StyleKielbik, Karolina, Aleksandra Pietras, Joanna Jablonska, Adrian Bakiera, Anna Borek, Grazyna Niedzielska, Michal Grzegorczyk, Ewelina Grywalska, and Izabela Korona-Glowniak. 2022. "Impact of Pneumococcal Vaccination on Nasopharyngeal Carriage of Streptococcus pneumoniae and Microbiota Profiles in Preschool Children in South East Poland" Vaccines 10, no. 5: 791. https://doi.org/10.3390/vaccines10050791
APA StyleKielbik, K., Pietras, A., Jablonska, J., Bakiera, A., Borek, A., Niedzielska, G., Grzegorczyk, M., Grywalska, E., & Korona-Glowniak, I. (2022). Impact of Pneumococcal Vaccination on Nasopharyngeal Carriage of Streptococcus pneumoniae and Microbiota Profiles in Preschool Children in South East Poland. Vaccines, 10(5), 791. https://doi.org/10.3390/vaccines10050791








