Assessing the Transmissibility of the New SARS-CoV-2 Variants: From Delta to Omicron
Abstract
:1. Introduction
2. Materials and Methods
2.1. Experimental Design
2.2. Estimation of Number of Variants
2.3. NPIs and Vaccination
2.4. Bayesian Model and Phylo-Dynamics Analysis
3. Results
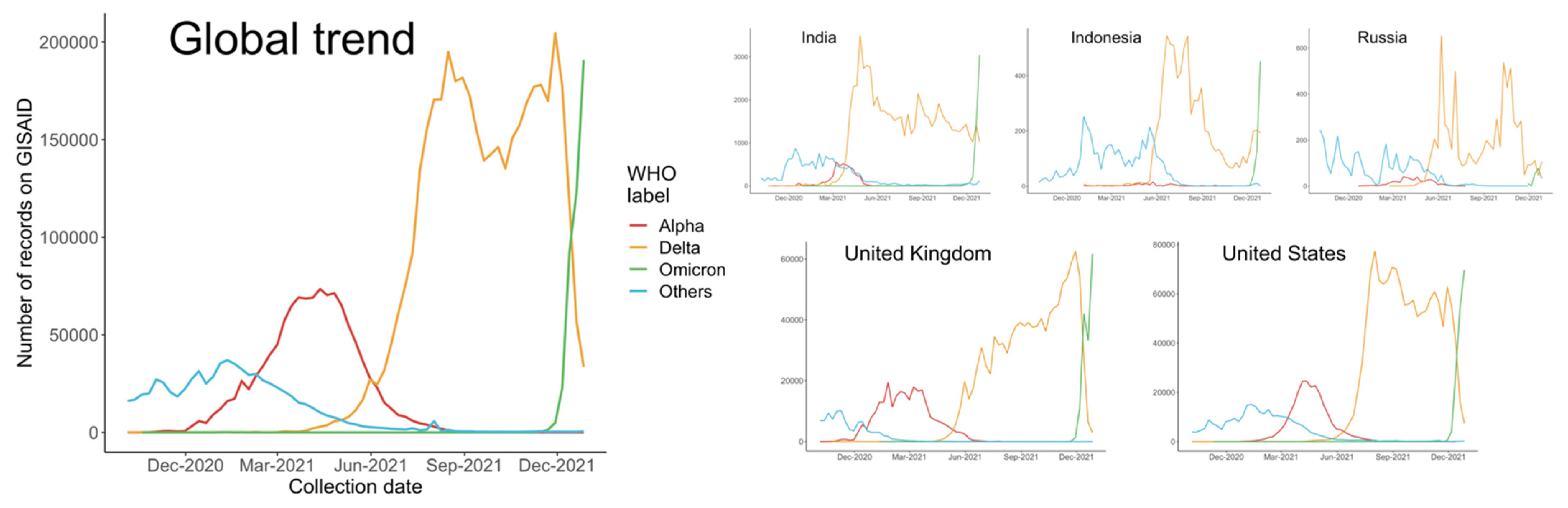
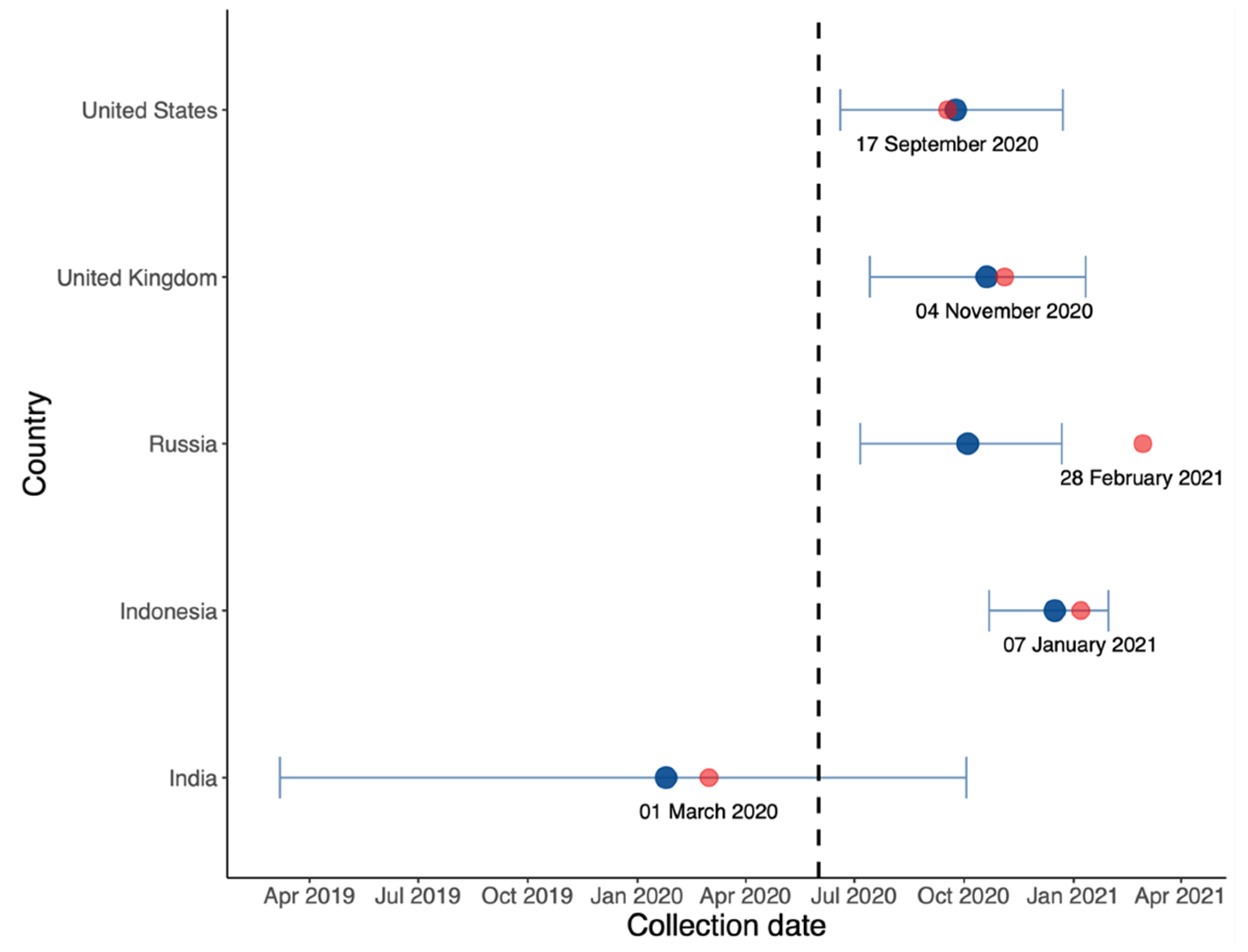
3.1. Global Trends and Major Countries
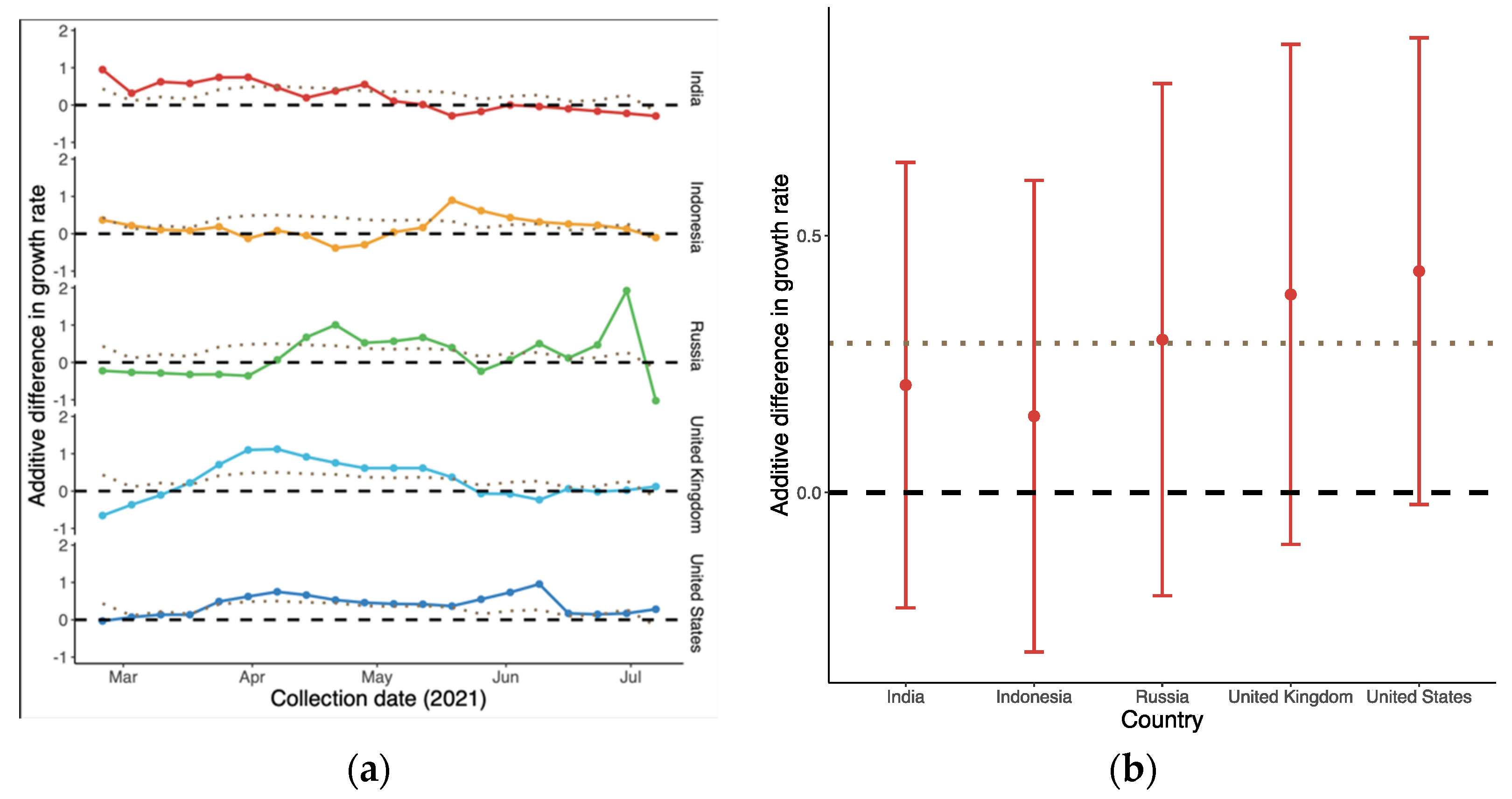
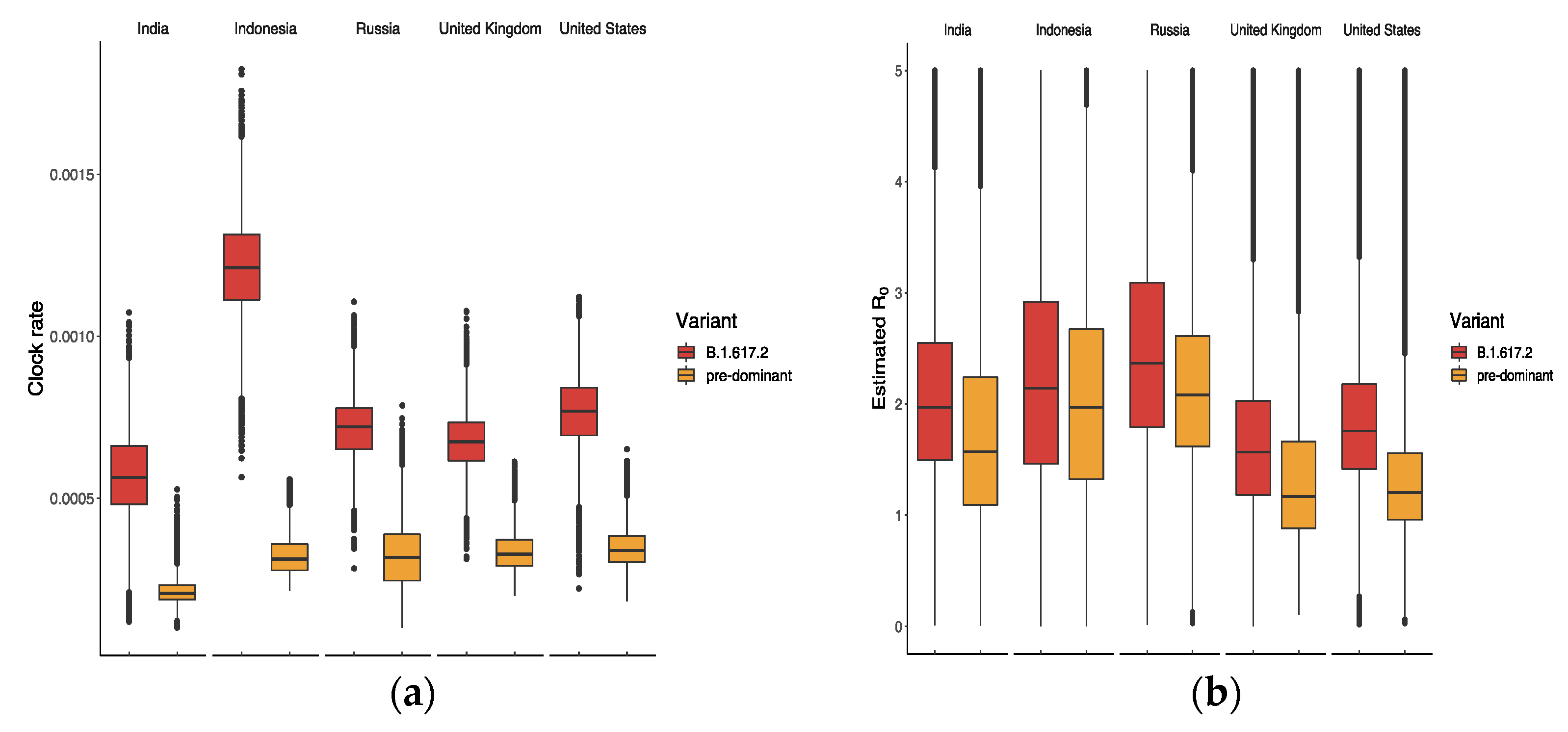
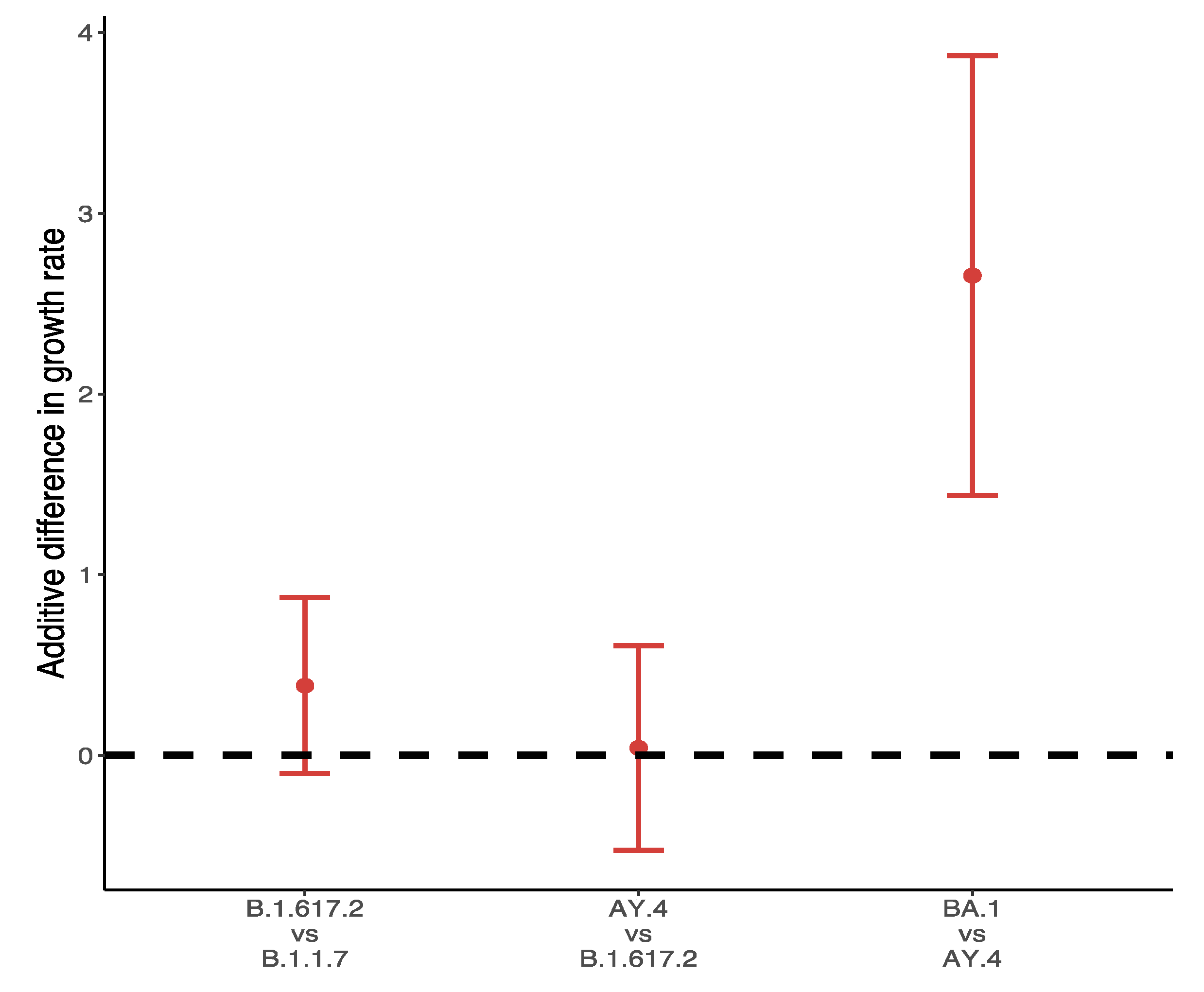
3.2. Relative Transmission Advantage of B.1.617.2 Compared to Predominant Variants
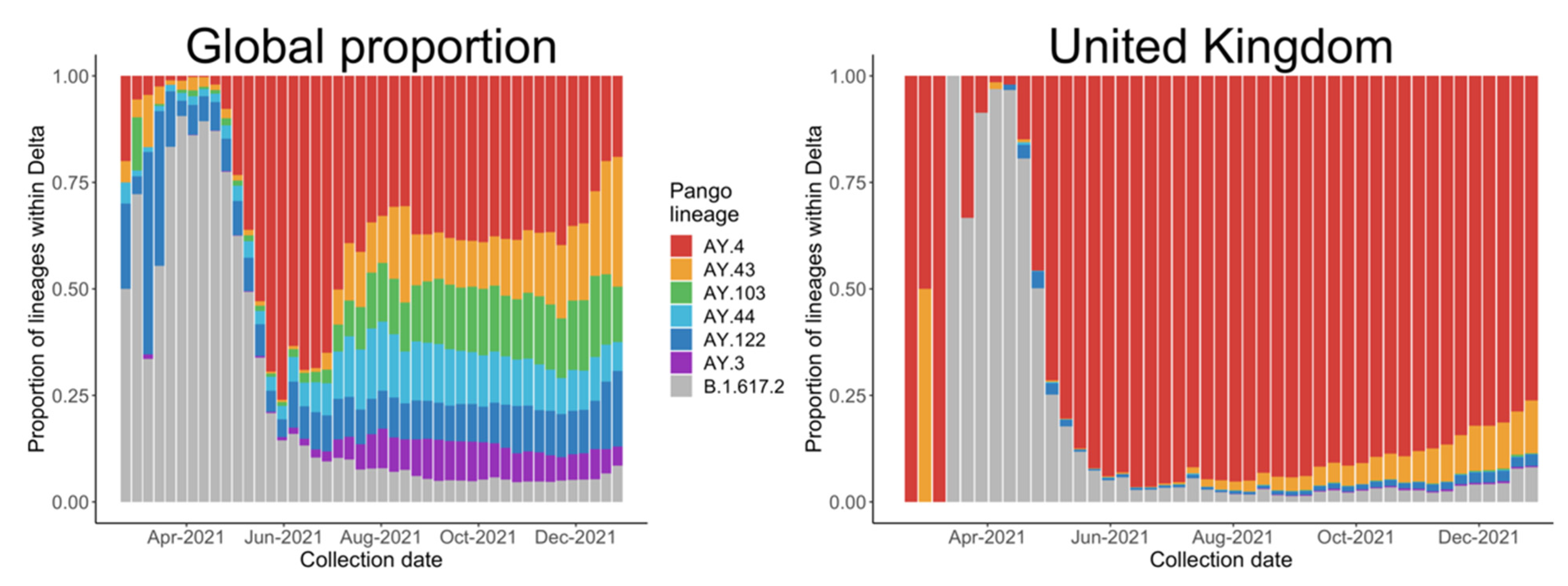
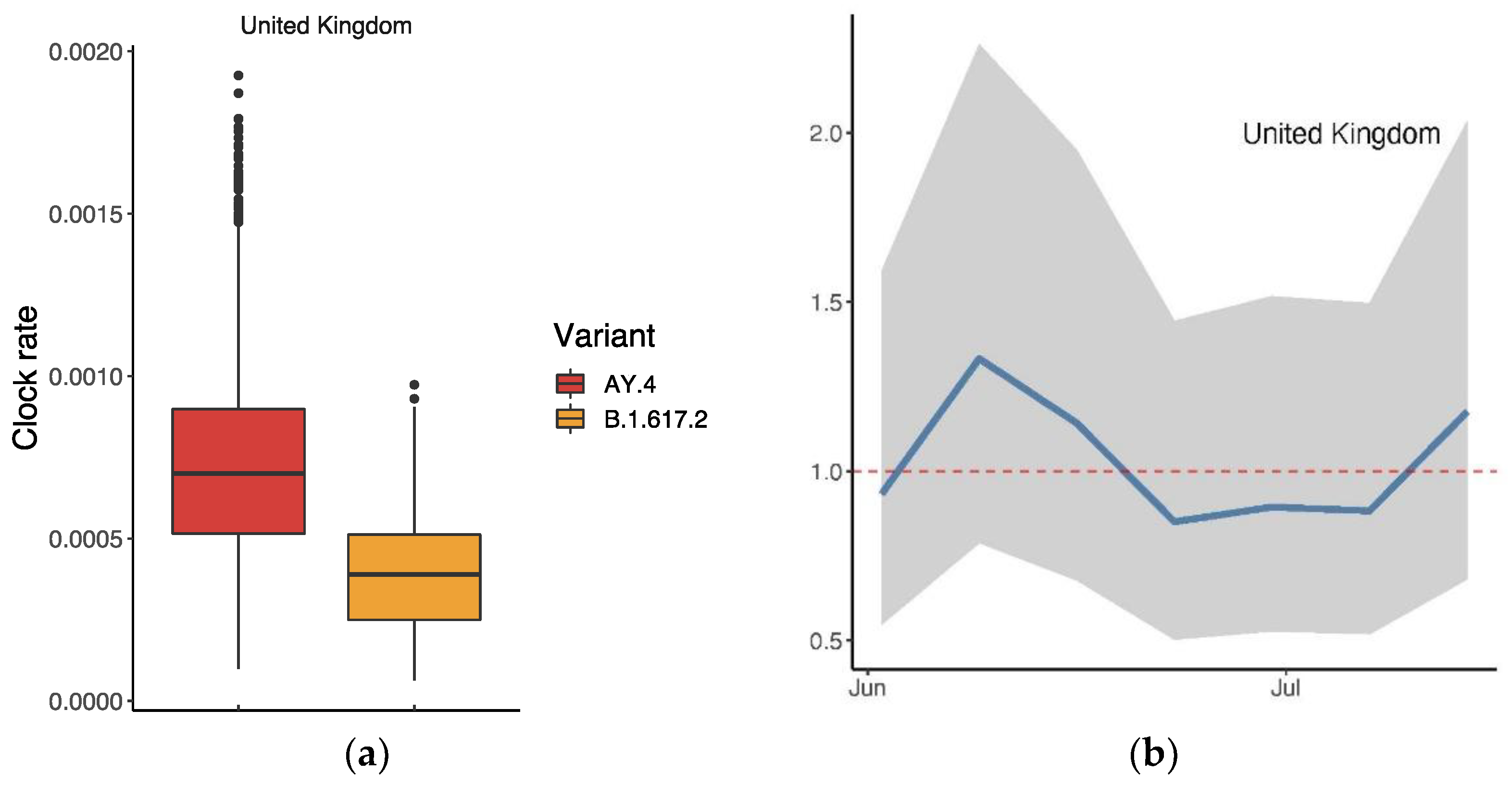
3.3. Relative Transmission Advantage of AY.4 compared to B.1.617.2
3.4. Differences in Growth Rates between Alpha, Delta and Omicron
3.5. Effect of NPIs and Vaccination
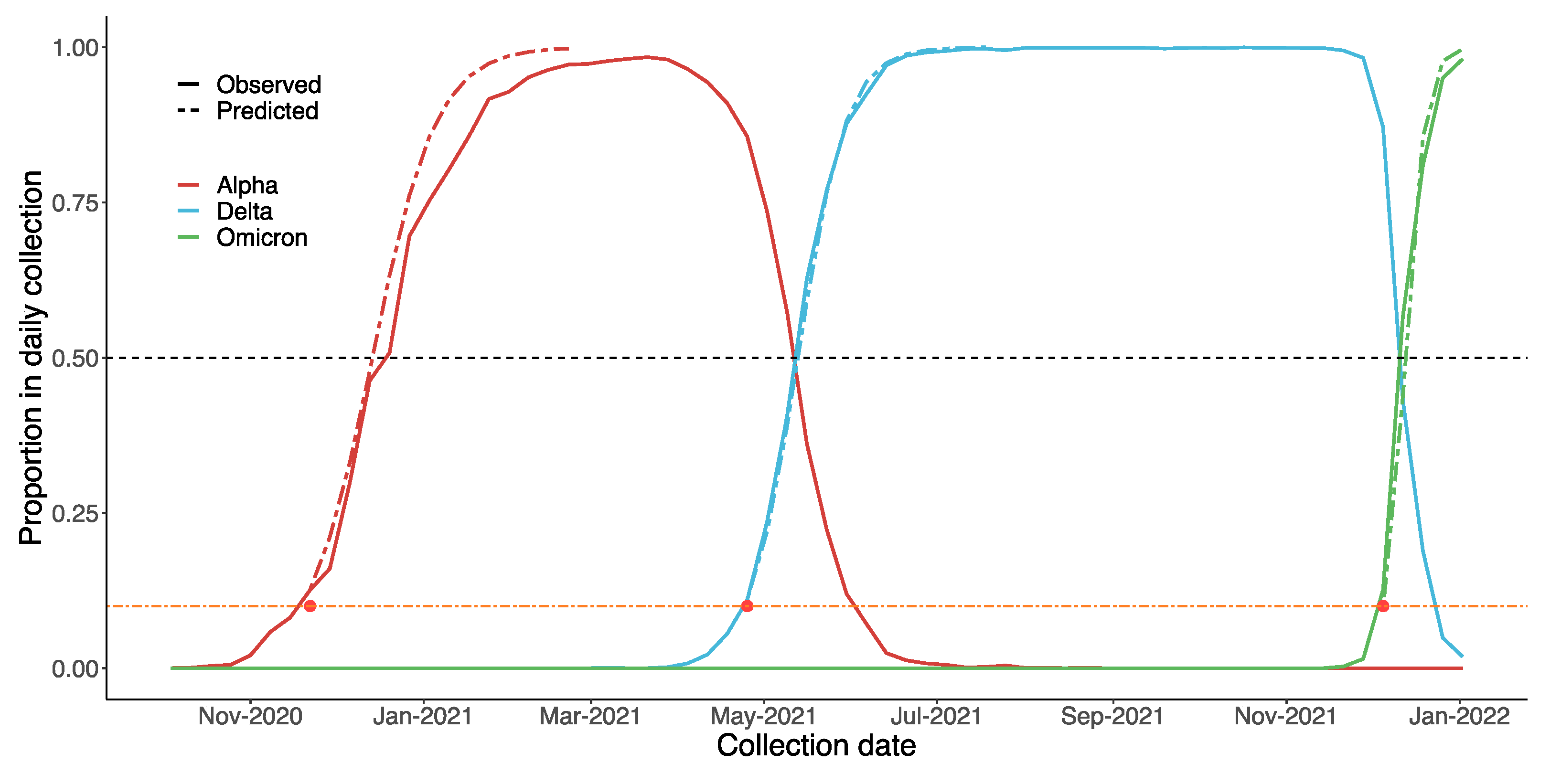
3.6. Estimating the Date on which a VOC Becomes Dominant
4. Conclusions
5. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Washington, N.L.; Gangavarapu, K.; Zeller, M.; Bolze, A.; Cirullli, E.T.; Barrett, K.M.S.; Larsen, B.B.; Anderson, C.; White, S.; Cassens, T.; et al. Emergence and rapid transmission of SARS-CoV-2 B.1.1.7 in the United States. Cell 2021, 184, 2587–2594.e7. [Google Scholar] [CrossRef] [PubMed]
- Volz, E.; Mishra, S.; Chand, M.; Barrett, J.C.; Johnson, R.; Geildergerg, R.; Hinsley, W.R.; Laydon, D.R.; Dabbrera, G.; O’Toole, A.; et al. Assessing transmissibility of SARS-CoV-2 lineage B.1.1.7 in England. Nature 2021, 593, 266–269. [Google Scholar] [CrossRef] [PubMed]
- Centers for Disease Control and Prevention. Variant Proportions. 2021. Available online: https://covid.cdc.gov/covid-data-tracker/#variant-proportions (accessed on 26 December 2021).
- Kannan, S.R.; Spratt, A.N.; Cohen, A.R.; Naqvi, S.H.; Chand, H.S.; Quinn, T.P.; Lorson, C.L.; Byrareddy, S.L.; Singh, K. Evolutionary analysis of the Delta and Delta Plus variants of the SARS-CoV-2 viruses. J. Autoimmun. 2021, 124, 102715. [Google Scholar] [CrossRef] [PubMed]
- Planas, D.; Veyer, D.; Baidaliuk, A.; Strapoli, I.; Guilvel-Benhanssine, F.; Rajah, M.M.; Planchais, C.; Porrot, F.; Robillard, N.; Puech, J.; et al. Reduced sensitivity of SARS-CoV-2 variant Delta to antibody neutralization. Nature 2021, 596, 276–280. [Google Scholar] [CrossRef] [PubMed]
- Suchard, M.A.; Lemey, P.; Baele, G.; Ayres, D.L.; Drummond, A.J.; Rambaut, A. Recent advances in SARS-CoV-2 Spike protein and RBD mutations comparison between new variants Alpha (B.1.1.7, United Kingdom), Beta (B.1.351, South Africa), Gamma (P.1, Brazil) and Delta (B.1.617.2, India). J. Virus Erad. 2021, 7, 100054. [Google Scholar]
- Pybus, O.G.; Rambaut, A. Evolutionary analysis of the dynamics of viral infectious disease. Nat. Rev. Genet. 2009, 10, 540–550. [Google Scholar] [CrossRef] [PubMed]
- Volz, E.M.; Koelle, K.; Bedford, T. Viral phylodynamics. PLoS Comput. Biol. 2013, 9, e103927. [Google Scholar] [CrossRef] [Green Version]
- Dong, E.; Du, H.; Gardner, L. An interactive web-based dashboard to track COVID-19 in real time. Lancet Infect. Dis. 2020, 20, 533–534. [Google Scholar] [CrossRef]
- Ritchie, H.; Mathieu, E.; Rodes-Guirao, L.; Appel, C.; Giattino, C.; Ortiz-Ospina, E.; Hasell, J.; Macdonald, B.; Dattani, S.; Roser, M. Coronavirus Pandemic (COVID-19). Available online: https://ourworldindata.org/coronavirus (accessed on 1 December 2021).
- Hale, T.; Angrist, N.; Goldszmidt, R.; Kira, B.; Patherick, A.; Phillips, T.; Webster, S.; Cameron-Blake, E.; Hallas, L.; Majumdar, S.; et al. A global panel database of pandemic policies (Oxford COVID-19 Government Response Tracker). Nat. Hum. Behav. 2021, 5, 529–538. [Google Scholar] [CrossRef] [PubMed]
- Drummond, A.J.; Rambaut, A. BEAST: Bayesian evolutionary analysis by sampling trees. BMC Evol. Biol. 2007, 7, 214. [Google Scholar] [CrossRef] [Green Version]
- Nguyen, L.T.; Schmidt, H.A.; Von Haeseler, A.; Minh, B.Q. IQ-TREE: A fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol. Biol. Evol. 2015, 32, 268–274. [Google Scholar] [CrossRef] [PubMed]
- Lyons, D.; Lauring, A.S. Evidence for the Selective Basis of Transition-to-Transversion Substitution Bias in Two RNA Viruses. Mol. Biol. Evol. 2017, 34, 3205–3215. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Koyama, T.; Platt, D.; Parida, L. Variant analysis of SARS-CoV-2 genomes. Bull. World Health Organ. 2020, 98, 495–504. [Google Scholar] [CrossRef] [PubMed]
- Kalia, K.; Saberwal, G.; Sharma, G. The lag in SARS-CoV-2 genome submissions to GISAID. Nat. Biotechnol. 2021, 39, 1058–1060. [Google Scholar] [CrossRef] [PubMed]
- Zelenova, M.; Ivanova, A.; Semyonov, S.; Gankin, Y. Analysis of 329,942 SARS-CoV-2 records retrieved from GISAID database. Comput. Biol. Med. 2021, 139, 104981. [Google Scholar] [CrossRef] [PubMed]
- Velazquez, A.; Bustria, M.; Ouyang, Y.; Moshiri, N. An analysis of clinical and geographical metadata of over 75,000 records in the GISAID COVID-19 database. medRxiv 2020. [Google Scholar] [CrossRef]
- Khare, S.; Gurry, C.; Freitas, L.; Schultz, M.B.; Bach, G.; Diallo, A.; Akite, N.; Ho, J.; Lee, R.T.; Yeo, W.; et al. GISAID’s role in pandemic response. China CDC Wkly. 2021, 3, 1049–1051. [Google Scholar] [CrossRef]
- Nonaka, C.; Franco, M.; Graf, T.; de Lorenzo Barcia, C.A.; de Avila Mendosa, A.V.; De Sousa, K.A.F.; Neiva, L.M.; Fosenca, V.; Mnedes, A.V.; de Aguiar, R.S.; et al. Genomic evidence of SARS-CoV-2 reinfection involving E484K spike mutation, Brazil. Emerg. Infect. Dis. 2021, 27, 1522–1524. [Google Scholar] [CrossRef]
- Hamed, S.M.; Elkhatib, W.F.; Khairalla, A.S.; Noreddin, A.M. Global dynamics of SARS-CoV-2 clades and the relation to COVID-19 epidemiology. Sci. Rep. 2021, 11, 8435. [Google Scholar] [CrossRef]
- Rockett, R.J.; Arnott, A.; Lam, C.; Sadsad, R.; Timms, V.; Gray, K.A.; Eden, J.S.; Chang, S.; Gall, M.; Draper, J.; et al. Revealing COVID-19 transmission in Australia by SARS-CoV-2 genomes sequencing and agent-based modeling. Nat. Med. 2020, 26, 1398–1404. [Google Scholar] [CrossRef]
- Vaid, S.; Cakan, C.; Bhandari, M. Using machine learning to estimate unobserved COVID-19 infections in North America. J. Bone Joint Surg. Am. 2020, 102, e70. [Google Scholar] [CrossRef] [PubMed]
- Car, Z.; Segota, S.B.; Andelic, N.; Lorencin, I.; Mirzjak, V. Modeling the spread of COVID-19 infection using a multilayer perceptron. Comput. Math. Methods Med. 2020, 2020, 5714714. [Google Scholar] [CrossRef] [PubMed]
- Velavan, T.P.; Meyer, C.G. The COVID-19 epidemic. Trop. Med. Int. Health 2020, 25, 278–280. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ghany, K.; Zawbaa, H.; Sabri, H. COVID-19 prediciton using LSTM algorithm: GCC case study. Inform. Med. Unlocked 2021, 23, 100566. [Google Scholar] [CrossRef]
- Muhareb, R.; Giacaman, R. Tracking COVID-19 responsibly. Lancet 2020, 3. [Google Scholar] [CrossRef] [Green Version]
- Chisadza, C.; Clance, M.; Gupta, R. Government effectiveness and the COVID-19 pandemic. Sustainability 2021, 13, 3042. [Google Scholar] [CrossRef]
- Koh, W.C.; Naing, L.; Wong, J. Estimating the impact of physical distancing measures in containing COVID-19: An empirical analysis. Int. J. Infect. Dis. 2020, 100, 42–49. [Google Scholar] [CrossRef] [PubMed]
- Celik, I.; Tallei, T.E. A computational comparative analysis of the binding mechanism of molnupiravir’s active metabolite to RND-dependent RNA polymerase of wild-type and Delta subvariant AY.4 of SARS-CoV-2. J. Cell Biochem. 2022. [Google Scholar] [CrossRef]
- Mazur-Panasiuk, N.; Rabalski, L.; Gromowski, T.; Nowicki, G.; Kowalski, W.; Wydmanski, W.; Szulc, P.; Kosincki, M.; Gackowska, K.; Drweska-Matelska, N.; et al. Expansion of a SARS-CoV-2 Delta variant with an 872 nt deletion encompassing ORF7a, ORF7b and ORF8, Poland, July to August 2021. Eurosurveillance 2021, 26, 2100902. [Google Scholar] [CrossRef]
- Umair, M.; Ikram, A.; Rehman, Z.; Haider, A.; Badar, N.; Ammar, M.; Ahad, A.; Suleman, R.; Salman, M. Genomic diversity of SARS-CoV-2 in Pakistan during fourth wave of pandemic. medRxiv 2021. [Google Scholar] [CrossRef]
- Alkhatib, M.; Bellocchi, M.C.; Marchegiani, G.; Grelli, S.; Micheli, V.; Stella, D.; Zerillo, B.; Carioti, L.; Svicher, V.; Rogliani, P.; et al. First case of a COVID-19 patient infected by Delta AY.4 with a rare deletion leading to a N gene target failure by a specific real time PCR assay: Novel Omicron VOC might be doing similar scenario? Microorganisms 2022, 10, 268. [Google Scholar] [CrossRef] [PubMed]
- Singanayagam, A.; Hakki, S.; Dunning, J.; Madon, K.J.; Crone, M.A.; Koycheva, A.; Derqui-Fernandez, N.; Barnett, J.L.; Whitfield, M.G.; Varo, R.; et al. Community transmission and viral load kinetics of the SARS-CoV-2 delta (B.1.617.2) variant in vaccinated and unvaccinated individuals in the UK: A prospective, longitudinal, cohort study. Lancet Infect. Dis. 2022, 22, 183–195. [Google Scholar] [CrossRef]
- Lassauniere, R.; Polacek, C.; Fonager, J.; Bennedbaek, M.; Boding, L.; Rasmussen, M.; Fomsgaard, A. Neutralisation of the SARS-CoV-2 Delta variant sub-lineages AY.4.2 and B.1.617.2 with the mutation E484K by Comirnaty (BNT162b2 mRNA) vaccine-elicited sera, Denmark, 1 to 26 November 2021. Eurosurveillance 2021, 26, 2101059. [Google Scholar] [CrossRef] [PubMed]
- Sonabend, R.; Whittles, L.K.; Imai, N.; Perez-Guzman, P.N.; Knock, E.S.; Rawson, T.; Gaythorpe, K.A.; Djaafara, B.A.; Hinsley, W.; FitzJohn, R.G.; et al. Non-pharmaceutical intervention, vaccination, and the SARS-CoV-2 delta variant in England: A mathematical modelling study. Lancet 2021, 398, 1825–1835. [Google Scholar] [CrossRef]
- Khatib, A.N.; McGuinness, S.; Wilder-Simth, A. COVID-19 transmission and the safety of air travel during the pandemic: A scoping review. Curr. Opin. Infect. Dis. 2021, 34, 415–422. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, T.P.; Wong, Z.S.Y.; Wang, L.; Thanh, T.T.; Van Nguyen, H.; Gilmour, S. Rapid impact assessments of COVID-19 control measures against the Delta variant and short-term projections of new confirmed cases in Vietnam. J. Glob. Health 2021, 11, 03118. [Google Scholar] [CrossRef] [PubMed]
- Aguiar, M.; Van-Dierdonck, J.B.; Mar, J.; Cusimano, N.; Knopof, D.; Anam, V.; Stollenwerk, N. Critical fluctuations in epidemic models explain COVID-19 post-lockdown dynamics. Sci. Rep. 2021, 11, 13839. [Google Scholar] [CrossRef]
- Holmes, E.C. Comparative studies of RNA virus evolution. In Origin and Evolution of Viruses, 2nd ed.; Chapter 5; Academic Press: Cambridge, MA, USA, 2008; pp. 119–134. [Google Scholar]
- Saad-Roy, C.M.; Wingreen, N.S.; Levin, S.A.; Grenfell, B.T. Dynamics in a simple evolutionary-epidemiological model for the evolution of an initial asymptomatic infection stage. Proc. Natl. Acad. Sci. USA 2020, 117, 11541–11550. [Google Scholar] [CrossRef] [PubMed]
- Calafiore, G.C.; Novara, C.; Possieri, C. A modified SIR model for the COVID-19 contagion in Italy 2020. In Proceedings of the 59th IEEE Conference on Decision and Control (CDC), Jeju, Korea, 14–18 December 2020; pp. 3889–3894. [Google Scholar]
- Ellison, G. Implications of Heterogeneous SIR Models for Analyses of COVID-19; NBER Working Papers Series; National Bureau of Economic Research: Cambridge, MA, USA, 2020; Volume 118, p. 27373. [Google Scholar]
- He, S.; Peng, Y.; Sun, K. SEIR modeling of the COVID-19 and its dynamics. Nonlinear Dyn. 2020, 101, 1667–1680. [Google Scholar] [CrossRef] [PubMed]

| n_GISAID 1 | n_estimated 2 | Name | n_GISAID 1 | n_estimated 2 | n_estimated 2 | Index | |
|---|---|---|---|---|---|---|---|
| 1 Februay 2021 to 17 July 2021 | B.1.617.2 | predominant variant | all | ||||
| India | B.1.617.1 | 0.0015 | |||||
| Indonesia | B.1.466.2 | 0.0021 | |||||
| Russia | B.1.1.523 | 0.0019 | |||||
| United Kingdom | B.1.1.7 | 0.2462 | |||||
| United States | B.1.1.7 | 0.0725 | |||||
| 30 May 2021 to 24 July 2021 | AY.4 | predominant variant | all | ||||
| India | B.1.617.2 | 0.0014 | |||||
| Indonesia | B.1.617.2 | 0.0013 | |||||
| Russia | B.1.617.2 | 0.0015 | |||||
| United Kingdom | B.1.617.2 | 0.1527 | |||||
| United States | B.1.617.2 | 0.0782 | |||||
| Intervention | Effective Size | 95% CI |
|---|---|---|
| Covid 19 testing policy | −0.64 | (−1.09, −0.19) |
| Covid contact tracing | 0.04 | (−0.3, 0.39) |
| Covid vaccination policy | 0.01 | (−0.17, 0.19) |
| Debt relief | −0.07 | (−0.28, 0.14) |
| Face covering policies | 0.42 | (0.05, 0.78) |
| Income support | −0.09 | (−0.48, 0.29) |
| International travel | −1.68 | (−2.32, −1.03) |
| Public campaigns | 2.24 | (1.07, 3.4) |
| Public events | 0.26 | (−0.1, 0.63) |
| Public gathering rules | 0.32 | (−0.13, 0.76) |
| Public transport | −0.2 | (−0.5, 0.11) |
| School closures | 0.05 | (−0.25, 0.35) |
| Stay at home | −0.13 | (−0.46, 0.2) |
| Workplace closures | −0.32 | (−0.69, 0.05) |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Dong, R.; Hu, T.; Zhang, Y.; Li, Y.; Zhou, X.-H. Assessing the Transmissibility of the New SARS-CoV-2 Variants: From Delta to Omicron. Vaccines 2022, 10, 496. https://doi.org/10.3390/vaccines10040496
Dong R, Hu T, Zhang Y, Li Y, Zhou X-H. Assessing the Transmissibility of the New SARS-CoV-2 Variants: From Delta to Omicron. Vaccines. 2022; 10(4):496. https://doi.org/10.3390/vaccines10040496
Chicago/Turabian StyleDong, Rui, Taojun Hu, Yunjun Zhang, Yang Li, and Xiao-Hua Zhou. 2022. "Assessing the Transmissibility of the New SARS-CoV-2 Variants: From Delta to Omicron" Vaccines 10, no. 4: 496. https://doi.org/10.3390/vaccines10040496
APA StyleDong, R., Hu, T., Zhang, Y., Li, Y., & Zhou, X.-H. (2022). Assessing the Transmissibility of the New SARS-CoV-2 Variants: From Delta to Omicron. Vaccines, 10(4), 496. https://doi.org/10.3390/vaccines10040496








