The Flavonoid Quercetin Induces AP-1 Activation in FRTL-5 Thyroid Cells
Abstract
1. Introduction
2. Materials and Methods
2.1. Materials
2.2. Cell Culture
2.3. Plasmids and Transfection
2.4. Luciferase and CAT Assay
2.5. Nuclear Extracts and Electrophoretic Mobility Shift Assay
2.6. Other Assays
2.7. Statistical Analysis
3. Results
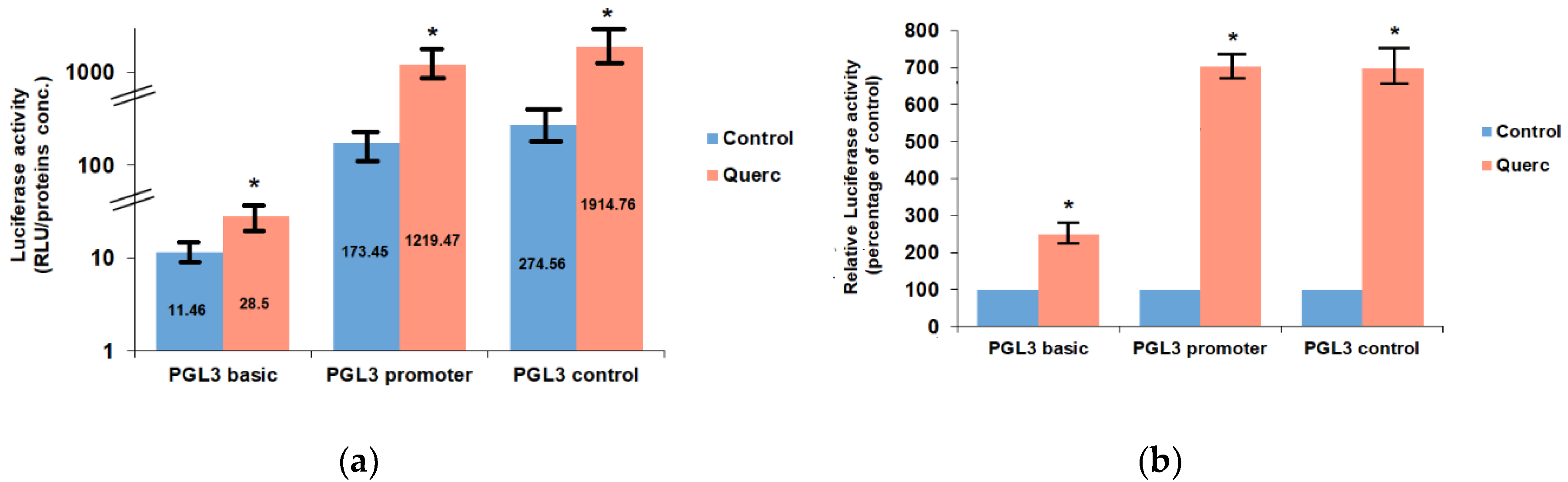
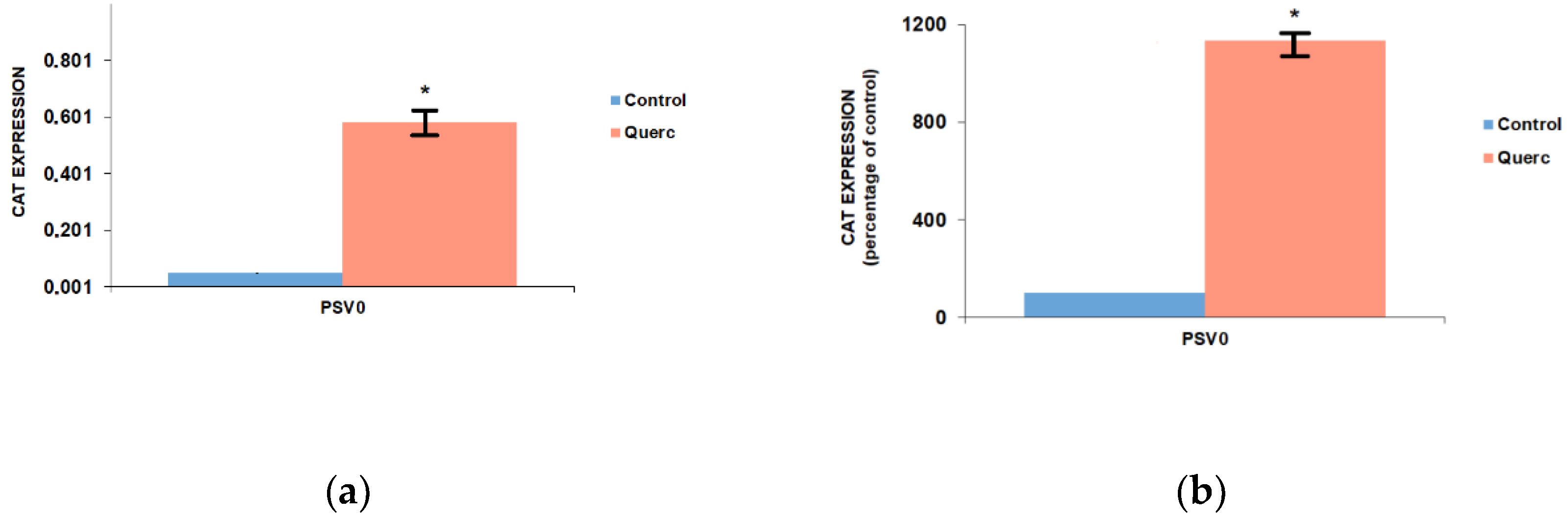
3.1. Quercetin Up-Regulates the Activity of the Following Reporter Vectors: PGL3 Basic, PGL3 Control, PGL3 Promoter, and PSV0-CAT
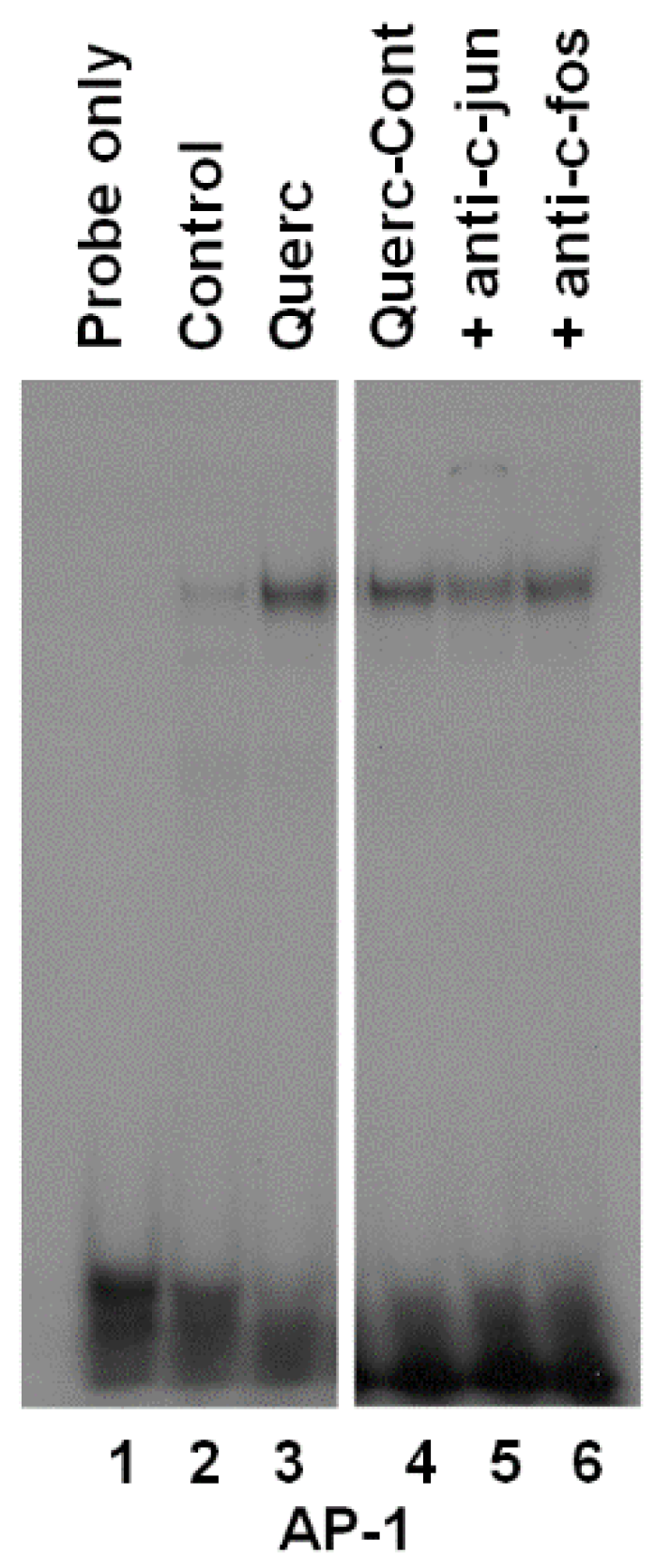
3.2. Quercetin Induces AP-1 Binding to DNA
4. Discussion
5. Conclusions
Funding
Acknowledgments
Conflicts of Interest
References
- Russo, M.; Spagnuolo, C.; Tedesco, I.; Bilotto, S.; Russo, G.L. The flavonoid quercetin in disease prevention and therapy: Facts and fancies. Biochem. Pharmacol. 2012, 83, 6–15. [Google Scholar] [CrossRef] [PubMed]
- Amor, S.; Chalons, P.; Aires, V.; Delmas, D. Polyphenol extracts from red wine and grapevine: Potential effects on cancers. Diseases 2018, 6, 106. [Google Scholar] [CrossRef]
- Santangelo, R.; Silvestrini, A.; Mancuso, C. Ginsenosides, catechins, quercetin and gut microbiota: Current evidence of challenging interactions. Food Chem. Toxicol. 2019, 123, 42–49. [Google Scholar] [CrossRef] [PubMed]
- Rajagopal, C.; Lankadasari, M.B.; Aranjani, J.M.; Harikumar, K.B. Targeting oncogenic transcription factors by polyphenols: A novel approach for cancer therapy. Pharmacol. Res. 2018, 130, 273–291. [Google Scholar] [CrossRef] [PubMed]
- Patel, R.V.; Mistry, B.M.; Shinde, S.K.; Syed, R.; Singh, V.; Shin, H.S. Therapeutic potential of quercetin as a cardiovascular agent. Eur. J. Med. Chem. 2018, 155, 889–904. [Google Scholar] [CrossRef] [PubMed]
- Miron, A.; Aprotosoaie, A.C.; Trifan, A.; Xiao, J. Flavonoids as modulators of metabolic enzymes and drug transporters. Ann. NY. Acad. Sci. 2017, 1398, 152–167. [Google Scholar] [CrossRef] [PubMed]
- Perez-Vizcaino, F.; Fraga, C.G. Research trends in flavonoids and health. Arch. Biochem. Biophys. 2018, 646, 107–112. [Google Scholar] [CrossRef]
- de Souza dos Santos, M.C.; Goncalves, C.F.L.; Vaisman, M.; Ferreira, A.C.F.; de Carvalho, D.P. Impact of flavonoids on thyroid function. Food Chem. Toxicol. 2011, 49, 2495–2502. [Google Scholar] [CrossRef] [PubMed]
- National Toxicology Program. Toxicology and carcinogenesis studies of Ginkgo biloba extract (CAS No. 90045-36-6) in F344/N rats and B6C3F1/N mice (Gavage studies). Natl. Toxicol. Program Tech. Rep. Ser. 2013, 578, 1–183. [Google Scholar]
- Giuliani, C.; Bucci, I.; Di Santo, S.; Rossi, C.; Grassadonia, A.; Piantelli, M.; Monaco, F.; Napolitano, G. The flavonoid quercetin inhibits thyroid-restricted genes expression and thyroid function. Food Chem. Toxicol. 2014, 66, 23–29. [Google Scholar] [CrossRef] [PubMed]
- Giuliani, C.; Noguchi, Y.; Harii, N.; Napolitano, G.; Tatone, D.; Bucci, I.; Piantelli, M.; Monaco, F.; Kohn, L.D. The flavonoid quercetin regulates growth and gene expression in rat FRTL-5 thyroid cells. Endocrinology 2008, 149, 84–92. [Google Scholar] [CrossRef]
- Ambesi Impiombato, F.S. Fast-growing thyroid cell strain. FRTL-5. US Patent 4,608,341, 26 August 1986. [Google Scholar]
- Kohn, L.D.; Valente, W.A. FRTL-5 manual: A current guide. In FRTL-5 Today; Ambesi Impiombato, F.S., Perrild, H., Eds.; Excerpta Medica: Amsterdam, The Netherlands, 1989; pp. 243–273. [Google Scholar]
- Giuliani, C.; Bucci, I.; Montani, V.; Singer, D.S.; Monaco, F.; Kohn, L.D.; Napolitano, G. Regulation of major histocompatibility complex gene expression in thyroid epithelial cells by methimazole and phenylmethimazole. J. Endocrinol. 2010, 204, 57–66. [Google Scholar] [CrossRef][Green Version]
- Lin, R.; Hogen, V.; Cannon, S.; Marion, K.M.; Fenton, M.S.; Hershman, J.M. Stability of recombinant human thyrotropin potency based on bioassay in FRTL-5 cells. Thyroid 2010, 20, 1139–1143. [Google Scholar] [CrossRef]
- Wen, G.; Ringseis, R.; Eder, K. Endoplasmic reticulum stress inhibits expression of genes involved in thyroid hormone synthesis and their key transcriptional regulators in FRTL-5 thyrocytes. PLoS ONE 2017, 12, e0187561. [Google Scholar] [CrossRef]
- Gorman, C.M.; Moffat, L.F.; Howard, B.H. Recombinant genomes which express chloramphenicol acetyltransferase in mammalian cells. Mol. Cell. Biol. 1982, 2, 1044–1051. [Google Scholar] [CrossRef]
- Lopata, M.A.; Cleveland, D.W.; Sollner-Webb, B. High level expression of a chloramphenicol acetyl transferase gene by DEAE-dextran mediated DNA transfections coupled with a dimetylsulfoxide or glycerol shock treatment. Nucleic Acids Res. 1984, 12, 5707–5717. [Google Scholar] [CrossRef]
- Napolitano, G.; Bucci, I.; Giuliani, C.; Massafra, C.; Di Petta, C.; Devangelio, E.; Singer, D.S.; Monaco, F.; Kohn, L.D. High glucose levels increase major histocompatibility complex class I gene expression in thyroid cells and amplify interferon-γ action. Endocrinology 2002, 143, 1008–1017. [Google Scholar] [CrossRef][Green Version]
- Dignam, J.; Lebowitz, R.; Roeder, R. Accurate transcription initiation by RNA polymerase II in a soluble extract from isolated mammalian nuclei. Necleic Acids Res. 1983, 11, 1475–1489. [Google Scholar] [CrossRef]
- Taniguchi, S.-I.; Shong, M.; Giuliani, C.; Napolitano, G.; Saji, M.; Montani, V.; Suzuki, K.; Singer, D.S.; Kohn, L.D. Iodide suppression of major histocompatibility class I gene expression in thyroid cells involves enhancer A and the transcription Factor NF-kB. Mol. Endocrinol. 1998, 12, 19–33. [Google Scholar] [CrossRef]
- pGL3 Luciferase Reporter Vectors. Available online: https://ita.promega.com/Products/Reporter-Assays-and-Transfection/Reporter-Vectors-and-Cell-Lines/pGL3-Luciferase-Reporter-Vectors/?fq=PGL%203%20basic&catNum=E1751 (accessed on 14 January 2019).
- Cloning Vector pGL3-Promoter, Complete Sequence. Available online: https://www.ncbi.nlm.nih.gov/nuccore/U47298.2/ (accessed on 29 January 2019).
- Lee, C.; Huang, C.-H. LASAGNA-Search 2.0: Integrated transcription factor binding site search and visualization in a browser. Bioinformatics 2014, 30, 1923–1925. [Google Scholar] [CrossRef]
- Grabe, N. AliBaba2: Context specific identification of transcription factor binding sites. In Silico Biology; IOS Press: Amsterdam, The Netherlands, 2002; Volume 2, pp. S1–S15. [Google Scholar]
- AliBaba2.1. Available online: http://gene-regulation.com/pub/programs/alibaba2/index.html? (accessed on 26 February 2019).
- Thierry, F.; Spyrou, G.; Yaniv, M.; Howley, P. Two AP1 sites binding JunB are essential for human papillomavirus type 18 transcription in keratinocytes. J. Virol. 1992, 66, 3740–3748. [Google Scholar]
- Delcuratolo, M.; Fertey, J.; Schneider, M.; Schuetz, J.; Leiprecht, N.; Hudjetz, B.; Brodbeck, S.; Corall, S.; Dreer, M.; Schwab, R.M.; et al. Papillomavirus-associated tumor formation critically depends on c-Fos expression induced by viral protein E2 and bromodomain protein Brd4. PLoS Pathol. 2016, 12(1), e1005366. [Google Scholar] [CrossRef]
- Ehrlich, R.; Maguire, J.E.; Singer, D.S. Identification of negative and positive regulatory elements associated with a class I major histocompatibility complex gene. Mol. Cell. Biol. 1988, 8, 695–703. [Google Scholar] [CrossRef]
- Araki, E.; Shimada, F.; Shichiri, M.; Mori, M.; Ebina, Y. pSV00CAT: Low background CAT plasmid. Nucleic. Acids Res. 1988, 16, 1627. [Google Scholar] [CrossRef][Green Version]
- Langner, K.D.; Weyer, U.; Doerfler, W. Trans effect of the El region of adenoviruses on the expression of a prokaryotic gene in mammalian cells: Resistance to 5′-CCGG-3′ methylation. Proc. Natl. Acad. Sci. USA 1986, 83, 1598–1602. [Google Scholar] [CrossRef] [PubMed]
- Phillips, B.W.; Sharma, R.; Leco, P.A.; Edwards, D.R. A Sequence-selective single-strand DNA-binding protein regulates basal transcription of the murine tissue inhibitor of Metalloproteinases-1 (Timp-1) gene. J. Biol. Chem. 1999, 274, 22197–22207. [Google Scholar] [CrossRef]
- Wang, W.-M.; Wu, S.-Y.; Lee, A.-Y.; Chiang, C.-M.; Phillips, B.W.; Sharma, R.; Leco, P.A.; Edwards, D.R. A Binding site specificity and factor redundancy in activator protein-1-driven human papillomavirus chromatin-dependent transcription. J. Biol. Chem. 2011, 286, 40974–40986. [Google Scholar] [CrossRef] [PubMed]
- Giuliani, C.; Bucci, I.; Napolitano, G. The role of the transcription factor Nuclear Factor-kappa B in thyroid autoimmunity and cancer. Front. Endocrinol. 2018, 9, 471. [Google Scholar] [CrossRef]
- Gupta, M.K.; Singh, D.B.; Shukla, R.; Misra, K. A comprehensive metabolic modeling of thyroid pathway in relation to thyroid pathophysiology and therapeutics. OMICS 2013, 17, 584–593. [Google Scholar] [CrossRef]
- Szabo-Freinas, N.; Blondeau, J.-P.; Pomerance, M. Activation of the cAMP pathway synergistically increases IL-1-induced IL-6 gene expression in FRTL-5 thyroid cells: Involvement of AP-1 transcription factors. Mol. Cell. Endocrinol. 2008, 284, 28–37. [Google Scholar] [CrossRef]
- Giuliani, C.; Saji, M.; Napolitano, G.; Palmer, L.A.; Taniguchi, S.I.; Shong, M.; Singer, D.S.; Kohn, L.D. Hormonal modulation of major histocompatibility complex class I gene expression involves an enhancer A-binding complex consisting of Fra-2 and the p50 subunit of NF-kappa B. J. Biol. Chem. 1995, 270, 11453–11462. [Google Scholar] [CrossRef] [PubMed]
- Lee, Y.-J.; Beak, S.-Y.; Choi, I.; Sung, J.-S. Quercetin and its metabolites protect hepatocytes against ethanol-induced oxidative stress by activation of Nrf2 and AP-1. Food Sci. Biotechnol. 2017, 27, 809–817. [Google Scholar] [CrossRef]
- Gopalakrishnan, A.; Xu, C.J.; Nair, S.S.; Chen, C.; Hebbar, V.; Kong, A.N. Modulation of activator protein-1 (AP-1) and MAPK pathway by flavonoids in human prostate cancer PC3 cells. Arch. Pharm. Res. 2006, 29, 633–644. [Google Scholar] [CrossRef]
- Kim, B.H.; Choi, J.S.; Yi, E.H.; Lee, J.K.; Won, C.; Ye, S.K.; Kim, M.H. Relative antioxidant activities of quercetin and its structurally related substances and their effects on NF-κB/CRE/AP-1 signaling in murine macrophages. Mol. Cells 2013, 35, 410–420. [Google Scholar] [CrossRef]
- Endale, M.; Park, S.C.; Kim, S.; Kim, S.H.; Yang, Y.; Cho, J.Y.; Rhee, M.H. Quercetin disrupts tyrosine-phosphorylated phosphatidylinositol 3-kinase and myeloid differentiation factor-88 association, and inhibits MAPK/AP-1 and IKK/NF-κB-induced inflammatory mediators production in RAW 264.7 cells. Immunobiology 2013, 218, 1452–1467. [Google Scholar] [CrossRef]
- Dougherty, D.C.; Sanders, M.M. Comparison of the responsiveness of the pGL3 and pGL4 luciferase reporter vectors to steroid hormones. Biotechniques 2005, 39, 203–207. [Google Scholar] [CrossRef]
- Jacobsen, L.B.; Calvin, S.A.; Lobenhofer, E.K. Transcriptional effects of transfection: The potential for misinterpretation of gene expression data generated from transiently transfected cells. Biotechniques 2009, 47, 617–624. [Google Scholar] [CrossRef] [PubMed]
- Martin, M.E.; Piette, J.; Yaniv, M.; Folk, R. Activation of the polyoma-virus enhancer by murine AP1 homolog and two contiguous proteins. Proc. Natl. Acad. Sci. USA 1988, 85, 5839–5843. [Google Scholar] [CrossRef] [PubMed]
- Hartl, M.; Reiter, F.; Bader, A.G.; Castellazzi, M.; Bister, K. JAC, a direct target of oncogenic transcription factor Jun, is involved in cell transformation and tumorigenesis. Proc. Natl. Acad. Sci. USA 2001, 98, 13601–13606. [Google Scholar] [CrossRef] [PubMed]
- Shih, H.; Pickwell, G.V.; Quattrochi, L.C. Differential Effects of Flavonoid Compounds on Tumor Promoter-Induced Activation of the Human CYP1A2 Enhancer. Arch. Biochem. Biophys. 2000, 373, 287–294. [Google Scholar] [CrossRef] [PubMed]
- Yuan, H.; Pan, Y.; Young, C.Y. Overexpression of c-Jun induced by quercetin and resveratrol inhibits the expression and function of the androgen receptor in human prostate cancer cells. Cancer Lett. 2004, 213, 155–163. [Google Scholar] [CrossRef] [PubMed]
- Xu, B.; Yang, H.; Sun, M.; Chen, H.; Jiang, L.; Zheng, X.; Ding, G.; Liu, Y.; Sheng, Y.; Cui, D.; et al. 2,3′,4,4′,5-Pentachlorobiphenyl induces inflammatory responses in the thyroid through JNK and aryl hydrocarbon Receptor-Mediated Pathway. Toxicol. Sci. 2016, 149, 300–311. [Google Scholar] [CrossRef] [PubMed]
- Napolitano, G.; Montani, V.; Giuliani, C.; Di Vincenzo, S.; Bucci, I.; Todisco, V.; Laglia, G.; Coppa, A.; Singer, D.S.; Nakazato, M.; et al. Transforming Growth Factor-β1 down-regulation of major histocompatibility complex class I in thyrocytes: Coordinate regulation of two separate elements by thyroid-specific as well as ubiquitous transcription factors. Mol. Endocrinol. 2000, 14, 486–505. [Google Scholar] [CrossRef] [PubMed][Green Version]
| Transcription Factor | Score | p Value |
|---|---|---|
| AP-1 | 13.33 | 0 |
| Oct-1 | 13.00 | 0 |
| Sp1 | 12.32 | 0.00025 |
© 2019 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Giuliani, C. The Flavonoid Quercetin Induces AP-1 Activation in FRTL-5 Thyroid Cells. Antioxidants 2019, 8, 112. https://doi.org/10.3390/antiox8050112
Giuliani C. The Flavonoid Quercetin Induces AP-1 Activation in FRTL-5 Thyroid Cells. Antioxidants. 2019; 8(5):112. https://doi.org/10.3390/antiox8050112
Chicago/Turabian StyleGiuliani, Cesidio. 2019. "The Flavonoid Quercetin Induces AP-1 Activation in FRTL-5 Thyroid Cells" Antioxidants 8, no. 5: 112. https://doi.org/10.3390/antiox8050112
APA StyleGiuliani, C. (2019). The Flavonoid Quercetin Induces AP-1 Activation in FRTL-5 Thyroid Cells. Antioxidants, 8(5), 112. https://doi.org/10.3390/antiox8050112




