S-allylmercaptoglutathione Is a Substrate for Glutathione Reductase (E.C. 1.8.1.7) from Yeast (Saccharomyces cerevisiae)
Abstract
1. Introduction
2. Materials & Methods
2.1. Allicin Synthesis
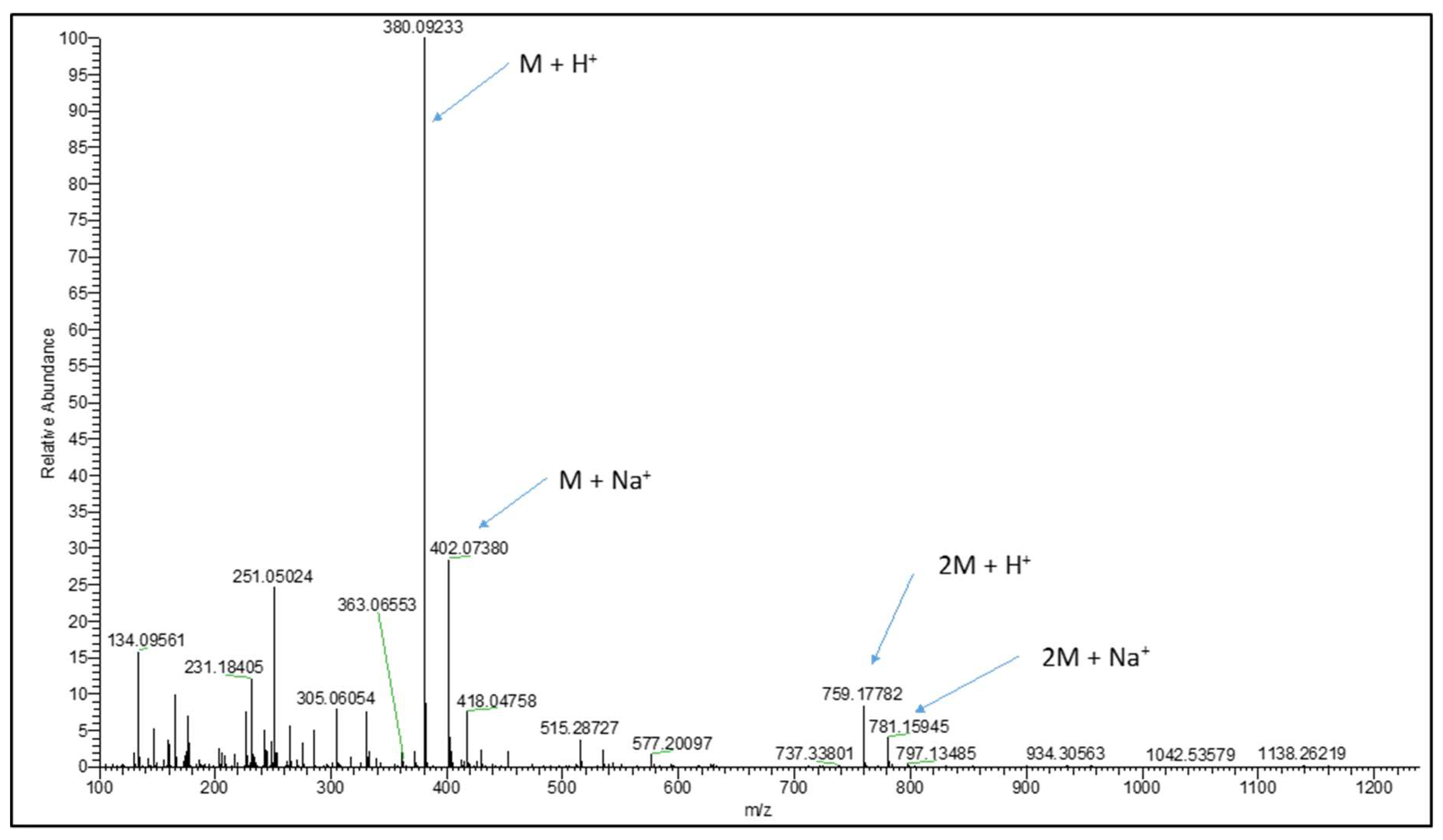
2.2. Synthesis and LC-MS of S-allylmercaptoglutathione
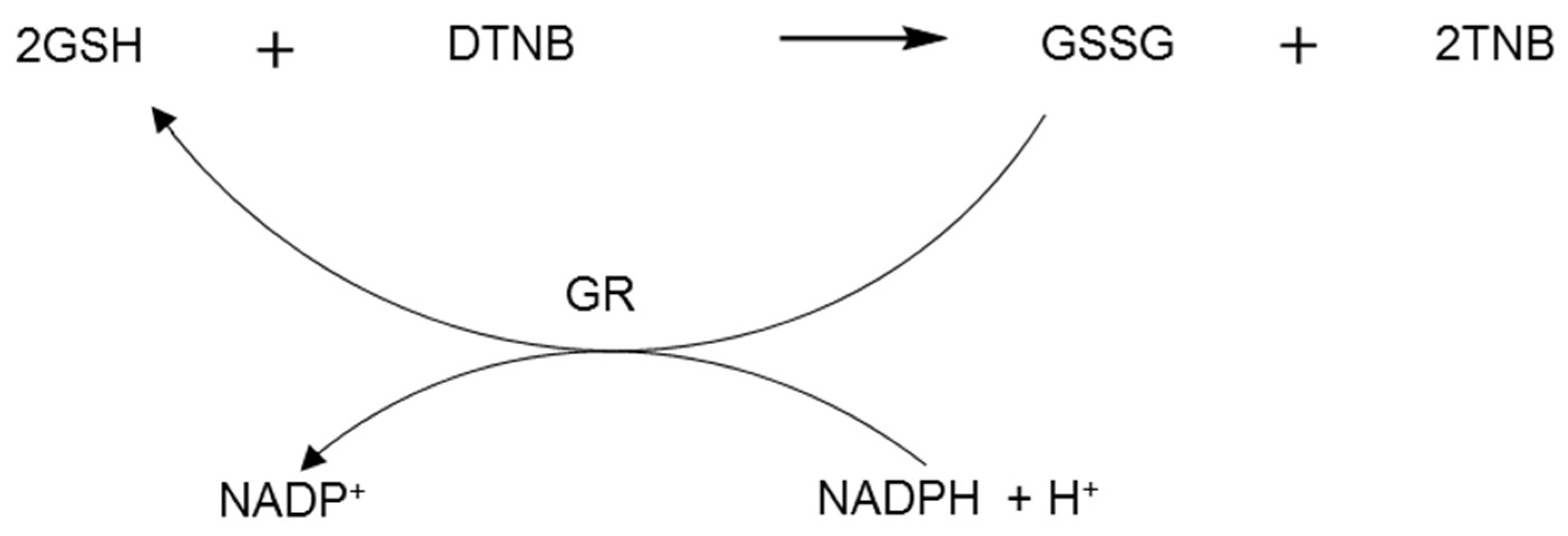
2.3. Glutathione Reductase Assay
2.4. Yeast
3. Results and Discussion
3.1. Synthesis of GSSA
3.2. Glutathione Reductase Assays
3.3. A Δgsh1 Mutant Can Grow on Medium without GSH When Supplemented with GSSA
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Noctor, G.; Foyer, C.H. Ascorbate and Glutathione: Keeping Active Oxygen Under Control. Annu. Rev. Plant Physiol. Plant Mol. Biol. 1998, 49, 249–279. [Google Scholar] [CrossRef] [PubMed]
- Schafer, F.Q.; Buettner, G.R. Redox environment of the cell as viewed through the redox state of the glutathione disulfide/glutathione couple. Free Radic. Biol. Med. 2001, 30, 1191–1212. [Google Scholar] [CrossRef]
- Morgan, B.; Ezerina, D.; Amoako, T.N.E.; Riemer, J.; Seedorf, M.; Dick, T.P. Multiple glutathione disulfide removal pathways mediate cytosolic redox homeostasis. Nat. Chem. Biol. 2013, 9, 119–125. [Google Scholar] [CrossRef] [PubMed]
- Winterbourn, C.C.; Hampton, M.B. Thiol chemistry and specificity in redox signaling. Free Radic. Biol. Med. 2008, 45, 549–561. [Google Scholar] [CrossRef] [PubMed]
- Karplus, P.A. A primer on peroxiredoxin biochemistry. Free Radic. Biol. Med. 2015, 80, 183–190. [Google Scholar] [CrossRef] [PubMed]
- Gruhlke, M.C.H.; Slusarenko, A.J. The biology of reactive sulfur species (RSS). Plant Physiol. Biochem. 2012, 59, 98–107. [Google Scholar] [CrossRef] [PubMed]
- Poole, L.B. The basics of thiols and cysteines in redox biology and chemistry. Free Radic. Biol. Med. 2015, 80, 148–157. [Google Scholar] [CrossRef] [PubMed]
- Griffith, O.W. Determination of Glutathione and Glutathione Disulfide Using Glutathione Reductase and 2-Vinylpyridine. Anal. Biochem. 1980, 106, 207–212. [Google Scholar] [CrossRef]
- Lawson, L.D.; Wood, S.G.; Hughes, B.G. HPLC Analysis of Allicin and Other Thiosulfinates in Garlic Clove Homogenates. Planta Med. 1991, 57, 263–270. [Google Scholar] [CrossRef] [PubMed]
- Cavallito, C.J.; Bailey, J.H. Allicin, the Antibacterial Principle of Allium sativum I. Isolation, Physical Properties and Antibacterial Action. J. Am. Chem. Soc. 1944, 66, 1950–1951. [Google Scholar] [CrossRef]
- Cavallito, C.J.; Buck, J.S.; Suter, C.M. Allicin, the Antibacterial Principle of Allium sativum. II. Determination of the Chemical Structure. J. Am. Chem. Soc. 1944, 66, 1952–1954. [Google Scholar] [CrossRef]
- Miron, T.; Rabinkov, A.; Mirelman, D.; Wilchek, M.; Weiner, L. The mode of action of allicin: Its ready permeability through phospholipid membranes may contribute to its biological activity. Biochim. Biophys. Acta (BBA) Biomembr. 2000, 1463, 20–30. [Google Scholar] [CrossRef]
- Rabinkov, A.; Miron, T.; Mirelman, D.; Wilchek, M.; Glozman, S.; Yavin, E.; Weiner, L. S-allylmercaptoglutathione: The reaction product of allicin with glutathione possesses SH-modifying and antioxidant properties. Biochim. Biophys. Acta (BBA) Mol. Cell Res. 2000, 1499, 144–153. [Google Scholar] [CrossRef]
- Gruhlke, M.C.H.; Portz, D.; Stitz, M.; Anwar, A.; Schneider, T.; Jacob, C.; Schlaich, N.L.; Slusarenko, A.J. Allicin disrupts the cell’s electrochemical potential and induces apoptosis in yeast. Free Radic. Biol. Med. 2012, 49, 1916–1924. [Google Scholar] [CrossRef] [PubMed]
- Müller, A.; Eller, J.; Albrecht, F.; Prochnow, P.; Kuhlmann, K.; Bandow, J.E.; Slusarenko, A.J.; Leichert, L.I. Allicin Induces Thiol Stress in Bacteria through S-Allylmercapto Modification of Protein Cysteines. J. Biol. Chem. 2016, 291, 11477–11490. [Google Scholar] [CrossRef] [PubMed]
- Gruhlke, M.C.H.; Schlembach, I.; Leontiev, R.; Uebachs, A.; Gollwitzer, P.; Weiss, A.; Delaunay, A.; Toledano, M.B.; Slusarenko, A.J. Yap1p, the central regulator of the S. cerevisiae oxidative stress response, is activated by allicin, a natural oxidant and defence substance of garlic. Free Radic. Biol. Med. 2017, 108, 793–802. [Google Scholar] [CrossRef] [PubMed]
- Leontiev, R.; Hohaus, N.; Jacob, C.; Gruhlke, M.C.H.; Slusarenko, A.J. A Comparison of the Antibacterial and Antifungal Activities of Thiosulfinate Analogues of Allicin. Sci. Rep. 2018, 8, 6763. [Google Scholar] [CrossRef] [PubMed]
- Albrecht, F.; Leontiev, R.; Jacob, C.; Slusarenko, A.J. An Optimized Facile Procedure to synthesize and purify Allicin. Molecules 2017, 22, 770. [Google Scholar] [CrossRef] [PubMed]
- Glutathione Reductase from Baker’s Yeast (Saccharomyces cerevisiae). Available online: https://www.sigmaaldrich.com/content/dam/sigma-aldrich/docs/Sigma/Product_Information_Sheet/2/g3664pis.pdf (accessed on 19 June 2018).
- Spector, D.; Labarre, J.; Toledano, M.B. A Genetic Investigation of the Essential Role of Glutathione: Mutations in the proline biosynthesis pathway are the only suppressors of glutathione auxotrophy in yeast. J. Biol. Chem. 2001, 276, 7011–7016. [Google Scholar] [CrossRef] [PubMed]
- Feldberg, R.S.; Chang, S.C.; Kotik, A.N.; Nadler, M.; Neuwirth, Z.; Sundstrom, D.C.; Thompson, N.H. In vitro mechanism of inhibition of bacterial cell growth by allicin. Antimicrob. Agents Chemother. 1988, 32, 1763–1768. [Google Scholar] [CrossRef] [PubMed]
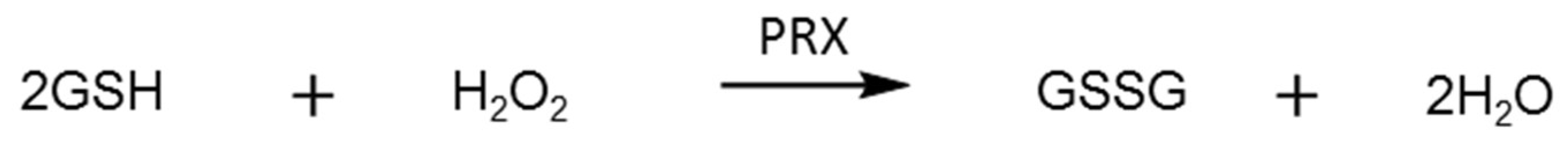
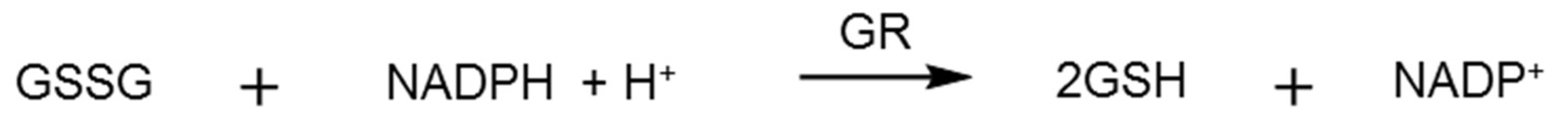




© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Horn, T.; Bettray, W.; Slusarenko, A.J.; Gruhlke, M.C.H. S-allylmercaptoglutathione Is a Substrate for Glutathione Reductase (E.C. 1.8.1.7) from Yeast (Saccharomyces cerevisiae). Antioxidants 2018, 7, 86. https://doi.org/10.3390/antiox7070086
Horn T, Bettray W, Slusarenko AJ, Gruhlke MCH. S-allylmercaptoglutathione Is a Substrate for Glutathione Reductase (E.C. 1.8.1.7) from Yeast (Saccharomyces cerevisiae). Antioxidants. 2018; 7(7):86. https://doi.org/10.3390/antiox7070086
Chicago/Turabian StyleHorn, Tobias, Wolfgang Bettray, Alan J. Slusarenko, and Martin C. H. Gruhlke. 2018. "S-allylmercaptoglutathione Is a Substrate for Glutathione Reductase (E.C. 1.8.1.7) from Yeast (Saccharomyces cerevisiae)" Antioxidants 7, no. 7: 86. https://doi.org/10.3390/antiox7070086
APA StyleHorn, T., Bettray, W., Slusarenko, A. J., & Gruhlke, M. C. H. (2018). S-allylmercaptoglutathione Is a Substrate for Glutathione Reductase (E.C. 1.8.1.7) from Yeast (Saccharomyces cerevisiae). Antioxidants, 7(7), 86. https://doi.org/10.3390/antiox7070086





