Jasminum sambac Cell Extract as Antioxidant Booster against Skin Aging
Abstract
1. Introduction
2. Materials and Methods
2.1. Plant Tissue Cultures and Extract Preparation
2.2. UPLC–MS/MS Analysis for JasHEx Chemical Characterization
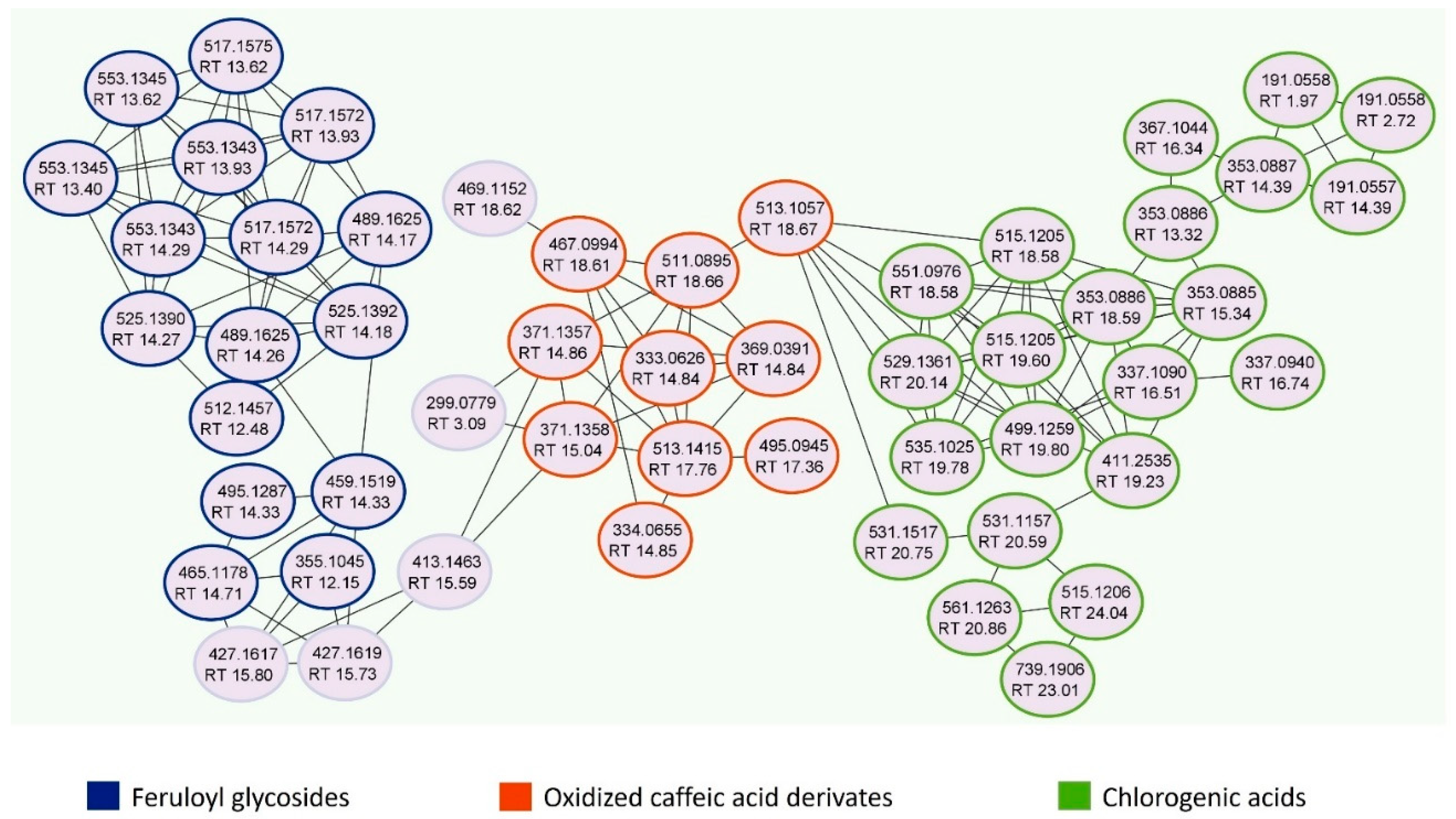
2.3. Global Natural Products Social Molecular Networking Analyses
2.4. Quantitative Analysis of Lignans and Triterpenes
2.5. Skin cell Cultures and Explants
2.6. Cytosolic ROS Assay in H2O2-Stressed HaCaT Cells
2.7. Enzyme-Linked Immunosorbent Assay (ELISA) for AGE Detection in Glyoxal Stressed Human Dermal Fibroblast (HDF) Cells
2.8. ImmunoHistoFluorescence Assay on Methyl-Glyoxal Stressed Skin Explants for Fibrillin-1 Detection
2.9. AlphaLISA Assay to Measure Procollagen Type I C-Peptide (PIP) Content
2.10. Nrf2 Luciferase-Based Transcription Activation Assay
2.11. Analysis of the Expression of SOD-1(NM_000454.5) and OH-1(NM_002133.3) Genes in HaCaT Cells
2.12. Nitric Oxide Assay in LPS-Stimulated RAW 264.7
3. Results
3.1. Qualitative and Quantitative Analysis of Jasminum sambac Cell Culture Hydro-Ethanolic Extract (JasHEx)
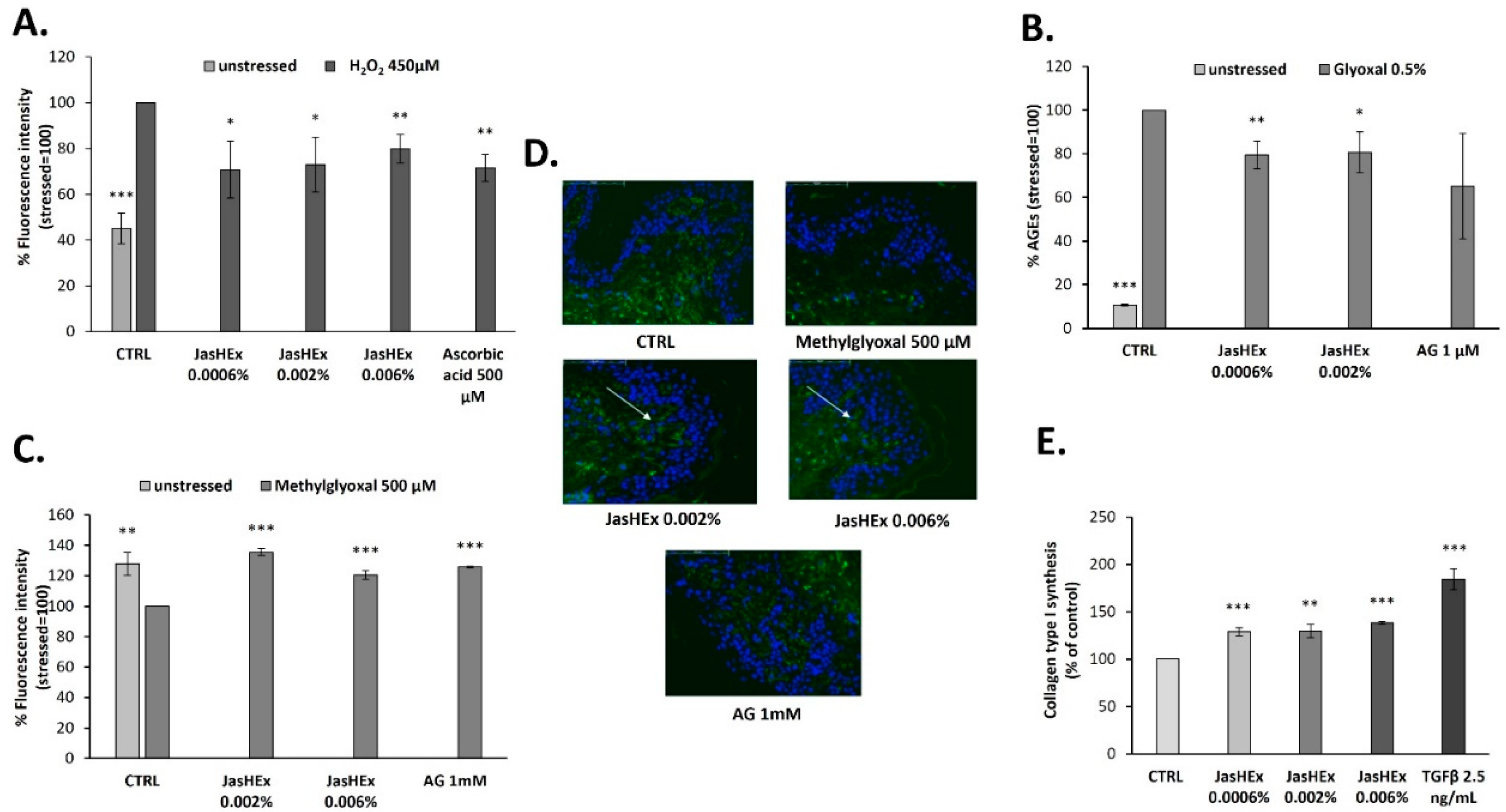
3.2. Cytosolic ROS Detection in H2O2-Stressed HaCaT Cells
3.3. AGE Detection in Glyoxal Treated HDF
3.4. Fibrillin-1 Detection in Methylglyoxal Stressed Skin Explants
3.5. Analysis of Collagen Type I Synthesis
3.6. Analysis of Nrf2/ARE Pathway in HaCaT Cells
3.7. Analysis of OH-1 and SOD-1 Gene Expression in HaCaT Cells
3.8. NO Determination in LPS-Stimulated RAW 264.7 Cells
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Bae, Y.S.; Oh, H.; Rhee, S.G.; Yoo, Y.D. Regulation of Reactive Oxygen Species Generation in Cell Signaling. Mol. Cells 2011, 32, 491–509. [Google Scholar] [CrossRef]
- Hasanuzzaman, M.; Bhuyan, M.H.M.B.; Zulfiqar, F.; Raza, A.; Mohsin, S.M.; Mahmud, J.A.; Fujita, M.; Fotopoulos, V. Reactive Oxygen Species and Antioxidant Defense in Plants under Abiotic Stress: Revisiting the Crucial Role of a Universal Defense Regulator. Antioxidants 2020, 9, 681. [Google Scholar] [CrossRef]
- Rinnerthaler, M.; Bischof, J.; Streubel, M.K.; Trost, A.; Richter, K. Oxidative Stress in Aging Human Skin. Biomolecules 2015, 5, 545–589. [Google Scholar] [CrossRef]
- Sárdy, M. Role of Matrix Metalloproteinases in Skin Ageing. Connect Tissue Res. 2009, 50, 132–138. [Google Scholar] [CrossRef]
- Masaki, H. Role of Antioxidants in the Skin: Anti-Aging Effects. J. Dermatol. Sci. 2010, 58, 85–90. [Google Scholar] [CrossRef]
- Perrone, A.; Giovino, A.; Benny, J.; Martinelli, F. Advanced Glycation End Products (AGEs): Biochemistry, Signaling, Analytical Methods, and Epigenetic Effects. Oxid. Med. Cell. Longev. 2020, 2020, 3818196. [Google Scholar] [CrossRef] [PubMed]
- Singh, V.P.; Bali, A.; Singh, N.; Jaggi, A.S. Advanced Glycation End Products and Diabetic Complications. Korean J. Physiol. Pharmacol. 2014, 18, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Cepas, V.; Collino, M.; Mayo, J.C.; Sainz, R.M. Redox Signaling and Advanced Glycation Endproducts (AGEs) in Diet-Related Diseases. Antioxidants 2020, 9, E142. [Google Scholar] [CrossRef] [PubMed]
- Twarda-Clapa, A.; Olczak, A.; Białkowska, A.M.; Koziołkiewicz, M. Advanced Glycation End-Products (AGEs): Formation, Chemistry, Classification, Receptors, and Diseases Related to AGEs. Cells 2022, 11, 1312. [Google Scholar] [CrossRef]
- Dyer, D.G.; Dunn, J.A.; Thorpe, S.R.; Bailie, K.E.; Lyons, T.J.; McCance, D.R.; Baynes, J.W. Accumulation of Maillard Reaction Products in Skin Collagen in Diabetes and Aging. J. Clin. Investig. 1993, 91, 2463–2469. [Google Scholar] [CrossRef] [PubMed]
- Verzijl, N.; DeGroot, J.; Oldehinkel, E.; Bank, R.A.; Thorpe, S.R.; Baynes, J.W.; Bayliss, M.T.; Bijlsma, J.W.; Lafeber, F.P.; Tekoppele, J.M. Age-Related Accumulation of Maillard Reaction Products in Human Articular Cartilage Collagen. Biochem. J. 2000, 350 Pt 2, 381–387. [Google Scholar] [CrossRef] [PubMed]
- Reiser, K.M. Nonenzymatic Glycation of Collagen in Aging and Diabetes. Proc. Soc. Exp. Biol. Med. 1998, 218, 23–37. [Google Scholar] [CrossRef] [PubMed]
- Toppo, A. A Review on Jasminum Sambac: A Potential Medicinal Plant. Int. J. Ind. Herbs. Drugs 2017, 2, 13–16. [Google Scholar]
- Barbulova, A.; Apone, F.; Colucci, G. Plant Cell Cultures as Source of Cosmetic Active Ingredients. Cosmetics 2014, 1, 94–104. [Google Scholar] [CrossRef]
- Ochoa-Villarreal, M.; Howat, S.; Jang, M.O.; Kim, I.S.; Jin, Y.-W.; Lee, E.-K.; Loake, G.J. Cambial Meristematic Cells: A Platform for the Production of Plant Natural Products. New Biotechnol. 2015, 32, 581–587. [Google Scholar] [CrossRef] [PubMed]
- Murashige, T.; Skoog, F. A Revised Medium for Rapid Growth and Bio Assays with Tobacco Tissue Cultures. Physiol. Plantarum. 1962, 15, 473–497. [Google Scholar] [CrossRef]
- Ceccacci, S.; De Lucia, A.; Tito, A.; Tortora, A.; Falanga, D.; Arciello, S.; Ausanio, G.; Di Cicco, C.; Monti, M.C.; Apone, F. An Oenothera Biennis Cell Cultures Extract Endowed with Skin Anti-Ageing Activity Improves Cell Mechanical Properties. Metabolites 2021, 11, 527. [Google Scholar] [CrossRef]
- Wang, M.; Carver, J.J.; Phelan, V.V.; Sanchez, L.M.; Garg, N.; Peng, Y.; Nguyen, D.D.; Watrous, J.; Kapono, C.A.; Luzzatto-Knaan, T.; et al. Sharing and Community Curation of Mass Spectrometry Data with GNPS. Nat. Biotechnol. 2016, 34, 828–837. [Google Scholar] [CrossRef]
- Pluskal, T.; Castillo, S.; Villar-Briones, A.; Orešič, M. MZmine 2: Modular Framework for Processing, Visualizing, and Analyzing Mass Spectrometry-Based Molecular Profile Data. BMC Bioinform. 2010, 11, 395. [Google Scholar] [CrossRef]
- Nothias, L.-F.; Petras, D.; Schmid, R.; Dührkop, K.; Rainer, J.; Sarvepalli, A.; Protsyuk, I.; Ernst, M.; Tsugawa, H.; Fleischauer, M.; et al. Feature-Based Molecular Networking in the GNPS Analysis Environment. Nat. Methods 2020, 17, 905–908. [Google Scholar] [CrossRef]
- Shannon, P.; Markiel, A.; Ozier, O.; Baliga, N.S.; Wang, J.T.; Ramage, D.; Amin, N.; Schwikowski, B.; Ideker, T. Cytoscape: A Software Environment for Integrated Models of Biomolecular Interaction Networks. Genome Res. 2003, 13, 2498–2504. [Google Scholar] [CrossRef] [PubMed]
- Peng, J.; Xie, J.; Shi, S.; Luo, L.; Li, K.; Xiong, P.; Cai, W. Diagnostic Fragment-Ion-Based for Rapid Identification of Chlorogenic Acids Derivatives in Inula Cappa Using UHPLC-Q-Exactive Orbitrap Mass Spectrometry. J. Anal. Methods Chem. 2021, 2021, 6393246. [Google Scholar] [CrossRef] [PubMed]
- Clifford, M.N.; Johnston, K.L.; Knight, S.; Kuhnert, N. Hierarchical Scheme for LC-MSn Identification of Chlorogenic Acids. J. Agric. Food Chem. 2003, 51, 2900–2911. [Google Scholar] [CrossRef] [PubMed]
- Quéméner, B.; Ralet, M.-C. Evidence for Linkage Position Determination in Known Feruloylated Mono- and Disaccharides Using Electrospray Ion Trap Mass Spectrometry. J. Mass Spectrom. 2004, 39, 1153–1160. [Google Scholar] [CrossRef]
- Xu, J.-G.; Hu, Q.-P.; Liu, Y. Antioxidant and DNA-Protective Activities of Chlorogenic Acid Isomers. J. Agric. Food Chem. 2012, 60, 11625–11630. [Google Scholar] [CrossRef]
- Touré, A.; Xueming, X. Flaxseed Lignans: Source, Biosynthesis, Metabolism, Antioxidant Activity, Bio-Active Components, and Health Benefits. Compr. Rev. Food Sci. Food Saf. 2010, 9, 261–269. [Google Scholar] [CrossRef]
- Agnieszka Loboda, E.R.-G.; Dulak, J. Targeting Nrf2-Mediated Gene Transcription by Triterpenoids and Their Derivatives. Biomol. Ther. 2012, 20, 499–505. [Google Scholar] [CrossRef]
- Gasser, P.; Arnold, F.; Peno-Mazzarino, L.; Bouzoud, D.; Luu, M.T.; Lati, E.; Mercier, M. Glycation Induction and Antiglycation Activity of Skin Care Ingredients on Living Human Skin Explants. Int. J. Cosmet. Sci. 2011, 33, 366–370. [Google Scholar] [CrossRef] [PubMed]
- Prockop, D.J. Mutations in Collagen Genes. Consequences for Rare and Common Diseases. J. Clin. Investig. 1985, 75, 783–787. [Google Scholar] [CrossRef]
- Huang, K.; Huang, J.; Xie, X.; Wang, S.; Chen, C.; Shen, X.; Liu, P.; Huang, H. Sirt1 Resists Advanced Glycation End Products-Induced Expressions of Fibronectin and TGF-Β1 by Activating the Nrf2/ARE Pathway in Glomerular Mesangial Cells. Free Radic. Biol. Med. 2013, 65, 528–540. [Google Scholar] [CrossRef]
- Tu, W.; Wang, H.; Li, S.; Liu, Q.; Sha, H. The Anti-Inflammatory and Anti-Oxidant Mechanisms of the Keap1/Nrf2/ARE Signaling Pathway in Chronic Diseases. Aging Dis. 2019, 10, 637–651. [Google Scholar] [CrossRef]
- Arias-Salvatierra, D.; Silbergeld, E.K.; Acosta-Saavedra, L.C.; Calderon-Aranda, E.S. Role of Nitric Oxide Produced by INOS through NF-ΚB Pathway in Migration of Cerebellar Granule Neurons Induced by Lipopolysaccharide. Cell. Signal. 2011, 23, 425–435. [Google Scholar] [CrossRef]
- Dong, H.; Zheng, L.; Yu, P.; Jiang, Q.; Wu, Y.; Huang, C.; Yin, B. Characterization and Application of Lignin–Carbohydrate Complexes from Lignocellulosic Materials as Antioxidants for Scavenging In Vitro and In Vivo Reactive Oxygen Species. ACS Sustain. Chem. Eng. 2020, 8, 256–266. [Google Scholar] [CrossRef]
- Pei, W.; Deng, J.; Wang, P.; Wang, X.; Zheng, L.; Zhang, Y.; Huang, C. Sustainable Lignin and Lignin-Derived Compounds as Potential Therapeutic Agents for Degenerative Orthopaedic Diseases: A Systemic Review. Int. J. Biol. Macromol. 2022, 212, 547–560. [Google Scholar] [CrossRef]
- Hemalatha, T.; Pulavendran, S.; Balachandran, C.; Manohar, B.M.; Puvanakrishnan, R. Arjunolic Acid: A Novel Phytomedicine with Multifunctional Therapeutic Applications. Indian J. Exp. Biol. 2010, 48, 238–247. [Google Scholar] [PubMed]
- Nagoor Meeran, M.F.; Goyal, S.N.; Suchal, K.; Sharma, C.; Patil, C.R.; Ojha, S.K. Pharmacological Properties, Molecular Mechanisms, and Pharmaceutical Development of Asiatic Acid: A Pentacyclic Triterpenoid of Therapeutic Promise. Front. Pharmacol. 2018, 9, 892. [Google Scholar] [CrossRef]
- Lozano-Mena, G.; Sánchez-González, M.; Juan, M.E.; Planas, J.M. Maslinic Acid, a Natural Phytoalexin-Type Triterpene from Olives—A Promising Nutraceutical? Molecules 2014, 19, 11538–11559. [Google Scholar] [CrossRef] [PubMed]
- Santiago, L.; Dayrit, K.; Correa, P.; Mayor, A.B. Comparison of Antioxidant and Free Radical Scavenging Activity of Triterpenes α-Amyrin, Oleanolic Acid and Ursolic Acid. J. Nat. Prod. 2014, 7, 29–36. [Google Scholar]
- Samsonowicz, M.; Kalinowska, M.; Gryko, K. Enhanced Antioxidant Activity of Ursolic Acid by Complexation with Copper (II): Experimental and Theoretical Study. Materials 2021, 14, E264. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Pan, X.; Jiang, L.; Chu, Y.; Gao, S.; Jiang, X.; Zhang, Y.; Chen, Y.; Luo, S.; Peng, C. The Biological Activity Mechanism of Chlorogenic Acid and Its Applications in Food Industry: A Review. Front. Nutr. 2022, 9. [Google Scholar] [CrossRef]
- Tonelli, C.; Chio, I.I.C.; Tuveson, D.A. Transcriptional Regulation by Nrf2. Antioxid. Redox Signal. 2018, 29, 1727–1745. [Google Scholar] [CrossRef] [PubMed]
- Hayes, J.D.; Dinkova-Kostova, A.T. The Nrf2 Regulatory Network Provides an Interface between Redox and Intermediary Metabolism. Trends Biochem. Sci. 2014, 39, 199–218. [Google Scholar] [CrossRef] [PubMed]
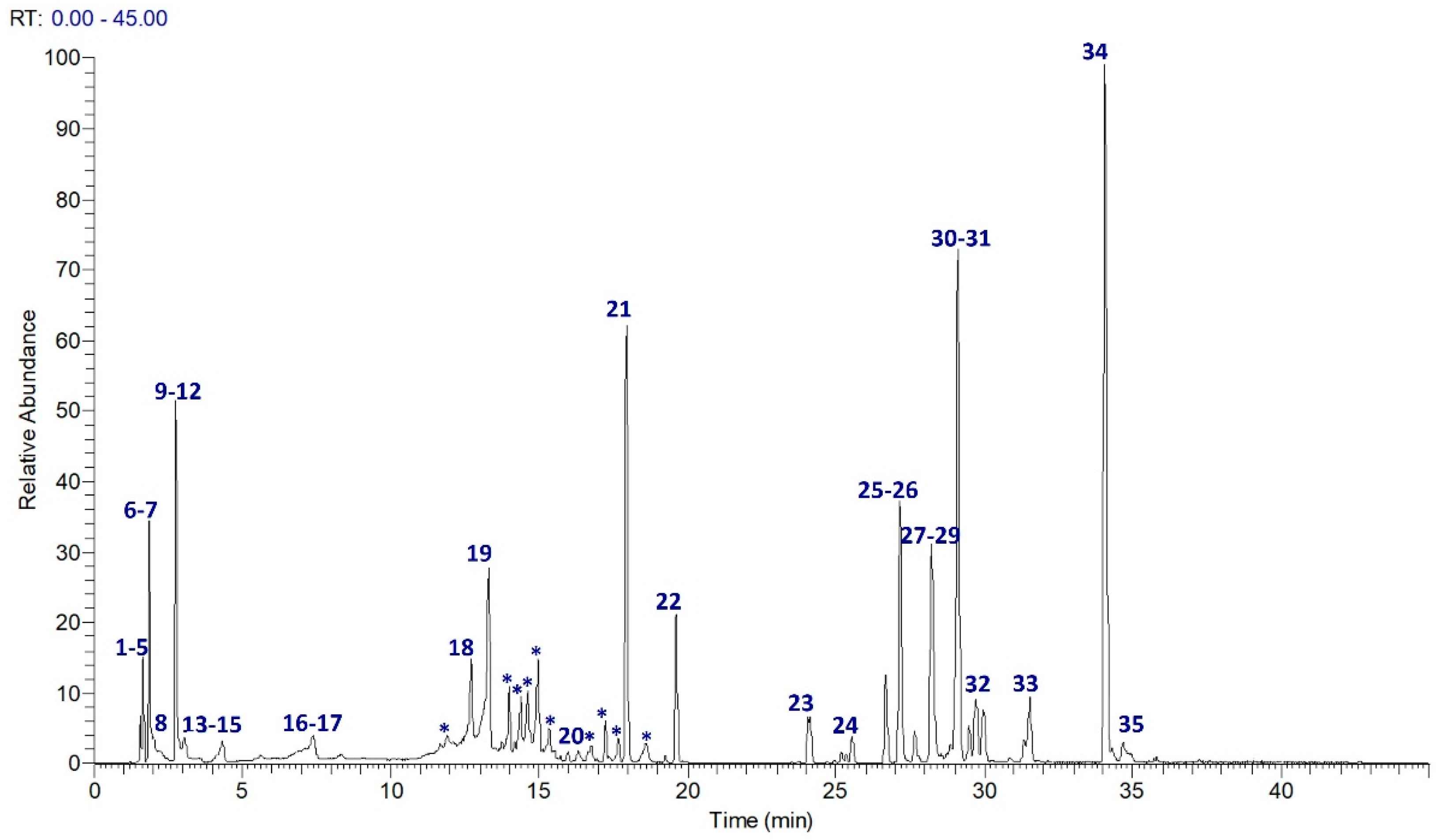

| Compound | MF (Mass Error ppm) | RT min | Precursor Ions m/z | MS2 Ions m/z (Relative Intensity %) | |
|---|---|---|---|---|---|
| 1 | L-Histidine | C6H9N3O2 (2.60 ppm) | 1.49 | 154.0615 | 137.0346 (37.67); 110.0712 (16.55); 93.0446 (40.25) |
| 2 | Pyridoxine | C8H11NO3 (2.98 ppm) | 1.56 | 168.0660 | 150.0551 (100); 122.0600 (46.71) |
| 3 | Pyridoxal | C8H9NO3 (3.01 ppm) | 1.56 | 166.0504 | 138.0551 (100); 108.0443 (56.53) |
| 4 | L-Asparagine | C4H8N2O3 (1.53 ppm) | 1.64 | 131.0453 | 114.0185 (100); 95.0239 (21.51); 70.0286 (38.76) |
| 5 | L-Glutamine | C5H10N2O3 (2.07 ppm) | 1.65 | 145.0611 | 127.0502 (93.52); 109.0396 (31.30); 84.0442 (21.65) |
| 6 | L-glutamic acid | C5H9NO4 (1.37 ppm) | 1.81 | 146.0450 | 128.0342 (52.95); 102.0548 (100) |
| 7 | Uridine | C9H12N2O6 (5.35 ppm) | 1.85 | 243.0625 | 200.0558 (9.58); 110.0236 (100) |
| 8 | Quinic acid | C7H12O6 (4.19 ppm) | 1.97 | 191.0558 | 173.0445 (2.07); 127.0389 (3.40) |
| 9 | Niacin | C6H5NO2 (0.82 ppm) | 2.73 | 122.0238 | 94.0286 (5.06); 78.0336 (3.99) |
| 10 | L-Tyrosine | C9H11NO3 (3.89 ppm) | 2.73 | 180.0662 | 163.0392 (100); 119.0491 (67.29); 93.0334 (24.76) |
| 11 | Malic acid | C4H6O5 (2.26 ppm) | 2.75 | 133.0134 | 115.0025 (100); 71.0126 (45.50) |
| 12 | Citric acid | C6H8O7 (4.19 ppm) | 2.82 | 191.0194 | 111.0076 (11.67); 85.0282 (100) |
| 13 | Guanosine | C10H13N5O5 (6.38 ppm) | 3.01 | 282.0851 | 150.0411 (100); 133.0145 (7.54) |
| 14 | Adenosine | C10H13N5O4 (6.39 ppm) | 3.23 | 266.0901 | 134.0461 (100) |
| 15 | L-Phenylalanine | C9H11NO2 (3.05 ppm) | 4.34 | 164.0711 | 147.0442 (100); 72.0079 (34.30) |
| 16 | Pantothenic Acid | C9H17NO5 (5.04 ppm) | 7.10 | 218.1034 | 146.0813 (73.48); 88.0392 (100) |
| 17 | L-Tryptophan | C11H12N2O2 (4.43 ppm) | 7.37 | 203.0824 | 142.0652 (27.99); 116.0494 (100); 74.0235 (43.29) |
| 18 | Caffeoylated monosaccharides [22] | C15H18O9 (5.28 ppm) | 11.67; 11.91; 12.70; 13.29 | 341.0885 | 179.0342; 161.0235; 135.0441 |
| 19 | Coumaroylated disaccharides | C21H28O13 (4.52 ppm) | 13.11; 13.54; 13.74; 14.00 | 487.1468 | 307.0828; 163.0392; 145.0285 |
| 20 | ‡ Secoisolariciresinol | C20H26O6 (6.09 ppm) | 16.76 | 361.1668 | 346.1422 (37.86); 179.0706 (26.43); 165.0548 (100) |
| 21 | ‡ Nortrachelogenin | C20H22O7 (4.82 ppm) | 17.94 | 373.1300 | 327.1242 (6.65); 312.1009 (3.98); 147.0442 (10.56) |
| 22 | ‡ Matairesinol | C20H22O6 (5.04 ppm) | 19.62 | 357.1351 | 342.1109 (8.61); 209.0816 (4.25); 122.0362 (14.23) |
| 23 | ‡ Arjunolic acid/Asiatic acid | C30H48O5 (4.72 ppm) | 24.09 | 487.3441 | 421.3155 (0.11); 409.3109 (0.44) |
| 24 | Glycerophosphocoline (18:3) | C26H48NO7P (4.98 ppm) | 25.69 | 562.3167 (M+HCOOH-H)- | 277.2173 (100); 224.0690 (13.23) |
| 25 | Glycerophosphoethanolamine (18:2) | C23H44NO7P (4.20 ppm) | 26.69; 27.17 | 476.2792 | 279.2328; 196.0375 |
| 26 | Glycerophosphocoline (18:2) | C26H50NO7P (4.08 ppm) | 26.75; 27.24 | 564.3319 (M+HCOOH-H)- | 279.2330; 224.0691 |
| 27 | Glycoglycerolipid (18:3) | C27H46O9 (3.90 ppm) | 27.78 | 513.3078 | 277.2171 (100); 253.0928 (5.07) |
| 28 | Glycerophosphoethanolamine (16:0) | C21H44NO7P (3.76 ppm) | 27.66; 28.20 | 452.2789 | 255.2328; 196.0374 |
| 29 | Glycerophosphocoline (16:0) | C24H50NO7P (3.89 ppm) | 27.73; 28.32 | 540.3317 (M+HCOOH-H)- | 255.2328; 224.0689 |
| 30 | Glycerophosphoethanolamine (18:1) | C23H46NO7P (4.39 ppm) | 28.54; 29.06 | 478.2949 | 281.2485; 196.0374 |
| 31 | Hydroxyoctadecadienoic acid | C18H32O3 (5.08 ppm) | 29.12 | 295.2283 | 277.2173 (100); 195.1384 (26.92); 171.1019 (45.17) |
| 32 | ‡ Maslinic acid | C30H48O4 (4.24 ppm) | 29.71 | 471.3489 | 423.3274 (0.39) |
| 33 | Glycerophosphoethanolamine (18:0) | C23H48NO7P (4.16 ppm) | 31.44 | 480.3105 | 283.2642 (100); 196.0373 (10.91) |
| 34 | Octadecatrienoic acid | C18H30O2 (5.41 ppm) | 34.05 | 277.2177 | 259.2071 (1.72) |
| 35 | ‡ Oleanolic acid/ Ursolic acid | C30H48O3 (5.05 ppm) | 34.93 | 455.3543 | 407.3312 (0.07) |
| Compound | Range (nM) | Calibration Curve | R2 | LOD (nM) | LOQ (nM) |
|---|---|---|---|---|---|
| Secoisolariciresinol | 50–25,000 | y = 2.00E + 08 x | 0.9951 | 3 | 10 |
| Nortrachelogenin | 50–25,000 | y = 2.00E + 08 x | 0.9938 | 15 | 50 |
| Matairesinol | 50–25,000 | y = 3.00E + 08 x | 0.9917 | 3 | 10 |
| Arjunolic acid | 100–25,000 | y = 2.00E + 07 x | 0.9929 | 15 | 50 |
| Asiatic acid | 100–25,000 | y = 2.00E + 07 x | 0.9865 | 8 | 25 |
| Maslinic acid | 100–25,000 | y = 4.00E + 07 x | 0.9957 | 8 | 25 |
| Oleanolic acid | 100–250,000 | y = 2.00E + 07 x | 0.9953 | 0.3 | 1 |
| Ursolic acid | 100–250,000 | y = 1.00E + 07 x | 0.9935 | 0.3 | 1 |
| Compound | Amount (µg/g of Extract) | % RSD |
|---|---|---|
| Secoisolariciresinol | 0.67 | 0.94 |
| Nortrachelogenin | 39.96 | 2.46 |
| Matairesinol | 6.75 | 0.65 |
| Arjunolic acid | 235.80 | 0.97 |
| Asiatic acid | 336.71 | 2.55 |
| Maslinic acid | 144.02 | 2.82 |
| Oleanolic acid | 214.00 | 3.57 |
| Ursolic acid | 542.80 | 2.17 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ceccacci, S.; Lucia, A.D.; Tortora, A.; Colantuono, A.; Carotenuto, G.; Tito, A.; Monti, M.C. Jasminum sambac Cell Extract as Antioxidant Booster against Skin Aging. Antioxidants 2022, 11, 2409. https://doi.org/10.3390/antiox11122409
Ceccacci S, Lucia AD, Tortora A, Colantuono A, Carotenuto G, Tito A, Monti MC. Jasminum sambac Cell Extract as Antioxidant Booster against Skin Aging. Antioxidants. 2022; 11(12):2409. https://doi.org/10.3390/antiox11122409
Chicago/Turabian StyleCeccacci, Sara, Adriana De Lucia, Assunta Tortora, Antonio Colantuono, Gennaro Carotenuto, Annalisa Tito, and Maria Chiara Monti. 2022. "Jasminum sambac Cell Extract as Antioxidant Booster against Skin Aging" Antioxidants 11, no. 12: 2409. https://doi.org/10.3390/antiox11122409
APA StyleCeccacci, S., Lucia, A. D., Tortora, A., Colantuono, A., Carotenuto, G., Tito, A., & Monti, M. C. (2022). Jasminum sambac Cell Extract as Antioxidant Booster against Skin Aging. Antioxidants, 11(12), 2409. https://doi.org/10.3390/antiox11122409







