1. Introduction
The human brain is an extremely large-scale network of neurons, whose function depends on mutual interactions continuously occurring between different sources of neuronal activity. Characterization of the brain function is often conducted through the analysis of functional connectivity, which reflects temporal correlations between the brain dynamics of spatially separated regions, and effective connectivity, which describes networks of directional effects of one neural element over another [
1]. This analysis is applied in particular through processing of electroencephalogram (EEG) signals, which record voltage differences between various locations on the head surface, through measuring the strength of interactions between close or remote brain areas. Brain connectivity analysis is often performed to investigate functional alterations of the brain activity in a wide variety of neurological disorders [
2,
3,
4]. In this context, the evaluation of connectivity patterns is of particular relevance to disorders of the central nervous system related to epilepsy, as seizure attacks typically produce redundant hyper-synchronous activity of neurons in the brain [
5]. In fact, estimating connectivity in the epileptic brain may lead to a better understanding of the occurrence and spreading of epileptiform activity, and is of relevance for different research and clinical applications [
3,
6,
7]. The importance of such approach is even more strengthened by recent increased evidence of seizure onset not in the entire brain (generalized seizures) or in a circumscribed region of the brain (focal seizures), but within a network of brain regions (the so-called “epileptic network” composed of several functionally connected cortical and subcortical brain structures and regions) [
8].
The assessment of brain connectivity is a challenging issue which has been addressed in the literature using a wide range of signal processing techniques including time domain methods, time/frequency approaches, parametric models, phase synchronization and nonlinear measures [
9,
10,
11]. Among them, the use of vector autoregressive (VAR) models is very popular, thanks to the simplicity of the linear parametric representation, to its flexibility (no assumption is made about the underlying neural mechanisms), and to the possibility to exploit it in the assessment of coupling and directed interactions in time, frequency and information-theoretic domains [
12,
13]. In particular, the connection of the VAR representation to the ubiquitous concept of Granger causality (GC) and to extensions of this concept developed in the framework of information dynamics [
14,
15] makes this approach eligible for the causal analysis of functional brain connectivity in several applicative contexts including the characterization of epileptic activity [
3]. It is worth remarking that ‘causal’ interpretations of Granger causality measures must be done cautiously, as these measures reflect a statistical and purely data-driven concept of causality. In particular, GC measures are designed to reflect the causal effect that the underlying physical mechanisms have on the analyzed neuroimaging data, and neither functional nor effective connectivity representations necessarily map univocally onto the underlying anatomical (structural) connectivity [
16].
The simplest and most intuitive way to study brain connectivity is to apply connectivity measures directly on the signals recorded from the scalp EEG sensors. However, scalp-level brain connectivity analysis suffers from several problems, including the fact that the position of scalp EEG electrodes do not relate trivially to the location of the underlying sources [
17], thus becoming mostly sensitive to activity correlated over large areas of the superficial cortical surface, with smaller contributions from deeper sources, the effects on the scalp recordings of the so-called normal EEG background consisting in a common electrical activity shared together with presumably different activity in comparable EEG classes, and the presence of recording artifacts which are generally assumed to be components of extracerebral origin [
18,
19]. Besides that, the scalp EEG signals are influenced by the well-known problem of volume conduction, which refers to the mixing effects which result from measuring electrical potentials at a distance from their source generators [
20]. Due to volume conduction, signals originating from the same source in the brain are detected by several EEG electrodes. As a result, VAR-based connectivity measures applied straightly to scalp EEG signals can lead to the detection of spurious causality and more generally of wrong connectivity patterns [
21,
22].
A solution to limit the adverse effects of volume conduction on EEG connectivity analysis is to transfer the connectivity estimation problem from the scalp sensor domain to the cortical source domain [
23]. This is typically achieved first reconstructing the source electrical activity and then applying the desired connectivity metrics on the estimated source time series. Although it is acknowledged that this approach is useful to limit the issue of volume conduction, it has also been criticized or even defined “elusive” in the past due to the lack of uniqueness of the reconstructed sources (e.g., depending on the method applied to reconstruct the sources, or the number of sources employed, or even on the portion of the time series used) since the relationship between each notion of functional connectivity and its underlying neural substrate is still unknown [
24]. Source reconstruction can be based on source localization methods, which allow spatial analysis but require accurate models of anatomy and electrical properties of the head [
25], or by blind decomposition methods such as Principal Component Analysis (PCA) or Independent Component Analysis (ICA), which do not require head modeling and yield source signals which can be interpreted as originating from cortical dipoles [
26]. A more accurate approach, which combines the steps of source signal estimation and connectivity analysis, is that proposed in [
27] who integrated ICA source reconstruction and VAR connectivity analysis. This approach, known as VARICA source connectivity analysis, has been further refined through its integration with the Common Spatial Patterns (CSP) technique for dimensionality reduction, leading to the CSPVARICA method [
28]. The use of CSP allows to retain only the EEG components which explain the difference in variance between two conditions, and it is thus particularly useful when differences between states (e.g., baseline/task or pre/post) are analyzed. The present work introduces a modified implementation of the SCoT toolbox for CSPVARICA EEG source connectivity analysis [
28] and reports its evaluation on simulations of interacting VAR cortical source processes mixed to obtain synthetic EEG signals, as well as its application to the characterization of brain connectivity in children manifesting episodes of generalized or focal seizures. The new implementation allows, besides the reconstruction of source EEG activities from scalp recordings based on improved CSP and ICA algorithms, the computation of the measures of information dynamics (information storage and information transfer with statistical significance assessment) performed in both sensor and source domains. Moreover, the comparison of scalp and cortical source information dynamics computed in epileptic children leads us to make inferences about the reorganization of the brain connectivity networks that occurs before and after epileptic seizures, distinguishing between generalized or focal seizures and addressing the role of volume conduction.
The algorithms of the framework for the quantification of EEG source information dynamics and evaluation on simulations presented in this work are collected and can be reproduced within the CSPVARICA Information Dynamics MATLAB toolbox, which is available as supplementary material of this article and can be freely downloaded from www.lucafaes.net/CSPVARICAInfoDyn.html.
2. Framework for the Quantification of EEG Source Information Dynamics
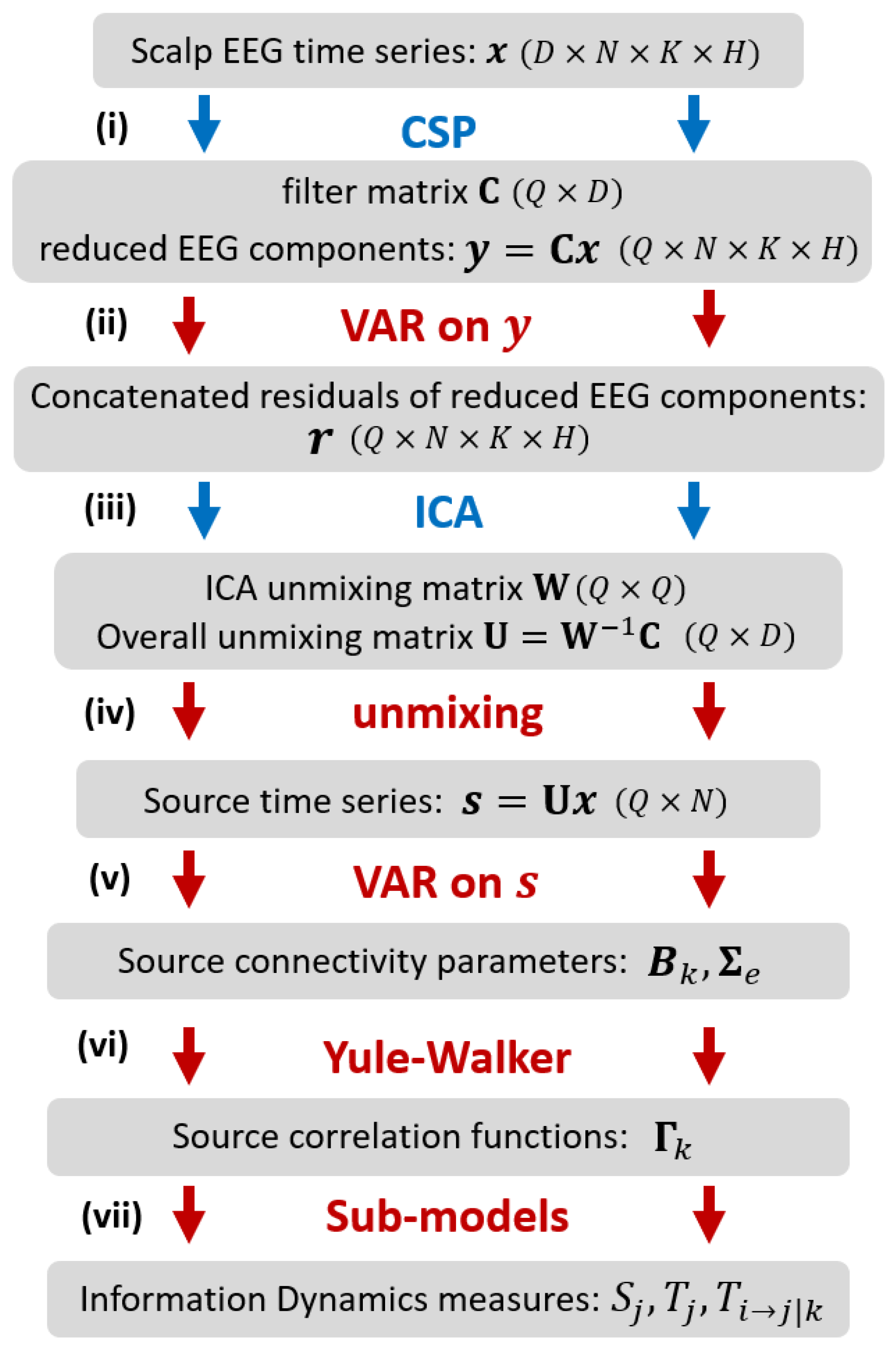
The methodology proposed in our work for assessing information dynamics is illustrated in
Figure 1. Considering two experimental conditions (classes
) and a given number of trials per condition (
), the starting point is a dataset of EEG time series of length
N acquired from
D scalp sensors for each trial and condition. The procedure consists of the following steps: (i) application of CSP [
29] to select, across all trials, the
Q reduced EEG components which better explain the difference in variance between the two conditions; (ii) VAR modeling [
13] of the reduced components, performed at a single trial level, to describe the time-lagged linear interactions among the EEG components; instantaneous interactions not explained by lagged effects are retained in the VAR residuals; (iii) application of ICA [
26] to the VAR residuals, concatenated over trials and classes to get an overall description of instantaneous EEG influences related to volume conduction; ICA returns the unmixing matrix that relates the instantaneously independent cortical sources to the reduced EEG components; (iv) unmixing of the EEG time series to get the source time series for each single trial; this step exploits the overall unmixing matrix relating the source and scalp dynamics, which is obtained combining the ICA unmixing matrix and the CSP filter matrix [
28]; (v) VAR modeling of the source time series, performed at a single trial level, to describe the time-lagged linear interactions among the instantaneously independent EEG sources [
27]; (vi) inverse solution of the Yule-Walker Equations for the VAR source parameters, to reconstruct the autocorrelation sequence of the source time series [
14]; (vii) definition of sub-models for each predefined source time series, to describe its predictability from its own dynamics or from the dynamics of the other sources in terms of partial variances [
30], and subsequent computation of the information dynamics measures.
2.1. Theoretical Formulation
In a stochastic signal processing setting, a set of EEG signals recorded with sampling frequency
from
D locations in the scalp are represented as a realization of a
D-dimensional discrete time random process,
, where the time index
n identifies the sample acquired at time
. To describe volume conduction, the process is modeled as an instantaneous mixing of
Q cortical sources
according to the linear equation:
where
M is a mixing matrix of dimension
. In turn, the sources are modeled as a VAR model of the following form:
where
are
coefficient matrices describing the source interactions at lag
k,
p is the VAR model order, and
is a
Q-dimensional vector of white and independent non-Gaussian noise processes with
diagonal covariance matrix
.
According to the CSPVARICA method proposed in [
28], the redundancy of the scalp recordings can be reduced applying the CSP technique to a set of EEGs collected in two conditions over multiple trials. Specifically, CSP results in
D spatial filters stored as the rows of a square transformation matrix that relates the original EEG scalp signals
to the reduced components
, and explains the differences in variance between the two conditions. A subset of
Q spatial filters is then selected, according to the procedure detailed in
Section 2.2, to form the
filter matrix
C. Filtering the
D original EEG signals
with such matrix yields the
Q reduced components
which retain the largest part of the EEG variance between the two analyzed conditions [
31]. The reduced components can be easily related to the source signals
using the
mixing matrix
[
28]:
Then, given that
and applying the source model (Equation (
2)), it can be easily shown that the following VAR representation holds for the reduced components:
with coefficient matrices
, and residuals
. Note that, if
W is known or estimated, the source processes can be obtained from the knowledge of the transformation matrix
C as follows:
Thus, the overall
unmixing matrix that allows the reconstruction of the sources starting from the scalp EEG signals has the form:
When both the source time series
and the VAR parameters (
,
) are known, they can be used to estimate the connectivity measures in the framework of information dynamics. This can be done according to the approach proposed in [
32,
33], which first determines the autocovariance sequence of the source process (Equation (
2)), then arranges the elements of the autocovariance matrix to compute the so-called “partial variances” which reflect the unpredictability of one source given its own dynamics or the dynamics of the other sources, and finally computes network information measures exploiting the relation between (partial) variance and (conditional) entropy. Specifically, the first step is accomplished in solving the inverse Yule-Walker equations [
14] for the process (Equation (
2)):
where
is the
autocovariance matrix of the sources evaluated at lag
k, and
is the Kronecker product. The second step is accomplished implicitly forming a VAR submodel having the
source
as the target and predicting it as a linear combination of its
q past values
and possibly of the
q past values of one or more other sources. All these past values are included in the regression vector
V, and then the partial variance of
given
V, quantifying the variance of the prediction error of the linear regression of of
on
V, is obtained as [
32]:
where
is the variance of
,
,
is the covariance between
and
V,
, and
is the covariance of
V,
. After separating the
Q sources
to evidence the target
, the driver
and the other sources
, the formulation of Equation (
8) is exploited to compute the partial variances of
given its own past
when
, given its past and the past of
when
, and given the past of all
Q sources
when
. Finally, these partial variances are used to compute information-theoretic measures of information storage, information transfer and conditional information transfer as [
32]:
The information storage
(Equation (
9)) can be considered as a measure of self-predictability for the
jth source, quantifying the amount of information shared between the present and the past dynamics of the source process. The information transfer
(Equation (
9)), or total Granger causality, is a measure of the amount of information contained in the present state of the
jth source that can be predicted by the past states of all the other sources. Finally, the conditional information transfer
(Equation (
9)), or conditional Granger causality, is a measure of the amount of information contained in the present state of the
jth source that can be predicted by the past states of the
ith source above and beyond the information that can predicted from the other sources. More details about information dynamics measures can be found in [
33] and in references therein.
2.2. Practical Estimation
The starting point of our analysis is a set of EEG signals collected in two different classes with presumably different brain activity over multiple sets (or trials); for each class and trial, D EEG signals are observed simultaneously recording N samples per signal, so that a period of duration secs is covered. The analysis steps to estimate the measures of information dynamics at the level of the cortical sources are described in the following.
2.2.1. Common Spatial Patterns (CSP)
The first step is the application of CSP [
29,
31] in order to find, from the original EEG data matrix of dimension
, the reduced data matrix of dimension
, with
, which explains the largest part of the difference in variance between the two classes
and
, as well as the spatial filter matrix
C that relates these two data matrices (Equation (
3)). Denoting the number of trials belonging to the two classes as
and
, the CSP algorithm is applied to all the trials of the same class at once.
From the
data matrix
arranged to contain as rows the EEG signals acquired for the trial
k of the class
(
), CSP estimates the
covariance matrices:
where
denotes the sum of the diagonal elements of the covariance matrix. Then, a spatial filtering matrix
is defined having the spatial filters
as columns (
). This matrix is obtained using a joint diagonalization of the class-related covariance matrices
and
, achieved solving the following generalized eigenvalue problem [
31]:
where
is the eigenvalue associated to each spatial filter
, and is related to the discriminative power of the filter: large eigenvalues correspond to spatial filters providing high variance in one class but low variance in the other, and vice versa. Thus, the spatial filters help to discriminate between the two classes.
When used without any dimensionality reduction, the CSP algorithm exploits all the
D spatial filters contained in the matrix
. However, it is usual to reduce dimensionality selecting a subset containing the
most relevant filters; the pruned spatial filtering matrix
C, applied to the EEG data matrices, leads to the reduced components that contribute most to the total EEG variance (see
Figure 1 and Equation (
3)). In this work,
is pruned manipulating the spatial covariance matrices
used by the CSP algorithm in a rigorous way exploiting Riemannian geometry [
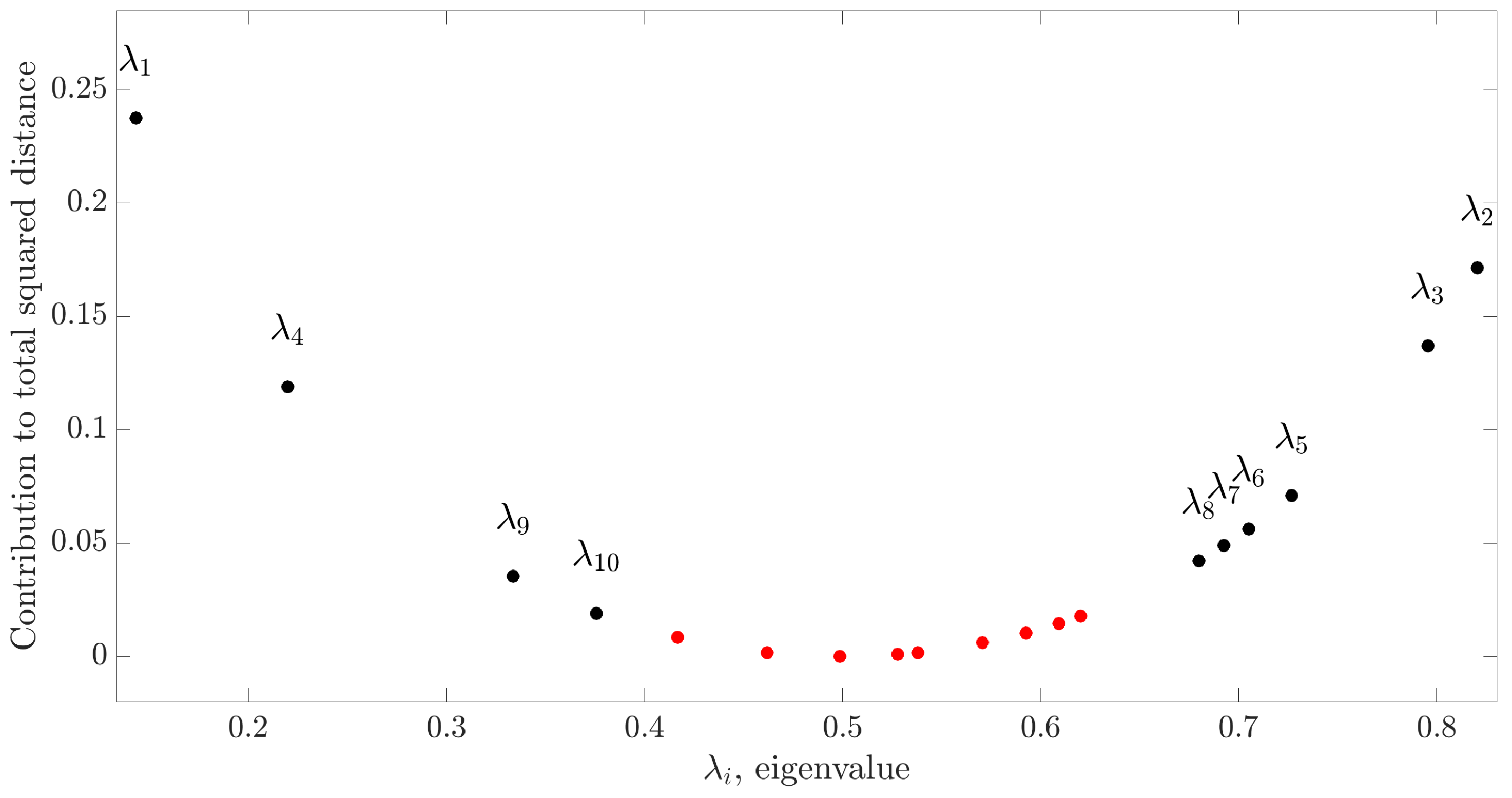
31]. This is achieved analyzing the contribution of the fractional distances
(
) to the total Riemannian distance between the two symmetric positive-definite covariance matrices
and
:
According to [
31], the fractional distance
and the percentage of total variance described by the spatial filter
directly depend on the corresponding eigenvalue
. Consequently, to choose the most appropriate number
Q of filters to discriminate the two classes, a viable criterion is to sum up the fractional distances in order to cover a predefined percentage of the total variance (greater than 90% in this study). After finding
Q and consequently the transformation matrix
C, we apply it to the EEG data
in order to reduce the dimensionality of the CSP-filtered signals
(
Figure 1, step (i)), and limit the number of sources.
2.2.2. VAR Modeling of Reduced EEG Components
The second analysis step consists in describing the interactions among the reduced EEG components in terms of VAR modeling. VAR identification is performed on each (
) data matrix
obtained for any trial
k of each class
(
). VAR identification allows to find estimates of the coefficients
and residuals
in Equation (
4), collected respectively in the
matrix
and in the
matrix
, after writing Equation (
4) in compact representation as:
where
and
(with
) are properly arranged observation matrices. Equation (
13) is typically solved through the multivariate least squares method [
13], finding the estimates
and
.
VAR identification is completed by selection of the model order and validation of the model assumption of whiteness of the residuals. In this work, the VAR model order is selected as the value of
p minimizing the cost function implemented by the Schwartz Bayesian Criterion (SBC) [
34]. Then, whiteness of the estimated residuals was tested using the multivariate Li-McLeod portmanteau test statistic Q [
35] which, under the null hypothesis of white residuals, returns a
p-value higher than 0.05 in case of whiteness [
28,
34].
2.2.3. Independent Component Analysis (ICA)
The VAR analysis performed on the CSP-filtered signals
explains the time-delayed cross-dependencies between these reduced EEG components but cannot describe their instantaneous (zero-lag) dependencies, which are thus retained in the residuals
. Such zero-lag dependencies are typically ascribed to volume conduction [
28], in a way such that the reduced EEG components result from the instantaneous mixing of the source signals and, following the VAR models (Equations (
2) and (
4)), the VAR residuals are subjected to the same mixing (i.e.,
and
). Therefore, under the assumption that the residuals
are non-Gaussian, ICA can be used to estimate the mixing matrix
W. In this work, the Infomax ICA algorithm included in EEGLab [
36,
37] was employed, which returns the estimate of the mixing matrix
W that minimizes the mutual information or maximizes the joint entropy between
and
in order to make them independent. The implemented algorithm is applied iteratively to overcome the randomness associated with the unsupervised estimation of unmixing matrix with ICA and thus to avoid the arbitrariness in the selection of the recovered sources [
38]. The decomposition of instantaneous source interactions achieved by ICA can be applied either separately to each set of residuals
R obtained solving (
13) for an individual EEG dataset, or collectively to the set of residuals concatenated for each trial and condition. While the first approach would return a different unmixing matrix for each trial and condition, in this work we adopted the second approach which assumes stationarity of the volume conduction effects across trials and conditions. This approach has the advantage of returning a single mixing matrix
W that is valid for each dataset.
2.2.4. Unmixing
After the decomposition of the residuals of the VAR model of the reduced EEG components by means of ICA, the whole unmixing matrix that relates the source and scalp signals (Equation (
5)) can be obtained combining the CSP filter matrix
C and the ICA unmixing matrix
simply as
(Equation (
6)). Then, the (
) data matrix of each set of source time series relevant to the trial
k of the class
is reconstructed applying the unmixing matrix to the corresponding set of scalp EEG signals, i.e.,
(
).
2.2.5. VAR Modeling of Source Signals
Once the source signals are estimated, their connectivity structure can be assessed from the VAR representation of Equation (
2). Note that, since such representation is linked to that of Equation (
4), there is no need to identify again a VAR model on the source time series; rather, the VAR parameters
and
can be obtained from the parameters
and
estimated from the reduced EEG components individually for each trial and condition, and from the mixing matrix
W estimated collectively for all trials and conditions, as
and
.
2.2.6. Yule-Walker Inverse Solution
The estimated parameters
and
are then exploited to compute the autocorrelation structure of the source signals through the solution of Equation (
7). Specifically, the correlation matrices
are estimated expressing Equation (
7) in a compact form corresponding to a discrete-time Lyapunov equation (see [
30,
32] for a detailed treatment). Then, the correlation matrices
can be obtained for any arbitrary lag
through iterative application of Equation (
7). In this work, the iteration was repeated up to a lag
to form the covariance and cross-covariance matrices appearing in Equation (
8), which are in turn used for the estimation of information dynamics.
2.2.7. Computation of Information Measures from VAR Sub-Models
The final step of the analysis pipeline is the computation of the measures of information dynamics from the VAR correlation structure of the EEG cortical sources. This step is performed, given a trial
k of a class
or
, for each selected pair of driver and target signals
and
and grouping the other drivers in the vector signal
, forming the submodels that describe
either from
, or from
and
, or from
and
. In either case, the prediction error variance (partial variance) relevant to each submodel is identified- without the need to perform the regression- exploiting Equation (
8), where the covariances appearing in the right-hand side of the equation are obtained arranging the correlation matrices found in the previous step. Finally, the partial variances are used to compute the measures of information storage
(Equation (
9)), information transfer
(Equation (
9)), and conditional information transfer
(Equation (
9)).
In this work we have also assessed the significant directed links between pairs of nodes in the network of source interactions, which can be defined taking the statistically significant values of the conditional information transfer [
14,
33]. To do this, we applied the Fisher F-test to compare the two nested regression models used in the computation of
, i.e., the full model describing
from
and the submodel describing
from
. The test statistic is computed from the partial variances
and
, and is compared with the
-th percentile of the Fisher distribution with
q and
degrees of freedom to reject or confirm the null hypothesis of equal partial variance between the full model and the submodel (here, a significance
was assumed).
3. Simulation Study
The proposed framework is first tested over simulated EEG signals. In the simulation, two different experimental scenarios (conditions) are designed to reproduce the dynamics of five cortical sources displaying alpha or beta rhythms, which are isolated in the first condition and interact realizing the propagation of alpha waves in the second condition. For each condition we generate, assuming a sampling frequency of 125 Hz, a VAR process of order
composed by
source processes, in which the diagonal elements of the matrix
are set to reproduce autonomous oscillations in each scalar source process, and the off-diagonal elements are set to impose specific connectivity patterns (see Ref. [
13] for details on the procedure to generate theoretical VAR processes). Specifically, in the first condition autonomous oscillations are set in the alpha band (center frequency
Hz, narrowband) for
,
, and
, and in the beta band (center frequency
Hz, broadband) for
and
; all off-diagonal elements of
are set to 0 in order to reproduce absence of connectivity. In the second condition, the autonomous oscillations are set in the alpha band for
and in the beta band for
,
,
, and
; in this case causal effects are imposed along the chain
, by setting
in order to reproduce the propagation of the alpha rhythm from
to
. Note that, given the theoretical values imposed for the VAR parameters, the exact values of the measures of information dynamics can be computed, and are used here as the ground truth for evaluating the performance of the algorithms. Then, 10 realizations (trials) of the source time series of length
samples are obtained for each condition feeding the simulated VAR model with non-gaussian innovations; the non-gaussian noise is generated applying the nonlinear transformation
to a gaussian white noise
(
to yield super-Gaussian distributions [
39]). The source time series are then multiplied with a mixing matrix
M to obtain the simulated scalp EEG signals (Equation (
1)). We choose a square mixing matrix (
) to allow the unmixing matrix to be full rank and thus invertible; the entries of
M are chosen to simulate the volume conduction effect of a source over two or three electrodes:
The analysis of information dynamics is first computed at the scalp level, i.e., identifying the VAR model directly on the
simulated EEG signals and computing the information measures from the estimated VAR parameters (steps (v, vi, vii) of the analysis, see
Figure 1). Then, the analysis is repeated applying the whole framework, i.e., applying in sequence CSP, VAR identification, ICA, reconstruction of the source model and computation of the information measures according to the whole pipeline depicted in
Figure 1.
Results are shown in
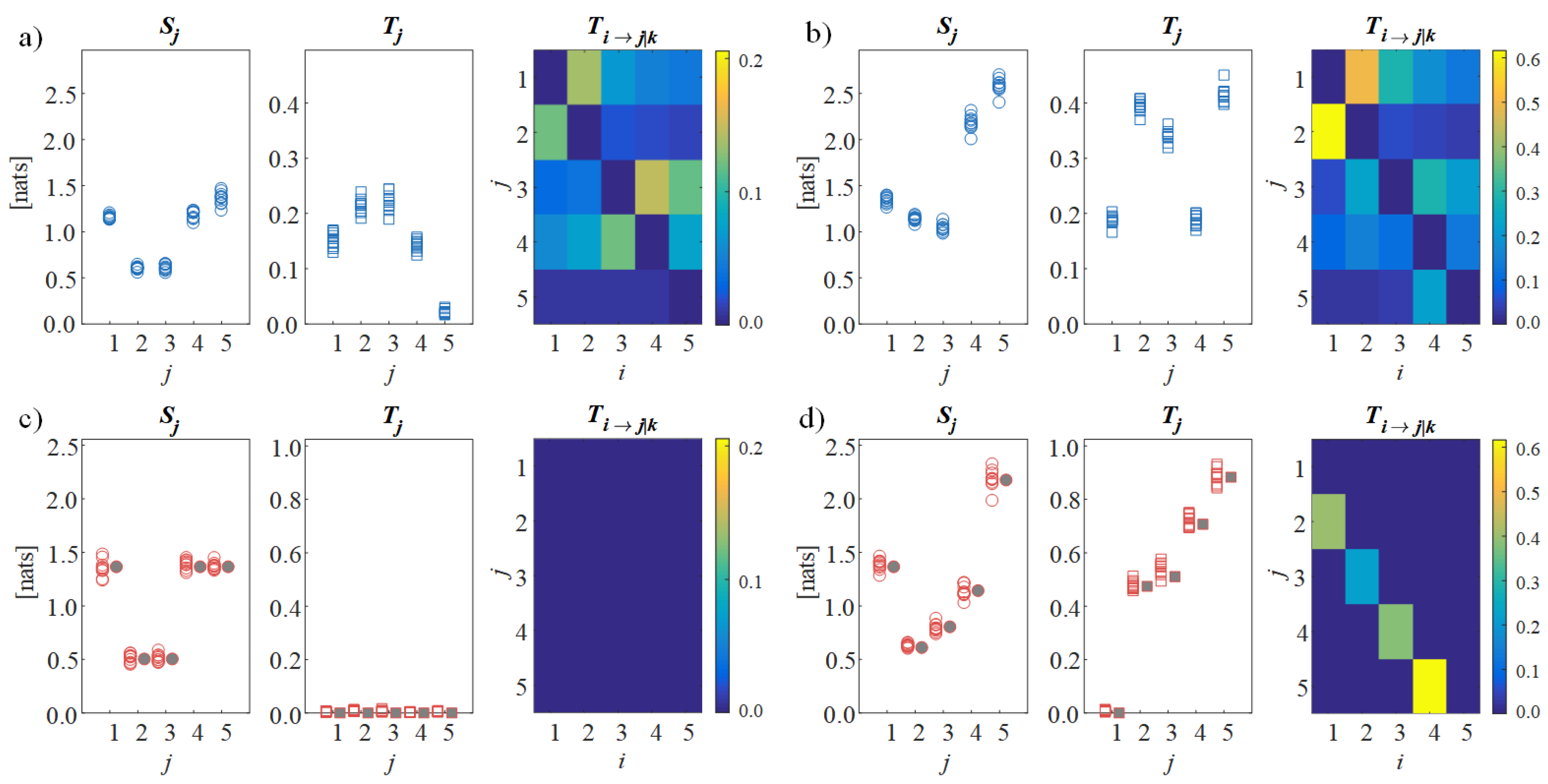
Figure 2, which depicts the values of the information measures computed from the simulated scalp signals
and from the reconstructed source signals
in the first condition of non-interacting sources
and in the second condition with directed connectivity among sources
. The analysis on the scalp signals shows that, even in the first condition with absence of causal interactions between the sources, the total and conditional information transfer assessed for the simulated EEG signals is substantial (high values of
and
in
Figure 2a), providing a false indication of directed connectivity. On the contrary, when the proposed CSPVARICA approach is applied, the lack of connectivity is clearly documented by the null values of the information transfer to each source and of the directed transfer between pairs of sources (null values of
and
in
Figure 2c). In the second condition, the scalp analysis remains confounding as the values of information transfer are scattered across the connectivity matrix (
Figure 2b), while the source analysis reveals the ability of the proposed framework to elicit the directional interactions that are imposed between the sources (i.e., significant directed connectivity from
to
, from
to
, from
to
, and from
to
, highlighted by the high values of
,
Figure 2d). Remarkably, the distributions across trials of information storage and information transfer are well aligned with the theoretical values resulting from the true VAR source parameters (filled symbols in
Figure 2c,d), thus documenting the accuracy of the reconstruction.
5. Discussion
The analysis of information dynamics, and in particular of the measures of information storage and information transfer, plays a fundamental role in neural computation, especially with regard to the network representation of the electrical activity of the brain [
45,
46]. The estimation of information dynamics, in particular information transfer, from scalp EEG signals is severely complicated by volume conduction effects which may dominate the transfer of information across EEG sensors and therefore blur the detection of meaningful patterns of information flow [
21,
22,
47]. While solutions to this problem have been proposed which keep working with scalp signals and try compensating volume conduction [
45,
47,
48], a more intuitive approach is to work on the source signals reconstructed from the sensor activity since it has been demonstrated that the source-based network representation constitutes a better approximation of the unknown true network structure [
17]. In this work we pursue this approach, integrating it into existing approaches for the linear parametric representation of the dynamics of cortical sources and of their mixing to produce scalp EEG signals [
27,
28]. The resulting framework constitutes, to the best of our knowledge, the first attempt to access the information dynamics of EEG sources through simultaneous modeling of time-lagged source interactions and instantaneous mixing effects. This is accomplished combining CSP which allows reducing the dimensionality of EEG signals while taking into account nearly the same amount of information in terms of variability, VAR for combined source and scalp EEG modeling, and ICA for obtaining instantaneously independent cortical sources.
The application of the proposed framework to a neural disorder characterized by abrupt changes in the dynamical state of the brain, i.e., epileptic seizures, led us to highlight the usefulness of source analysis of information dynamics in comparison to scalp analysis. In particular, we found similar information-theoretic measures computed from scalp and source signals in the case of generalized seizures, while differences between the two approaches were revealed for focal seizures. The different results are likely due to the characteristics of brain activity for different seizure types [
49]. In the case of generalized seizures, the entire brain volume is entangled in the seizure activity, which is thus observed everywhere in the brain; as a consequence, one may expect that volume conduction does not alter substantially the patterns of interaction among sources, which should be observed similarly at the level of the scalp. On the contrary, in focal epileptic seizures, the activity is more localized in a brain subvolume nearby the seizure origin, and volume conduction causes its spreading to other locations and appearance at scalp sensors far away from the actual seizure focus location; this is expected to have an effect on how connectivity patterns are detected at the scalp compared with the source level. In fact, analyzing focal seizure episodes we did not detect any remarkable variation across conditions in the measures of information dynamics computed at the scalp, while the changes in the information transfer were noticed when considering the EEG sources. Remarkably, the profiles across conditions of the measure of information storage were similar when analyzing scalp and sensor signals, also for the focal seizures. These results are in line with previous findings [
47] evidencing that, in response to an action which evokes localized occipital brain reactions like the eyes closure, information storage is informative also when computed at the scalp level, likely because of its univariate nature, while information transfer is blurred to uniform values due to the adverse effects of volume conduction on connectivity measures. To highlight the importance of a distinction between different type of seizures, we remark that differences have been evidenced also in relation to autonomic nervous regulation, e.g., probed by measures (including information-theoretic ones) of heart rate variability computed before and after focal and generalized seizures [
40].
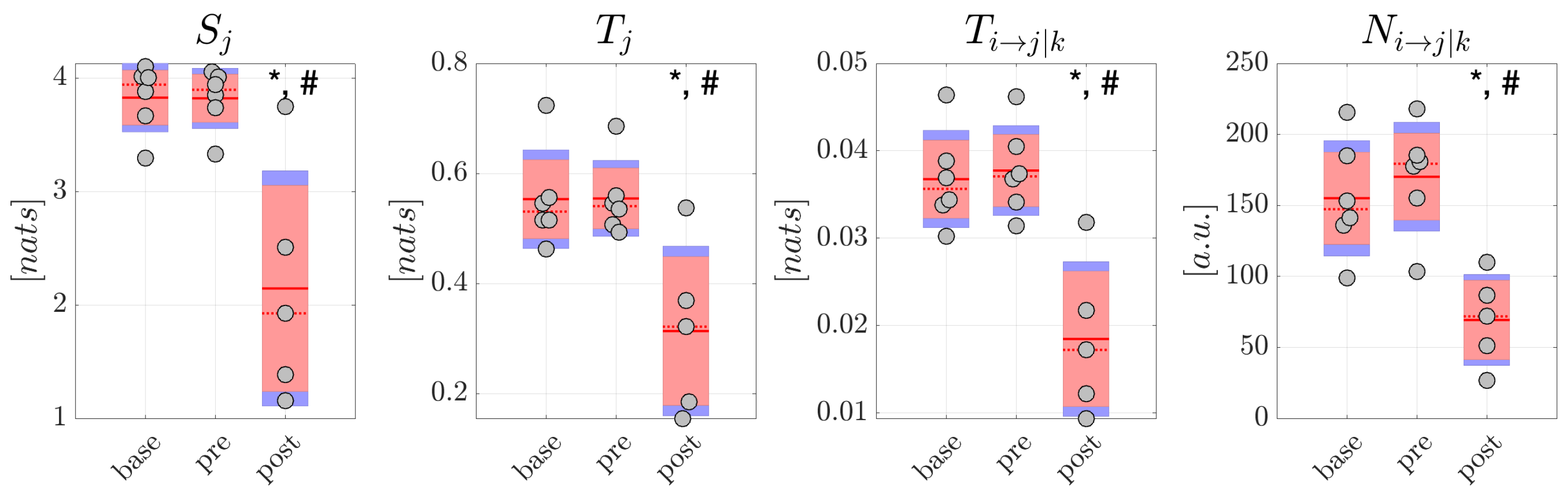
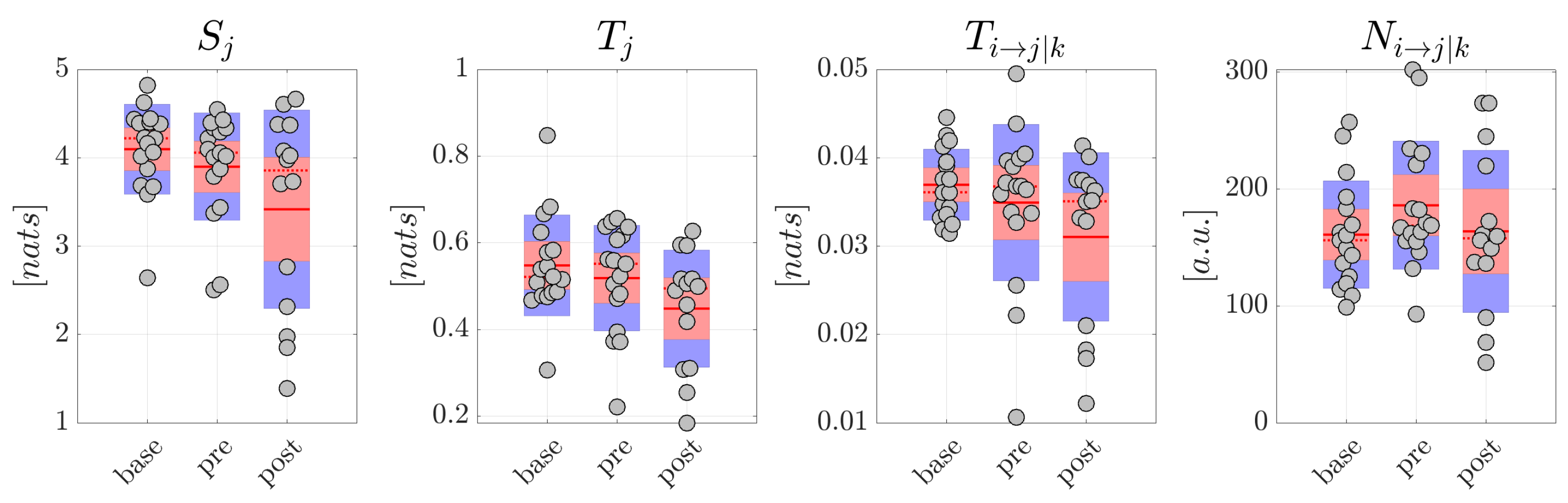
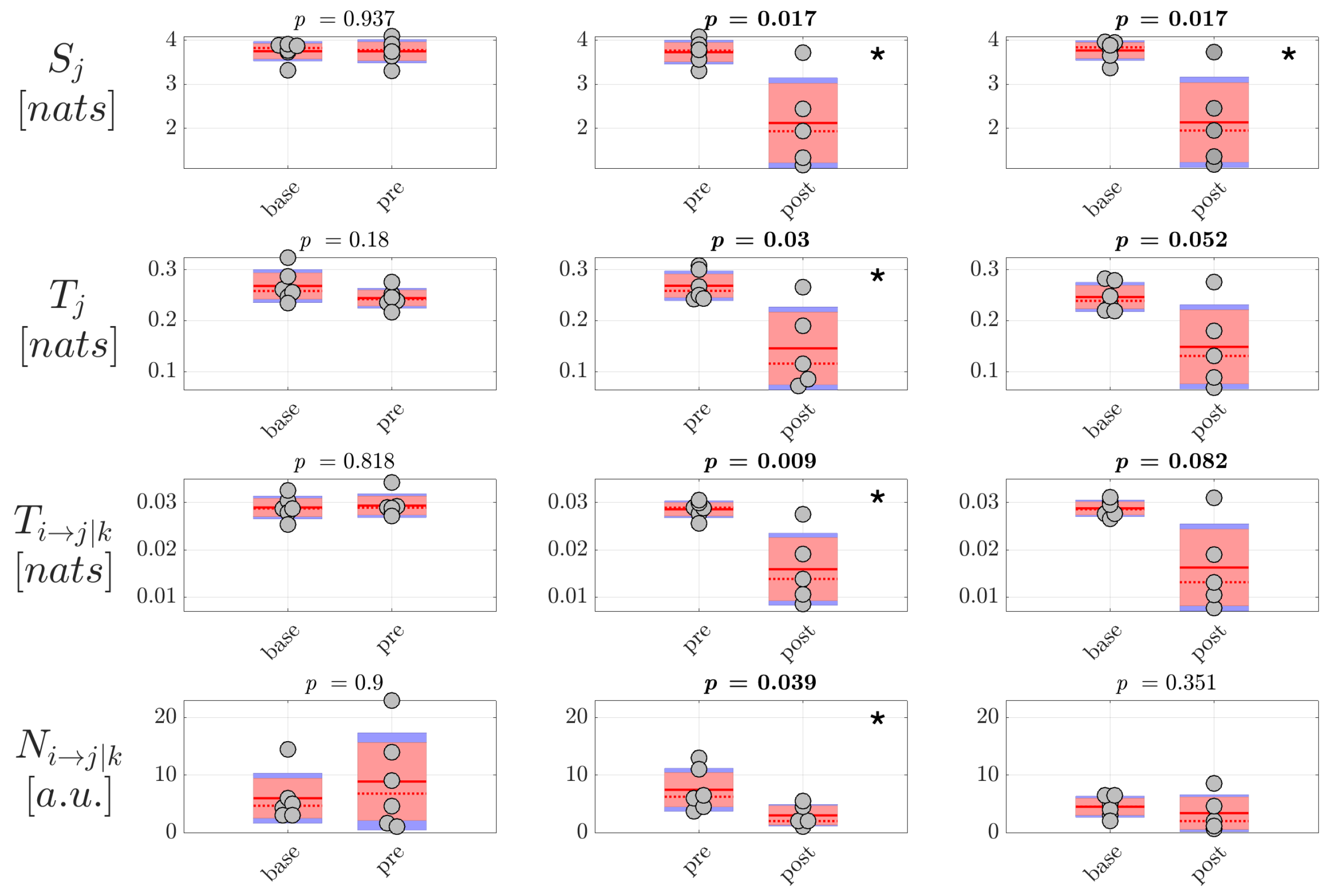
Although obtained from a small number of patients, the analysis of generalized seizures evidenced clear patterns of alteration of the measures of information dynamics when comparing the epochs preceding and following the ictal episodes. In particular, both the amounts of information stored in the EEG signals and in the reconstructed sources, and the amounts of information transferred within the scalp network and within the source network, decreased significantly moving from the to the window. This finding suggests that a reorganization of the brain network underlying spontaneous brain activity in non-ictal conditions takes place after the ictal episodes, with decreased ability of the network nodes to actively store information and to send information between each other. On the other hand, the absence of significant differences in the comparison between the and conditions suggests the lack of predictive value for the measures of information dynamics, at least regarding the onset of generalized seizures.
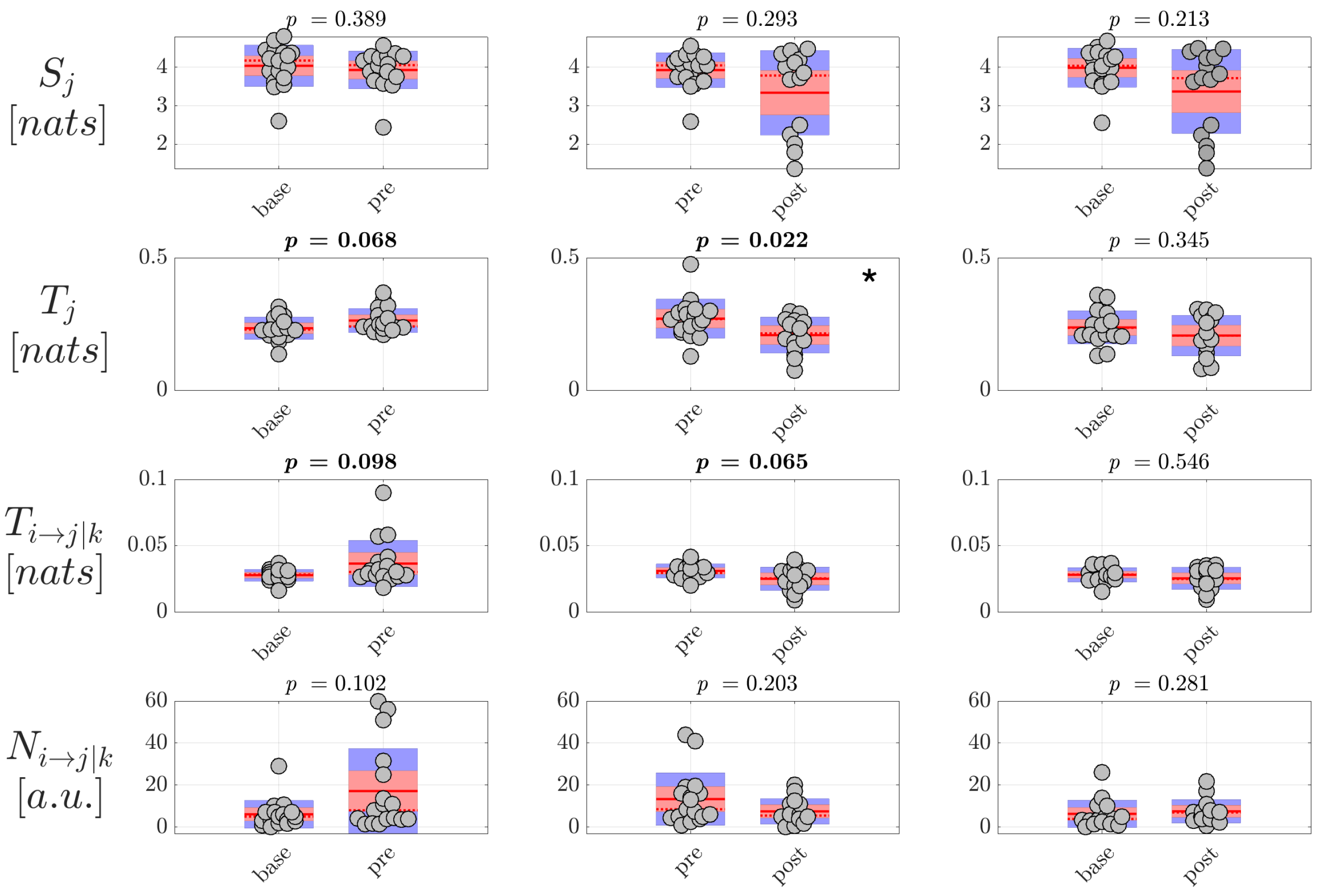
As regards focal seizures, the analysis performed through our approach at the level of the cortical sources evidenced changes of directed connectivity within the source signals in both the comparisons
/
and
/
, with the conditional and particularly the total information transferred to the network nodes increasing just before the seizure and returning to basal values just after the seizure. Such trends suggest alterations in the connectivity of the source EEG network occurring before the onset of focal epileptic seizures. The use of information-theoretic measures as an indicator of phase transitions has been demonstrated using classic spin model in physics [
50], as well as in financial systems [
51] and in the brain: recent studies have shown that the maximal amount of information transfer among units is representative of the critical state on a brain network [
52], and that synergy measures derived from information transfer in a multivariate fashion peak before the transition from disordered to ordered phases [
53]. In this context, the use of measures of information transfer like those presented in this study, possibly integrated with multivariate measures obtained in the context of information dynamics [
54], may have important implications in seizure prediction. If computed within the EEG source network, such measures are indeed good candidate tools to identify peaks of information flow located away from the time of transition between non-ictal and ictal brain activity, being thus predictive of the transition itself.
A limitation of our analysis consists in the fact that the time windows chosen as baseline might not necessarily be free of epileptic activity and already reflect an altered state of the brain, since often a seizure is preceded by long periods of seizure-readiness of the brain. This may explain the little difference found between
and
conditions. Future studies on more complete databases will analyze interictal periods where the difference with the seconds preceding the seizure onset is expected to be larger. Another potential limitation of the present study is that the proposed methodology is tested on a quite restricted database containing EEG acquired on children of different genders and type of seizure, and substantial variability in age. The reported variability across episodes of the information measures, especially in the post-ictal phase of generalized seizures, also suggests that larger and more homogeneous databases should be employed in future works. The high variability after the seizure is likely related to the difficulty in establishing clearly and objectively when the seizure is over, as seizures can terminate across brain areas via a variety of spatiotemporal patterns [
55]. These aspects might jeopardize the acceptance of the methodology and question its generalization ability in a clinical setting. In addition, using the EEG signals from subjects with wide age range one might expect developmental changes in characteristics (e.g., seizure semiology [
56] or interictal discharges [
57]) that should be evaluated separately in narrower age groups during thorough studies, including the developmental characteristics of the information dynamics measures. On the other hand, comparable group sizes are used in other works on the dynamics of epilepsy, such as in [
58] for recurrence quantification analysis, or in [
59] for studying heart rate variability as an indicator of autonomic dysregulation. While application on larger benchmarking datasets is a primary focus of future work, the aim of this paper was to present the proposed framework addressing all technical aspects and its evaluation in a simulation setting, while also showing its potential in a clinical application. Besides, the proposed framework might be further developed and improved, as some open questions emerged during the research. First of all, since the present study demonstrated promising results of the application of selected information-theoretic measures to the EEG sources, other measures should be added to the framework, to enable studying of brain activity characteristics in two conditions. Exploiting both available and new measures requires the investigation of their robustness with respect of the initial data for computation of EEG sources, i.e., duration of the EEG recordings, number of seizure instances per class and per subject, stationarity of the signal, etc. This should unveil any existing limitations in the applicability of the proposed approach to study rare EEG phenomena, or dealing with small number of subjects, which is often the case in EEG research, as well as evaluation of the results of the analysis in subject-specific vs. non-subject specific settings. A further important extension of the proposed framework would be adding the time and spatial dimensions to the description of the information-theoretic characteristics of brain activity. Proper assessment of time-varying measures of EEG sources requires the study of the VAR model fitting and finding the unmixing matrices for different durations and time positions of trials in classes under comparison to find optimal combination of these settings to compute the informative EEG sources. Also, since CSP provides the spatial distribution of activity patterns, the proposed framework could be extended to account the localization of sources in combination with assessing their entropy and connectivity characteristics. Interpretation of the CSP spatial characteristics and relation of those with the actual localization over the scalp and in the brain tissue volume might be an important advancement of the framework, as it would open the way to the use of directed information measures for the localization of the drivers of epileptiform activity [
60].