Remediation of Organically Contaminated Soil Through the Combination of Assisted Phytoremediation and Bioaugmentation
Abstract
1. Introduction
2. Materials and Methods
2.1. Experimental Design
2.2. Soil Physicochemical Properties
2.3. Soil Biological Properties
2.4. Statistical Analyses
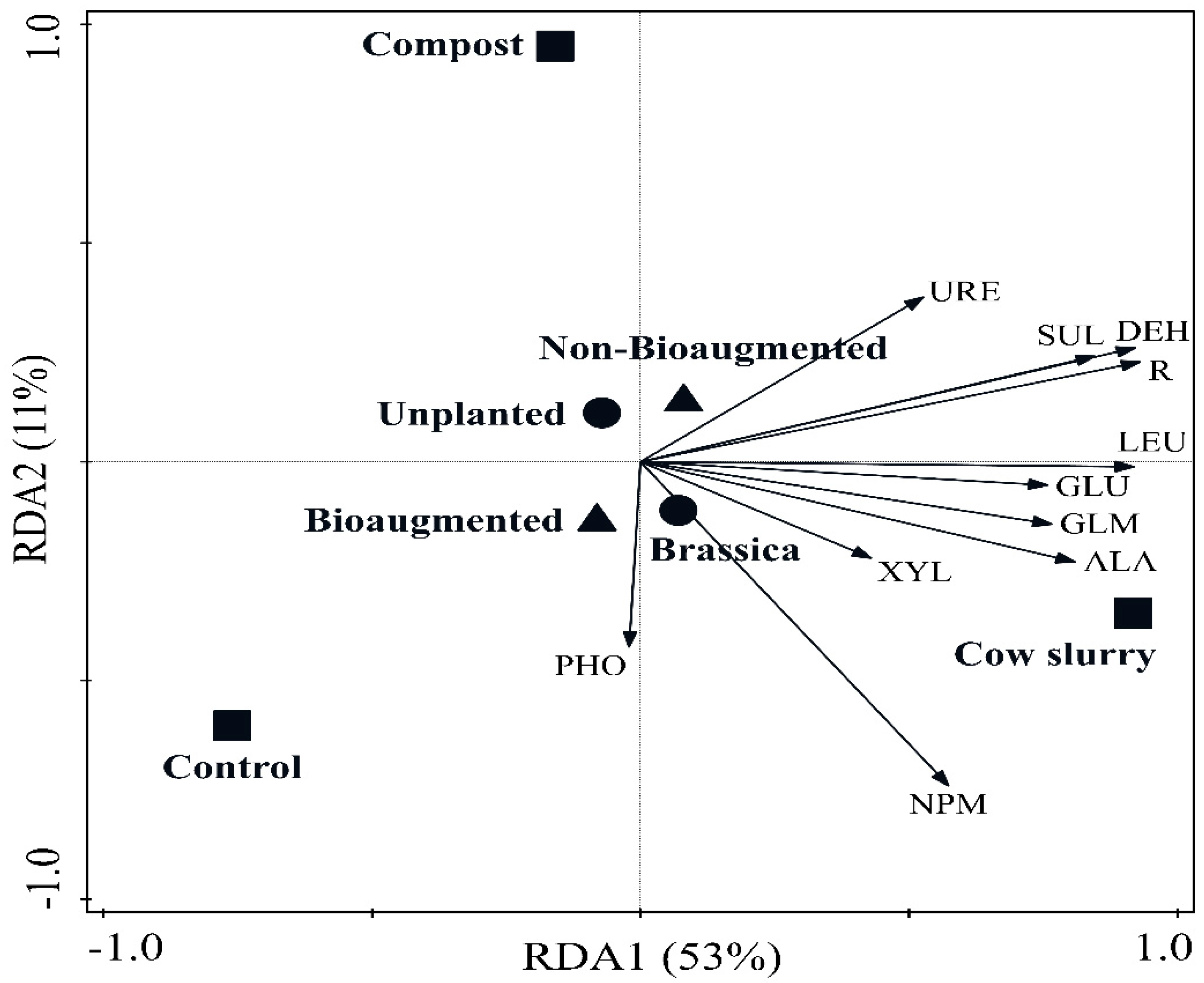
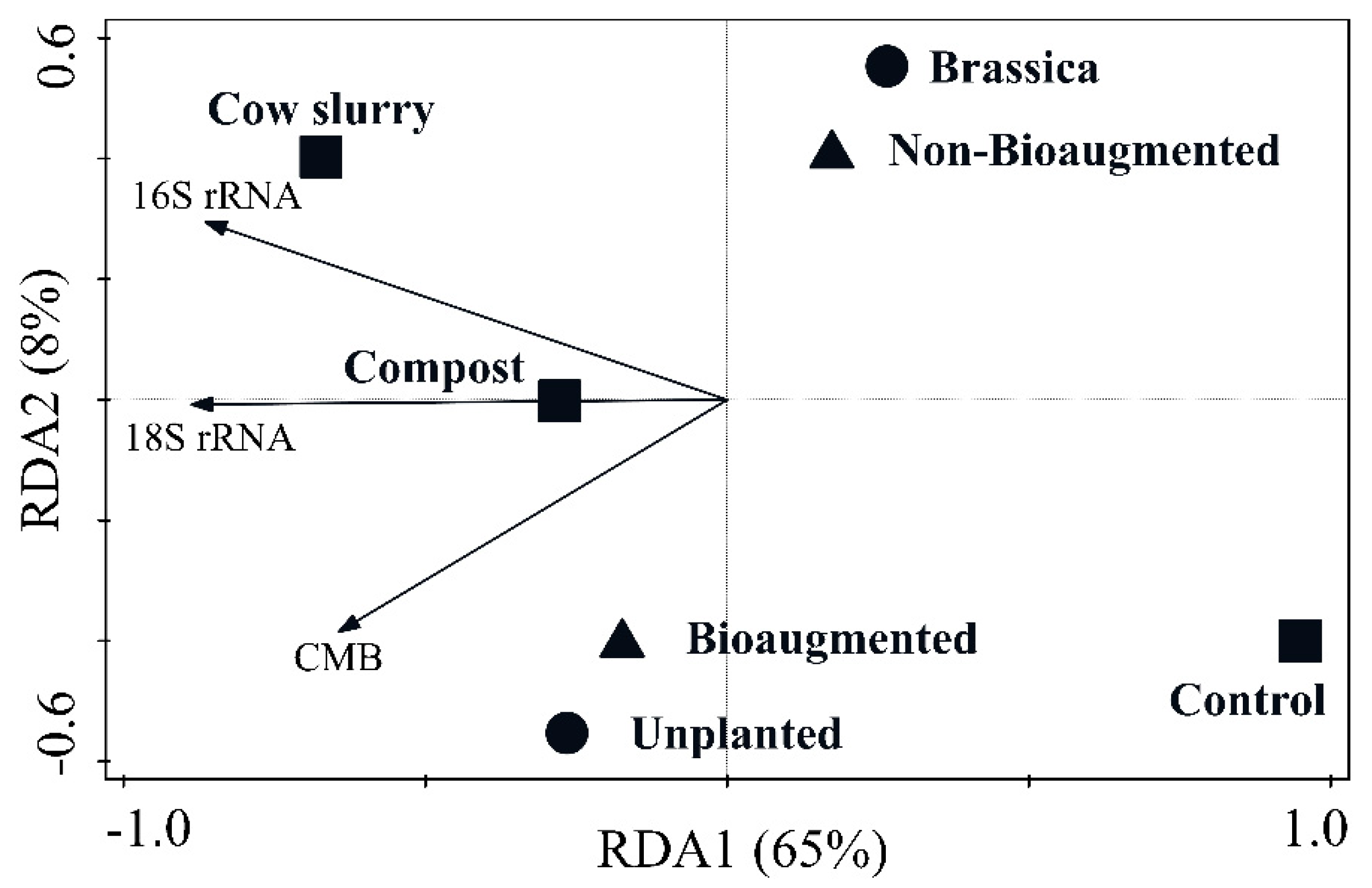
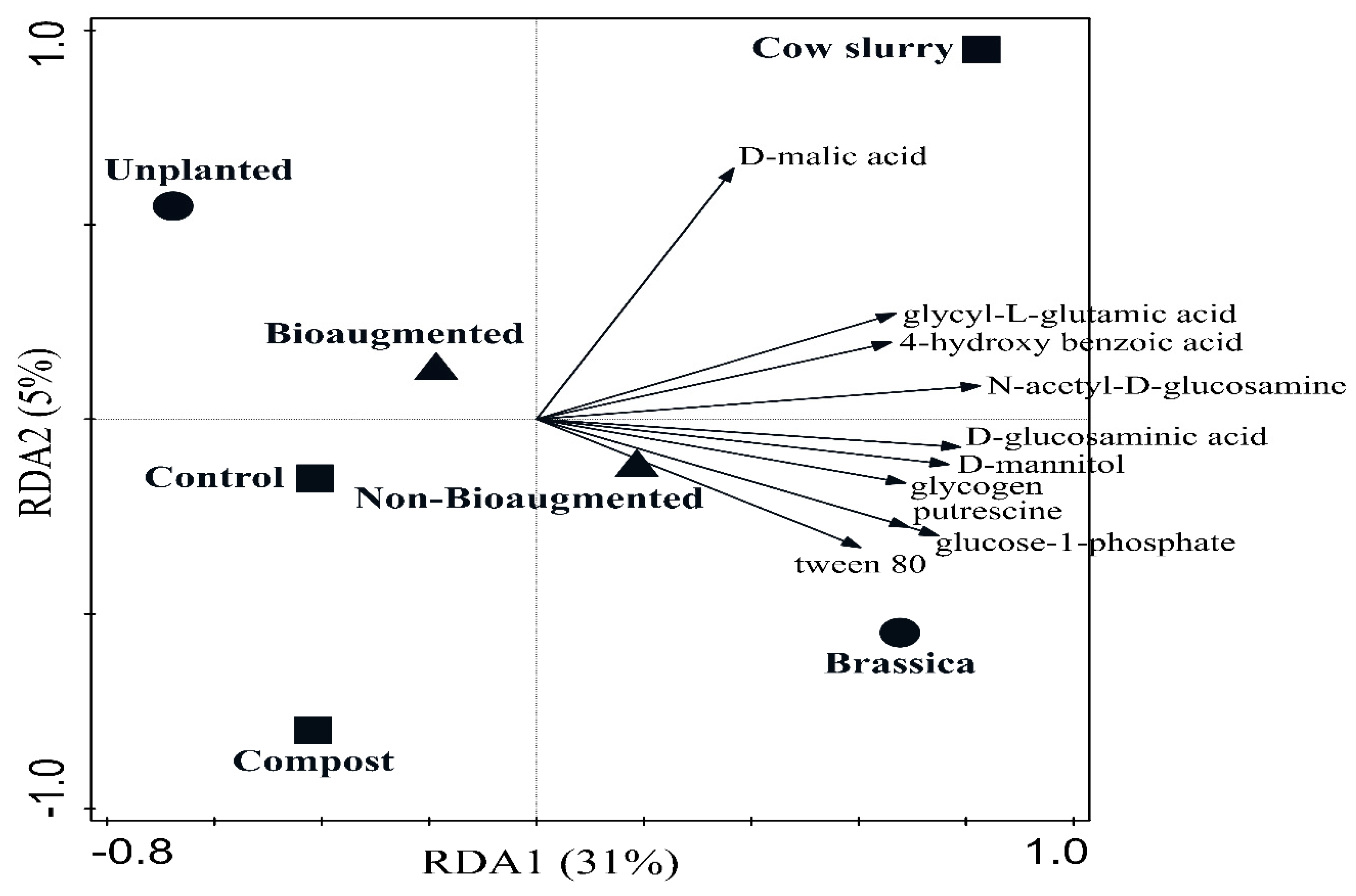
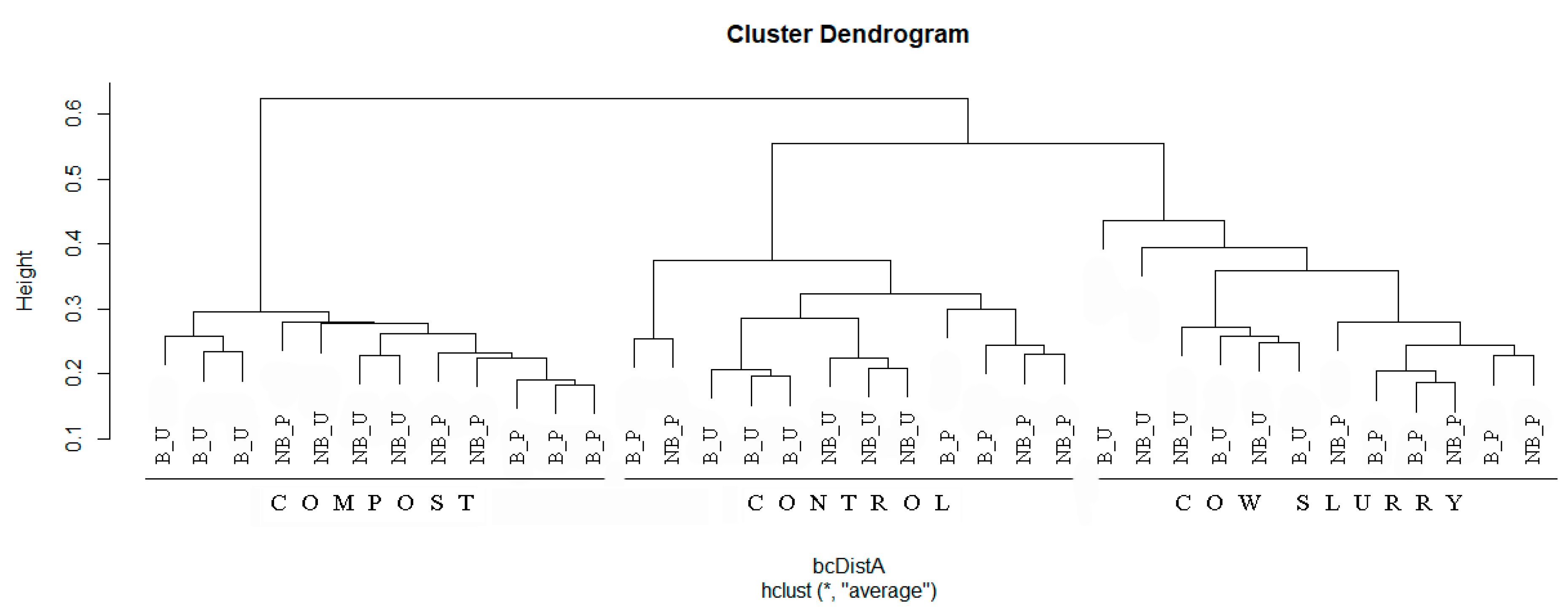
3. Results
4. Discussion
4.1. Contaminant Concentrations
4.2. Soil Health
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Garbisu, C.; Alkorta, I. Bioremediation: Principles and future. J. Clean Technol. Environ. Toxicol. Occup. Med. 1997, 6, 351–366. [Google Scholar]
- Garbisu, C.; Alkorta, I. Utilization of genetically engineered microorganisms (GEMs) for bioremediation. J. Clean Technol. Biotechnol. 1999, 74, 599–606. [Google Scholar] [CrossRef]
- Garbisu, C.; Alkorta, I. Basic concepts on heavy metal soil bioremediation. Eur. J. Miner. Process. Environ. Prot. 2003, 3, 58–66. [Google Scholar]
- Alkorta, I.; Becerril, J.M.; Garbisu, C. Recovery of soil health: The ultimate goal of soil remediation processes. In Trends in Bioremediation and Phytoremediation; Plaza, G., Ed.; Research Signpost: Kerala, India, 2010; Chapter 1; pp. 1–9. ISBN 978-81-308-0424-8. [Google Scholar]
- Garbisu, C.; Garaiyurrebaso, O.; Epelde, L.; Grohmann, E.; Alkorta, I. Plasmid-mediated bioaugmentation for the bioremediation of contaminated soils. Front. Microbiol. 2017, 8, 1966. [Google Scholar] [CrossRef]
- Ferraro, A.; Dottorini, G.; Massini, G.; Miritana, V.M.; Signorini, A.; Lembo, G.; Fabbricino, M. Combined bioaugmentation with anaerobic ruminal fungi and fermentative bacteria to enhance biogas production from wheat straw and mushroom spent straw. Bioresour. Technol. 2018, 260, 364–373. [Google Scholar] [CrossRef] [PubMed]
- Correa-García, S.; Pande, P.; Séguin, A.; St-Arnaud, M.; Yergeau, E. Rhizoremediation of petroleum hydrocarbons: A model system for plant microbiome manipulation. Microb. Biotechnol. 2018, 11, 819–832. [Google Scholar] [CrossRef]
- Idowu, O.; Semple, K.T.; Ramadass, K.; Connor, W.O.; Hansbro, P.; Thavamani, P. Beyond the obvious: Environmental health implications of polar polycyclic aromatic hydrocarbons. Environ. Int. 2019, 123, 543–557. [Google Scholar] [CrossRef]
- Arthur, E.L.; Rice, P.J.; Rice, P.J.; Anderson, T.A.; Baladi, S.M. Phytoremediation—An overview. Crit. Rev. Plant Sci. 2005, 24, 109–122. [Google Scholar] [CrossRef]
- Gaskin, S.E.; Bentham, R.H. Rhizoremediation of hydrocarbon contaminated soil using Australian native grasses. Sci. Total Environ. 2010, 408, 3683–3688. [Google Scholar] [CrossRef]
- Kowalchuk, G.A.; Buma, D.S.; Boer, W.D.; Klinkhamer, P.G.L.; van Veen, J.A. Effects of above-ground plant species composition and diversity on the diversity of soil-borne microorganisms. Antonie Van Leeuwenhoek 2002, 81, 509–520. [Google Scholar] [CrossRef]
- Smalla, K.; Wieland, G.; Buchner, A.; Zock, A.; Parzy, J.; Kaiser, S.; Roskot, N.; Heuer, H.; Berg, G. Bulk and rhizosphere soil bacterial communities studied by denaturing gradient gel electrophoresis: Plant-dependent enrichment and seasonal shifts revealed. Appl. Environ. Microbiol. 2001, 67, 4742–4751. [Google Scholar] [CrossRef]
- Yateem, A.; Al-Sharrah, T.; Bin-Haji, A. Investigation of microbes in the rhizosphere of selected grasses for rhizoremediation of hydrocarbon-contaminated soils. Soil Sediment Contam. Int. J. 2007, 16, 269–280. [Google Scholar] [CrossRef]
- Heinaru, E.; Merimaa, M.; Viggor, S.; Lehiste, M.; Leito, I.; Truu, J. Biodegradation efficiency of functionally important populations selected for bioaugmentation in phenol- and oil-polluted area. FEMS Microbiol. Ecol. 2005, 51, 363–373. [Google Scholar] [CrossRef]
- Chen, L.; Luo, S.; Xiao, X.; Guo, H.; Chen, J.; Wan, Y.; Li, B.; Xu, T.; Xi, Q.; Rao, C.; et al. Application of plant growth-promoting endophytes (PGPE) isolated from Solanum nigrum L. for phytoextraction of Cd-polluted soils. Appl. Soil Ecol. 2010, 46, 383–389. [Google Scholar] [CrossRef]
- Zhang, Y.F.; He, L.Y.; Chen, Z.J.; Zhang, W.H.; Wang, Q.Y.; Qian, M.; Sheng, X.F. Characterization of lead-resistant and ACC deaminase-producing endophytic bacteria and their potential in promoting lead accumulation of rape. J. Hazard. Mater. 2011, 186, 1720–1725. [Google Scholar] [CrossRef]
- Burges, A.; Epelde, L.; Benito, G.; Artetxe, U.; Becerril, J.M.; Garbisu, C. Enhancement of ecosystem services during endophyte-assisted aided phytostabilization of metal contaminated mine soil. Sci. Total Environ. 2016, 562, 480–492. [Google Scholar] [CrossRef]
- Das, S.; Jeong, S.T.; Das, S.; Kim, P.J. Composted cattle manure increases microbial activity and soil fertility more than composted swine manure in a submerged rice paddy. Front. Microbiol. 2017, 8, 1702. [Google Scholar] [CrossRef]
- Fließbach, A.; Oberholzer, H.; Gunst, L.; Mäder, P. Soil organic matter and biological soil quality indicators after 21 years of organic and conventional farming. Agric. Ecosyst. Environ. 2007, 118, 273–284. [Google Scholar] [CrossRef]
- Evanylo, G.; Sherony, C.; Spargo, J.; Starner, D.; Brosius, M.; Haering, K. Agriculture, ecosystems and environment soil and water environmental effects of fertilizer-, manure-, and compost-based fertility practices in an organic vegetable cropping system. Agric. Ecosyst. Environ. 2008, 127, 50–58. [Google Scholar] [CrossRef]
- Epelde, L.; Becerril, J.M.; Mijangos, I.; Garbisu, C. Evaluation of the efficiency of a phytostabilization process with biological indicators of soil health. J. Environ. Qual. 2009, 38, 2041–2049. [Google Scholar] [CrossRef]
- Epelde, L.; Burges, A.; Mijangos, I.; Garbisu, C. Microbial properties and attributes of ecological relevance for soil quality monitoring during a chemical stabilization field study. Appl. Soil Ecol. 2014, 75, 1–12. [Google Scholar] [CrossRef]
- Hernández-Allica, J.; Becerril, J.M.; Zárate, O.; Garbisu, C. Assessment of the efficiency of a metal phytoextraction process with biological indicators of soil health. Plant Soil 2006, 281, 147–158. [Google Scholar] [CrossRef]
- Barrutia, O.; Garbisu, C.; Epelde, L.; Sampedro, M.C.; Goicolea, M.A.; Becerril, J.M. Plant tolerance to diesel minimizes its impact on soil microbial characteristics during rhizoremediation of diesel-contaminated soils. Sci. Total Environ. 2011, 409, 4087–4093. [Google Scholar] [CrossRef] [PubMed]
- Galende, M.A.; Becerril, J.M.; Barrutia, O.; Artetxe, U.; Garbisu, C.; Hernández, A. Field assessment of the effectiveness of organic amendments for aided phytostabilization of a Pb-Zn contaminated mine soil. J. Geochem. Explor. 2014, 145, 181–189. [Google Scholar] [CrossRef]
- Mijangos, I.; Albizu, I.; Epelde, L.; Amezaga, I.; Mendarte, S.; Garbisu, C. Effects of liming on soil properties and plant performance of temperate mountainous grasslands. J. Environ. Manag. 2010, 91, 2066–2074. [Google Scholar] [CrossRef]
- ISO. ISO 18287, 2006: Soil Quality—Determination of Polycyclic Aromatic Hydrocarbons (PAH)—Gas Chromatographic Method with Mass Spectrometric Detection (GC-MS); ISO: Geneva, Switzerland, 2006. [Google Scholar]
- ISO. ISO 16703, 2004: Soil Quality—Determination of Content of Hydrocarbon in the Range C10 to C40 by Gas Chromatography; ISO: Geneva, Switzerland, 2004. [Google Scholar]
- ISO. ISO 16558-1, 2015: Soil Quality—Risk-Based Petroleum Hydrocarbons. Part 1: Determination of Aliphatic and Aromatic Fractions of Volatile Petroleum Hydrocarbons Using Gas Chromatography (Static Headspace Method); ISO: Geneva, Switzerland, 2015. [Google Scholar]
- Lacalle, R.G.; Gómez-Sagasti, M.T.; Artetxe, U.; Garbisu, C.; Becerril, J.M. Brassica napus has a key role in the recovery of the health of soils contaminated with metals and diesel by nano-rhizoremediation. Sci. Total Environ. 2018, 618, 347–356. [Google Scholar] [CrossRef]
- Nelson, D.W.; Sommers, L.E. Total carbon, organic carbon and organic matter. In Methods of Soil Analysis; Part 3. Chemical Methods; Sparks, D.L., Page, A.L., Helmke, P.A., Loeppert, R.H., Soltanpour, P.N., Tabatabai, M.A., Johnston, C.T., Sumner, M.E., Eds.; SSSA and ASA: Madison, WI, USA, 1996; pp. 961–1010. [Google Scholar]
- Wu, J.; Joergensen, R.G.; Pommerening, B.; Chaussod, R.; Brookes, P.C. Measurement of soil microbial biomass C by fumigation-extraction: An automated procedure. Soil Biol. Biochem. 1990, 22, 1167–1169. [Google Scholar] [CrossRef]
- ISO. ISO 10694, 1995: Soil Quality—Determination of Organic and Total Carbon after Dry Combustión (Elementary Analysis); ISO: Geneva, Switzerland, 1995. [Google Scholar]
- ISO. ISO 13878, 1998: Soil Quality—Determination of Total Nitrogen Content by Dry Combustión (Elementary Analysis); ISO: Geneva, Switzerland, 1998. [Google Scholar]
- Hechmi, N.; Bosso, L.; El-Bassia, L.; Scelza, R.; Testa, A.; Jedidi, N.; Rao, M.A. Depletion of pentachlorophenol in soil microcosms with Byssochlamys nivea and Scopulariopsis brumptii as detoxification agents. Chemosphere 2016, 165, 547–554. [Google Scholar] [CrossRef]
- Wang, H.; Kuang, S.; Lang, Q.; Yu, W. Effects of aged oil sludge on soil physicochemical properties and fungal diversity revealed by high-throughput sequencing analysis. Archaea 2018, 9264259. [Google Scholar] [CrossRef]
- Vance, E.D.; Brookes, P.C.; Jenkinson, D.S. An extraction method for measuring soil microbial biomass C. Soil Biol. Biochem. 1987, 19, 703–707. [Google Scholar] [CrossRef]
- Kowalchuk, G.A.; Stephen, J.R.; DeBoer, W.; Prosser, J.I.; Embley, T.M.; Woldendorp, J.W. Analysis of ammonia-oxidizing bacteria of the beta subdivision of the class Proteobacteria in coastal sand dunes by denaturing gradient gel electrophoresis and sequencing of PCR-amplified 16S ribosomal DNA fragments. Appl. Environ. Microbiol. 1997, 63, 1489–1497. [Google Scholar]
- Dhanasekaran, S.; Doherty, T.M.; Kenneth, J. Comparison of different standards for real-time PCR-based absolute quantification. J. Immunol. Methods 2010, 354, 34–39. [Google Scholar] [CrossRef]
- Vainio, E.J.; Hantula, J. Direct analysis of wood-inhabiting fungi using denaturing gradient gel electrophoresis of amplified ribosomal DNA. Mycol. Res. 2000, 104, 927–936. [Google Scholar] [CrossRef]
- Lueders, T.; Manefield, M.; Friedrich, M.W. Enhanced sensitivity of DNA- and rRNA-based stable isotope probing by fractionation and quantitative analysis of isopycnic centrifugation gradients. Environ. Microbiol. 2004, 6, 73–78. [Google Scholar] [CrossRef]
- ISO. ISO-16072, 2002: Soil Quality—Laboratory Methods for Determination of Microbial Soil Respiration; ISO: Geneva, Switzerland, 2002. [Google Scholar]
- Powers, R.F. Mineralizable soil nitrogen as an index of nitrogen availability to forest trees. Soil Sci. Soc. Am. J. 1980, 44, 1314–1320. [Google Scholar] [CrossRef]
- ISO. ISO/TS 22939, 2010: Soil Quality—Measurement of Enzyme Activity Patterns in Soil Samples Using Fluorogenic Substrates in Micro-Well Plates; ISO: Geneva, Switzerland, 2010. [Google Scholar]
- Dick, R.P. Soil enzyme activities as integrative indicators of soil health. In Biological Indicators of Soil Health; Pankhurst, C.E., Doube, B.M., Gupta, V., Eds.; CAB International: Wallingford, UK, 1997; pp. 121–156. [Google Scholar]
- Kandeler, E.; Gerber, H. Short-term assay of soil urease activity using colorimetric determination of ammonium. Biol. Fertil. Soils 1988, 6, 68–72. [Google Scholar] [CrossRef]
- ISO. ISO-23753-2, 2005: Soil Quality—Determination of Dehydrogenase Activity in Soils—Part 2: Method Using Iodotetrazolium Chloride (INT); ISO: Geneva, Switzerland, 2005. [Google Scholar]
- Epelde, L.; Becerril, J.M.; Hernandez-Allica, J.; Barrutia, O.; Garbisu, C. Functional diversity as indicator of the recovery of soil health derived from Thlaspi caerulescens growth and metal phytoextraction. Appl. Soil Ecol. 2008, 39, 299–310. [Google Scholar] [CrossRef]
- Preston-Mafham, J.; Boddy, L.; Randerson, P.F. Analysis of microbial community functional diversity using sole-carbon-source utilisation profiles—A critique. FEMS Microbiol. Ecol. 2002, 42, 1–14. [Google Scholar]
- Lanzén, A.; Epelde, L.; Blanco, F.; Martin, I.; Artetxe, U.; Garbisu, C. Multi-targeted metagenetic analysis of the influence of climate and environmental parameters on soil microbial communities along an elevational gradient. Sci. Rep. 2016, 6, 28257. [Google Scholar] [CrossRef]
- Øvreås, L.; Forney, L.; Daae, F.L. Distribution of bacterioplankton in meromictic Lake Saelenvannet, as determined by denaturing gradient gel electrophoresis of PCR-amplified gene fragments coding for 16S rRNA. Appl. Environ. Microbiol. 1997, 63, 3367–3373. [Google Scholar]
- Caporaso, J.G.; Lauber, C.L.; Walters, W.A.; Berg-Lyons, D.; Huntley, J.; Fierer, N.; Owens, S.M.; Betley, J.; Fraser, L.; Bauer, M.; et al. Ultra-high-throughput microbial community analysis on the Illumina HiSeq and MiSeq platforms. ISME J. 2012, 6, 1621–1624. [Google Scholar] [CrossRef]
- Lanzén, A.; Jørgensen, S.L.; Huson, D.H.; Gorfer, M.; Grindhaug, S.H.; Jonassen, I.; Øvreås, L.; Urich, T. CREST—Classification Resources for Environmental Sequence Tags. PLoS ONE 2012, 7, e49334. [Google Scholar] [CrossRef]
- Cui, Z.; Zhang, X.; Yang, H.; Sun, L. Bioremediation of heavy metal pollution utilizing composite microbial agent of Mucor circinelloides, Actinomucor sp. and Mortierella sp. J. Environ. Chem. Eng. 2017, 5, 3616–3621. [Google Scholar] [CrossRef]
- Zhang, T.; He, J.; Feng, H.; Zhan, X. Improvement of soil nutrient and biological properties and establishment of Lycium barbarum L. in an impermeable saline-sodic soil using drip irrigation. Soil Res. 2019, 57, 75–84. [Google Scholar] [CrossRef]
- Ter Braak, C.J.F.; Šmilauer, P. CANOCO Reference Manual and CanoDraw for Windows User’s Guide: Software for Canonical Community Ordination; Microcomputer Power: Ithaca, NY, USA, 2002. [Google Scholar]
- Oksanen, J.; Blanchet, F.G.; Kindt, R.; Legendre, P.; Minchin, P.R.; O’Hara, R.B.; Simpson, J.L.; Solymos, P.; Stevens, M.H.H.; Szoecs, E.; et al. Vegan: Community Ecology Package. R Package Version 2.3-1. Available online: https://rdrr.io/rforge/vegan/ (accessed on 21 October 2019).
- Wang, L.; Li, F.; Zhan, Y.; Zhu, L. Shifts in microbial community structure during in situ surfactant-enhanced bioremediation of polycyclic aromatic hydrocarbon-contaminated soil. Environ. Sci. Pollut. Res. 2016, 23, 14451–14461. [Google Scholar] [CrossRef]
- Bai, N.; Wang, S.; Abuduaini, R.; Zhang, M.; Zhu, X.; Zhao, Y. Rhamnolipid-aided biodegradation of carbendazim by Rhodococcus sp. D-1: Characteristics, products, and phytotoxicity. Sci. Total Environ. 2017, 590–591, 343–351. [Google Scholar] [CrossRef]
- Szulc, A.; Ambrozewicz, D.; Sydow, M.; Lawniczak, L.; Piotrowska-Cyplik, A.; Marecik, R.; Chrzanowski, L. The influence of bioaugmentation and biosurfactant addition on bioremediation efficiency of diesel-oil contaminated soil: Feasibility during field studies. J. Environ. Manag. 2014, 132, 121–128. [Google Scholar] [CrossRef]
- Galende, M.A.; Becerril, J.M.; Gómez-Sagasti, M.T.; Barrutia, O.; Epelde, L.; Garbisu, C.; Hernández, A. Chemical stabilization of metal-contaminated mine soil: Early short-term soil-amendment interactions and their effects on biological and chemical parameters. Water Air Soil Pollut. 2014, 225, 1863. [Google Scholar] [CrossRef]
- Masy, T.; Demanèche, S.; Tromme, O.; Thonart, P.; Jacques, P.; Hiligsmann, S.; Vogel, T.M. Hydrocarbon biostimulation and bioaugmentation in organic carbon and clay-rich soils. Soil Biol. Biochem. 2016, 99, 66–74. [Google Scholar] [CrossRef]
- Koshlaf, E.; Shahsavari, E.; Haleyur, N.; Osborn, A.M.; Ball, A.S. Effect of biostimulation on the distribution and composition of the microbial community of a polycyclic aromatic hydrocarbon-contaminated landfill soil during bioremediation. Geoderma 2019, 338, 216–225. [Google Scholar] [CrossRef]
- Agnello, A.C.; Bagard, M.; van Hullebusch, E.D.; Esposito, G.; Huguenot, D. Comparative bioremediation of heavy metals and petroleum hydrocarbons co-contaminated soil by natural attenuation, phytoremediation, bioaugmentation and bioaugmentation-assisted phytoremediation. Sci. Total Environ. 2016, 563–564, 693–703. [Google Scholar] [CrossRef] [PubMed]
- Mertens, B.; Boon, N.; Verstraete, W. Slow-release inoculation allows sustained biodegradation of gamma-hexachlorocyclohexane. Appl. Environ. Microbiol. 2006, 72, 622–627. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Power, B.; Liu, X.; Germaine, K.J.; Ryan, D.; Brazil, D.; Dowling, D.N. Alginate beads as a storage, delivery and containment system for genetically modified PCB degrader and PCB biosensor derivatives of Pseudomonas fluorescens F113. J. Appl. Microbiol. 2011, 110, 1351–1358. [Google Scholar] [CrossRef]
- Van Elsas, J.D.; Heijnen, C.E. Methods for the introduction of bacteria into soil: A review. Biol. Fertil. Soils 1990, 10, 127–133. [Google Scholar] [CrossRef]
- Epelde, L.; Becerril, J.M.; Kowalchuk, G.A.; Deng, Y.; Zhou, J.Z.; Garbisu, C. Impact of metal pollution and Thlaspi caerulescens growth on soil microbial communities. Appl. Environ. Microbiol. 2010, 76, 7843–7853. [Google Scholar] [CrossRef]

| Rhizoremediation | Bioaugmentation | Biostimulation |
|---|---|---|
| Brassica napus | Bioaugmented | Control |
| Dried cow slurry | ||
| Composted horse manure | ||
| Non-bioaugmented | Control | |
| Dried cow slurry | ||
| Composted horse manure | ||
| Unplanted | Bioaugmented | Control |
| Dried cow slurry | ||
| Composted horse manure | ||
| Non-bioaugmented | Control | |
| Dried cow slurry | ||
| Composted horse manure |
| Treatment | TPHs | Total-PAHs | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | ||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Brassica | Non-bioaugmented | Control | 4367 ± 1514 | 2467 ± 58 | 62 ± 4 | 41 ± 3 | 42 ± 3 | 257 ± 15 | 68 ± 3 | 400 ± 17 | 273 ± 12 | 250 ± 10 | 227 ± 6 |
| Compost | 3253 ± 979 | 2237 ± 254 | 61 ± 6 | 38 ± 3 | 39 ± 4 | 240 ± 31 | 65 ± 4 | 366 ± 44 | 248 ± 25 | 224 ± 25 | 199 ± 25 | ||
| Cow slurry | 3496 ± 921 | 2698 ± 348 | 81 ± 19 | 47 ± 8 | 49 ± 8 | 300 ± 51 | 78 ± 12 | 441 ± 57 | 300 ± 40 | 277 ± 29 | 243 ± 26 | ||
| Bioaugmented | Control | 2900 ± 361 | 2633 ± 306 | 72 ± 6 | 44 ± 6 | 49 ± 6 | 290 ± 35 | 84 ± 12 | 440 ± 56 | 300 ± 40 | 267 ± 32 | 217 ± 40 | |
| Compost | 2277 ± 141 | 2806 ± 211 | 79 ± 12 | 51 ± 7 | 54 ± 7 | 321 ± 31 | 92 ± 13 | 476 ± 44 | 317 ± 32 | 289 ± 25 | 232 ± 12 | ||
| Cow slurry | 2888 ± 348 | 3420 ± 635 | 94 ± 23 | 69 ± 19 | 70 ± 19 | 407 ± 86 | 106 ± 24 | 574 ± 106 | 388 ± 71 | 342 ± 63 | 274 ± 34 | ||
| Unplanted | Non-bioaugmented | Control | 3400 ± 400 | 3067 ± 115 | 84 ± 11 | 53 ± 6 | 56 ± 7 | 340 ± 36 | 89 ± 10 | 507 ± 31 | 347 ± 21 | 310 ± 10 | 267 ± 23 |
| Compost | 3172 ± 440 | 3009 ± 373 | 89 ± 15 | 54 ± 8 | 57 ± 9 | 342 ± 49 | 92 ± 14 | 488 ± 68 | 333 ± 43 | 305 ± 37 | 260 ± 39 | ||
| Cow slurry | 3268 ± 237 | 2432 ± 263 | 61 ± 8 | 41 ± 7 | 44 ± 8 | 262 ± 41 | 71 ± 11 | 395 ± 43 | 270 ± 33 | 251 ± 30 | 198 ± 24 | ||
| Bioaugmented | Control | 3767 ± 1415 | 2767 ± 115 | 76 ± 4 | 48 ± 1 | 50 ± 1 | 303 ± 6 | 87 ± 10 | 457 ± 12 | 310 ± 10 | 283 ± 15 | 257 ± 12 | |
| Compost | 3375 ± 373 | 2928 ± 122 | 86 ± 12 | 63 ± 23 | 57 ± 8 | 317 ± 24 | 86 ± 5 | 468 ± 19 | 317 ± 12 | 297 ± 7 | 244 ± 21 | ||
| Cow slurry | 3192 ± 342 | 2622 ± 228 | 65 ± 9 | 43 ± 5 | 45 ± 6 | 277 ± 35 | 75 ± 10 | 426 ± 40 | 293 ± 29 | 270 ± 29 | 236 ± 35 | ||
| Plant (P) | ns | ns | ns | ns | ns | ns | ns | ns | ns | ns | ns | ||
| Bioaugmentation (B) | ns | 0.041 | ns | 0.035 | 0.030 | 0.044 | 0.009 | 0.026 | 0.038 | 0.039 | ns | ||
| Amendment (A) | ns | ns | ns | ns | ns | ns | ns | ns | ns | ns | ns | ||
| P x B | 0.030 | 0.010 | ns | ns | 0.009 | 0.004 | 0.005 | 0.004 | 0.005 | 0.012 | ns | ||
| P x A | ns | 0.001 | 0.000 | 0.003 | 0.003 | 0.001 | 0.006 | 0.001 | 0.001 | 0.001 | 0.001 | ||
| B x A | ns | ns | ns | ns | ns | ns | ns | ns | ns | ns | ns | ||
| P x B x A | ns | ns | ns | ns | ns | ns | ns | ns | ns | ns | ns | ||
| 10 | 11 | 12 | 13 | 14 | 15 | 16 | 17 | 18 | 19 | 20 | |||
| Brassica | Non-bioaugmented | Control | 277 ± 6 | 120 ± 0 | 193 ± 6 | 41 ± 2 | 99 ± 1 | 117 ± 6 | 1767 ± 58 | 84 ± 7 | 273 ± 32 | 1367 ± 379 | 2667 ± 1079 |
| Compost | 244 ± 32 | 107 ± 14 | 171 ± 24 | 36 ± 2 | 89 ± 13 | 105 ± 12 | 1667 ± 186 | 83 ± 5 | 232 ± 49 | 1057 ± 391 | 1871 ± 550 | ||
| Cow slurry | 293 ± 33 | 125 ± 11 | 205 ± 20 | 42 ± 3 | 105 ± 8 | 125 ± 11 | 2014 ± 237 | 108 ± 25 | 266 ± 56 | 1227 ± 383 | 1900 ± 475 | ||
| Bioaugmented | Control | 287 ± 32 | 127 ± 15 | 193 ± 21 | 43 ± 5 | 96 ± 8 | 120 ± 10 | 1933 ± 208 | 89 ± 2 | 243 ± 25 | 950 ± 132 | 1600 ± 200 | |
| Compost | 305 ± 21 | 130 ± 14 | 211 ± 19 | 44 ± 4 | 107 ± 7 | 125 ± 8 | 2074 ± 211 | 85 ± 10 | 211 ± 19 | 732 ± 88 | 1253 ± 78 | ||
| Cow slurry | 361 ± 63 | 156 ± 29 | 247 ± 40 | 47 ± 7 | 121 ± 18 | 148 ± 23 | 2470 ± 475 | 116 ± 26 | 285 ± 59 | 996 ± 166 | 1482 ± 114 | ||
| Unplanted | Non-bioaugmented | Control | 333 ± 6 | 147 ± 6 | 230 ± 0 | 46 ± 1 | 113 ± 6 | 137 ± 6 | 2233 ± 58 | 102 ± 8 | 287 ± 21 | 1200 ± 200 | 1800 ± 200 |
| Compost | 317 ± 37 | 138 ± 19 | 224 ± 19 | 47 ± 6 | 113 ± 9 | 134 ± 12 | 2196 ± 244 | 111 ± 15 | 277 ± 51 | 1126 ± 249 | 1627 ± 141 | ||
| Cow slurry | 274 ± 30 | 119 ± 15 | 190 ± 24 | 44 ± 7 | 101 ± 12 | 119 ± 16 | 1786 ± 237 | 92 ± 10 | 258 ± 17 | 1121 ± 122 | 1786 ± 132 | ||
| Bioaugmented | Control | 303 ± 23 | 133 ± 6 | 210 ± 17 | 44 ± 5 | 107 ± 12 | 127 ± 12 | 2033 ± 58 | 97 ± 14 | 273 ± 59 | 1260 ± 468 | 2133 ± 839 | |
| Compost | 317 ± 12 | 138 ± 7 | 220 ± 0 | 50 ± 5 | 120 ± 3 | 138 ± 7 | 2115 ± 70 | 124 ± 18 | 338 ± 93 | 1200 ± 154 | 1708 ± 122 | ||
| Cow slurry | 293 ± 24 | 125 ± 11 | 205 ± 11 | 43 ± 4 | 107 ± 5 | 125 ± 11 | 1900 ± 174 | 89 ± 15 | 251 ± 20 | 1045 ± 119 | 1786 ± 237 | ||
| Plant (P) | ns | ns | ns | 0.041 | 0.029 | ns | ns | ns | ns | ns | ns | ||
| Bioaugmentation (B) | 0.046 | ns | ns | ns | ns | ns | ns | ns | ns | ns | ns | ||
| Amendment (A) | ns | ns | ns | ns | ns | ns | ns | ns | ns | ns | ns | ||
| P × B | 0.021 | 0.024 | 0.032 | ns | ns | ns | 0.011 | ns | ns | ns | 0.012 | ||
| P × A | 0.003 | 0.005 | 0.002 | 0.048 | 0.005 | 0.004 | 0.001 | 0.001 | 0.036 | ns | ns | ||
| B × A | ns | ns | ns | ns | ns | ns | ns | ns | ns | ns | ns | ||
| P × B × A | ns | ns | ns | ns | ns | ns | ns | ns | ns | ns | ns | ||
| Treatment | pH | EC | CWS | C | N | ||
|---|---|---|---|---|---|---|---|
| Brassica | Non-bioaugmented | Control | 7.5 ± 0.2 | 1.6 ± 0.3 | 344 ± 39 | 13.1 ± 1.5 | 0.26 ± 0.02 |
| Compost | 7.8 ± 0.0 | 1.6 ± 0.4 | 680 ± 70 | 12.8 ± 1.5 | 0.38 ± 0.04 | ||
| Cow slurry | 7.6 ± 0.1 | 1.6 ± 0.1 | 568 ± 34 | 18.3 ± 6.8 | 0.40 ± 0.07 | ||
| Bioaugmented | Control | 7.6 ± 0.1 | 1.4 ± 0.2 | 328 ± 83 | 14.1 ± 1.6 | 0.28 ± 0.02 | |
| Compost | 7.7 ± 0.0 | 1.5 ± 0.4 | 650 ± 97 | 16.5 ± 2.6 | 0.39 ± 0.01 | ||
| Cow slurry | 7.7 ± 0.1 | 1.7 ± 0.4 | 490 ± 73 | 14.2 ± 0.7 | 0.37 ± 0.01 | ||
| Unplanted | Non-bioaugmented | Control | 7.7 ± 0.0 | 1.3 ± 0.0 | 316 ± 39 | 15.2 ± 4.6 | 0.28 ± 0.04 |
| Compost | 7.7 ± 0.0 | 1.9 ± 0.3 | 645 ± 110 | 14.3 ± 1.8 | 0.38 ± 0.02 | ||
| Cow slurry | 7.6 ± 0.0 | 1.9 ± 0.2 | 648 ± 34 | 13.4 ± 0.5 | 0.36 ± 0.00 | ||
| Bioaugmented | Control | 7.6 ± 0.1 | 1.5 ± 0.1 | 278 ± 134 | 16.9 ± 2.6 | 0.31 ± 0.04 | |
| Compost | 7.7 ± 0.1 | 1.4 ± 0.4 | 572 ± 79 | 16.9 ± 2.5 | 0.40 ± 0.04 | ||
| Cow slurry | 7.5 ± 0.0 | 1.7 ± 0.1 | 553 ± 122 | 13.8 ± 2.4 | 0.34 ± 0.03 | ||
| Plant (P) | ns | ns | ns | ns | ns | ||
| Bioaugmentation (B) | ns | ns | ns | ns | ns | ||
| Amendment (A) | 0.003 | ns | 0.000 | ns | 0.000 | ||
| P × B | ns | ns | ns | ns | ns | ||
| P × A | 0.041 | ns | ns | ns | ns | ||
| B × A | ns | ns | ns | ns | ns | ||
| P × B × A | ns | ns | ns | ns | ns | ||
| Treatment | R | NPM | GLU | GLM | PHO | XYL | LEU | ALA | DEH | SUL | URE | ||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Brassica | Non-bioaugmented | Control | 1.8 ± 0.2 f | 31 ± 4 | 349 ± 60 | 223 ± 19 fg | 2054 ± 381 bc | 72 ± 10 e | 330 ± 48 | 497 ± 52 | 17 ± 4 | 49 ± 6 | 17 ± 6 |
| Compost | 3.4 ± 0.1 c | 3 ± 0 | 478 ± 51 | 337 ± 16 de | 2091 ± 98 bc | 123 ± 19 de | 443 ± 20 | 650 ± 60 | 99 ± 4 | 71 ± 7 | 19 ± 1 | ||
| Cow slurry | 4.1 ± 0.1 a | 130 ± 14 | 728 ± 109 | 515 ± 65 b | 2387 ± 181 b | 221 ± 4 b | 787 ± 32 | 1239 ± 92 | 278 ± 21 | 106 ± 5 | 20 ± 3 | ||
| Bioaugmented | Control | 1.9 ± 0.3 f | 35 ± 4 | 327 ± 37 | 205 ± 17 g | 2117 ± 120 bc | 71 ± 6 e | 343 ± 13 | 568 ± 20 | 24 ± 3 | 46 ± 20 | 9 ± 3 | |
| Compost | 2.9 ± 0.2 d | 10 ± 7 | 382 ± 45 | 293 ± 59 ef | 1628 ± 109 bc | 94 ± 8 de | 370 ± 76 | 558 ± 77 | 96 ± 12 | 76 ± 7 | 13 ± 0 | ||
| Cow slurry | 3.7 ± 0.1 b | 111 ± 8 | 620 ± 121 | 480 ± 43 bc | 2431 ± 341 b | 189 ± 24 bc | 736 ± 21 | 1239 ± 57 | 290 ± 36 | 99 ± 10 | 16 ± 4 | ||
| Unplanted | Non-bioaugmented | Control | 1.3 ± 0.1 g | 25 ± 1 | 322 ± 46 | 181 ± 31 g | 1143 ± 84 c | 61 ± 6 e | 269 ± 25 | 467 ± 44 | 16 ± 8 | 48 ± 2 | 13 ± 2 |
| Compost | 2.6 ± 0.1 de | 3 ± 2 | 406 ± 54 | 325 ± 24 e | 1521 ± 90 bc | 122 ± 17 de | 433 ± 8 | 640 ± 52 | 85 ± 6 | 75 ± 2 | 18 ± 2 | ||
| Cow slurry | 4.2 ± 0.2 a | 108 ± 4 | 568 ± 78 | 684 ± 86 a | 1771 ± 154 bc | 220 ± 37 b | 866 ± 168 | 1380 ± 310 | 198 ± 46 | 85 ± 11 | 21 ± 2 | ||
| Bioaugmented | Control | 1.3 ± 0.1 g | 16 ± 9 | 356 ± 152 | 411 ± 76 cd | 4703 ± 184 a | 407 ± 100 a | 207 ± 97 | 827 ± 566 | 13 ± 7 | 48 ± 2 | 13 ± 2 | |
| Compost | 2.4 ± 0.1 e | 7 ± 1 | 304 ± 63 | 177 ± 39 g | 1217 ± 113 c | 66 ± 22 e | 341 ± 54 | 536 ± 29 | 81 ± 14 | 64 ± 3 | 22 ± 3 | ||
| Cow slurry | 4.3 ± 0.1 a | 73 ± 36 | 401 ± 24 | 485 ± 17 bc | 1566 ± 35 bc | 141 ± 11 cd | 718 ± 40 | 987 ± 73 | 198 ± 28 | 82 ± 4 | 21 ± 7 | ||
| Plant (P) | 0.000 | 0.001 | 0.003 | 0.034 | ns | 0.001 | ns | ns | 0.000 | 0.015 | 0.046 | ||
| Bioaugmentation (B) | 0.005 | ns | 0.008 | 0.033 | 0.024 | 0.036 | 0.005 | ns | ns | ns | ns | ||
| Amendment (A) | 0.000 | 0.000 | 0.000 | 0.000 | 0.003 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | ||
| P × B | 0.026 | ns | ns | ns | 0.006 | 0.000 | ns | ns | ns | ns | 0.007 | ||
| P × A | 0.000 | 0.027 | 0.023 | 0.001 | 0.004 | 0.000 | ns | ns | 0.000 | 0.022 | ns | ||
| B × A | 0.023 | 0.009 | 0.091 | 0.000 | 0.000 | 0.000 | ns | 0.040 | ns | ns | ns | ||
| P × B × A | 0.021 | ns | ns | 0.000 | 0.001 | 0.000 | ns | ns | ns | ns | ns | ||
| Treatment | CMB | Bacterial Abundance | Fungal Abundance | ||
|---|---|---|---|---|---|
| Brassica | Non-bioaugmented | Control | 960 ± 202 | 0.9 × 108 ± 3 × 107 | 0.9 × 105 ± 0.3 × 105 e |
| Compost | 1159 ± 260 | 2.1 × 108 ± 4.7 × 107 | 2.4 × 105 ± 0.5 × 105 cde | ||
| Cow slurry | 1505 ± 226 | 2.7 × 108 ± 5.4 × 107 | 8.2 × 105 ± 1.6 × 105 cde | ||
| Bioaugmented | Control | 994 ± 255 | 1.1 × 108 ± 3.1 × 107 | 1.7 × 105 ± 1.7 × 105 de | |
| Compost | 1672 ± 106 | 2.4 × 108 ± 2.5 × 107 | 8.6 × 105 ± 0.1 × 105 cd | ||
| Cow slurry | 1882 ± 304 | 3.9 × 108 ± 7.7 × 107 | 30 × 105 ± 2.7 × 105 a | ||
| Unplanted | Non-bioaugmented | Control | 1410 ± 312 | 0.9 × 108 ± 1.6 × 107 | 2.3 × 105 ± 0.4 × 105 cde |
| Compost | 1855 ± 221 | 3.5 × 108 ± 9.4 × 107 | 16.9 × 105 ± 4.7 × 105 b | ||
| Cow slurry | 1630 ± 122 | 3.4 × 108 ± 6.5 × 107 | 25.2 × 105 ± 9.7 × 105 a | ||
| Bioaugmented | Control | 1894 ± 292 | 1.0 × 108 ± 2.1 × 107 | 4.6 × 105 ± 1.4 × 105 cde | |
| Compost | 2639 ± 463 | 2.3 × 108 ± 4.2 × 107 | 9.5 × 105 ± 3.1 × 105 c | ||
| Cow slurry | 1908 ± 498 | 3.0 × 108 ± 4.4 × 107 | 17.7 × 105 ± 5.9 × 105 b | ||
| Plant (P) | 0.000 | ns | 0.004 | ||
| Bioaugmentation (B) | 0.000 | ns | 0.048 | ||
| Amendment (A) | 0.001 | 0.000 | 0.000 | ||
| P × B | ns | 0.008 | 0.000 | ||
| P × A | 0.011 | ns | ns | ||
| B × A | ns | ns | ns | ||
| P × B × A | ns | ns | 0.000 | ||
| Treatment | lag | t1/2 | Slope | AWCDt1/2 | AWCDmax | NUSt1/2 | NUSmax | H’ | ||
|---|---|---|---|---|---|---|---|---|---|---|
| Brassica | Non-bioaugmented | Control | 19 ± 0.0 | 45.1 ± 3.4 | 2.46 ± 0.3 ab | 0.54 ± 0.0 | 1.43 ± 0.0 a | 20 ± 1.1 | 30 ± 0.0 ab | 4.15 ± 0.08 |
| Compost | 19 ± 0.0 | 44.1 ± 1.0 | 2.10 ± 0.1 bc | 0.49 ± 00.0 | 1.19 ± 0.1 cde | 17 ± 1.0 | 29 ± 0.0 bc | 3.90 ± 0.10 | ||
| Cow slurry | 15 ± 0.0 | 36.4 ± 0.8 | 2.60 ± 0.3 a | 0.44 ± 0.0 | 1.16 ± 0.0 de | 18 ± 0.5 | 30 ± 0.6 a | 4.01 ± 0.05 | ||
| Bioaugmented | Control | 19 ± 0.0 | 47.3 ± 1.8 | 2.04 ± 0.1 c | 0.47 ± 0.0 | 1.25 ± 0.1 bcde | 19 ± 0.7 | 30 ± 0.6 ab | 4.14 ± 0.06 | |
| Compost | 19 ± 0.0 | 43.1 ± 2.4 | 2.23 ± 0.2 bc | 0.47 ± 0.0 | 1.21 ± 0.0 cde | 18 ± 1.0 | 29 ± 1.0 bc | 3.93 ± 0.19 | ||
| Cow slurry | 15 ± 0.0 | 34.1 ± 0.8 | 2.64 ± 0.3 a | 0.42 ± 0.0 | 1.12 ± 0.0 ef | 16 ± 0.8 | 30 ± 0.0 ab | 3.84 ± 0.07 | ||
| Unplanted | Non-bioaugmented | Control | 20 ± 0.0 | 60.1 ± 6.4 | 1.58 ± 0.1 d | 0.52 ± 0.0 | 1.33 ± 0.0 abc | 20 ± 2.1 | 30 ± 0.6 ab | 4.12 ± 0.21 |
| Compost | 20 ± 0.0 | 57.8 ± 1.8 | 1.65 ± 0.1 d | 0.54 ± 0.0 | 1.38 ± 0.1 ab | 21 ± 0.7 | 29 ± 0.0 bc | 4.23 ± 0.01 | ||
| Cow slurry | 15 ± 0.0 | 38.8 ± 3.9 | 2.18 ± 0.3 bc | 0.49 ± 0.0 | 1.28 ± 0.0 bcd | 18 ± 1.7 | 30 ± 0.6 ab | 4.00 ± 0.15 | ||
| Bioaugmented | Control | 20 ± 0.0 | 55.3 ± 6.1 | 1.39 ± 0.2 de | 0.39 ± 0.1 | 1.01 ± 0.1 f | 18 ± 2.2 | 28 ± 0.6 c | 3.99 ± 0.13 | |
| Compost | 20 ± 0.0 | 64.1 ± 3.1 | 1.16 ± 0.1 e | 0.40 ± 0.0 | 1.00 ± 0.1 f | 19 ± 1.9 | 27 ± 1.0 d | 4.10 ± 0.15 | ||
| Cow slurry | 15 ± 0.0 | 37.3 ± 3.5 | 2.34 ± 0.1 abc | 0.47 ± 0.0 | 1.25 ± 0.0 bcde | 18 ± 1.1 | 30 ± 0.0 ab | 3.98 ± 0.08 | ||
| Plant (P) | 0.001 | 0.000 | 0.000 | ns | ns | 0.041 | 0.001 | ns | ||
| Bioaugmentation (B) | ns | ns | ns | 0.000 | 0.000 | ns | 0.003 | ns | ||
| Amendment (A) | 0.000 | 0.000 | 0.000 | ns | ns | 0.003 | 0.000 | 0.026 | ||
| P × B | ns | ns | ns | 0.010 | 0.001 | ns | 0.045 | ns | ||
| P × A | 0.018 | 0.000 | 0.024 | 0.002 | 0.001 | ns | ns | ns | ||
| B × A | ns | ns | 0.050 | 0.018 | 0.004 | ns | ns | ns | ||
| P × B × A | ns | ns | 0.027 | ns | 0.006 | ns | 0.023 | ns | ||
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Anza, M.; Salazar, O.; Epelde, L.; Becerril, J.M.; Alkorta, I.; Garbisu, C. Remediation of Organically Contaminated Soil Through the Combination of Assisted Phytoremediation and Bioaugmentation. Appl. Sci. 2019, 9, 4757. https://doi.org/10.3390/app9224757
Anza M, Salazar O, Epelde L, Becerril JM, Alkorta I, Garbisu C. Remediation of Organically Contaminated Soil Through the Combination of Assisted Phytoremediation and Bioaugmentation. Applied Sciences. 2019; 9(22):4757. https://doi.org/10.3390/app9224757
Chicago/Turabian StyleAnza, Mikel, Oihane Salazar, Lur Epelde, José María Becerril, Itziar Alkorta, and Carlos Garbisu. 2019. "Remediation of Organically Contaminated Soil Through the Combination of Assisted Phytoremediation and Bioaugmentation" Applied Sciences 9, no. 22: 4757. https://doi.org/10.3390/app9224757
APA StyleAnza, M., Salazar, O., Epelde, L., Becerril, J. M., Alkorta, I., & Garbisu, C. (2019). Remediation of Organically Contaminated Soil Through the Combination of Assisted Phytoremediation and Bioaugmentation. Applied Sciences, 9(22), 4757. https://doi.org/10.3390/app9224757






