Genomic and Microscopic Analysis of Ballast Water in the Great Lakes Region
Abstract
1. Introduction
- Size class greater than 50 µM in minimum dimension (normally assumed to be zooplankton) Less than 10 viable organisms per cubic metre.
- Size class between 10 and 50 µM in minimum dimension. Less than 10 viable organisms per mL.
- Indicator bacteria:
- ○
- Vibrio cholera; Less than 1 colony forming unit per 100 mL.
- ○
- Escherichia coli; Less than 250 colony forming units per 100 mL.
- ○
- Intestinal Enterococci; Less than 100 colony forming unit per 100 mL.
2. Materials and Methods
2.1. Collection and Shipping of Water Samples
2.2. Processing of Samples
- An assessment of the total numbers of phytoplankton,
- The taxonomic groupings of phytoplankton present and their numbers,
- Size of the specific algal species present,
- Molecular analyses to provide an estimate of relative abundance of phytoplankton and bacteria including rare species.
2.3. Light Microscopy
2.4. High Throughput Sequencing (HTS) for Phytoplankton (and Bacterial) Species
- 16S Amplicon PCR Forward Primer = 5′TCGTCGGCAGCGTCAGATGTGTATAAGAGACAGCCTACGGGNGGCWGCAG
- 16S Amplicon PCR Reverse Primer = 5′GTCTCGTGGGCTCGGAGATGTGTATAAGAGACAGGACTACHVGGGTATCTAATCC
- 18S Algal Amplicon PCR Forward Primer-5′P73 (forward primer)—AAT CAG TTA TAG TTT ATT TGR TGG TACC
- 18S Algal Amplicon PCR Reverse Primer-5′P47 (reverse primer)—TCT CAG GCT CCC TCT CCG GA
2.5. Determination of Biovolume
2.6. Comparison of Taxonomic Groups Determined by Microscopy and Molecular Sequencing
3. Results
3.1. Microscopic Counts
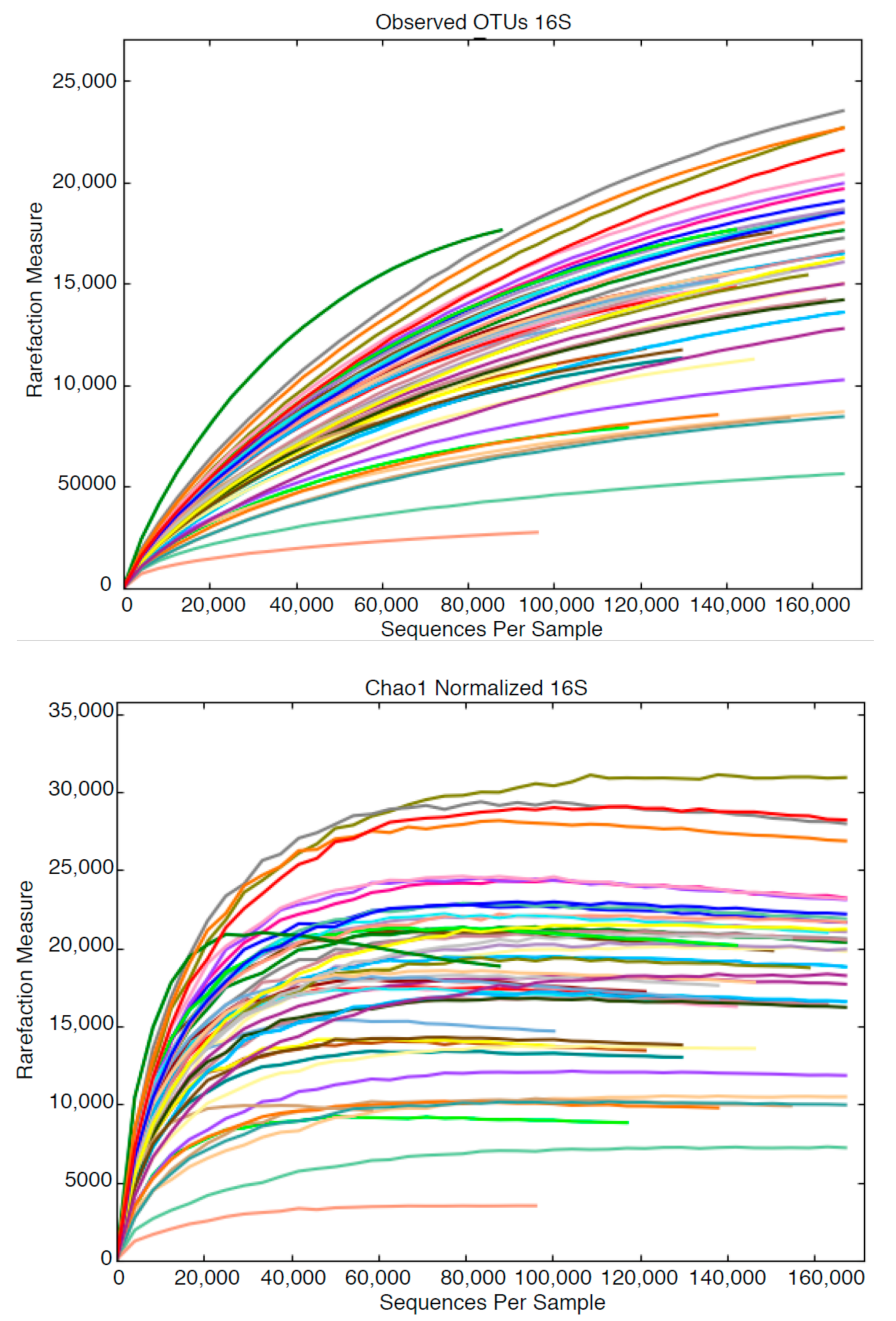
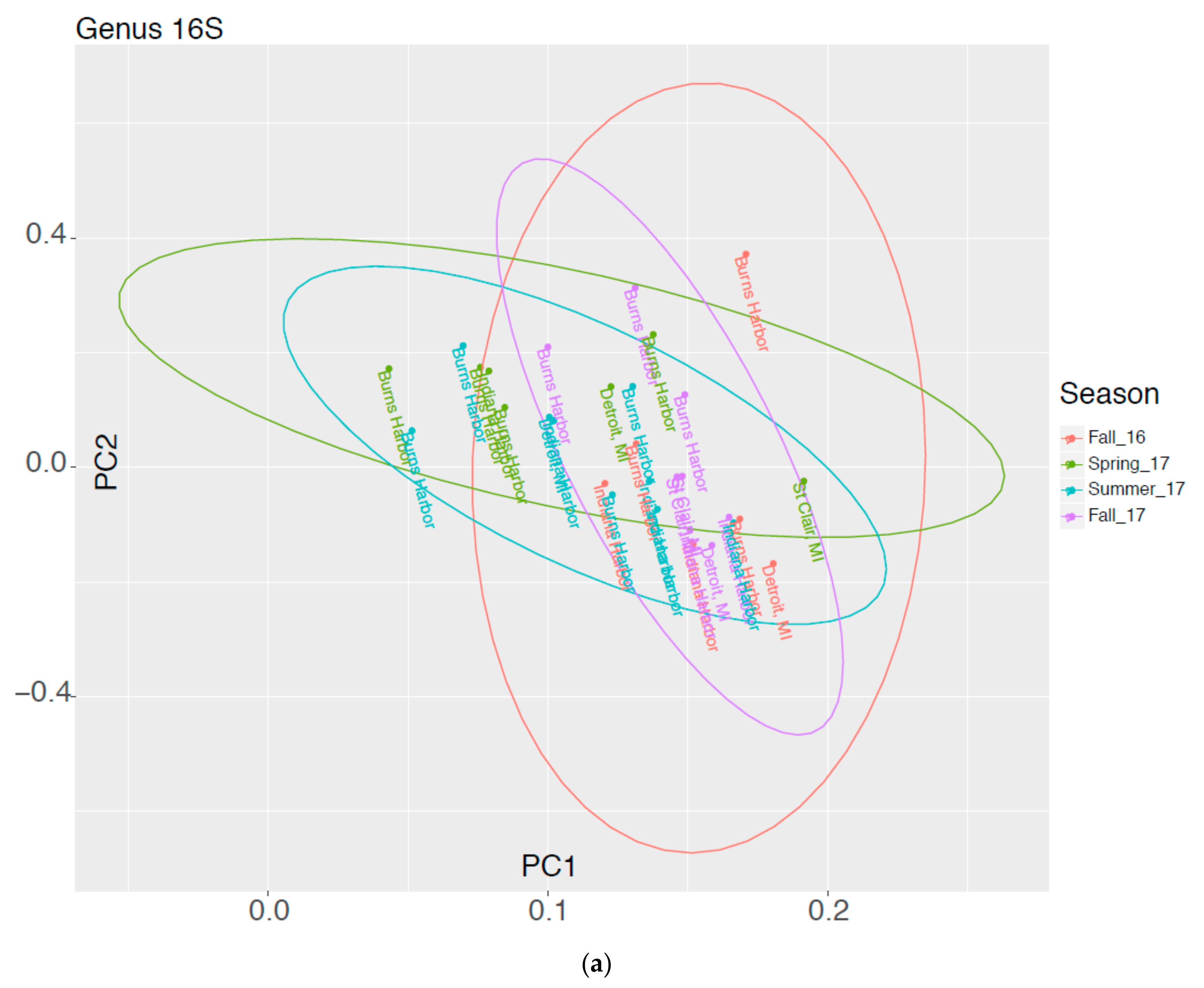
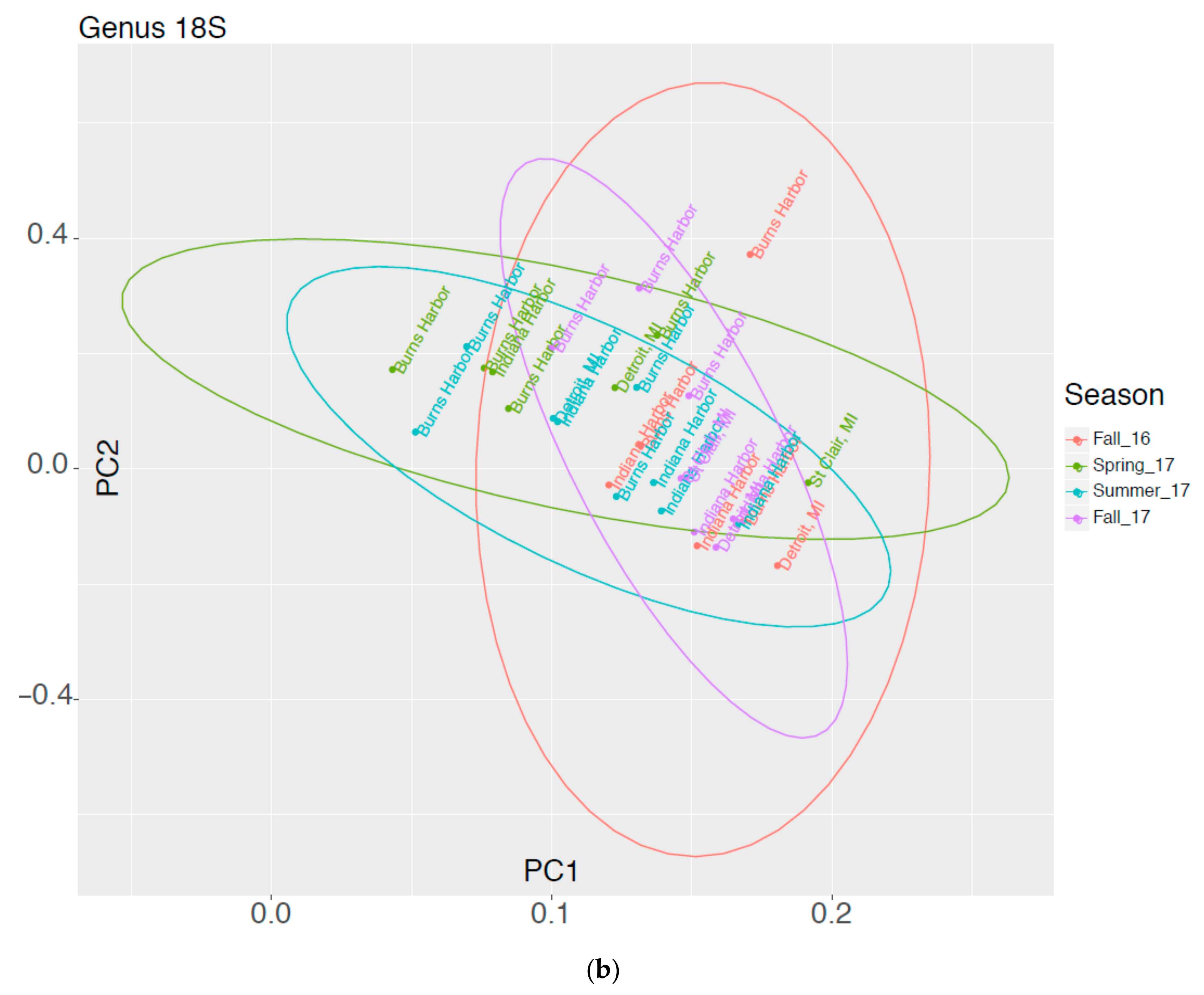
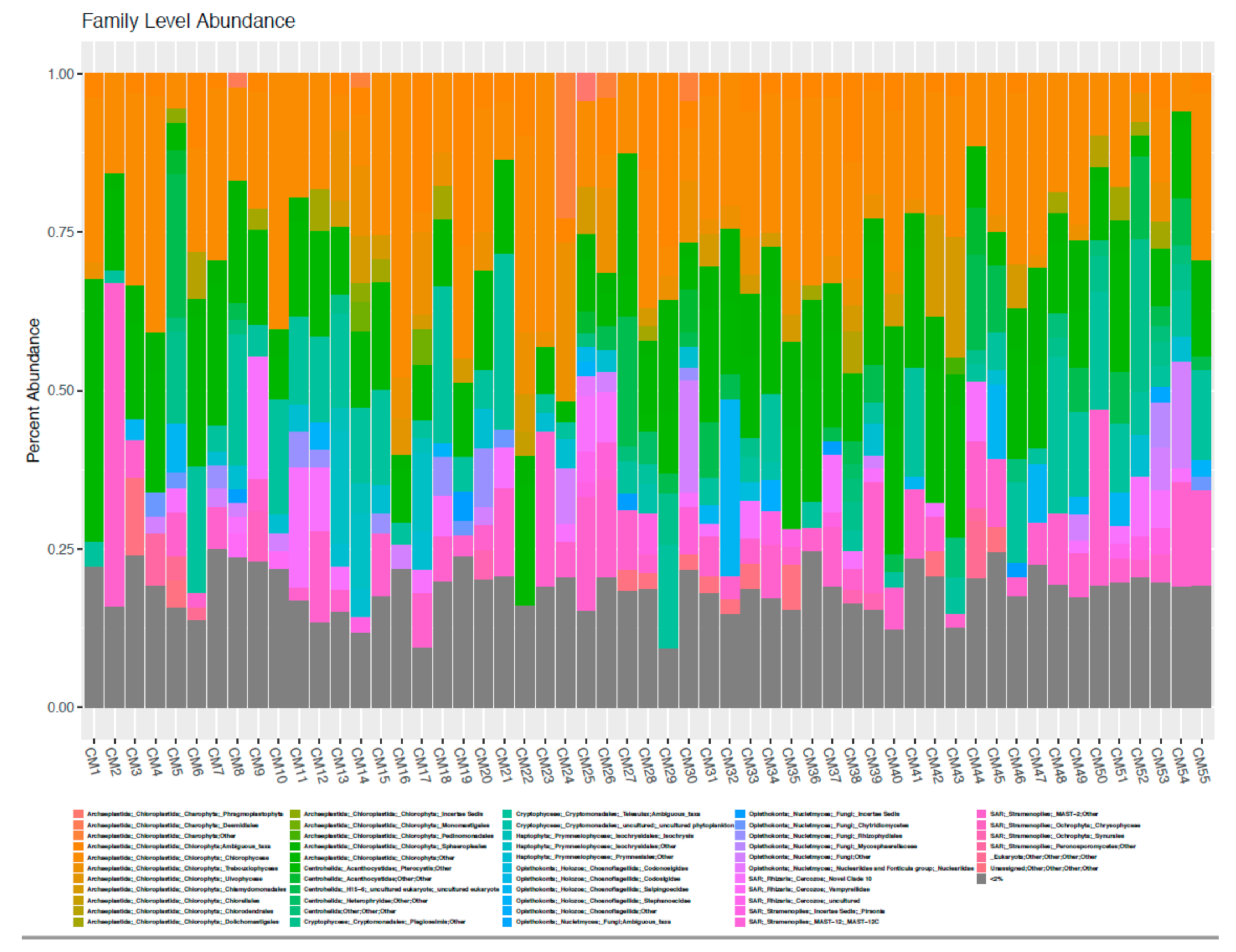
3.2. High Throughput Nucleic Acid Sequencing
3.3. Comparison of Taxonomic Profiles Derived from Microscopy and Molecular Methods
4. Discussion
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Carlton, J.T.; Geller, J.B. Ecological roulette—The global transport of nonindigenous marine organisms. Science 1993, 261, 78–82. [Google Scholar] [CrossRef] [PubMed]
- Matej, D.; Gollasch, S. Global Maritime Transport and Ballast Water Management; Issues and Solutions—Springer Series in Invasion Ecology; Springer: Dordrecht, The Netherlands, 2015; Volume 8. [Google Scholar]
- Pimentel, D.; McNair, S.; Janecka, J.; Wightman, J.; Simmonds, C.; O’Connell, C.; Wong, E.; Russel, L.; Zern, J.; Aquino, A.; et al. Economic and environmental threats of alien plant, animal, and microbe invasions. Agric. Ecosyst. Environ. 2001, 84, 1–20. [Google Scholar] [CrossRef]
- Orieska, M.P.J.; Aldridge, D.C. Estimating the economic costs of invasive alien species as a tool for prioritization. In CAB Reviews: Perspectives in Agriculture, Veterinary Science, Nutrition and Natural Resources; Centre for Agriculture and Bioscience International: Oxfordshire, UK, 2011; Volume 6, ISSN 1749-8848. [Google Scholar]
- Williams, F.; Eschen, R.; Harris, A.; Djeddour, D.; Pratt, C.; Shaw, R.S.; Varia, S.; Lamontagne-Godwin, J.; Thomas, S.E.; Murphy, S.T. The Economic Cost of Invasive Non-Native Species on Great Britain; Report to the Scottish Government, DEFRA & Department for Economy & Transport, Welsh Assembly Government, CAB/001/09; CABI: Wallingford, UK, 2010. [Google Scholar]
- IMO (International Maritime Organization). International Convention for the Control and Management of Ships’ Ballast Water and Sediments; International Maritime Organization: London, UK, 2004. [Google Scholar]
- Elskus, A.; Mitchelmore, C.; Wright, D.; Henquinet, J.; Welschmeyer, N.; Flynn, C.; Watten, B. Efficacy and residual toxicity of a sodium hydroxide based ballast water treatment system for freshwater bulk freighters. J. Great Lakes Res. 2017, 43, 744–754. [Google Scholar] [CrossRef]
- Castro, M.; Veldhuis, M. Temporal changes in phytoplankton biomass and cellular properties; implications for the IMO ballast water convention. Environ. Technol. 2018, 40, 1455–1466. [Google Scholar] [CrossRef] [PubMed]
- Bradie, J.; Broeg, K.; Gianoli, C.; Heitmüller, S.; Lo Curto, A.; Nakata, A.; Rolke, M.; Schillak, L.; Stehouwer, P.; Byllaart, J.V.; et al. A shipboard comparison of analytic methods for ballast water compliance monitoring. J. Sea Res. 2018, 133, 11–19. [Google Scholar] [CrossRef]
- Wright, D.A. Problems associated with performance and compliance testing for ballast water treatment. J. Mar. Des. Oper. 2007, 12, 25–38. [Google Scholar]
- Hess-Erga, O.-K.; Moreno-Andrés, J.; Enger, Ǿ.; Vadstein, O. Microorganisms in ballast water. Disinfection, community dynamics and implications for management. Sci. Total Environ. 2019, 657, 704–716. [Google Scholar] [CrossRef] [PubMed]
- Kagami, M.; Urabe, J. Phytoplankton growth rate as a function of cell size: An experimental test in Lake Biwa. Limnology 2001, 2, 111–117. [Google Scholar]
- Liebich, V.; Stehouwer, P.; Veldhuis, M. Regrowth of potential invasive phytoplankton following UV-based ballast water treatment. Aquat. Invasions 2012, 7, 29–36. [Google Scholar] [CrossRef]
- Jerde, C.L.; Chadderton, W.I.; Mahon, A.R.; Renshaw, M.A.; Corush, J.; Budny, M.L.; Mysorekar, S.; Lodge, D.M. Detection of Asian Carp DNA as part of a Great Lakes surveillance program. Can. J. Fish. Aquat. Sci. 2013, 70, 522–526. [Google Scholar] [CrossRef]
- Darling, J.A.; Frederick, R.M. Nucleic acids-based tools for ballast water surveillance, monitoring, and research. J. Sea Res. 2018, 133, 43–52. [Google Scholar] [CrossRef] [PubMed]
- Pagenkopp Lohan, K.M.; Fleischer, R.C.; Carney, K.J.; Kolzer, K.K.; Ruiz, G.M. Molecular characterization of protistan species and communities in ships’ ballast water across three U.S. coasts. Biodivers. Res. 2017, 23, 680–691. [Google Scholar] [CrossRef]
- Gerhard, W.A.; Gunsch, C.K. Metabarcoding and machine learning analysis of environmental DNA in ballast water arriving to hub ports. Environ. Int. 2019, 124, 312–319. [Google Scholar] [CrossRef] [PubMed]
- Xaoi, X.; Sogge, H.; Lagesen, K.; Tooming-Klunderud, A.; Jakobsen, K.S.; Rohrlak, T. Use of high throughput sequencing and light microscopy show contrasting results in a study of phytoplankton occurrence in a freshwater environment. PLoS ONE 2014, 9, e106510. [Google Scholar] [CrossRef] [PubMed]
- Klindworth, A.; Pruesse, E.; Schweer, T.; Peplles, J.; Quast, C.; Horn, M.; Glockner, F.O. Evaluation of general 16S ribosomal RNA gene PCR primers for classical and nextgeneration sequencing-based diversity studies. Nucleic Acids Res. 2013, 41, e1. [Google Scholar] [CrossRef] [PubMed]
- Bérard, A.; Dorigo, U.; Humbert, J.F.; Martin-Laurent, F. Microalgae community structure analysis based on 18S rDNA amplification from DNA extracted directly from soil as a potential soil bioindicator. Agron. Sustain. Dev. 2005, 25, 285–291. [Google Scholar] [CrossRef]
- Kovala, P.E.; Larrance, J.P. Computation of Phytoplankton Cell Numbers, Cell Volume, Cell Surface Area and Plasma Volume per Litre, from Microscopic Counts; Special Report; University of Washington: Seattle, WA, USA, 1966; Volume 38, pp. 1–91. [Google Scholar]
- Edler, L. (Ed.) Recommendations on the Methods for Marine Biological Studies in the Baltic Seas; Phytoplankton and Chlorophyll. The Baltic Marine Biologists Publication no. 5; Springer: Stockholm, Sweden, 1979. [Google Scholar]
- Hillebrand, H.; Durselen, C.-D.; Kikrschtel, D.; Pollingher, U.; Zohary, T. Biovolume calculation for pelagic and benthic microalgae. J. Phycol. 1999, 35, 403–424. [Google Scholar] [CrossRef]
- Sun, J.; Liu, D. Geometric model for calculating cell biovolume and surface area for plankton. J. Plankton Res. 2003, 25, 1331–1346. [Google Scholar] [CrossRef]
- Olenina, I.; Hajdu, S.; Edler, L.; Andersson, A.; Wasmund, N.; Busch, S.; Göbel, J.; Gromisz, S.; Huseby, S.; Huttunen, M.; et al. Biovolumes and size-classes of phytoplankton in the Baltic Sea. In HELCOM Baltic Sea Environment Proceedings; Helsinki Commission: Helsinki, Finland, 2006; 144p. [Google Scholar]
- Vadrucci, M.R.; Cabrini, M.; Basset, A. Biovolume determination of phytoplankton guilds in transitional water ecosystems of Mediterranean Ecoregion. Transit. Waters Bull. 2007, 2, 83–102. [Google Scholar]
- Smayda, T.J. What to count. In Phytoplankton Manual; Sournia, A., Ed.; UNESCO: Paris, France, 1978; p. 337. [Google Scholar]
- Gollasch, S.; David, M. Ballast water sampling and sample analysis for compliance. In Global Maritime Transport and Ballast Water Management; Invading Nature—Springer Series on Invasion Ecology; Springer: Dordrecht, The Netherlands, 2015; pp. 171–223. [Google Scholar]
- Hou, Y.; Lin, S. Distinct gene number-genome size relationships for eukaryotes and non-eukaryotes: Gene content estimation for dinoflagellate genomes. PLoS ONE 2009, 4, e6978. [Google Scholar] [CrossRef] [PubMed]


| Ratio of Total Cell Area to Total Cell Volume | Cytoplasmic Layer Thickness |
|---|---|
| X < 0.35 | 2 μm |
| 0.35 ≤ X > 0.50 | 1.5 μm |
| 0.50–0.89 | 1 μm |
| >0.90 | PV = Total Volume |
| Code | Shape | Volume | Surface Area |
|---|---|---|---|
| 1 | SPHERE | π/6*D3 | π*D2 |
| 2 | CYLINDER | π/4*D2*H | π*D*(D/2 + H) |
| 3 | ELLIPSEOID | π/6*D*H*W | |
| 4 | CONE | π/12*D2*H | |
| 5 | TRAPEZOIDAL PRISM | 1/2 *H*DE*(W + W2) | SHAPE NOT IN CURRENT USE |
| 6 | CUBE | W3 | 6*W2 |
| 7 | RETANGULAR BOX | H*W*DE | (2*H*W) + (2*W*DE) + (2**H*DE) |
| 8 | TRUNCATED CONE | π/12*H*(D2 + (D*D2) + D22 | π/4*(D22 + D2 + 2H*(D2 + D) |
| 9 | TRIANGULAR PRISM | 1/2 *H*W*D | (W*DE) + (3*H*W) |
| 10 | ELLIPTICAL PRISM | π/4*H*W*DE | π/2*(H*W + (H + W)*DE |
| 11 | CONE-HALF SPHERE | π/12*D2*(H + D) | 1/2*π*D*(L + D) |
| 12 | NOT ASSIGNED | ||
| 13 | NOT ASSIGNED | ||
| 14 | DUMBELL | 2*(π/6*D3) | 2*π*D2 |
| 15 | PRISM ON PARALLELOGRAM | π/2*H*W*DE | *DE |
| 16 | PYRAMID | 1/3H*W*DE | SHAPE NOT IN CURRENT USE |
| 17 | CYLINER-2 HALF SPHERES | π*D2*((H/4 + D/6) | π*D*(D + H) |
| 18 | PROLATE SPHERE | π/6*D2*H | |
| 19 | 2 CONES | π/12*D2*H | |
| 20 | HALF SPHERE | π/12*D3 | 3π/4*D2 |
| 21 | CYLINDER + CONE | ||
| 22 | CYMBELLOID | USE ELLIPESOID EQUATIONS | USE ELLIPESOID EQUATIONS |
| 23 | 2 ELLIPOSEOID | π/6*D*H*W*2 | |
| 24 | SICKLE-SHAPED PRISM | π/4*H*DE*W | π/4*(H + W + DE*W + H*DE) + H*DE |
| Indiana Harbor, Indiana | ||||||||||||||
| 2 October 2016 | 4 October 2016 | 25 November 2016 | 3 April 2017 | 25 April 2017 | 27 June 2017 | |||||||||
| Cell No. | % | Cell No. | % | Cell No. | % | Cell No. | % | Cell No. | % | Cell No. | % | |||
| Diatoms | 109.3 | 5.1 | 28.3 | 1.9 | 219.3 | 78 | 36.3 | 18.8 | 109.6 | 77.2 | 0 | 0 | ||
| Green Algae | 1734 | 82.1 | 1188 | 78.8 | 27 | 9.6 | 34.5 | 17.9 | 17.6 | 12.4 | 46.2 | 61.8 | ||
| Cyanobact eria | 268.0 | 12.7 | 270.9 | 18 | 31.0 | 11 | 120.2 | 62.2 | 14.8 | 10.4 | 28.5 | 38.2 | ||
| Dinoflagell ates | 1.3 | 0.1 | 20 | 1.3 | 4.0 | 1.4 | 2.1 | 1.1 | 0 | 0 | 0 | 0 | ||
| 27 July 2017 | 22 August 17 | 6 September 17 | 13 September 17 | 20 September 17 | 25 September 17 | |||||||||
| Cell No. | % | Cell No. | % | Cell No. | % | Cell No. | % | Cell No. | % | Cell No. | % | |||
| Diatoms | 28 | 7.7 | 200.1 | 24.6 | 187.3 | 5.5 | 325 | 41 | 225.7 | 21.3 | 42.2 | 1.8 | ||
| Green Algae | 88.0 | 24.2 | 109 | 13.4 | 288.2 | 8.4 | 162 | 20.4 | 74.3 | 7.7 | 138.0 | 5.9 | ||
| Cyano bacteria | 244.0 | 67.2 | 498.5 | 61.4 | 2926 | 85.8 | 304 | 38.4 | 750.0 | 71.4 | 2142 | 92.2 | ||
| Dino flagellates | 3.0 | 0.9 | 4.0 | 0.6 | 9.7 | 0.3 | 12.5 | 0.2 | 1.0 | 0 | 0 | 0 | ||
| Burns Harbor, Indiana | ||||||||||||||
| 1 October 2016 | 30 October 2016 | 17 November 2016 | 24 December 16 | 3 December 16 | 3 April 2017 | 9 May 17 | ||||||||
| Cell No. | % | Cell No. | % | Cell No. | % | Cell No. | % | Cell No. | % | Cell No. | % | Cell No. | % | |
| Diatoms | 130.1 | 51 | 261.0 | 61.6 | 1042 3 | 78.6 | 341.4 | 80.9 | 96.2 | 83.1 | 2.8 | 13.4 | 857. 1 | 98.8 |
| Green Algae | 106.5 | 41.8 | 96.0 | 22.6 | 24.5 | 1.8 | 40.5 | 9.6 | 12.7 | 11 | 16.4 | 78.8 | 5.36 | 0.6 |
| Cyanobact | 11 | 4.3 | 65.5 | 15.4 | 256 | 19.3 | 36.0 | 8.5 | 6.4 | 5.5 | 1.2 | 5.8 | 0.9 | 0.1 |
| eria | ||||||||||||||
| Dinoflagellates | 7.4 | 2.9 | 1.3 | 0.4 | 3.0 | 0.3 | 4.0 | 0.5 | 0.4 | 0.4 | 0.4 | 2 | 3.7 | 0.5 |
| 17 May 17 | 26 July 2017 | 10 August 17 | 13 September 17 | 12 October 2017 | 13 October 2017 | |||||||||
| Cell No. | % | Cell No. | % | Cell No. | % | Cell No. | % | Cell No. | % | Cell No. | % | |||
| Diatoms | 100.4 | 79.3 | 220.5 | 49.3 | 294.0 | 81.5 | 326.5 | 29.9 | 120.0 | 40 | 297.0 | 41.3 | ||
| Green Algae | 2.1 | 1.7 | 142.0 | 31.8 | 55.0 | 15.2 | 69.5 | 6.3 | 133 | 44.4 | 111.5 | 15.5 | ||
| Cyanobact eria | 24 | 19 | 67.0 | 15 | 6.7 | 1.8 | 683.0 | 62.5 | 45.4 | 15.1 | 303.0 | 42.2 | ||
| Dinoflagellates | 0 | 0 | 17.0 | 3.9 | 5.0 | 1.5 | 14.2 | 2.3 | 1.0 | 0.5 | 7.0 | 1 | ||
| Monroe, Michigan | ||||||||||||||
| 9 September 16 | 5 October 2016 | 13 October 2016 | 28 September 16 | 17 May 17 | 23 June 2017 | |||||||||
| Cell No. | % | Cell No. | % | Cell No. | % | Cell No. | % | Cell No. | % | Cell No. | % | |||
| Diatoms | 208.9 | 13 | 501.6 | 46.6 | 119.4 | 37.3 | 241.8 | 71.6 | 227.7 | 58.7 | 76.8 | 31.5 | ||
| Green Algae | 92.0 | 5.7 | 185.0 | 17.2 | 24.3 | 7.6 | 10.3 | 3 | 20.5 | 5.3 | 26.7 | 10.9 | ||
| Cyanobact | 1066 | 66.5 | 292.0 | 27.2 | 176.5 | 55 | 85.7 | 25.4 | 139.6 | 36 | 76.0 | 31.2 | ||
| eria | ||||||||||||||
| Dinoflagellates | 235.7 | 14.7 | 96.3 | 9 | 0 | 0 | 0 | 0 | 0 | 0 | 64.3 | 26.4 | ||
| St Clair, Michigan | ||||||||||||||
| 1 May 17 | 14 May 17 | 6 June 2017 | 15 October 2017 | 22 October 2017 | ||||||||||
| Cell No. | % | Cell No. | % | Cell No. | % | Cell No. | % | Cell No. | % | |||||
| Diatoms | 212. 6 | 92.2 | 573.3 | 93 | 69.0 | 58 | 43.8 | 39.5 | 45.3 | 17.8 | ||||
| Green Algae | 14.8 | 6.4 | 40.7 | 6.6 | 47.0 | 39.5 | 11.7 | 10.5 | 33.0 | 13 | ||||
| Cyano bacteria | 3.2 | 1.4 | 0 | 0 | 0.7 | 0.8 | 54.4 | 49 | 175.4 | 69.2 | ||||
| Dinoflagellates | 0 | 0 | 2.0 | 0.4 | 2.3 | 1.7 | 1.0 | 1 | 0 | 0 | ||||
| Detroit, Michigan | ||||||||||||||
| 1 December 16 | 17 May 17 | 4 July 17 | 7 August 17 | 9 October 2017 | ||||||||||
| Cell No. | % | Cell No. | % | Cell No. | % | Cell No. | % | Cell No. | % | |||||
| Diatoms | 55.7 | 72.2 | 500.0 | 98.6 | 99.0 | 14.6 | 66.5 | 1.5 | 82.0 | 29.6 | ||||
| Green Algae | 1.8 | 2.3 | 3.0 | 0.6 | 77.0 | 11.3 | 17.0 | 0 | 54.0 | 19.5 | ||||
| Cyano bacteria | 19.6 | 25.4 | 0 | 0 | 500.0 | 73.6 | 4299. 0 | 98.5 | 135.0 | 48.7 | ||||
| Dinoflagellates | 0 | 0 | 4.0 | 0.8 | 3.0 | 0.5 | 3.0 | 0 | 6.3 | 2.2 | ||||
| Ashtabula, Ohio | ||||||||||||||
| 14 August 17 | 30 August 17 | 27 September 17 | 2 October 2017 | |||||||||||
| Cell No. | % | Cell No. | % | Cell No. | % | Cell No. | % | |||||||
| Diatoms | 121. 1 | 12.8 | 75.2 | 33.7 | 570.2 | 13.5 | 166.0 | 15 | ||||||
| Green Algae | 70.8 | 7.4 | 132.5 | 59.3 | 394 | 9.3 | 151.5 | 13.7 | ||||||
| Cyanobact | 746 | 78.6 | 16.0 | 7 | 3235. 0 | 76.5 | 787.5 | 71 | ||||||
| eria | 0 | |||||||||||||
| Dinoflagellates | 11.3 | 1.2 | 0 | 0 | 30.0 | 0.7 | 3.0 | 0.3 | ||||||
| ESSEXVILLE, MICHIGAN | TWO HARBORS, MINNESOTA | SUPERIOR, WISCONSIN | ||||||||||||
| 7 June 2017 | 23 October 2017 | 31 October 2017 | ||||||||||||
| Cell No. | % | Cell No. | % | Cell No. | % | |||||||||
| Diatoms | 366 | 14.3 | 10.0 | 3.1 | 98.0 | 21.6 | ||||||||
| Green Algae | 105 4.0 | 41.3 | 13.0 | 4.1 | 76.5 | 16.5 | ||||||||
| Cyanobacteria | 110 8.0 | 43.4 | 237.0 | 92.8 | 284.5 | 61.4 | ||||||||
| Dinoflagellates | 26.0 | 1 | 0 | 0 | 4.5 | 0.5 | ||||||||
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wright, D.A.; Mitchelmore, C.L.; Place, A.; Williams, E.; Orano-Dawson, C. Genomic and Microscopic Analysis of Ballast Water in the Great Lakes Region. Appl. Sci. 2019, 9, 2441. https://doi.org/10.3390/app9122441
Wright DA, Mitchelmore CL, Place A, Williams E, Orano-Dawson C. Genomic and Microscopic Analysis of Ballast Water in the Great Lakes Region. Applied Sciences. 2019; 9(12):2441. https://doi.org/10.3390/app9122441
Chicago/Turabian StyleWright, David A., Carys L. Mitchelmore, Allen Place, Ernest Williams, and Celia Orano-Dawson. 2019. "Genomic and Microscopic Analysis of Ballast Water in the Great Lakes Region" Applied Sciences 9, no. 12: 2441. https://doi.org/10.3390/app9122441
APA StyleWright, D. A., Mitchelmore, C. L., Place, A., Williams, E., & Orano-Dawson, C. (2019). Genomic and Microscopic Analysis of Ballast Water in the Great Lakes Region. Applied Sciences, 9(12), 2441. https://doi.org/10.3390/app9122441






