Method-Induced Errors in Fractal Analysis of Lung Microscopic Images Segmented with the Use of HistAENN (Histogram-Based Autoencoder Neural Network)
Abstract
Featured Application
Abstract
1. Introduction
- This work assumes hierarchical design using autoencoder neural network (the first hierarchy level) and the fractal-based estimator for complex structure analysis (the second hierarchy level) of segmented images and shows how to select or preselect algorithms in the second hierarchy level algorithm using small data sets and the semisupervised training principle. The choice of the best algorithm can be automated. Manual segmentation of entire images, required for supervised learning for creating training pairs, is not required in this case. It is the main contribution of this paper.
- This paper demonstrates a different approach to the design of image segmentation algorithms, because in majority of papers’ single results of neural networks are provided. There are numerous reasons why single results are provided, such as learning time, but it leads to false final conclusions about the architecture of the neural network. Neural network should be learned multiple times, because such empirical verification leads to different non-optimal networks, and the distribution for most quality parameters is achieved. Single learning gives a single value of neural network quality parameters, so the comparison of two different neural network architectures leads to significant errors.
- This paper addresses the problem of lung autopsy microscopic image analysis that is completely different from examination of regular histological slides of lung tissue. The issue of result variability due to the selection of image segmentation, and image analysis algorithms are discussed. The outcome has good potential for further designing of classification algorithms, which is essential not only for researchers and software developers but also for the forensic pathologist community. Moreover, methods described and discussed in this paper are appropriate for different types of digital images.
- The segmentation algorithm with the use of machine learning approach and comparison of results for two fractal-based algorithms—2D box-counting and fractals related lacunarity—are discussed in this paper. Binary images from the classification algorithm should be achieved, but the inherited properties of histological slides do not allow the discovery of an exact solution. The segmentation algorithm introduces errors and could be considered as noise. Segmented images are processed by fractal algorithms and input data including noise influence on the final variability of estimated fractal descriptors. Low variability of the system is especially important for semisupervised learning, because this type of learning is preferred for the processing of large images with some control of this process by specialists (patomorphologists or cytomorphologists.
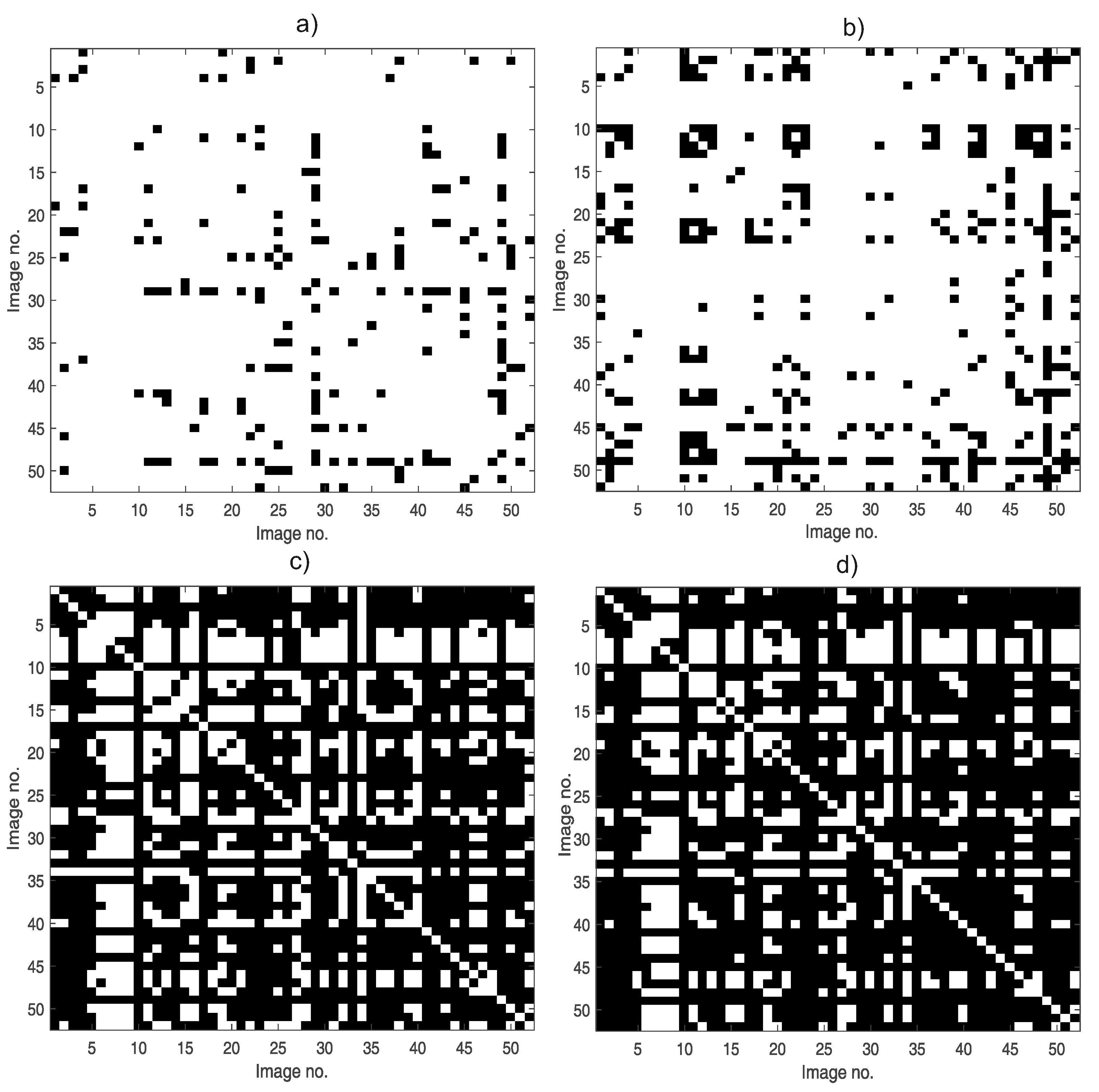
- Method-induced errors could be estimated using the Monte Carlo approach. This work uses 100 HistAENNs trained for every image for the determination of algorithm influence on results. Overlapping tables could be achieved and analyzed for the determination of variability. The selection of a possibly more acceptable algorithm (e.g., fractal) and the selection of parameters for particular algorithm could be attained. The analysis of variability which may be applied for data sets with very raw manual segmentation is most important. Moreover, providing the expected classification results for the selection of segmentation and fractal analysis algorithms is not necessary.
- This paper shows the viability of the designing of segmentation algorithms with the use of neural networks if the appropriate rotational invariance algorithm is applied. It is possible by the application of the Sliding Window Local Histogram (SWLH) to achieve desired invariance. The training of such rotational invariance inside much deeper and larger CNN (Convolutional Neural Network [6,7,8]) is feasible, but SWLH that is a part of HistAENN simplifies training. SWLH reduces the size of the neural network as well as training time, so Monte Carlo tests are possible with a few days of processing.
2. Related Works
3. Data
4. Methods
4.1. Variability of Estimators
4.2. Manual Segmentation Techniques in Semisupervised Learning
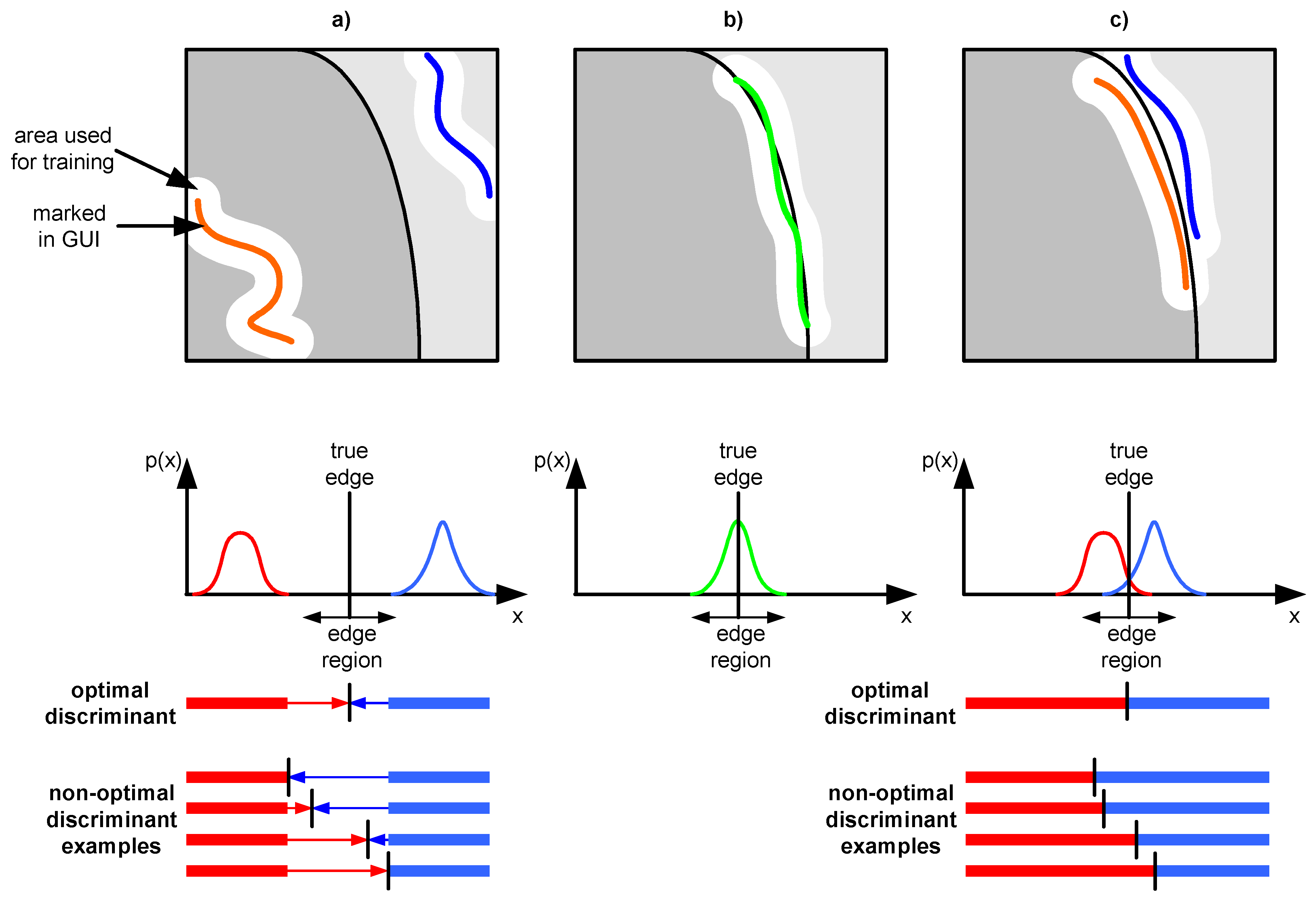
- The inner part of region labeling is straightforward for the user but leads to numerous problems. Separation between two regions could be significant and both distributions do not overlap (Figure 3a). There are numerous possible discriminants which could be achieved during the training of a classifier using typical neural network training algorithms. Optimal discriminant is between both distributions and it is usually not achieved.
- The near-to-edge selection uses previously mentioned properties of distributions to achieve better discrimination. Both selections in this method overlap and both distributions overlap too. This means that the distance between them is much smaller compared to the first method (Figure 3a), but full overlapping is not achieved as in the distribution shown in Figure 3b. The application of typical neural network training algorithms leads to positioning of the discriminant between both overlapped distributions. There is no gap between them, thus an optimal or very close to optimal solution is achieved.
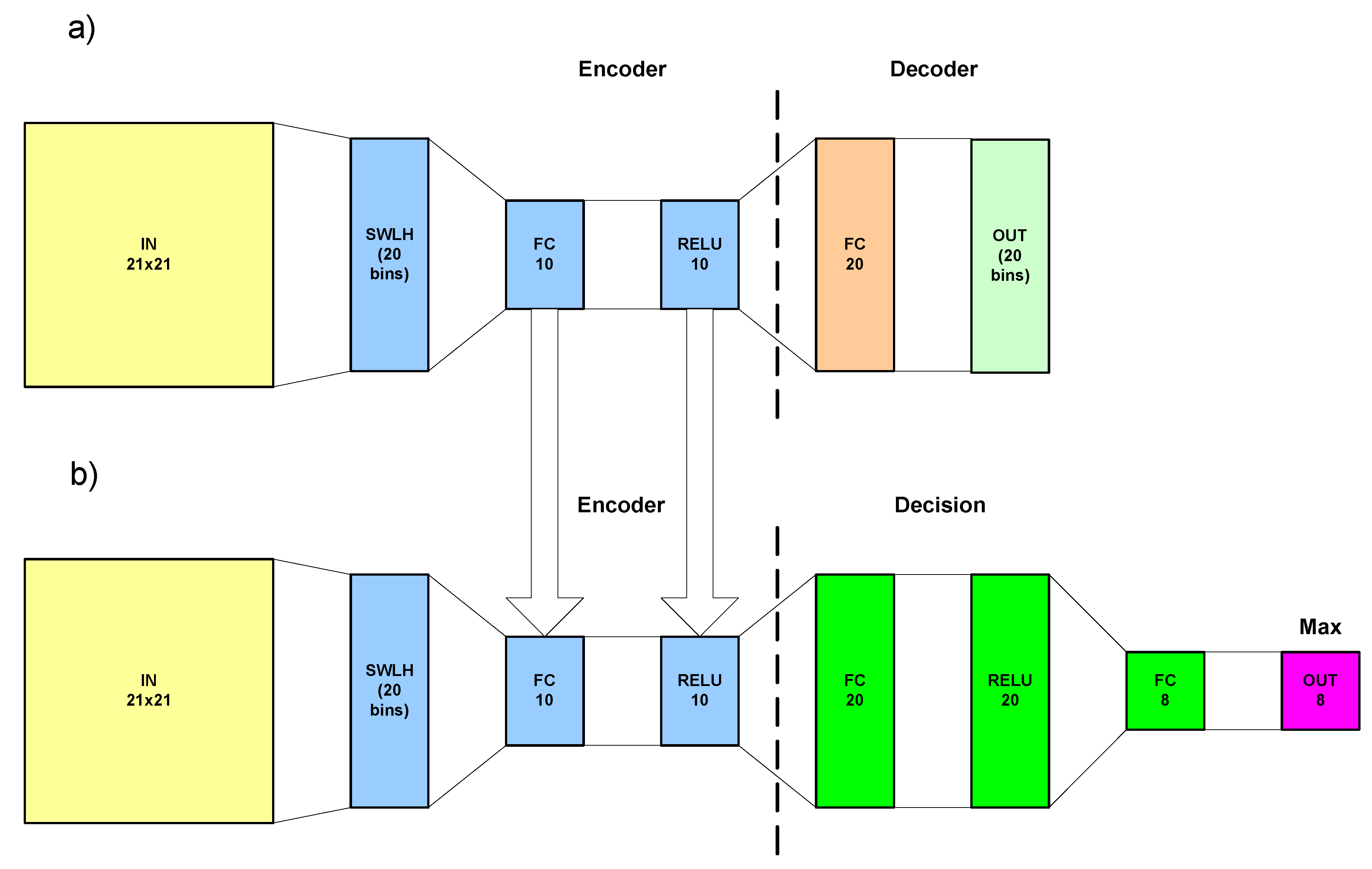
4.3. Architecture of HistAENN and Two-Step Learning
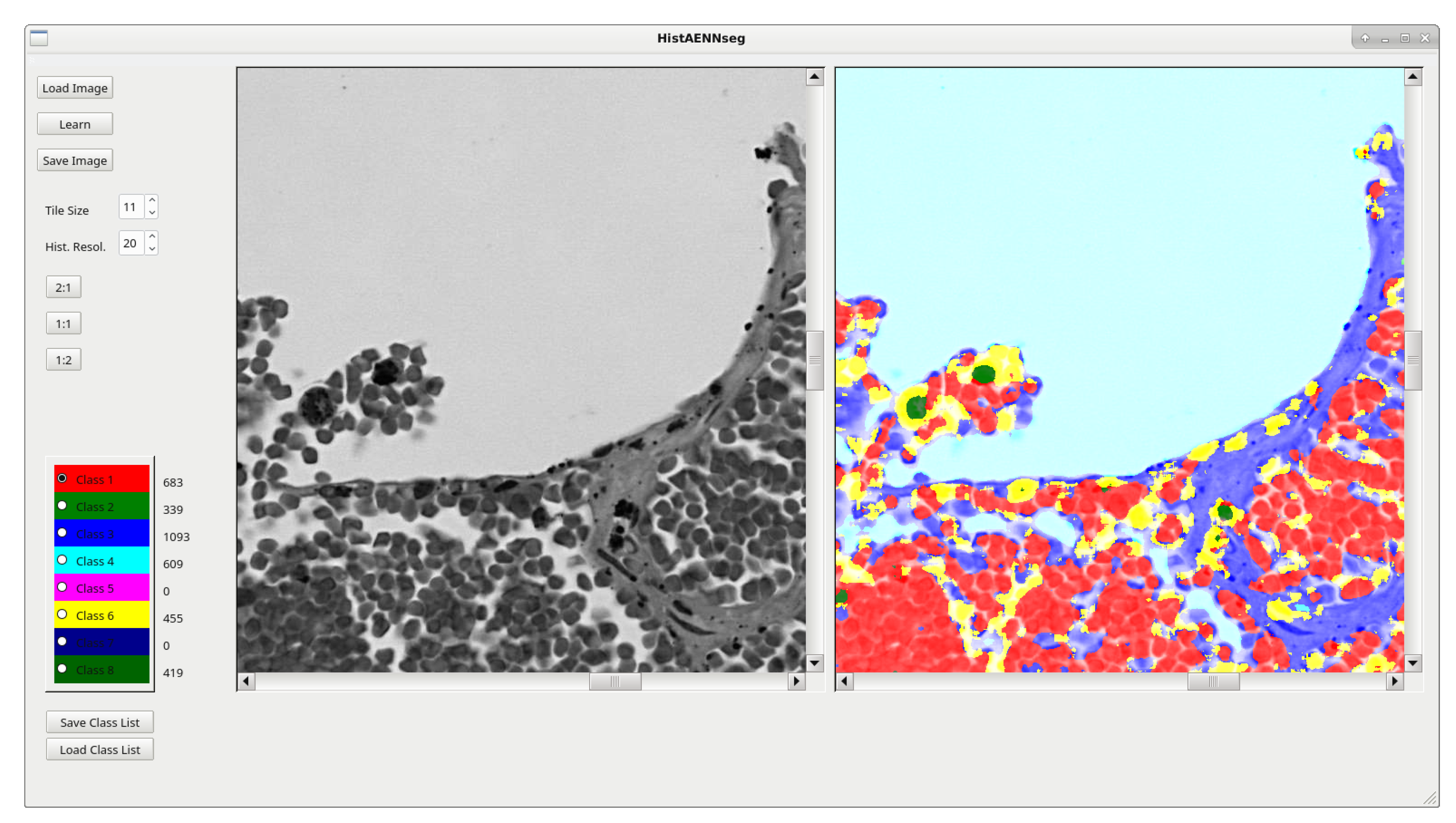
4.4. HistAENNseg—Image Segmentation Software
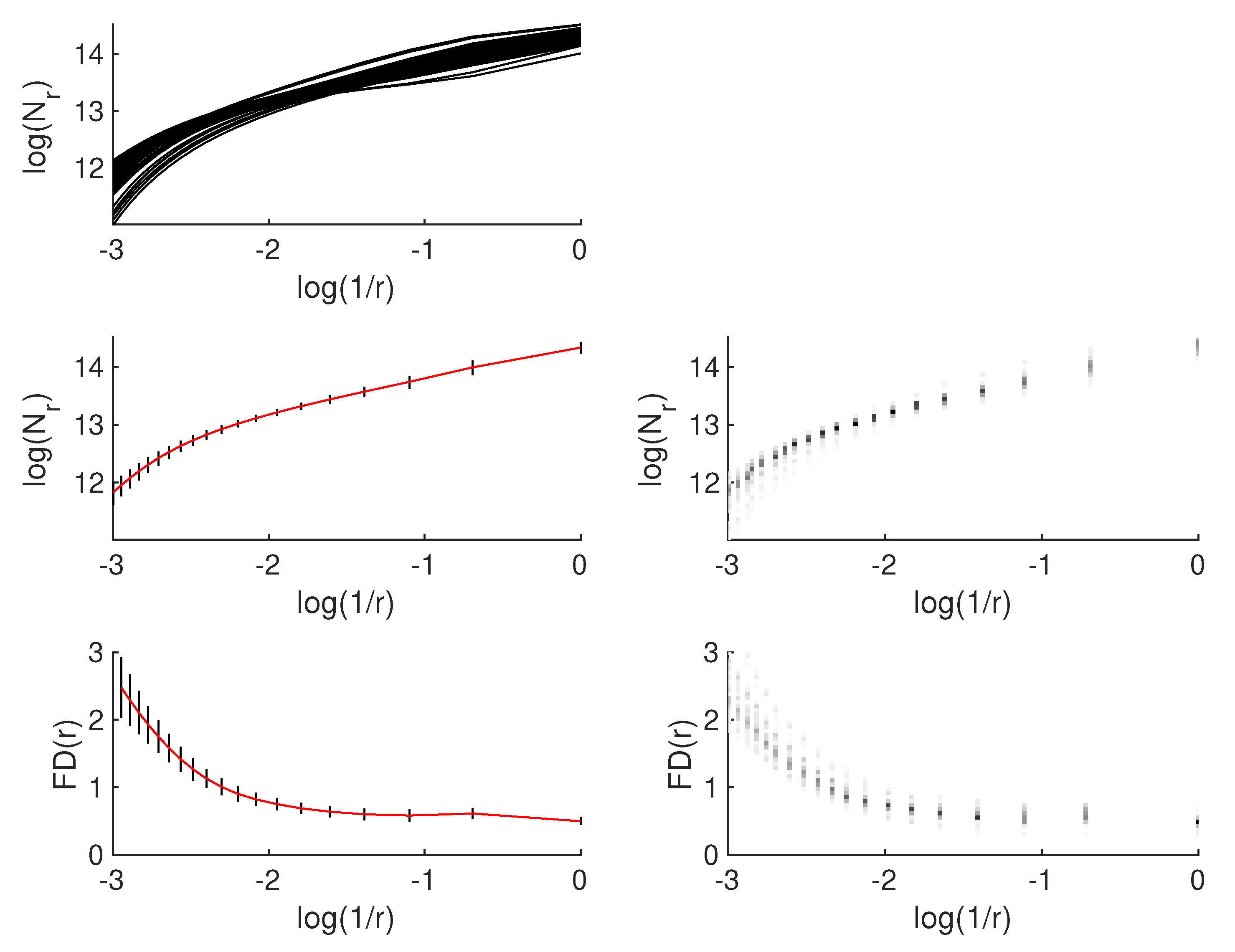
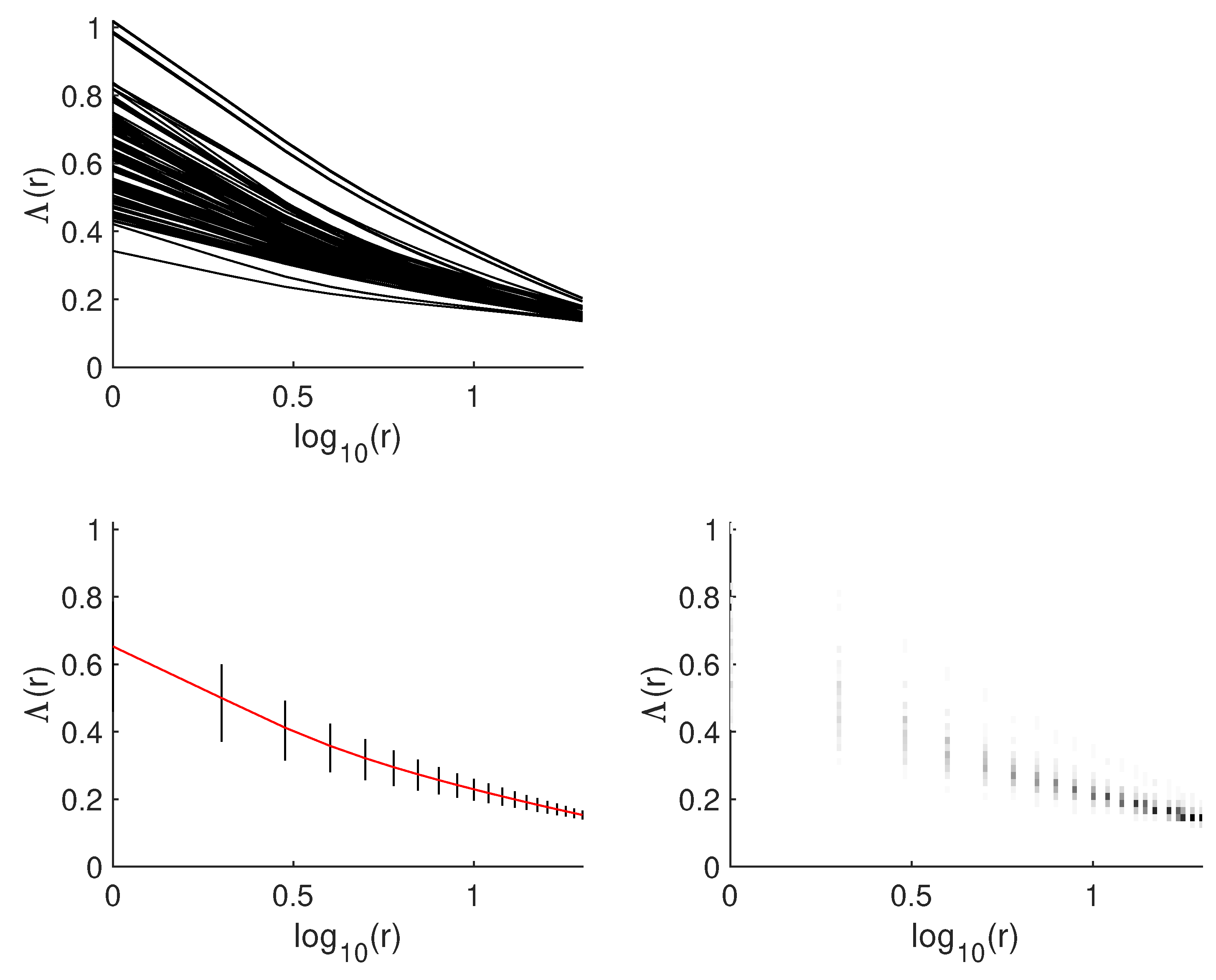
4.5. Fractal-Based Analysis of Microscopic Lung Images
5. Results
6. Discussion
6.1. All-in-One and Hierarchical Segmentations
- All-in-One approach, where a single machine learning algorithm is used for overall image processing. Such an approach could also be used for the final classification purposes, so the input is the image and the output is the recognized class.
- Hierarchical approach, with well-defined processing stages where some of them could be machine learning-based and others could be typical image processing algorithms, such as filters.
6.2. Selection of Learning Principles
- Supervised learning requires input and output image pairs. The input image from each pair is available directly, but the desired output image should be manually segmented. Large slide images and their large data set lead to extremely high costs of design. The advantage is the possibility of designing a fully automatic CADx system.
- Unsupervised learning requires input images only, so the problem of manual segmentation of desired output images is avoided. This approach is based on automatic clustering. The number of classes could be arbitrarily selected or could be estimated automatically [76,77]. Unsupervised learning could be applied for well separated classes directly, but in most cases requires manual fitting of algorithms for a particular data set. This additional effort depends on the image content.
- Semisupervised learning is a promising solution for cost reduction of manual segmentation. This process assumes labeling of very small parts of images that belong to the specific classes. Semisupervised learning could be used for CADx system development or as a CADx working principle. In the latter case, semisupervised learning is used by patomorphologists during the analysis of a particular image. Obtained results are checked and corrected iteratively (Human in The Loop) for segmentation improving to the desired level. A very significant advantage of this learning principle is the possibility of an image segmentation system design without access to all possible image variants.
6.3. Invariant Image Representation and Neural Networks
- Invariant transformations guaranteed by machine learning,
- Invariant transformations guaranteed by preprocessing algorithms.
6.4. Selection of Fractal Descriptors for Image Analysis
7. Final Conclusions and Further Work
Author Contributions
Acknowledgments
Conflicts of Interest
References
- Doi, K. Computer-Aided Diagnosis in Medical Imaging: Historical Review, Current Status and Future Potential. Comput. Med. Imaging Graph. Off. J. Comput. Med. Imaging Soc. 2007, 31, 198–211. [Google Scholar] [CrossRef] [PubMed]
- Li, Q.; Nishikawa, R.M. (Eds.) Computer-Aided Detection and Diagnosis in Medical Imaging; CRC Press: Boca Raton, FL, USA, 2015. [Google Scholar]
- Yu, K.; Hyun, N.; Fetterman, B.; Lorey, T.; Raine-Bennett, T.R.; Zhang, H.; Stamps, R.E.; Poitras, N.E.; Wheeler, W.; Befano, B.; et al. Automated Cervical Screening and Triage, Based on HPV Testing and Computer-Interpreted Cytology. J. Natl. Cancer Inst. 2018, 110, 1222–1228. [Google Scholar] [CrossRef] [PubMed]
- Ceelie, H.; Dinkelaar, R.B.; van Gelder, W. Examination of peripheral blood films using automated microscopy; evaluation of Diffmaster Octavia and Cellavision DM96. J. Clin. Pathol. 2007, 60, 72–79. [Google Scholar] [CrossRef] [PubMed]
- Stouten, K.; Riedl, J.A.; Levin, M.D.; Gelder, W. Examination of peripheral blood smears: Performance evaluation of a digital microscope system using a large-scale leukocyte database. Int. J. Lab. Hematol. 2015, 37, e137–e140. [Google Scholar] [CrossRef] [PubMed]
- LeCun, Y.; Kavukcuoglu, K.; Farabet, C. Convolutional networks and applications in vision. In Proceedings of the 2010 IEEE International Symposium on Circuits and Systems, Paris, France, 30 May–2 June 2010; pp. 253–256. [Google Scholar]
- LeCun, Y.; Bengio, Y.; Hinton, G. Deep learning. Nature 2015, 521, 436–444. [Google Scholar] [CrossRef] [PubMed]
- Goodfellow, I.; Bengio, Y.; Courville, A. Deep Learning (Adaptive Computation and Machine Learning); MIT Press: Cambridge, MA, USA, 2016. [Google Scholar]
- Sun, W.; Xu, G.; Gong, P.; Liang, S. Fractal analysis of remotely sensed images: A review of methods and applications. Int. J. Remote Sens. 2006, 27, 4963–4990. [Google Scholar] [CrossRef]
- Metropolis, N. Monte Carlo Method. In From Cardinals to Chaos: Reflection on the Life and Legacy of Stanislaw Ulam; CUP Archive: Cambridge, UK, 1989; p. 125. [Google Scholar]
- Kroese, D.P.; Brereton, T.; Taimre, T.; Botev, Z.I. Why the Monte Carlo method is so important today. Wiley Interdiscip. Rev. Comput. Stat. 2014, 6, 386–392. [Google Scholar] [CrossRef]
- Lee, D.H. Pseudo-Label: The Simple and Efficient Semi-Supervised Learning Method for Deep Neural Networks. In Proceedings of the ICML 2013 Workshop: Challenges in Representation Learning (WREPL), Atlanta, GA, USA, 16–21 June 2013. [Google Scholar]
- Gan, H.; Sang, N.; Huang, R.; Tong, X.; Dan, Z. Using clustering analysis to improve semi-supervised classification. Neurocomputing 2013, 101, 290–298. [Google Scholar] [CrossRef]
- Schwenker, F.; Trentin, E. Pattern classification and clustering: A review of partially supervised learning approaches. Pattern Recognit. Lett. 2014, 37, 4–14. [Google Scholar] [CrossRef]
- Feng, Z.; Nie, D.; Wang, L.; Shen, D. Semi-supervised learning for pelvic MR image segmentation based on multi-task residual fully convolutional networks. In Proceedings of the 2018 IEEE 15th International Symposium on Biomedical Imaging (ISBI 2018), Washington, DC, USA, 4–7 April 2018; pp. 885–888. [Google Scholar]
- Lopes, R.; Betrouni, N. Fractal and multifractal analysis: A review. Med. Image Anal. 2009, 13, 634–649. [Google Scholar] [CrossRef] [PubMed]
- Xiao, X.; Zhao, J.; Qiang, Y.; Wang, H.; Xiao, Y.; Zhang, X.; Zhang, Y. An Automated Segmentation Method for Lung Parenchyma Image Sequences Based on Fractal Geometry and Convex Hull Algorithm. Appl. Sci. 2018, 8, 832. [Google Scholar] [CrossRef]
- Wen, R.; Sinding-Larsen, R. Uncertainty in Fractal Dimension Estimated from Power Spectra and Variogram. Math. Geol. 1997, 29, 727–753. [Google Scholar] [CrossRef]
- Kendall, A.; Badrinarayanan, V.; Cipolla, R. Bayesian SegNet: Model Uncertainty in Deep Convolutional Encoder-Decoder Architectures for Scene Understanding. arXiv, 2015; arXiv:1511.02680. [Google Scholar]
- Gal, Y. Uncertainty in Deep Learning. Ph.D. Thesis, University of Cambridge, Cambridge, UK, 2016. [Google Scholar]
- Polychronaki, G.E.; Ktonas, P.Y.; Gatzonis, S.; Siatouni, A.; Asvestas, P.A.; Tsekou, H.; Sakas, D.; Nikita, K.S. Comparison of fractal dimension estimation algorithms for epileptic seizure onset detection. J. Neural Eng. 2010, 7, 046007. [Google Scholar] [CrossRef] [PubMed]
- Shi, C.T. Signal Pattern Recognition Based on Fractal Features and Machine Learning. Appl. Sci. 2018, 8, 1327. [Google Scholar] [CrossRef]
- Li, J.; Sun, C.; Du, Q. A New Box-Counting Method for Estimation of Image Fractal Dimension. In Proceedings of the 2006 International Conference on Image Processing, Atlanta, GA, USA, 8–11 October 2006; pp. 3029–3032. [Google Scholar]
- Sanghera, B.; Banerjee, D.; Khan, A.; Simcock, I.; Stirling, J.J.; Glynne-Jones, R.; Goh, V. Reproducibility of 2D and 3D Fractal Analysis Techniques for the Assessment of Spatial Heterogeneity of Regional Blood Flow in Rectal Cancer. Radiology 2012, 263, 865–873. [Google Scholar] [CrossRef] [PubMed]
- Zio, E. Practical Applications of Monte Carlo Simulation for System Reliability Analysis. In The Monte Carlo Simulation Method for System Reliability and Risk Analysis; Chapter: The Monte Carlo Simulation Method for System Reliability and Risk Analysis; Springer: London, UK, 2013; pp. 83–107. [Google Scholar]
- Singh, H.; Pal, P. Article: Software Reliability Testing using Monte Carlo Methods. Int. J. Comput. Appl. 2013, 69, 41–44. [Google Scholar]
- Thompson, N.A.; Weiss, D.J. A Framework for the Development of Computerized Adaptive Tests. Pract. Assess. Res. Eval. 2011, 16, 1–9. [Google Scholar]
- Zhou, B.; Okamura, H.; Dohi, T. Markov Chain Monte Carlo Random Testing. In Advances in Computer Science and Information Technology; Kim, T.H., Adeli, H., Eds.; Springer: Berlin/Heidelberg, Germany, 2010; pp. 447–456. [Google Scholar]
- Schatzmann, E.; Gerrard, R.; Barbour, A.D. Measures of Niche Overlap, I. Math. Med. Biol. J. IMA 1986, 3, 99–113. [Google Scholar] [CrossRef]
- Cornell, H. Encyclopedia of Theoretical Ecology; Chapter: Niche Overlap; University of California Press: Berkeley, CA, USA, 2011; pp. 489–497. [Google Scholar]
- Fasel, B.; Gatica-Perez, D. Rotation-Invariant Neoperceptron. In Proceedings of the 18th International Conference on Pattern Recognition (ICPR’06), Hong Kong, China, 20–24 August 2006; Volume 3, pp. 336–339. [Google Scholar]
- Dieleman, S.; Willett, K.W.; Dambre, J. Rotation-invariant convolutional neural networks for galaxy morphology prediction. MNRAS 2015, 450, 1441–1459. [Google Scholar] [CrossRef]
- Cheng, G.; Zhou, P.; Han, J. RIFD-CNN: Rotation-Invariant and Fisher Discriminative Convolutional Neural Networks for Object Detection. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Las Vegas, NV, USA, 27–30 June 2016. [Google Scholar]
- Cheng, G.; Zhou, P.; Han, J. Learning Rotation-Invariant Convolutional Neural Networks for Object Detection in VHR Optical Remote Sensing Images. IEEE Trans. Geosci. Remote Sens. 2016, 54, 7405–7415. [Google Scholar] [CrossRef]
- Marcos, D.; Volpi, M.; Tuia, D. Learning rotation invariant convolutional filters for texture classification. In Proceedings of the 23rd International Conference on Pattern Recognition, Cancun, Mexico, 4–8 December 2016; pp. 2012–2017. [Google Scholar]
- Levinshtein, A.; Stere, A.; Kutulakos, K.N.; Fleet, D.J.; Dickinson, S.J.; Siddiqi, K. TurboPixels: Fast Superpixels Using Geometric Flows. IEEE Trans. Pattern Anal. Mach. Intell. 2009, 31, 2290–2297. [Google Scholar] [CrossRef] [PubMed]
- Stutz, D.; Hermans, A.; Leibe, B. Superpixels: An evaluation of the state-of-the-art. Comput. Vis. Image Underst. 2018, 166, 1–27. [Google Scholar] [CrossRef]
- Cousty, J.; Bertrand, G.; Najman, L.; Couprie, M. Watershed Cuts: Thinnings, Shortest Path Forests, and Topological Watersheds. IEEE Trans. Pattern Anal. Mach. Intell. 2010, 32, 925–939. [Google Scholar] [CrossRef] [PubMed]
- Ciecholewski, M. River channel segmentation in polarimetric SAR images: Watershed transform combined with average contrast maximisation. Expert Syst. Appl. 2017, 82, 196–215. [Google Scholar] [CrossRef]
- Ciecholewski, M. Malignant and Benign Mass Segmentation in Mammograms Using Active Contour Methods. Symmetry 2017, 9, 277. [Google Scholar] [CrossRef]
- Ding, K.; Xiao, L.; Weng, G. Active contours driven by region-scalable fitting and optimized Laplacian of Gaussian energy for image segmentation. Signal Process. 2017, 134, 224–233. [Google Scholar] [CrossRef]
- Panchonia, A. Histopathological Evaluation of Lung Autopsy: 100 Cases Study. J. Res. Med. Dent. Sci. 2018, 6, 109–112. [Google Scholar] [CrossRef]
- Chauhan, G.; Agrawal, M.; Thakkar, N.; Parghi, B. Spectrum of histopathological lesions in lung autopsy. J. Res. Med. Dent. Sci. 2015, 3, 109. [Google Scholar] [CrossRef]
- Lennon, F.E.; Cianci, G.C.; Cipriani, N.A.; Hensing, T.A.; Zhang, H.J.; Chen, C.T.; Murgu, S.D.; Vokes, E.E.; Vannier, M.; Salgia, R. Lung cancer—A fractal viewpoint. Nat. Rev. Clin. Oncol. 2015, 12, 664–675. [Google Scholar] [CrossRef] [PubMed]
- Kurawar, R.R.; Vasaikar, M.S. Spectrum of Histomorphological Changes in Lungs at Autopsy: A 5 Year Study. Ann. Pathol. Lab. Med. 2017, 4, A106–A112. [Google Scholar] [CrossRef]
- Khare, P.; Gupta, R.; Ahuja, M.; Khare, N.; Agarwal, S.; Bansal, D. Prevalence of Lung Lesions at Autopsy: A Histopathological Study. J. Clin. Diagn. Res. 2017, 11, EC13–EC16. [Google Scholar] [CrossRef] [PubMed]
- Andra Cocariu, E.; Mageriu, V.; Staniceanu, F.; Bastian, A.; Socoliuc, C.; Zurac, S. Correlations Between the Autolytic Changes and Postmortem Interval in Refrigerated Cadavers. Rom. J. Internal Med. 2016, 54, 105–112. [Google Scholar] [CrossRef] [PubMed]
- Goode, A.; Gilbert, B.; Harkes, J.; Jukic, D.; Satyanarayanan, M. OpenSlide: A Vendor-Neutral Software Foundation for Digital Pathology. J. Pathol. Informat. 2013, 4, 8. [Google Scholar]
- Martinez, K.; Cupitt, J. VIPS—A highly tuned image processing software architecture. In Proceedings of the IEEE International Conference on Image Processing, Genova, Italy, 14 September 2005; pp. 574–577. [Google Scholar]
- Baddeley, A.; Jensen, E.B.V. Stereology for Statisticians; Chapman & Hall/CRC: London, UK, 2005. [Google Scholar]
- Mei, S.; Wang, Y.; Wen, G. Automatic Fabric Defect Detection with a Multi-Scale Convolutional Denoising Autoencoder Network Model. Sensors 2018, 18, 1064. [Google Scholar] [CrossRef] [PubMed]
- Liang, P.; Shi, W.; Zhang, X. Remote Sensing Image Classification Based on Stacked Denoising Autoencoder. Remote Sens. 2018, 10, 16. [Google Scholar] [CrossRef]
- MathWorks. Image Processing Toolbox. User’s Guide; MathWorks: Natick, MA, USA, 2018. [Google Scholar]
- King, D.E. Dlib-ml: A Machine Learning Toolkit. J. Mach. Learn. Res. 2009, 10, 1755–1758. [Google Scholar]
- NVIDIA. cuDNN Developer Guide; NVDIA: Santa Clara, CA, USA, 2018. [Google Scholar]
- Stroustrup, B. Programming: Principles and Practical Using C++, 2nd ed.; Addison-Wesley: Boston, MA, USA, 2014. [Google Scholar]
- Mandelbrot, B. The Fractal Geometry of the Nature; W. H. Freeman and Company: Stuttgart, Germany, 1983. [Google Scholar]
- Peitgen, H.; Jürgens, H.; Saupe, D. Fractals for the Classrooms: Part One: Introduction to Fractals and Chaos; Springer-Verlag: Berlin/Heidelberg, Germany, 1991. [Google Scholar]
- Peitgen, H.; Jürgens, H.; Saupe, D. Fractals for the Classrooms: Part Two: Complex Systems and Mandelbrot; Springer-Verlag: Berlin/Heidelberg, Germany, 1992. [Google Scholar]
- Plotnick, R.; Gardner, R.; Hargrove, W.; Prestegaard, K.; Perlmutter, M. Lacunarity analysis: A general technique for the analysis of spatial patterns. Phys. Rev. E 1996, 53, 5461–5468. [Google Scholar] [CrossRef]
- Matheron, G. Principles of geostatistics. Econ. Geol. 1963, 58, 1246–1266. [Google Scholar] [CrossRef]
- Wackernagel, H. Multivariate Geostatistics. A Introduction with Applications; Springer: Berlin/Heidelberg, Germany, 2003. [Google Scholar]
- Clarke, K. Computation of the Fractal Dimension of Topographic Surfaces using the Triangular Prism Surface Area Method. Comput. Geosci. 1986, 12, 713–722. [Google Scholar] [CrossRef]
- Sun, W. Three New Implementations of the Triangular Prism Method for Computing the Fractal Dimension of Remote Sensing Images. Photogramm. Eng. Remote Sens. 2006, 72, 372–382. [Google Scholar] [CrossRef]
- Mandelbrot, B.; Passoja, D.; Paullay, A. Fractal character of fracture surfaces of metals. Nature 1984, 308, 721–722. [Google Scholar] [CrossRef]
- Mazurek, P.; Oszutowska-Mazurek, D. From Slit–Island Method to Ising Model—Analysis of Grayscale Images. Int. J. Appl. Math. Comput. Sci. 2014, 24, 49–63. [Google Scholar] [CrossRef]
- Harte, D. Multifractals. Theory and Applications; Chapman & Hall/CRC: London, UK, 2001. [Google Scholar]
- Seuront, L. Fractals and Multifractals in Ecology and Aquatic Science; CRC Press: Boca Raton, FL, USA, 2010. [Google Scholar]
- Kaye, B. A Random Walk Through Fractal Dimensions; Wiley-VCH: Weinheim, Germany, 1994. [Google Scholar]
- Robbins, H.; Monro, S. A Stochastic Approximation Method. Ann. Math. Stat. 1951, 22, 400–407. [Google Scholar] [CrossRef]
- Mei, S.; Montanari, A.; Nguyen, P.M. A mean field view of the landscape of two-layer neural networks. Proc. Natl. Acad. Sci. USA 2018, 1–7. [Google Scholar] [CrossRef] [PubMed]
- El-Baz, A.; Jilang, X.; Suri, J.S. (Eds.) Biomedical Image Segmentation: Advances and Trends; CRC Press: Boca Raton, FL, USA, 2016. [Google Scholar]
- Fernández-Delgado, M.; Cernadas, E.; Barro, S.; Amorim, D. Do we Need Hundreds of Classifiers to Solve Real World Classification Problems? J. Mach. Learn. Res. 2014, 15, 3133–3181. [Google Scholar]
- Chapella, O.; Schölkopf, B.; Zien, A. (Eds.) Semi–Supervised Learning; MIT Press: Cambridge, MA, USA, 2006. [Google Scholar]
- Albalate, A.; Minker, W. Semi–Supervised and Unsupervised Machine Learning; Wiley: Hoboken, NJ, USA, 2011. [Google Scholar]
- Gan, G.; Ma, C.; Wu, J. Data Clustering. Theory, Algorithms, and Applications; SIAM: Philadelphia, PA, USA, 2007. [Google Scholar]
- Aggarwal, C.C.; Reddy, C.K. (Eds.) Data Clustering. Algorithms and Applications; Chapman & Hall/CRC: London, UK, 2014. [Google Scholar]
| Case | Type | Scales List | |
|---|---|---|---|
| 1 | box-counting | ||
| 2 | box-counting | ||
| 3 | lacunarity | ||
| 4 | lacunarity |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Oszutowska-Mazurek, D.; Mazurek, P.; Parafiniuk, M.; Stachowicz, A. Method-Induced Errors in Fractal Analysis of Lung Microscopic Images Segmented with the Use of HistAENN (Histogram-Based Autoencoder Neural Network). Appl. Sci. 2018, 8, 2356. https://doi.org/10.3390/app8122356
Oszutowska-Mazurek D, Mazurek P, Parafiniuk M, Stachowicz A. Method-Induced Errors in Fractal Analysis of Lung Microscopic Images Segmented with the Use of HistAENN (Histogram-Based Autoencoder Neural Network). Applied Sciences. 2018; 8(12):2356. https://doi.org/10.3390/app8122356
Chicago/Turabian StyleOszutowska-Mazurek, Dorota, Przemyslaw Mazurek, Miroslaw Parafiniuk, and Agnieszka Stachowicz. 2018. "Method-Induced Errors in Fractal Analysis of Lung Microscopic Images Segmented with the Use of HistAENN (Histogram-Based Autoencoder Neural Network)" Applied Sciences 8, no. 12: 2356. https://doi.org/10.3390/app8122356
APA StyleOszutowska-Mazurek, D., Mazurek, P., Parafiniuk, M., & Stachowicz, A. (2018). Method-Induced Errors in Fractal Analysis of Lung Microscopic Images Segmented with the Use of HistAENN (Histogram-Based Autoencoder Neural Network). Applied Sciences, 8(12), 2356. https://doi.org/10.3390/app8122356






