DNA Barcoding and Molecular Phylogenetics Revealed a New Cryptic Bamboo Aphid Species of the Genus Takecallis (Hemiptera: Aphididae)
Abstract
:1. Introduction
2. Materials and Methods
2.1. Specimen Collection and Identification
2.2. DNA Sequencing and Phylogenetic Analysis
2.3. Species Delimitation
3. Results
3.1. Molecular Phylogeny
3.2. Species Delimitation
3.3. Taxonomy
3.3.1. Differential Diagnosis
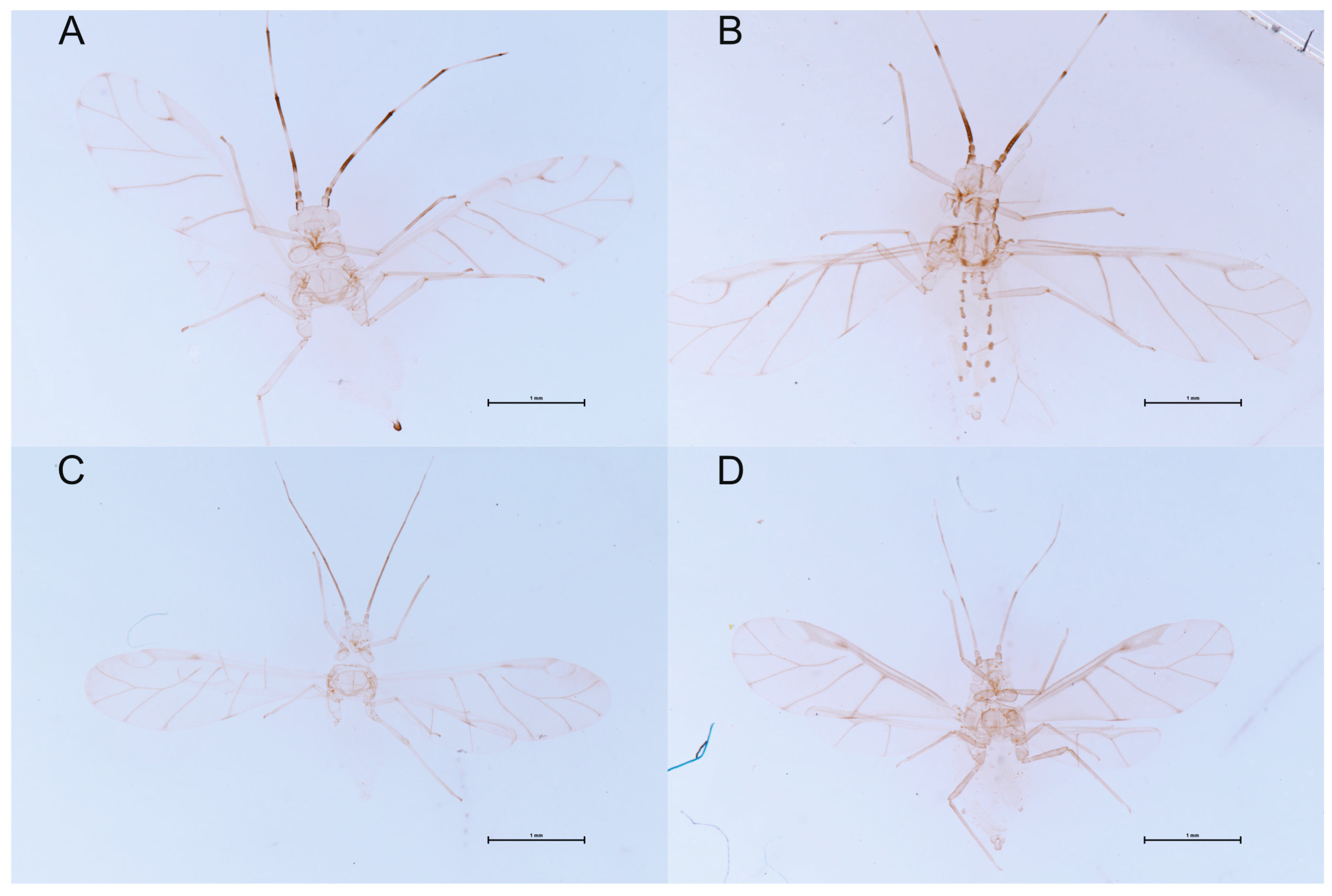
3.3.2. Species Description
3.3.3. Key to Species of the Genus Takecallis
- Antennae longer than body..................................................................................2
- –
- Antennae shorter than body.............................................................................8
- Ultimate rostral segment without any accessory setae. Abdominal tergites 1–7 with paired dark circular spots around spinal setae...............................................T. assumenta
- –
- Ultimate rostral segment with 3–4 accessory setae. Abdominal tergites 1–7 with or without dark spots....................................................................................3
- Antennal segment I with (4-) 5–7 setae. Antennal segments III–VI dark, except for basal part of antennal segment III. Antennal segment III with 4–15 secondary rhinaria................................................................................................4
- –
- Antennal segment I usually with 4, rarely with 3 or 5, setae. Antennal segments III–VI or IV–VI base usually pale basally and dark apically. Antennal segment III with 3–9 secondary rhinaria.....................................................................................6
- Antennal segment III with 10–15 secondary rhinaria, marginal seta on abdominal tergite VI not positioned on base of siphunculus.................................................T. affinis s. str.
- –
- Antennal segment III with 4–7 (often 5) secondary rhinaria, marginal seta on abdominal tergite VI positioned on base of siphunculus............................................5
- Antennae 1.3 times body length, antennal segment I with 5–7 setae, cauda with 18 setae, each lobe of anal plate with 15 setae, spinal abdominal setae on low elevations..........................................................T. affinis ssp. niitakayamensis
- –
- Antennae 0.95–1.14 times body length, antennal segment I with 4 setae, cauda with 8–11 setae, each lobe of anal plate with 5–9 setae, spinal abdominal setae barely visible................................................................ T. nigroantennatus sp. nov.
- Thorax and abdomen usually without dark dorsal markings, never with paired elongate dark abdominal patches. Cauda slightly dusky or blackish...............................................7
- –
- Thorax with variably-developed longitudinal dark stripes, and abdomen with a pair of elongate dark patches on each tergite. Cauda pale or dusky....................T. arundinariae
- Antennae 3.36–4.00 mm, secondary rhinaria densely concentrated on very short dark
- –
- Section of proximal third of antennal segment III...........................................T. alba
- –
- Antennae 2.36–2.51 mm, secondary rhinaria spread over longer dark section oc cupying most of proximal third of antennal segment III...........................................T. arundicolens
- Secondary rhinaria confined to basal third of antennal segment III. Each abdominal tergite bearing only 2 spinal setae .................................................................T. taiwana
- –
- Secondary rhinaria extending about half of length of antennal segment III. Each abdominal tergite bearing at least 4 setae besides the marginal ones...................T. sasae
4. Discussion
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Foottit, R.G.; Maw, H.E.; von Dohlen, C.D.; Hebert, P.D. Species identification of aphids (Insecta: Hemiptera: Aphididae) through DNA barcodes. Mol. Ecol. Resour. 2008, 8, 1189–1201. [Google Scholar] [CrossRef]
- Lozier, J.D.; Foottit, R.G.; Miller, G.L.; Mills, N.J.; Roderick, G.K. Molecular and morphological evaluation of the aphid genus Hyalopterus Koch (Insecta: Hemiptera: Aphididae), with a description of a new species. Zootaxa 2008, 1688, 1–9. [Google Scholar] [CrossRef] [Green Version]
- Théry, T.; Kanturski, M.; Favret, C. Molecular data and species diagnosis in Essigella Del Guercio, 1909 (Sternorrhyncha, Aphididae, Lachninae). ZooKeys 2018, 765, 103–122. [Google Scholar] [CrossRef] [PubMed]
- Barbagallo, S.; Cocuzza, G.E.M. Description of a new Myzocallis (Hemiptera Aphididae) living on Valonia oak in Southern Italy with DNA barcoding accounts on allied species-group. Zootaxa 2022, 5183, 187–202. [Google Scholar] [CrossRef] [PubMed]
- Barjadze, S.; Halbert, S.E.; Ben-Schlomo, R. A new gall-producing species of Geoica Hart, 1894 (Hemiptera: Aphididae: Eriosomatinae) from Israel. Zootaxa 2022, 5183, 343–354. [Google Scholar] [CrossRef]
- Lee, Y.; Lee, W.; Kanturski, M.; Foottit, R.G.; Akimoto, S.I.; Lee, S. Cryptic diversity of the subfamily Calaphidinae (Hemiptera: Aphididae) revealed by comprehensive DNA barcoding. PLoS ONE 2017, 12, e0176582. [Google Scholar] [CrossRef] [Green Version]
- Kanturski, M.; Lee, Y.; Choi, J.; Lee, S. DNA barcoding and a precise morphological comparison revealed a cryptic species in the Nippolachnus piri complex (Hemiptera: Aphididae: Lachninae). Sci. Rep. 2018, 8, 1–16. [Google Scholar] [CrossRef]
- Li, Q.; Wu, L.; Cheng, Z.; Liu, Z.; Huang, X. Two new cryptic species within Periphyllus koelreuteriae (Hemiptera: Aphididae) feeding on Koelreuteria (Sapindaceae) in China. Zootaxa 2022, 5183, 220–238. [Google Scholar] [CrossRef]
- Lee, Y.; Kanturski, M.; Foottit, R.G.; Kim, S.; Lee, S. Molecular phylogeny and evolution of Calaphidinae (Hemiptera: Aphididae). Cladistics 2022, 38, 159–186. [Google Scholar] [CrossRef]
- Favret, C. Aphid Species File. Available online: http://aphid.speciesfile.org/HomePage/Aphid/HomePage.aspx (accessed on 28 March 2023).
- Matsumura, S. A list of the Aphididae of Japan, with description of new species and genera. J. Coll. Agric. Tohoku Imp. Univ. 1917, 7, 351–414. [Google Scholar]
- Higuchi, H. A revision of the genus Takacallis Matsumura. Insecta Matsumurana 1968, 31, 25–34. [Google Scholar]
- Quednau, F.W. Atlas of the Drepanosiphine aphid Part II: Panaphidini Oestlund, 1923 Panaphidina Oestlund, 1923 (Hemiptera: Aphididae: Calaphidinae); American Entomological Institute: Gainesville, FL, USA, 2003; 301p. [Google Scholar]
- Qiao, G.-X.; Zhang, G.-X. Review of the genus Takecallis Matsumura (Homoptera: Aphididae: Myzocallidinae) from China and description of one new species. Raffles Bull. Zool. 2004, 52, 373–378. [Google Scholar]
- Lee, S.; Lee, Y.; Kim, S. Insects Fauna of Korea. Calaphidinae (Arthropoda: Insecta: Hemiptera: Aphididae: Calaphidinae); National Institute of Biological Resources: Incheon, Republic of Korea, 2018; Volume 9, pp. 1–246. [Google Scholar]
- Blackman, R.L.; Eastop, V.F. Aphids of the World’s Plants: An Online Identification and Information Guide. 2023. Available online: http://www.aphidsonworldsplants.info (accessed on 3 April 2023).
- Alford, D.V. Pests of Ornamental Trees, Shrubs and Flowers: A Color Handbook, 2nd ed.; Academic Press: London, UK, 2012. [Google Scholar]
- Wieczorek, K. The first detection of the alien, invasive bamboo aphid species of the genus Takecallis (Hemiptera: Aphididae) in Poland. J. Plant Prot. Res. 2023, 63, 233–238. [Google Scholar] [CrossRef]
- Wieczorek, K.; Chłond, D. Hop-on, hop-off: The first record of the alien species crescent-marked lily aphid (Neomyzus circumflexus) (Insecta, Hemiptera, Aphididae) in Greenland. Polar Res. 2020, 39. [Google Scholar] [CrossRef]
- Ilharco, F.A.; van Harten, A. Systematics. In Aphids: Their Biology, Natural Enemies and Control; Minks, A.K., Harrewijn, P., Eds.; Elsevier Science Publishers: Amsterdam, The Netherlands, 1987; pp. 51–77. [Google Scholar]
- Folmer, O.; Black, M.; Hoeh, W.; Lutz, R.; Vrijenhoek, R. DNA primers for amplification of mitochondrial cytochrome c oxidase subunit I from diverse metazoan invertebrates. Mol. Mar. Biol. Biotechnol. 1994, 3, 294–299. [Google Scholar] [PubMed]
- Palumbi, S.R. Nucleic acids, II: The polymerase chain reaction. In Molecular Systematics; Hillis, D.M., Moritz, C., Mable, B.K., Eds.; Sinauer Associates, Inc.: Sunderland, MA, USA, 1996; pp. 205–247. [Google Scholar]
- von Dohlen, C.D.; Kurosu, U.; Aoki, S. Phylogenetics and evolution of the eastern Asian-eastern North American disjunct aphid tribe, Hormaphidini (Hemiptera: Aphididae). Mol. Phylogenet. Evol. 2002, 23, 257–267. [Google Scholar] [CrossRef]
- Choi, H.; Shin, S.; Jung, S.; Clarke, D.J.; Lee, S. Molecular phylogeny of Macrosiphini (Hemiptera: Aphididae): An evolutionary hypothesis for the Pterocomma-group habitat adaptation. Mol. Phylogenet. Evol. 2018, 121, 12–21. [Google Scholar] [CrossRef] [PubMed]
- Sanger, F.; Nicklen, S.; Coulson, A.R. DNA sequencing with chain-terminating inhibitor. Proc. Natl. Acad. Sci. USA 1977, 74, 5463–5467. [Google Scholar] [CrossRef]
- Geneious R10.2.6. Available online: https://www.geneious.com (accessed on 15 February 2023).
- Bensasson, D.; Zhang, D.X.; Hartl, D.L.; Hewitt, G.M. Mitochondrial pseudogenes: Evolution’s misplaced witnesses. Trends Ecol. Evol. 2001, 16, 314–321. [Google Scholar] [CrossRef]
- Song, H.; Buhay, J.E.; Whiting, M.F.; Crandall, K.A. Many species in one: DNA barcoding overestimates the number of species when nuclear mitochondrial pseudogenes are coamplified. Proc. Natl. Acad. Sci. USA 2008, 105, 13486–13491. [Google Scholar] [CrossRef]
- Katoh, K.; Misawa, K.; Kuma, K.I.; Miyata, T. MAFFT: A novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Res. 2002, 30, 3059–3066. [Google Scholar] [CrossRef] [Green Version]
- Li, Q.; Deng, J.; Chen, C.; Zeng, L.; Lin, X.; Cheng, Z.; Qiao, G.-X.; Huang, X. DNA Barcoding subtropical aphids and implications for population differentiation. Insects 2019, 11, 11. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gwiazdowski, R.A.; Foottit, R.G.; Maw, H.E.; Hebert, P.D. The Hemiptera (Insecta) of Canada: Constructing a reference library of DNA barcodes. PLoS ONE 2015, 10, e0125635. [Google Scholar] [CrossRef] [Green Version]
- Hebert, P.D.; Ratnasingham, S.; Zakharov, E.V.; Telfer, A.C.; Levesque-Beaudin, V.; Milton, M.A.; Pedersen, S.; Jannetta, P.; de Waard, J.R. Counting animal species with DNA barcodes: Canadian insects. Phil. Trans. R Soc. B 2016, 371, 20150333. [Google Scholar] [CrossRef] [Green Version]
- Valenzuela, I.; Boulton, A.; Malipatil, M.B. First record of Takecallis arundinariae (Essig) (Hemiptera: Aphidididae) from Australia. Gen. Appl. Ent. 2010, 39, 23–25. [Google Scholar]
- Nylander, J.A.A. MrModeltest; Version 2; Program Distributed by the Author; Evolutionary Biology Centre, Uppsala University: Uppsala, Sweden, 2004. [Google Scholar]
- Ronquist, F.; Teslenko, M.; van der Mark, P.; Ayres, D.L.; Darling, A.; Höhna, S.; Larget, B.; Liu, L.; Suchard, M.A.; Huelsenbeck, J.P. MrBayes 3.2: Efficient Bayesian phylogenetic inference and model choice across a large model space. Syst. Biol. 2012, 61, 539–542. [Google Scholar] [CrossRef] [Green Version]
- Rambaut, A.; Drummond, A.J.; Xie, D.; Baele, G.; Suchard, M.A. Posterior summarisation in Bayesian phylogenetics using Tracer 1.7. Syst. Biol. 2018, 67, 901–904. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nguyen, L.T.; Schmidt, H.A.; von Haeseler, A.; Minh, B.Q. IQ-TREE: A Fast and Effective Stochastic Algorithm for Estimating Maximum-Likelihood Phylogenies. Mol. Biol. Evol. 2015, 32, 268–274. [Google Scholar] [CrossRef] [PubMed]
- Hoang, D.T.; Chernomor, O.; von Haeseler, A.; Minh, B.Q.; Vinh, L.S. UFBoot2: Improving the Ultrafast Bootstrap Approximation. Mol. Biol. Evol. 2018, 35, 518–522. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular Evolutionary Genetics Analysis across computing platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef]
- Zhang, J.; Kapli, P.; Pavlidis, P.; Stamatakis, A. A general species delimitation method with applications to phylogenetic placements. Bioinformatics 2013, 29, 2869–2876. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Puillandre, N.; Lambert, A.; Brouillet, S.; Achaz, G.J.M.E. ABGD, Automatic Barcode Gap Discovery for primary species delimitation. Mol. Ecol. 2012, 21, 1864–1877. [Google Scholar] [CrossRef]
- Puillandre, N.; Brouillet, S.; Achaz, G. ASAP: Assemble species by automatic partitioning. Mol. Ecol. Resour. 2021, 21, 609–620. [Google Scholar] [CrossRef] [PubMed]
- Dransfield, R.D.; Brightwell, R. InfluentialPoints. Takecallis arundicolens. 2023. Available online: https://influentialpoints.com/Gallery/Takecallis_arundicolens_Black-tailed_bamboo_aphid.htm (accessed on 2 April 2023).
- Stapleton, C. An Online Bamboo Identification. Pests of Bamboos Cultivated in Western Gardens. 2023. Available online: http://bamboo-identification.co.uk/html/pests.html (accessed on 21 March 2023).
- Scheurer, S.; Binazzi, A. Notes on bio-ecology and ethology of Cinara curvipes (Patch), a newly introduced species into Europe (Aphididae Lachninae). Redia 2004, 87, 61–65. [Google Scholar]
- Borowiak-Sobkowiak, B.; Durak, R. Biology and ecology of Appendiseta robiniae (Hemiptera: Aphidoidea)—An alien species in Europe. Cent. Eur. J. Biol. 2012, 7, 487–494. [Google Scholar] [CrossRef] [Green Version]
- Gostel, M.R.; Kress, W.J. The Expanding Role of DNA Barcodes: Indispensable Tools for Ecology, Evolution, and Conservation. Diversity 2022, 14, 213. [Google Scholar] [CrossRef]
- Song, N.; Lin, X.; Zhao, T. Description of the Three Complete Mitochondrial Genomes of Click Beetles (Coleoptera, Elateridae) with Phylogenetic Implications. Taxonomy 2023, 3, 204–220. [Google Scholar] [CrossRef]





| Sample ID | COI | EF1 | Reference | |
|---|---|---|---|---|
| Takecallis nigroantennatus sp. nov—isolate 1BAA | OR001758 | Present Study | OR039806 | Present Study |
| Takecallis nigroantennatus sp. nov—isolate 2BAN | OR001759 | Present Study | OR039807 | Present Study |
| Takecallis nigroantennatus sp. nov—isolate 3BAN | OR001760 | Present Study | OR039808 | Present Study |
| Takecallis nigroantennatus sp. nov—isolate 4BAA | OR001761 | Present Study | OR039809 | Present Study |
| Takecallis nigroantennatus sp. nov—isolate 5BAN | OR001762 | Present Study | OR039810 | Present Study |
| Takecallis nigroantennatus sp. nov—isolate 6BAA | OR001763 | Present Study | OR039811 | Present Study |
| Takecallis nigroantennatus sp. nov—isolate 7BAN | OR001764 | Present Study | OR039812 | Present Study |
| Takecallis taiwana | MH820965 | [30] | MT039369 | Direct submission |
| MH820989 | [30] | |||
| MH820958 | [30] | |||
| MH820978 | [30] | |||
| MH820985 | [30] | |||
| MH820961 | [30] | |||
| MH820975 | [30] | |||
| MH820966 | [30] | |||
| MH820962 | [30] | |||
| MH820968 | [30] | |||
| MH820980 | [30] | |||
| MH820981 | [30] | |||
| MH820996 | [30] | |||
| MH820990 | [30] | |||
| MH820973 | [30] | |||
| MH820993 | [30] | |||
| MH820974 | [30] | |||
| MH820977 | [30] | |||
| MH820970 | [30] | |||
| MH820987 | [30] | |||
| MH820964 | [30] | |||
| MH820988 | [30] | |||
| MH820979 | [30] | |||
| MH820963 | [30] | |||
| MH820984 | [30] | |||
| MH820991 | [30] | |||
| MH820967 | [30] | |||
| MH820982 | [30] | |||
| MH820976 | [30] | |||
| MH820997 | [30] | |||
| MH820992 | [30] | |||
| MH820969 | [30] | |||
| MH820995 | [30] | |||
| MH820959 | [30] | |||
| MH820951 | [30] | |||
| Takecallis sasae | KY307057 | [6] | MT262445 | Direct submission |
| KY307060 | [6] | |||
| KY307059 | [6] | |||
| KY307063 | [6] | |||
| KY307061 | [6] | |||
| KY307062 | [6] | |||
| KY307058 | [6] | |||
| KY307056 | [6] | |||
| Takecallis aroundicolens | EU701915 | [1] | MT262446 | Direct submission |
| KY307036 | [6] | |||
| KY307039 | [6] | |||
| KY307037 | [6] | |||
| KY307038 | [6] | |||
| KY307035 | [6] | |||
| KY307047 | [6] | |||
| KY307049 | [6] | |||
| KY307051 | [6] | |||
| KY307050 | [6] | |||
| KY307034 | [6] | |||
| KY307048 | [6] | |||
| KY307029 | [6] | |||
| KY307030 | [6] | |||
| KY307043 | [6] | |||
| KY307031 | [6] | |||
| KY307053 | [6] | |||
| KY307032 | [6] | |||
| KY307028 | [6] | |||
| KY307033 | [6] | |||
| KY307046 | [6] | |||
| KY307052 | [6] | |||
| KY307045 | [6] | |||
| KY307044 | [6] | |||
| KY307040 | [6] | |||
| KY307041 | [6] | |||
| KY307042 | [6] | |||
| KY307027 | [6] | |||
| KY307022 | [6] | |||
| KY307024 | [6] | |||
| KY307025 | [6] | |||
| KY307023 | [6] | |||
| KY307026 | [6] | |||
| KY307049 | [6] | |||
| Takecallis arundinariae | KR039454 | [31] | MT262444 | Direct submission |
| MH820948 | [30] | |||
| MH820950 | [30] | |||
| MH820955 | [30] | |||
| MH820956 | [30] | |||
| MH820947 | [30] | |||
| MH820954 | [30] | |||
| MH820957 | [30] | |||
| MH820952 | [30] | |||
| KR579007 | [32] | |||
| MH820951 | [30] | |||
| JX205217 | Direct submission | |||
| KR034713 | [31] | |||
| MH820953 | [30] | |||
| KR034488 | [31] | |||
| KY307054 | [6] | |||
| KF639650 | Direct submission | |||
| KY307055 | [6] | |||
| KR043148 | [31] | |||
| GU135641 | [33] | |||
| Euceraphis lineata | KR033551 | [31] | ||
| Calaphis magnoliae | KY306818 | [6] | MT262369 | [9] |
| Glyphinaphis bambusae | - | MN167448 | Direct submission |
| No. | Body | Antenna | Antennal Segments | Ultimate Rostral Segment | Second Segment of Hind Tarsus | |||
|---|---|---|---|---|---|---|---|---|
| III | IV | V | VIbase + VI Processus Terminalis | |||||
| 1 Holotype | 1.79 | 1.88 | 0.64 | 0.39 | 0.34 | 0.21 + 0.20 | 0.07 | 0.10 |
| 2 | 1.42 | 1.62 | 0.52 | 0.35 | 0.30 | 0.19 + 0.20 | 0.07 | 0.11 |
| 3 | 1.40 | 1.57 | 0.47 | 0.35 | 0.29 | 0.19 + 0.19 | 0.07 | 0.12 |
| 4 | 1.75 | 1.89 | 0.59 | 0.41 | 0.35 | 0.22 + 0.21 | 0.08 | 0.12 |
| 5 | 1.74 | 1.87 | 0.64 | 0.45 | 0.40 | 0.21 + 0.22 | 0.07 | 0.10 |
| 6 | 1.70 | 1.87 | 0.62 | 0.39 | 0.33 | 0.20 + 0.21 | 0.08 | 0.10 |
| 7 | 1.74 | 1.85 | 0.63 | 0.56 | 0.33 | 0.22 + 0.21 | 0.08 | 0.11 |
| 8 | 1.95 | 1.87 | 0.64 | 0.45 | 0.38 | 0.22 + 0.22 | 0.07 | 0.10 |
| 9 | 1.74 | 1.90 | 0.63 | 0.43 | 0.36 | 0.25 + 0.20 | 0.07 | 0.09 |
| 10 | 1.86 | 1.89 | 0.63 | 0.48 | 0.33 | 0.20 + 0.21 | 0.08 | 0.11 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wieczorek, K.; Sawka-Gądek, N. DNA Barcoding and Molecular Phylogenetics Revealed a New Cryptic Bamboo Aphid Species of the Genus Takecallis (Hemiptera: Aphididae). Appl. Sci. 2023, 13, 7798. https://doi.org/10.3390/app13137798
Wieczorek K, Sawka-Gądek N. DNA Barcoding and Molecular Phylogenetics Revealed a New Cryptic Bamboo Aphid Species of the Genus Takecallis (Hemiptera: Aphididae). Applied Sciences. 2023; 13(13):7798. https://doi.org/10.3390/app13137798
Chicago/Turabian StyleWieczorek, Karina, and Natalia Sawka-Gądek. 2023. "DNA Barcoding and Molecular Phylogenetics Revealed a New Cryptic Bamboo Aphid Species of the Genus Takecallis (Hemiptera: Aphididae)" Applied Sciences 13, no. 13: 7798. https://doi.org/10.3390/app13137798






