Classification of the Microstructural Elements of the Vegetal Tissue of the Pumpkin (Cucurbita pepo L.) Using Convolutional Neural Networks
Abstract
1. Introduction
2. Materials and Methods
2.1. Obtaining Micrographs
- Pumpkin fruits (Cucurbita pepo L.) were collected and stored at 15–20 C. Cylinders (25 mm length, 15 mm diameter) from the mesocarp’s middle zone, parallel to the fruit’s major axis, were taken.
- A rectangular slab of 0.5–1.0 mm of thickness was gently cut parallel to cylinder’s height at the maximum section area. The slab was then divided into four symmetrical cuts, and each quarter was newly divided into six parts.
- These parts were fixed in 2.5% glutaraldehyde in 1.25% PIPES buffer at pH 7.0–7.2 during 24 h at room temperature. The parts were then dehydrated in a water/ethanol series and embedded in LR White resin (London Resin Co., Basingstoke, UK). After the samples were embedded in resin, semi-thin sections (0.6 μm) of the resin blocks were obtained with a microtome (model Reichert-Supernova, Leica, Wien, Austria).
- The sections were stained with an aqueous solution Azure II 0.5%, Methylene Blue 0.5%, Borax 0.5% during 30 s. They were then washed in distilled water and mounted on a glass slide.
- Micrographs of the stained samples were obtained under a stereomicroscope (Olympus SZ-11, Tokyo, Japan) that was attached to a digital color video camera (SONY SSC-DC50AP, Tokyo, Japan) and a computer.
2.2. Digital Treatment of the Micrographs
2.2.1. Improvement and Enhancement
2.2.2. Image Processing
2.3. Data Extraction
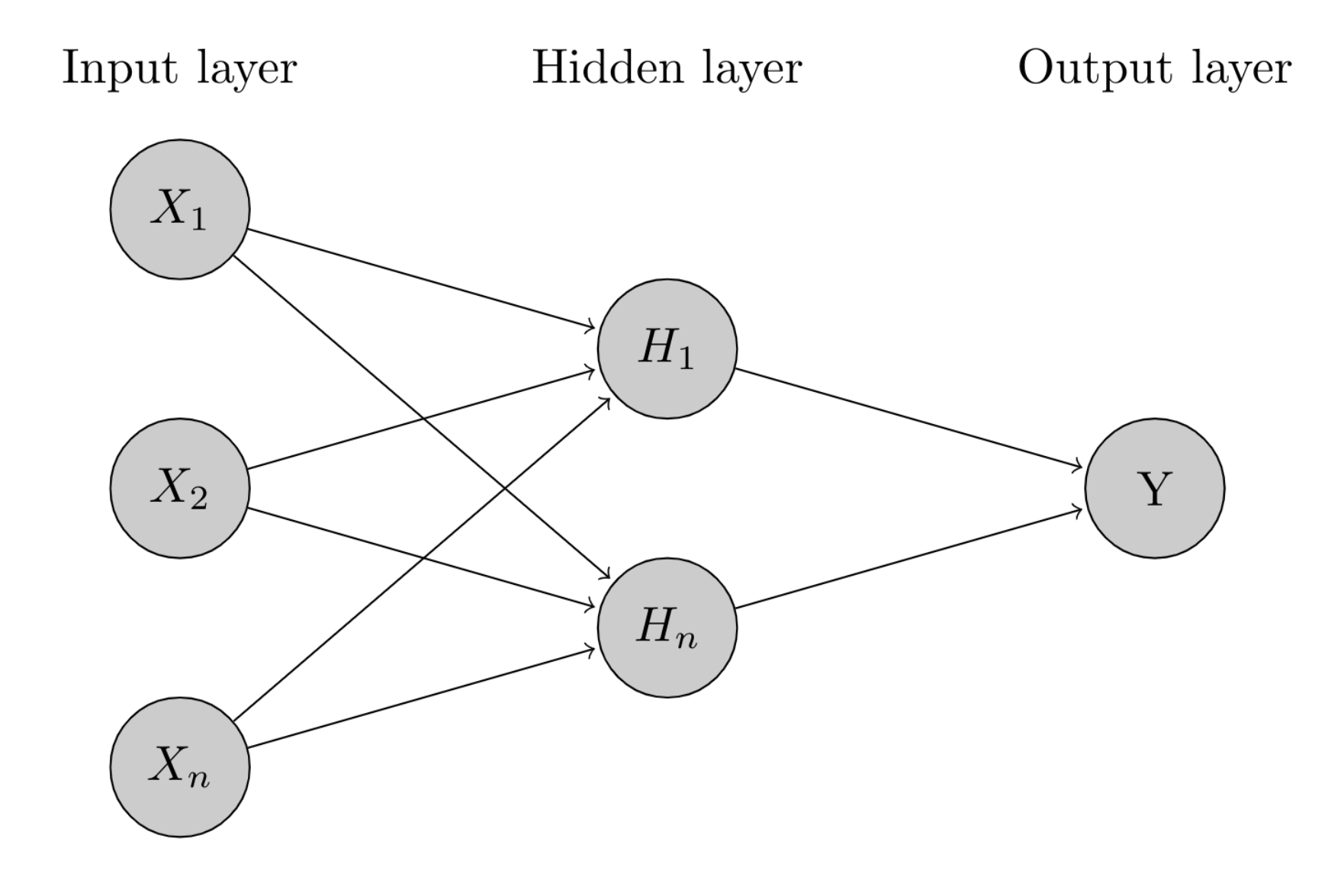
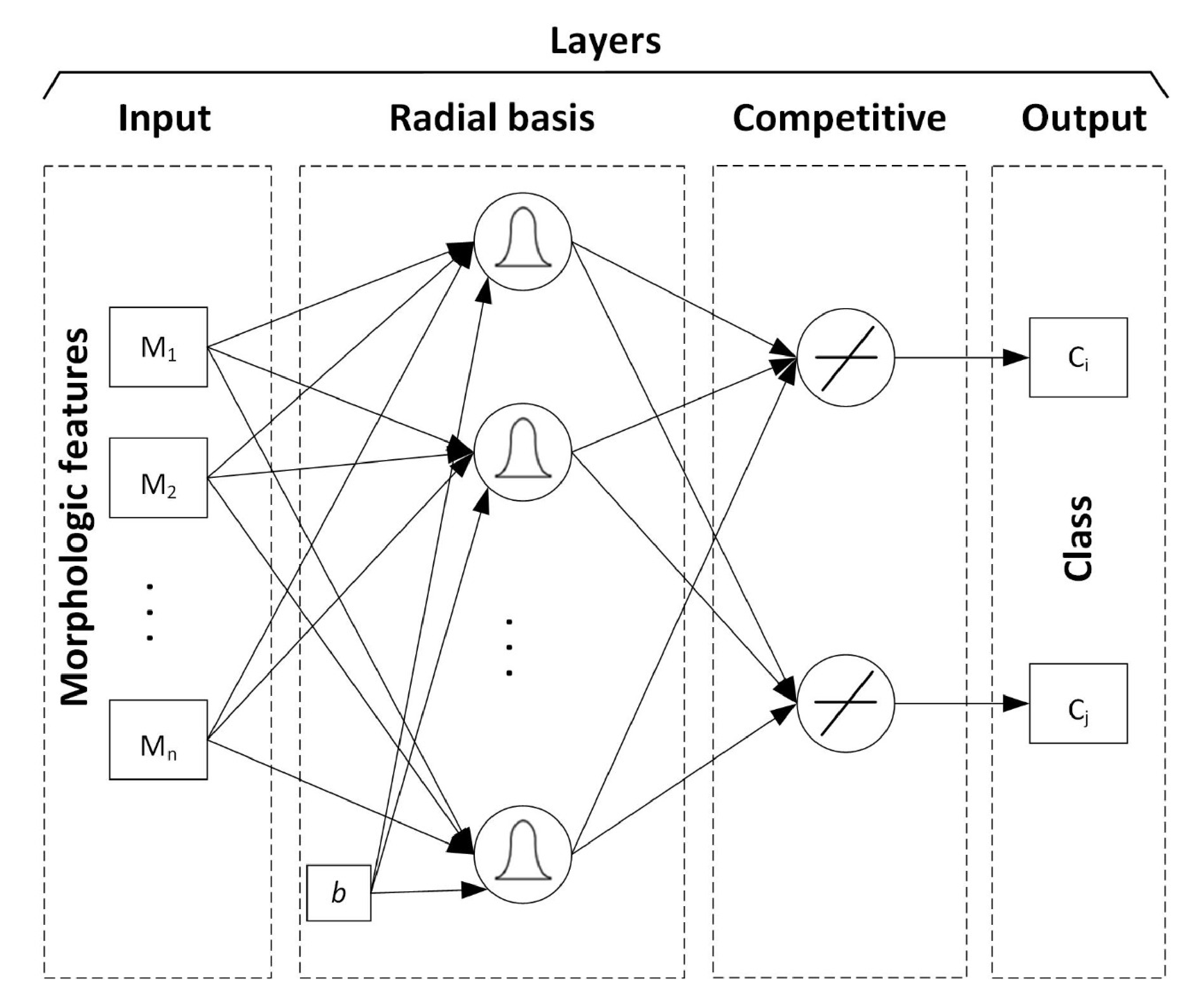
2.4. Radial Basis Neural Network—RBNN
2.5. Convolutional Neural Network—CNN
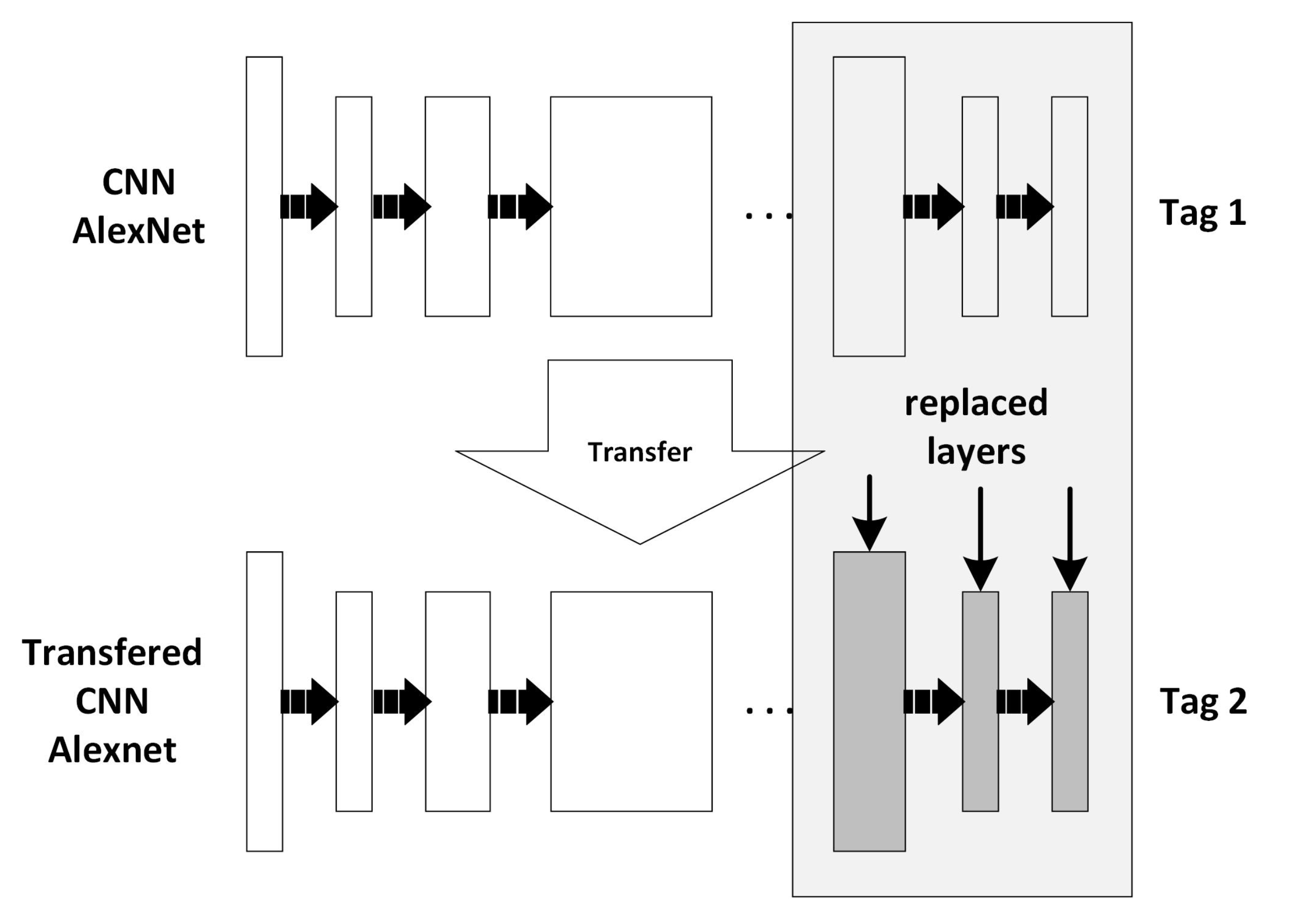
2.6. Learning Transfer
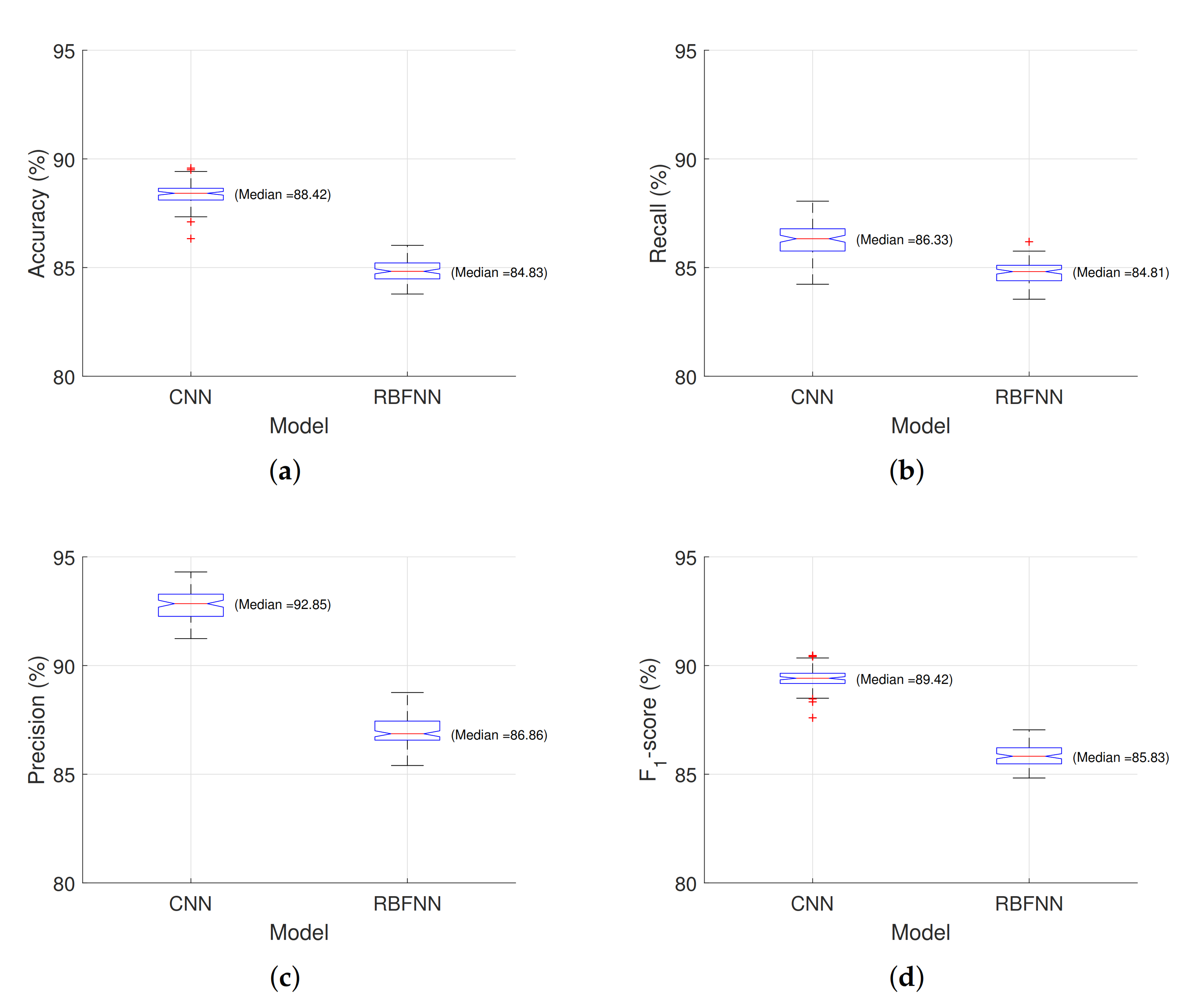
2.7. Statistical Comparison of Models
- Accuracy: This measures how many observations, both positive and negative, were correctly classified and it is defined by Equation (7).
- Recall: This measures how many observations out of all of the positive observations were classified as positive (see Equation (8)).
- Precision: This measures how many observations predicted as positive are, in fact, positive (see Equation (9)).
- F1–score: This combines precision and recall into one metric (harmonic mean, see Equation (10)).
3. Results and Discussions
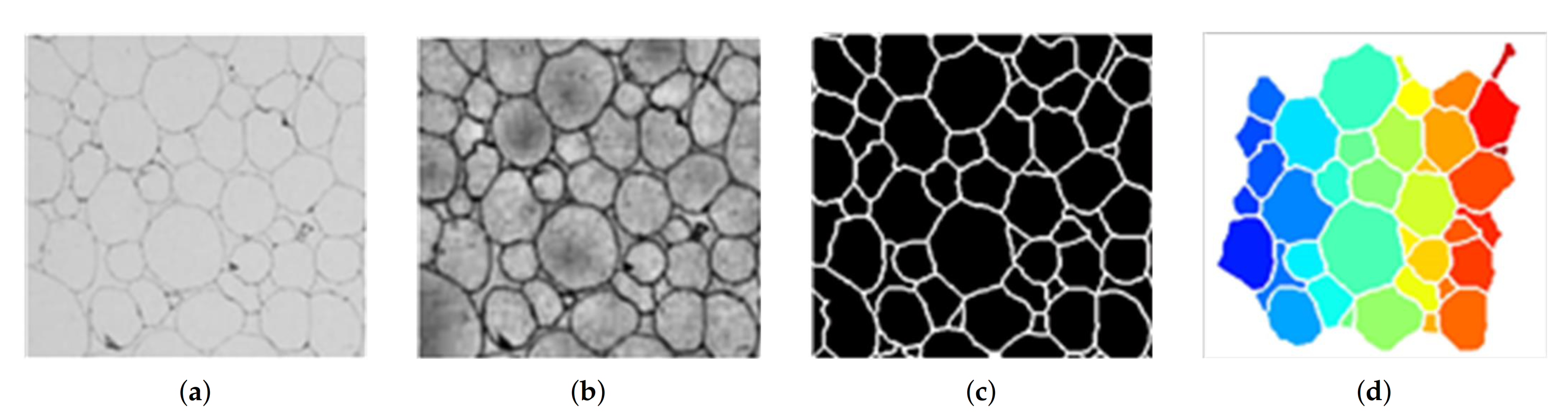
3.1. Digital Micrograph Processing
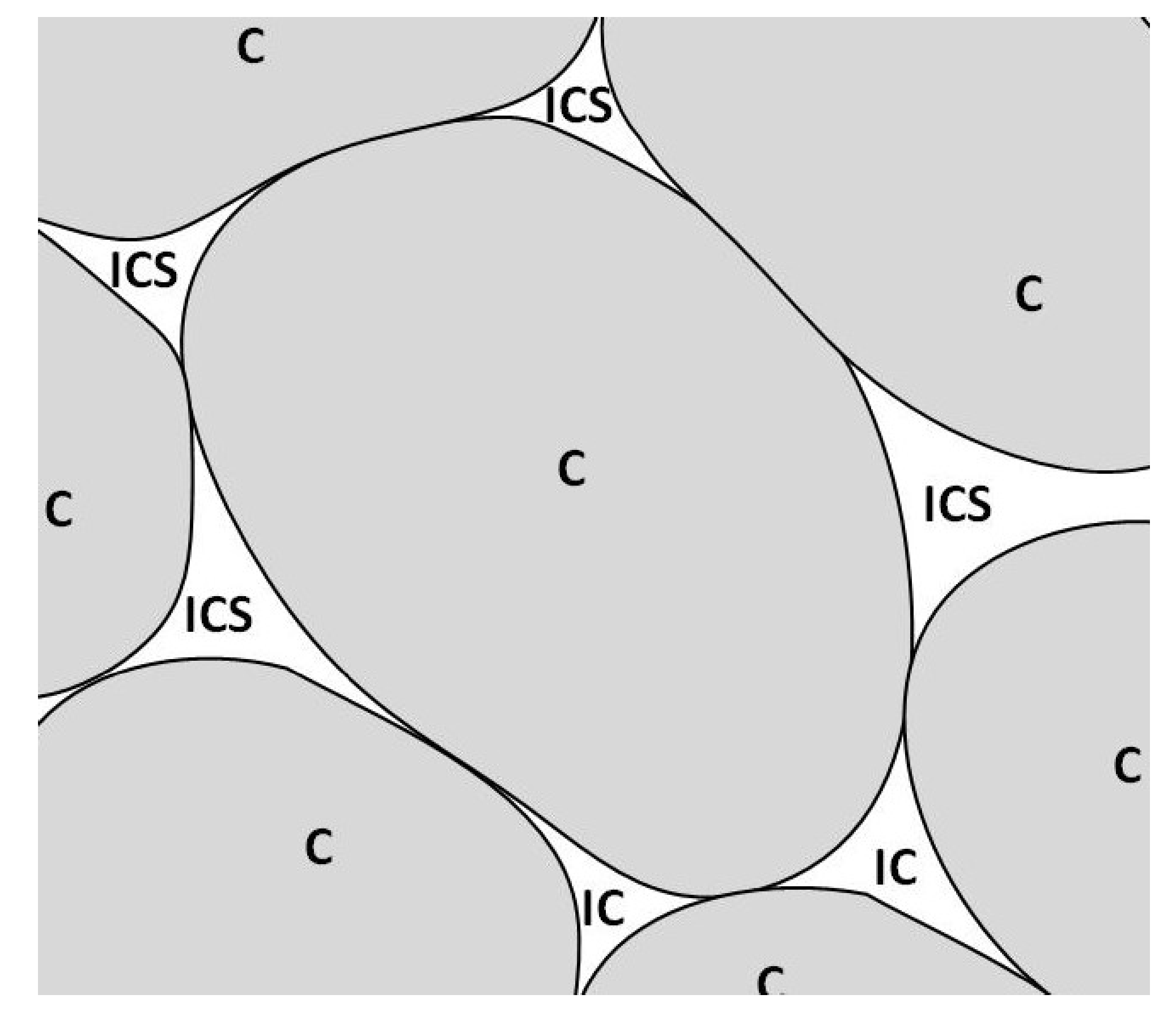
3.2. Microstructural Elements
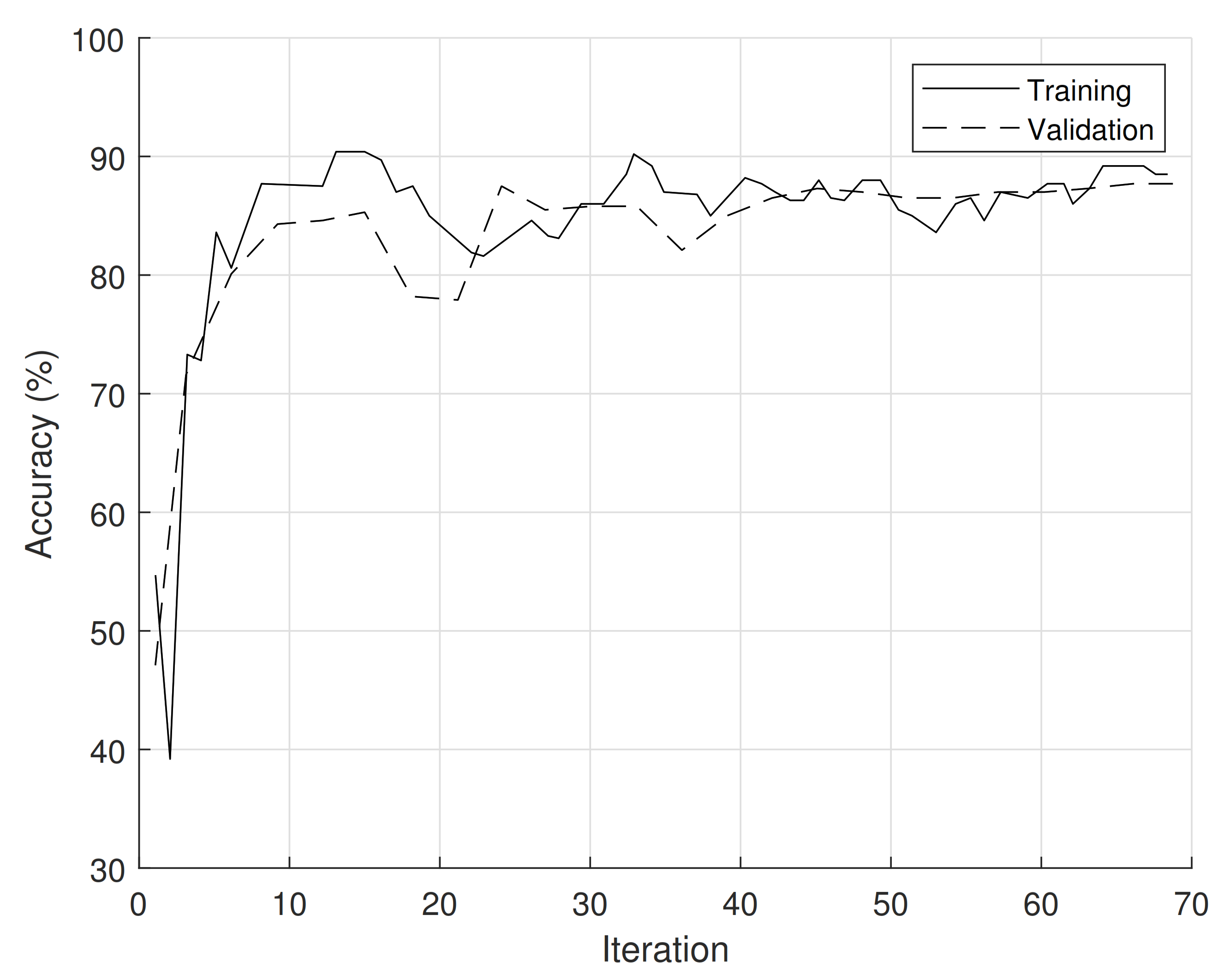
3.3. CNN Implementation
3.4. Statistical Analysis
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Conflicts of Interest
References
- Betoret, E.; Betoret, N.; Rocculi, P.; Dalla Rosa, M. Strategies to improve food functionality: Structure–property relationships on high pressures homogenization, vacuum impregnation and drying technologies. Trends Food Sci. Technol. 2015, 46, 1–12. [Google Scholar] [CrossRef]
- Fito, P.; LeMaguer, M.; Betoret, N.; Fito, P.J. Advanced food process engineering to model real foods and processes: The SAFES methodology. J. Food Eng. 2007, 83, 173–185. [Google Scholar] [CrossRef]
- Topete-Betancourt, A.; Figueroa-Cárdenas, J.d.D.; Morales-Sánchez, E.; Arámbula-Villa, G.; Pérez-Robles, J.F. Evaluation of the mechanism of oil uptake and water loss during deep-fat frying of tortilla chips. Rev. Mex. Ing. QuíMica 2020, 19, 409–422. [Google Scholar] [CrossRef]
- Silva-Jara, J.M.; López-Cruz, R.; Ragazzo-Sánchez, J.A.; Calderón-Santoyo, M. Antagonistic microorganisms efficiency to suppress damage caused by Colletotrichum gloeosporioides in papaya crop: Perspectives and challenges. Rev. Mex. Ing. QuíMica 2020, 19, 839–849. [Google Scholar] [CrossRef]
- Aguilera, J.M. Why food microstructure? J. Food Eng. 2005, 67, 3–11. [Google Scholar] [CrossRef]
- Pieczywek, P.; Zdunek, A. Automatic classification of cells and intercellular spaces of apple tissue. Comput. Electron. Agric. 2012, 81, 72–78. [Google Scholar] [CrossRef]
- Mayor, L.; Pissarra, J.; Sereno, A.M. Microstructural changes during osmotic dehydration of parenchymatic pumpkin tissue. J. Food Eng. 2008, 85, 326–339. [Google Scholar] [CrossRef]
- Mebatsion, H.; Verboven, P.; Verlinden, B.; Ho, Q.; Nguyen, T.; Nicolaï, B. Microscale modelling of fruit tissue using Voronoi tessellations. Comput. Electron. Agric. 2006, 52, 36–48. [Google Scholar] [CrossRef]
- Oblitas, J.; Castro, W.; Mayor, L. Effect of different combinations of size and shape parameters in the percentage error of classification of structural elements in vegetal tissue of the pumpkin Cucurbita pepo L. using probabilistic neural networks. Rev. Fac. Ing. Univ. Antioq. 2016, 30–37. [Google Scholar] [CrossRef]
- Meng, N.; Lam, E.; Tsia, K.; So, H. Large-scale multi-class image-based cell classification with deep learning. IEEE J. Biomed. Health Inform. 2018, 23. [Google Scholar] [CrossRef]
- Adeshina, S.; Adedigba, A.; Adeniyi, A.; Aibinu, A. Breast Cancer Histopathology Image Classification with Deep Convolutional Neural Networks. In Proceedings of the IEEE 14th International Conference on Electronics Computer and Computation, Kaskelen, Kazakhstan, 29 Novermber–1 December 2018; pp. 206–212. [Google Scholar] [CrossRef]
- Aliyu, H.; Sudirman, R.; Razak, M.; Wahab, M. Red Blood Cell Classification: Deep Learning Architecture Versus Support Vector Machine. In Proceedings of the IEEE 2nd International Conference on BioSignal Analysis, Processing and Systems, Kuching, Malaysia, 24–26 July 2018; pp. 142–147. [Google Scholar] [CrossRef]
- Reddy, A.; Juliet, D. Transfer Learning with ResNet-50 for Malaria Cell-Image Classification. In Proceedings of the IEEE International Conference on Communication and Signal Processing, Weihai, China, 28–30 September 2019; pp. 945–949. [Google Scholar] [CrossRef]
- Mayor, L.; Moreira, R.; Sereno, A. Shrinkage, density, porosity and shape changes during dehydration of pumpkin (Cucurbita pepo L.) fruits. J. Food Eng. 2011, 103, 29–37. [Google Scholar] [CrossRef]
- Castro, W.; Oblitas, J.; De-La-Torre, M.; Cotrina, C.; Bazán, K.; Avila-George, H. Classification of cape gooseberry fruit according to its level of ripeness using machine learning techniques and different color spaces. IEEE Access 2019, 7, 27389–27400. [Google Scholar] [CrossRef]
- Valdez-Morones, T.; Pérez-Espinosa, H.; Avila-George, H.; Oblitas, J.; Castro, W. An Android App for detecting damage on tobacco (Nicotiana tabacum L) leaves caused by blue mold (Penospora tabacina Adam). In Proceedings of the 2018 7th International Conference On Software Process Improvement (CIMPS), Jalisco, Mexico, 17–19 October 2018; pp. 125–129. [Google Scholar] [CrossRef]
- Castro, W.; Oblitas, J.; Chuquizuta, T.; Avila-George, H. Application of image analysis to optimization of the bread-making process based on the acceptability of the crust color. J. Cereal Sci. 2017, 74, 194–199. [Google Scholar] [CrossRef]
- De-la Torre, M.; Zatarain, O.; Avila-George, H.; Muñoz, M.; Oblitas, J.; Lozada, R.; Mejía, J.; Castro, W. Multivariate Analysis and Machine Learning for Ripeness Classification of Cape Gooseberry Fruits. Processes 2019, 7, 928. [Google Scholar] [CrossRef]
- Saha, P.; Borgefors, G.; di Baja, G. Skeletonization: Theory, Methods and Applications; Academic Press: Cambridge, MA, USA, 2017. [Google Scholar]
- González, R.; Woods, R.; Eddins, S. Digital Image Processing Using MATLAB; Pearson Education: Chennai, India, 2004. [Google Scholar]
- Fernández-Navarro, F.; Hervás-Martínez, C.; Gutiérrez, P.; Carbonero-Ruz, M. Evolutionary q-Gaussian radial basis function neural networks for multiclassification. Neural Netw. 2011, 24, 779–784. [Google Scholar] [CrossRef]
- Huang, W.; Oh, S.; Pedrycz, W. Design of hybrid radial basis function neural networks (HRBFNNs) realized with the aid of hybridization of fuzzy clustering method (FCM) and polynomial neural networks (PNNs). Neural Netw. 2014, 60, 166–181. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.; Nejati, H.; Do, T.; Cheung, N.; Cheah, L. Image-based vehicle analysis using deep neural network: A systematic study. In Proceedings of the IEEE International Conference on Digital Signal Processing, Beijing, China, 16–18 October 2016; pp. 276–280. [Google Scholar] [CrossRef]
- Toliupa, S.; Tereikovskyi, I.; Tereikovskyi, O.; Tereikovska, L.; Nakonechnyi, V.; Kulakov, Y. Keyboard Dynamic Analysis by Alexnet Type Neural Network. In Proceedings of the 2020 IEEE 15th International Conference on Advanced Trends in Radioelectronics, Telecommunications and Computer Engineering (TCSET), Lviv-Slavske, Ukraine, 25–29 February 2020; pp. 416–420. [Google Scholar] [CrossRef]
- Lu, S.; Lu, Z.; Zhang, Y. Pathological brain detection based on AlexNet and transfer learning. J. Comput. Sci. 2019, 30, 41–47. [Google Scholar] [CrossRef]
- Kraus, O.; Ba, J.; Frey, B. Classifying and segmenting microscopy images with deep multiple instance learning. Bioinformatics 2016, 32, i52–i59. [Google Scholar] [CrossRef] [PubMed]
- Du, C.; Sun, D. Learning techniques used in computer vision for food quality evaluation: A review. J. Food Eng. 2006, 72, 39–55. [Google Scholar] [CrossRef]
- Brosnan, T.; Sun, D. Improving quality inspection of food products by computer vision—A review. J. Food Eng. 2004, 61, 3–16. [Google Scholar] [CrossRef]
- Baker, N.; Lu, H.; Erlikhman, G.; Kellman, P. Deep convolutional networks do not classify based on global object shape. PLoS Comput. Biol. 2018, 14, e1006613. [Google Scholar] [CrossRef] [PubMed]
- Rohmatillah, M.; Pramono, S.; Suyono, H.; Sena, S. Automatic Cervical Cell Classification Using Features Extracted by Convolutional Neural Network. In Proceedings of the IEEE 2018 Electrical Power, Electronics, Communications, Controls and Informatics Seminar (EECCIS), Batu, Indonesia, 9–11 October 2018; pp. 382–386. [Google Scholar] [CrossRef]
- Sadanandan, S.; Ranefall, P.; Wählby, C. Feature augmented deep neural networks for segmentation of cells. In Proceedings of the European Conference on Computer Vision, Amsterdam, The Netherlands, 8–16 October 2016; pp. 231–243. [Google Scholar] [CrossRef]
- Sharma, M.; Bhave, A.; Janghel, R. White Blood Cell Classification Using Convolutional Neural Network. In Soft Computing and Signal Processing; Springer: Berlin, Germany, 2019; pp. 135–143. [Google Scholar] [CrossRef]
- Song, W.; Li, S.; Liu, J.; Qin, H.; Zhang, B.; Zhang, S.; Hao, A. Multitask Cascade Convolution Neural Networks for Automatic Thyroid Nodule Detection and Recognition. IEEE J. Biomed. Health Inform. 2018, 23, 1215–1224. [Google Scholar] [CrossRef] [PubMed]
- Akram, S.; Kannala, J.; Eklund, L.; Heikkilä, J. Cell proposal network for microscopy image analysis. In Proceedings of the 2016 IEEE International Conference on Image Processing (ICIP), Phoenix, AZ, USA, 25–28 September 2016; pp. 3199–3203. [Google Scholar] [CrossRef]






| Comparison | Performance Measure | |||||
|---|---|---|---|---|---|---|
| Accuracy | Recall | Precision | ||||
| Means | t-test | −47.4999 | −25.9556 | −36.9701 | −14.8146 | |
| p-value | 0 | 0 | 0 | 0 | ||
| Standard deviation | F-test | 0.5036 | 0.6453 | 2.0789 | 1.2957 | |
| p-value | 0.0007 | 0.0304 | 0.0003 | 0.1992 | ||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Oblitas, J.; Mejia, J.; De-la-Torre, M.; Avila-George, H.; Seguí Gil, L.; Mayor López, L.; Ibarz, A.; Castro, W. Classification of the Microstructural Elements of the Vegetal Tissue of the Pumpkin (Cucurbita pepo L.) Using Convolutional Neural Networks. Appl. Sci. 2021, 11, 1581. https://doi.org/10.3390/app11041581
Oblitas J, Mejia J, De-la-Torre M, Avila-George H, Seguí Gil L, Mayor López L, Ibarz A, Castro W. Classification of the Microstructural Elements of the Vegetal Tissue of the Pumpkin (Cucurbita pepo L.) Using Convolutional Neural Networks. Applied Sciences. 2021; 11(4):1581. https://doi.org/10.3390/app11041581
Chicago/Turabian StyleOblitas, Jimy, Jezreel Mejia, Miguel De-la-Torre, Himer Avila-George, Lucía Seguí Gil, Luis Mayor López, Albert Ibarz, and Wilson Castro. 2021. "Classification of the Microstructural Elements of the Vegetal Tissue of the Pumpkin (Cucurbita pepo L.) Using Convolutional Neural Networks" Applied Sciences 11, no. 4: 1581. https://doi.org/10.3390/app11041581
APA StyleOblitas, J., Mejia, J., De-la-Torre, M., Avila-George, H., Seguí Gil, L., Mayor López, L., Ibarz, A., & Castro, W. (2021). Classification of the Microstructural Elements of the Vegetal Tissue of the Pumpkin (Cucurbita pepo L.) Using Convolutional Neural Networks. Applied Sciences, 11(4), 1581. https://doi.org/10.3390/app11041581











