A Novel Embedded Feature Selection and Dimensionality Reduction Method for an SVM Type Classifier to Predict Periventricular Leukomalacia (PVL) in Neonates
Abstract
1. Introduction
2. Methods
2.1. Raw Data
2.2. Machine Learning Data
2.3. Feature Selection
2.3.1. Filter Method
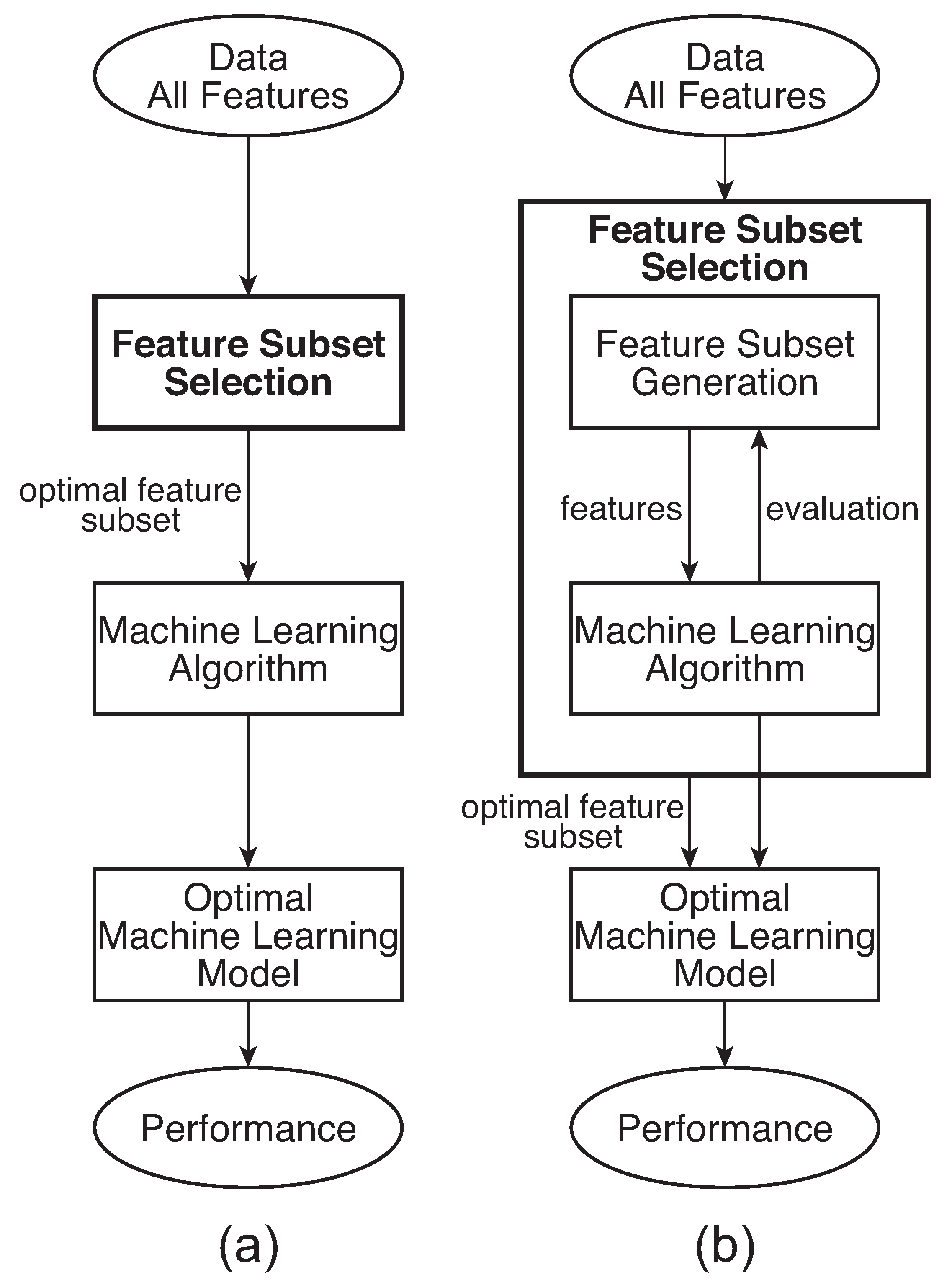
2.3.2. Wrapper Method
| Algorithm 1 Embedded Feature Subset Optimization Algorithm | |
| functionGenetic-Algorithm(population, Fitness-Function) | |
| inputs: population, set of c random feature subsets | ▹ c: number of chromosomes |
| repeat | |
| newpopulation empty set | |
| for to Size(population) do | |
| Random-Selection(population, Fitness-Function) | ▹ x is selected random w.r.t. fitness-score as its probability |
| Random-Selection(population, Fitness-Function) | ▹ y is selected random w.r.t. fitness-score as its probability |
| child ← Reproduce() | ▹ are chromosomes and subsets of the feature set |
| if small random probability then child ← Mutate(child) | ▹ this probability is defined by a selected mutation-rate |
| add child to newpopulation | |
| until solution is found that satisfies minimum criteria, or enough have elapsed | |
| return the best set in population, according to Fitness-Function | ▹ best feature subset, according to the Fitness-Function |
| functionReproduce() | |
| inputs: , two chromosomes from the population | ▹ evaluated by the Fitness-Function |
| Length(x); | |
| number from 1 to n | ▹ l is defined by a selected crossover-rate |
| Append(Substring(), Substring()) | ▹ new chromosome |
| returnchild | |
| functionFitness-Function(population) | ▹ user defined |
| inputs: population, a set of c random feature subsets | ▹ c: number of chromosomes |
| for to Size(population) do | |
| Size(population(j)) | ▹ is the dimensionality of chromosome |
| SVM-model ←Train-SVM(Training-Samples(population(j)) | |
| accuracy of the SVM-model on the Training-Samples | |
| Decision-Boundary(SVM-model) | |
| Magnitude() | ▹ the 2-norm of decision boundary margin |
| ▹ a is a user defined weight parameter of | |
| ▹ b is a user defined weight parameter of | |
| ▹ fitness-score of the chromosome | |
| fitness-score | ▹ fitness-scores for all chromosomes in the population |
| returnfitness-score | |
3. Results
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Holzinger, A. Biomedical Informatics: Descovering Knowledge in Big Data, 1st ed.; Springer International Publishing Switzerland: Graz, Austria, 2014. [Google Scholar]
- Holzinger, A. Machine Learning for Health Informatics; Springer: Berlin/Heidelberg, Germany, 2016; pp. 1–24. [Google Scholar]
- Bender, D.; Nadkarni, V.M.; Nataraj, C. A machine learning algorithm to improve patient-centric pediatric cardiopulmonary resuscitation. Inform. Med. Unlocked 2020, 19, 100339. [Google Scholar] [CrossRef]
- Tan, K.C.; Yu, Q.; Heng, C.M.; Lee, T.H. Evolutionary computing for knowledge discovery in medical diagnosis. Artif. Intell. Med. 2003, 27, 129–154. [Google Scholar] [CrossRef]
- Holzinger, A. Interactive machine learning for health informatics: When do we need the human-in-the-loop? Brain Inform. 2016, 3, 119–131. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.K.; Blaine Easley, R.; Brady, K.M. Neurocognitive monitoring and care during pediatric cardiopulmonary bypass-current and future directions. Curr. Cardiol. Rev. 2008, 4, 123–139. [Google Scholar] [CrossRef] [PubMed]
- Vapnik, V. Statistical Learning Theory; Springer: London, UK, 1998. [Google Scholar]
- Licht, D.J.; Shera, D.M.; Clancy, R.R.; Wernovsky, G.; Montenegro, L.M.; Nicolson, S.C.; Zimmerman, R.A.; Spray, T.L.; Gaynor, J.W.; Vossough, A. Brain maturation is delayed in infants with complex congenital heart defects. J. Thorac. Cardiovasc. Surg. 2009, 137, 529–537. [Google Scholar] [CrossRef] [PubMed]
- Samanta, B.; Bird, G.L.; Kuijpers, M.; Zimmerman, R.A.; Jarvik, G.P.; Wernovsky, G.; Clancy, R.R.; Licht, D.J.; Gaynor, J.W.; Nataraj, C. Prediction of periventricular leukomalacia. Part II: Selection of hemodynamic features using computational intelligence. Artif. Intell. Med. 2009, 46, 217–231. [Google Scholar] [CrossRef] [PubMed]
- Licht, D.J.; Wang, J.; Silvestre, D.W.; Nicolson, S.C.; Montenegro, L.M.; Wernovsky, G.; Tabbutt, S.; Durning, S.M.; Shera, D.M.; Gaynor, J.W.; et al. Preoperative cerebral blood flow is diminished in neonates with severe congenital heart defects. J. Thorac. Cardiovasc. Surg. 2004, 128, 841–849. [Google Scholar] [CrossRef] [PubMed]
- Samanta, B.; Bird, G.L.; Kuijpers, M.; Zimmerman, R.A.; Jarvik, G.P.; Wernovsky, G.; Clancy, R.R.; Licht, D.J.; Gaynor, J.W.; Nataraj, C. Prediction of periventricular leukomalacia. Part I: Selection of hemodynamic features using logistic regression and decision tree algorithms. Artif. Intell. Med. 2009, 46, 201–215. [Google Scholar] [CrossRef] [PubMed][Green Version]
- McCarthy, A.L.; Winters, M.E.; Busch, D.R.; Gonzalez-Giraldo, E.; Ko, T.S.; Lynch, J.M.; Schwab, P.J.; Xiao, R.; Buckley, E.M.; Vossough, A.; et al. Scoring system for periventricular leukomalacia in infants with congenital heart disease. Pediatr. Res. 2015, 78, 304–309. [Google Scholar] [CrossRef] [PubMed]
- Jalali, A.; Berg, R.; Nadkarni, V.; Nataraj, C. Improving Cardiopulmonary Resuscitation (CPR) by Dynamic Variation of CPR Parameters. In Proceedings of the Dynamic Systems and Control Conference, ASME, Palo Alto, CA, USA, 21–23 October 2013. [Google Scholar]
- Jalali, A.; Simpao, A.F.; Galvez, J.A.; Licht, D.J.; Nataraj, C. Prediction of Periventricular Leukomalacia in Neonates after Cardiac Surgery Using Machine Learning Algorithms. J. Med. Syst. 2018, 42, 177. [Google Scholar] [CrossRef] [PubMed]
- Bender, D.; Jalali, A.; Licht, D.J.; Nataraj, C. Prediction of periventricular leukomalacia occurrence in neonates using a novel support vector machine classifier optimization method. In Proceedings of the ASME 2015 Dynamic Systems and Control Conference, Columbus, OH, USA, 28–30 October 2015; Volume 1, p. 74. [Google Scholar]
- Bender, D.; Jalali, A.; Licht, D.J.; Nataraj, C. Prediction of periventricular leukomalacia occurrence in neonates using a novel unsupervised learning method. In Proceedings of the Dynamics Systems and Control Conference, Washington, DC, USA, 17–19 June 2014. [Google Scholar]
- Jalali, A.; Buckley, E.M.; Lynch, J.M.; Schwab, P.J.; Licht, D.J.; Nataraj, C. Prediction of periventricular leukomalacia occurrence in neonates after heart surgery. IEEE J. Biomed. Health Inform. 2014, 18, 1453–1460. [Google Scholar] [CrossRef] [PubMed]
- Jalali, A.; Licht, D.J.; Nataraj, C. Application of decision tree in the prediction of periventricular leukomalacia (PVL) occurrence in neonates after heart surgery. In Proceedings of the 2012 Annual International Conference of the IEEE Engineering in Medicine and Biology Society, San Diego, CA, USA, 28 August–1 September 2012; pp. 5931–5934. [Google Scholar] [CrossRef]
- Settles, B. Active Learning. In Synthesis Lectures on Artificial Intelligence and Machine Learning; Morgan and Claypool Publishers: Williston, VT, USA, 2012; Volume 6. [Google Scholar]
- Supratak, A.; Wu, C.; Dong, H.; Sun, K.; Guo, Y. Survey on Feature Extraction and Applications of Biosignals. In Machine Learning for Health Informatics: State-of-the-Art and Future Challenges; Holzinger, A., Ed.; Springer: Berlin/Heidelberg, Germany, 2016; pp. 161–182. [Google Scholar]
- Wernovsky, G.; Licht, D.J. Neurodevelopmental Outcomes in Children with Congenital Heart Disease—What Can We Impact? Pediatr. Crit. Care Med. 2016, 17, S232–S242. [Google Scholar] [CrossRef] [PubMed]
- Flach, P. Machine Learning—The Art and Science of Algorithms that Make Sense of Data, 1st ed.; Cambridge University Press: Cambridge, UK, 2012. [Google Scholar]
- Russell, S.J.; Norvig, P. Artificial Intelligence a Modern Approach, 3rd ed.; Pearson Higher Education: London, UK, 2010. [Google Scholar]
- Saghapour, E.; Kermani, S.; Sehhati, M. A novel feature ranking method for prediction of cancer stages using proteomics data. PLoS ONE 2017, 12, e0184203. [Google Scholar] [CrossRef] [PubMed]
- Kohavi, R.; John, G.H. Wrappers for feature subset selection. Artif. Intell. 1997, 97, 273–324. [Google Scholar] [CrossRef]
- Wang, G.; Lochovsky, F.H.; Yang, Q. Feature Selection with Conditional Mutual Information MaxiMin in Text Categorization. In Proceedings of the Thirteenth ACM International Conference on Information and Knowledge Management 2004, Queensland, Australia, 1–5 November 2004; pp. 342–349. [Google Scholar]
- Novovicova, J.; Somol, P.; Haindl, M.; Pudil, P. Conditional Mutual Information Based Feature Selection for Classification Task. In Proceedings of the Iberoamerican Congress on Pattern Recognition 2007, Valparaíso, Chile, 7–10 November 2007; pp. 417–426. [Google Scholar]
- Vergara, J.R.; Estevez, P.A. A review of feature selection methods based on mutual information. Neural Comput. Appl. 2014, 24, 175–186. [Google Scholar] [CrossRef]
- Rostami, M.; Berahmand, K.; Forouzandeh, S. A novel community detection based genetic algorithm for feature selection. J. Big Data 2021, 8, 2. [Google Scholar] [CrossRef]
- Burges, C.J.C. A Tutorial on Support Vector Machines for Pattern Recognition. Data Min. Knowl. Discov. 1998, 2, 121–167. [Google Scholar] [CrossRef]
- Xie, Y.; Zhang, T. A fault diagnosis approach using SVM with data dimension reduction by PCA and LDA method. In Proceedings of the 2015 Chinese Automation Congress (CAC), Wuhan, China, 27–29 November 2015; pp. 869–874. [Google Scholar] [CrossRef]
- Nguyen, T.D.; Lee, H. A New SVM Method for an Indirect Matrix Converter With Common-Mode Voltage Reduction. IEEE Trans. Ind. Inform. 2014, 10, 61–72. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bender, D.; Licht, D.J.; Nataraj, C. A Novel Embedded Feature Selection and Dimensionality Reduction Method for an SVM Type Classifier to Predict Periventricular Leukomalacia (PVL) in Neonates. Appl. Sci. 2021, 11, 11156. https://doi.org/10.3390/app112311156
Bender D, Licht DJ, Nataraj C. A Novel Embedded Feature Selection and Dimensionality Reduction Method for an SVM Type Classifier to Predict Periventricular Leukomalacia (PVL) in Neonates. Applied Sciences. 2021; 11(23):11156. https://doi.org/10.3390/app112311156
Chicago/Turabian StyleBender, Dieter, Daniel J. Licht, and C. Nataraj. 2021. "A Novel Embedded Feature Selection and Dimensionality Reduction Method for an SVM Type Classifier to Predict Periventricular Leukomalacia (PVL) in Neonates" Applied Sciences 11, no. 23: 11156. https://doi.org/10.3390/app112311156
APA StyleBender, D., Licht, D. J., & Nataraj, C. (2021). A Novel Embedded Feature Selection and Dimensionality Reduction Method for an SVM Type Classifier to Predict Periventricular Leukomalacia (PVL) in Neonates. Applied Sciences, 11(23), 11156. https://doi.org/10.3390/app112311156





