Antisense-Mediated Down-Regulation of Factor V-Short Splicing in a Liver Cell Line Model
Abstract
1. Introduction
2. Materials and Methods
2.1. Bioinformatics Analysis
2.2. Antisense Oligonucleotides
2.3. Characterisation of the F5 Gene in HepG2 Cells
2.3.1. Cell Model
2.3.2. DNA Isolation
2.3.3. Multiplex Ligation-Dependent Probe Amplification (MLPA)
2.3.4. Amplification and Sequencing of F5 exon 13
2.4. Cell Culture and Treatment
2.5. F5 Transcript Analysis
2.5.1. RNA Isolation and Reverse Transcription
2.5.2. F5 mRNA Analysis by PCR and Gel Electrophoresis
2.5.3. Quantification of Alternatively Spliced F5 Transcripts by qPCR
2.5.4. Statistical Analysis
3. Results
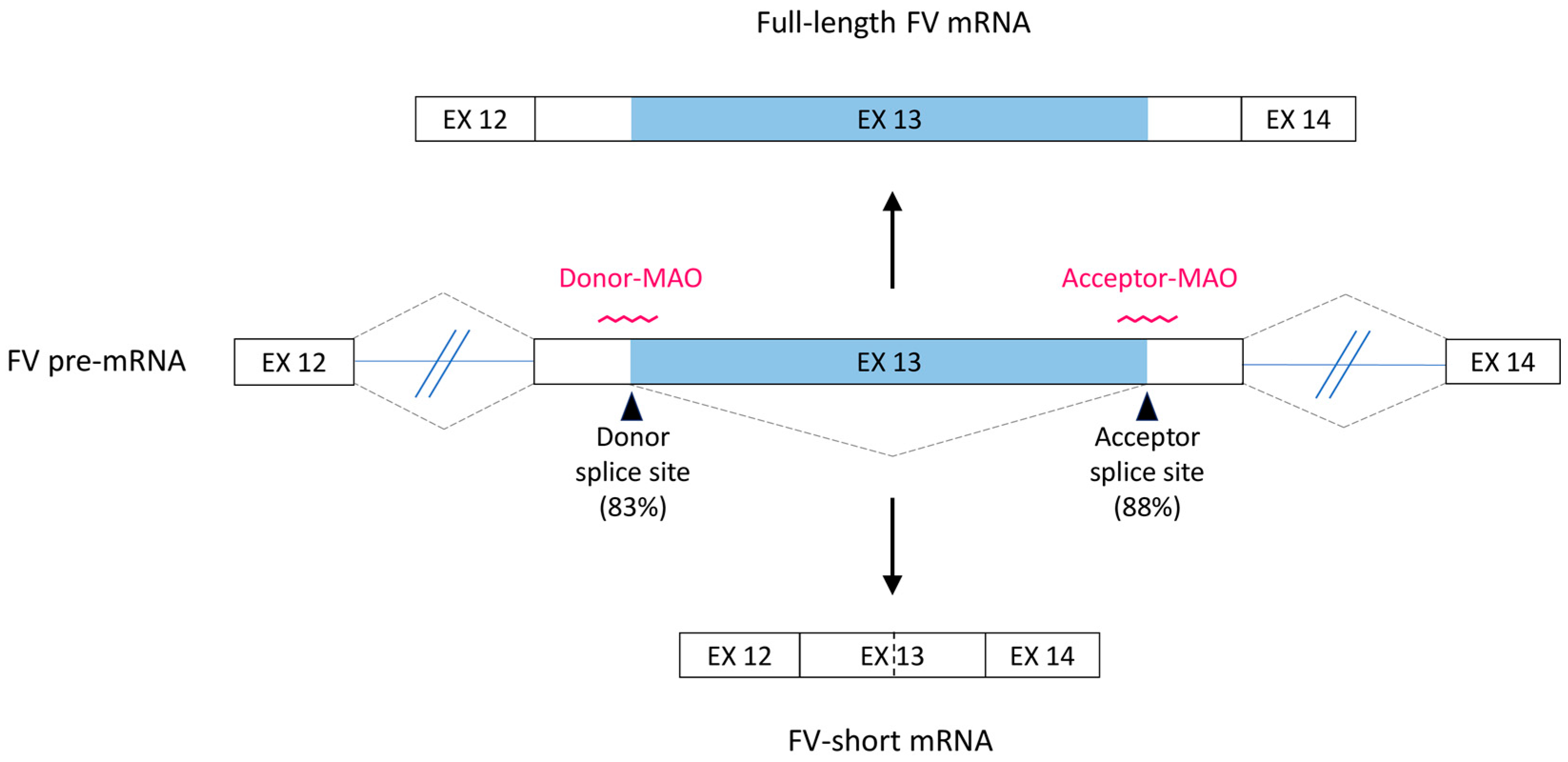
3.1. Bioinformatics Analysis and Antisense Strategy
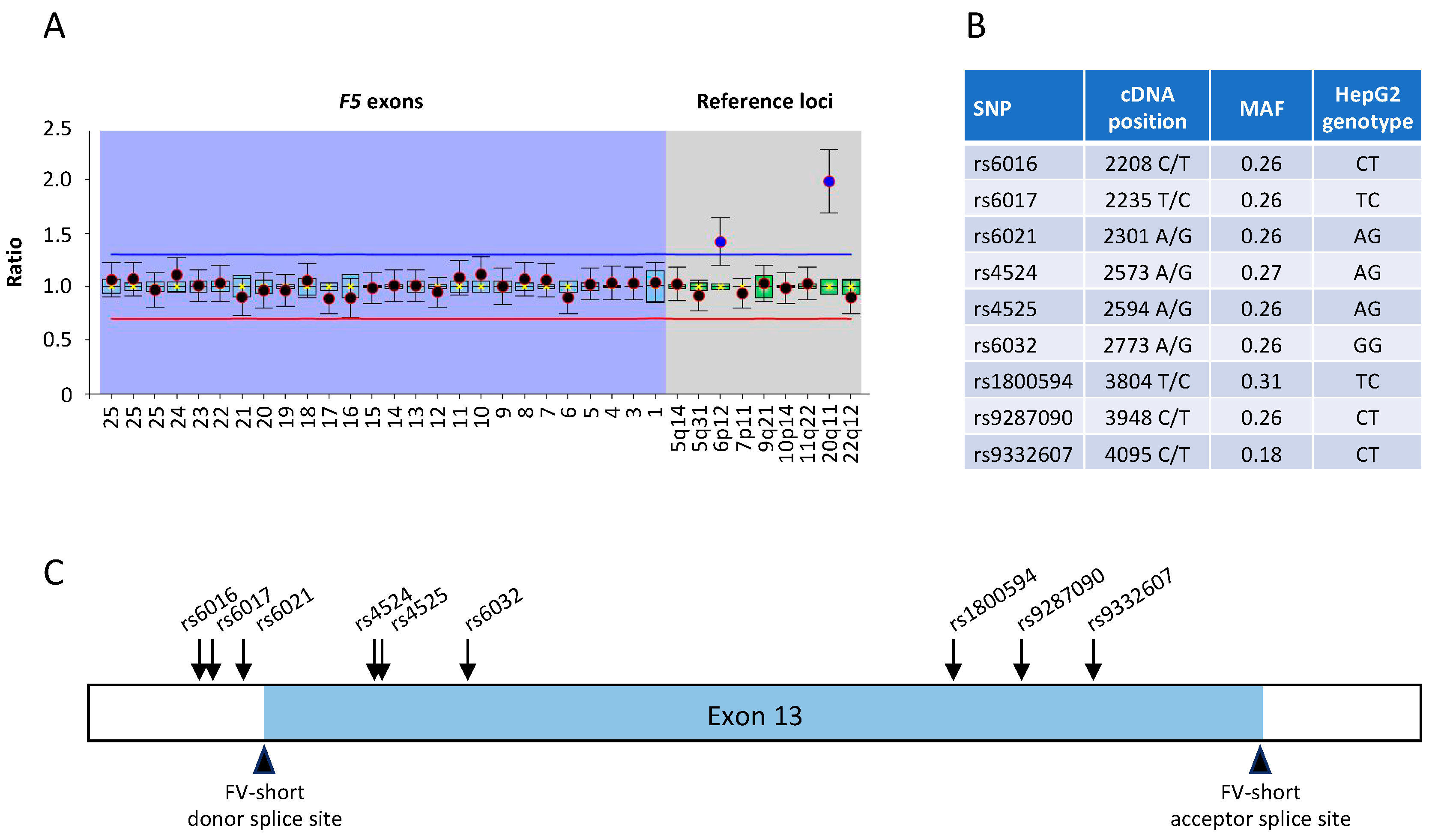
3.2. Characterisation of the F5 Gene in HepG2 Cells
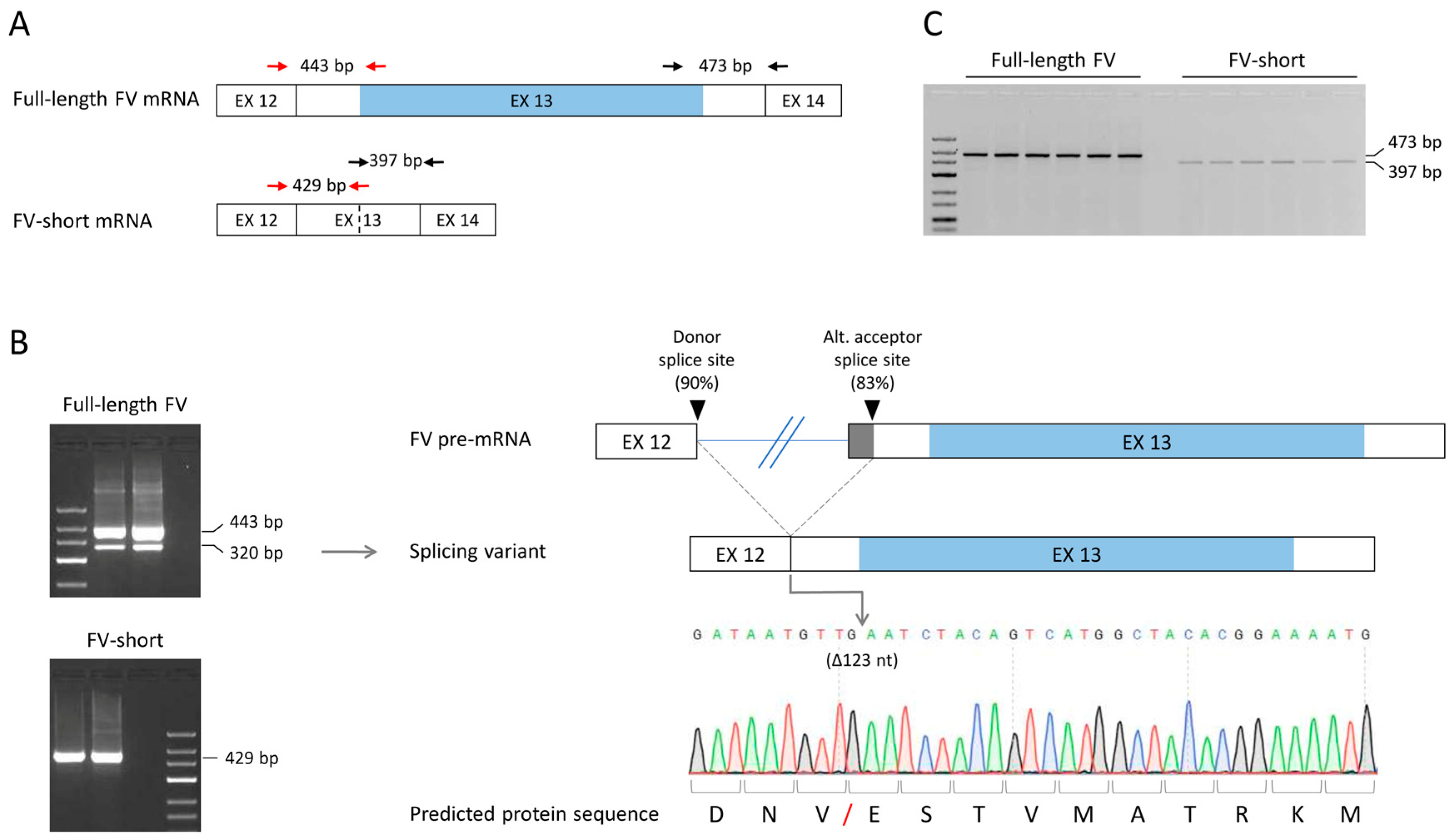
3.3. Optimisation of Full-Length FV and FV-Short Transcript Amplification and Identification of a New Alternatively Spliced F5 Transcript
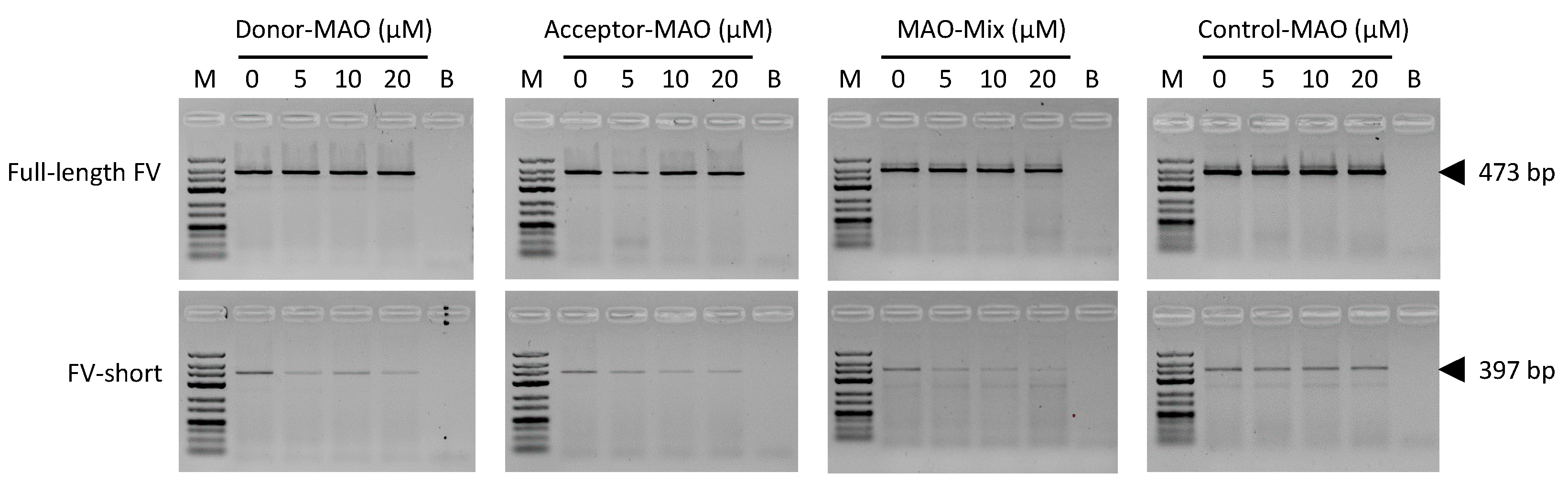
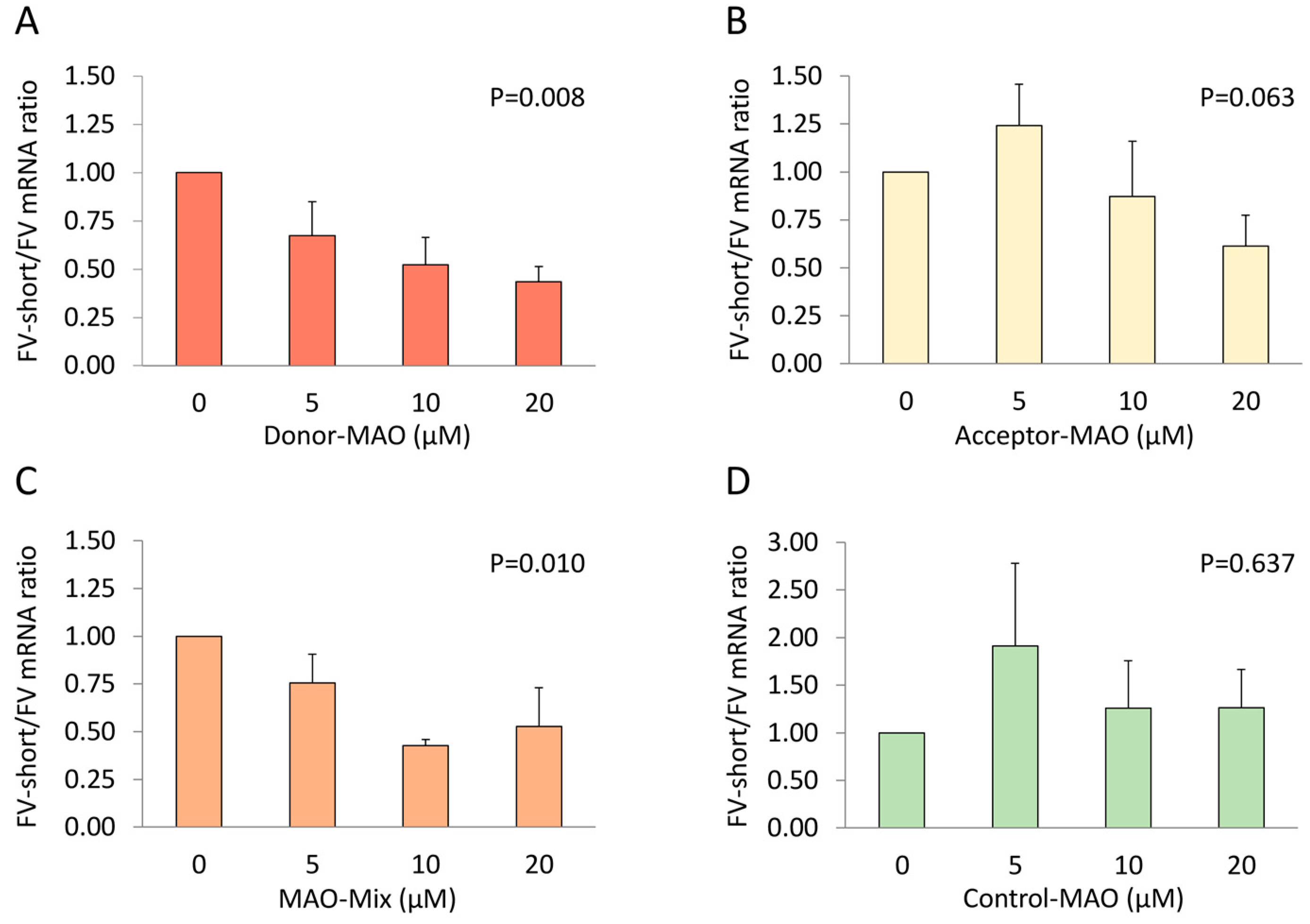
3.4. F5 Splicing Modulation in HepG2 Cells
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Wang, Z.; Burge, C.B. Splicing regulation: From a parts list of regulatory elements to an integrated splicing code. RNA 2008, 14, 802–813. [Google Scholar] [CrossRef] [PubMed]
- Kornblihtt, A.R.; Schor, I.E.; Allo, M.; Dujardin, G.; Petrillo, E.; Munoz, M.J. Alternative splicing: A pivotal step between eukaryotic transcription and translation. Nat. Rev. Mol. Cell Biol. 2013, 14, 153–165. [Google Scholar] [CrossRef] [PubMed]
- Nilsen, T.W.; Graveley, B.R. Expansion of the eukaryotic proteome by alternative splicing. Nature 2010, 463, 457–463. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.I.; van de Geijn, B.; Raj, A.; Knowles, D.A.; Petti, A.A.; Golan, D.; Gilad, Y.; Pritchard, J.K. RNA splicing is a primary link between genetic variation and disease. Science 2016, 352, 600–604. [Google Scholar] [CrossRef] [PubMed]
- Lord, J.; Baralle, D. Splicing in the Diagnosis of Rare Disease: Advances and Challenges. Front. Genet. 2021, 12, 689892. [Google Scholar] [CrossRef] [PubMed]
- Hammond, S.M.; Wood, M.J. Genetic therapies for RNA mis-splicing diseases. Trends Genet. 2011, 27, 196–205. [Google Scholar] [CrossRef] [PubMed]
- Black, A.J.; Gamarra, J.R.; Giudice, J. More than a messenger: Alternative splicing as a therapeutic target. Biochim. Biophys. Acta Gene Regul. Mech. 2019, 1862, 194395. [Google Scholar] [CrossRef] [PubMed]
- Balestra, D.; Branchini, A. Molecular Mechanisms and Determinants of Innovative Correction Approaches in Coagulation Factor Deficiencies. Int. J. Mol. Sci. 2019, 20, 3036. [Google Scholar] [CrossRef] [PubMed]
- Havens, M.A.; Hastings, M.L. Splice-switching antisense oligonucleotides as therapeutic drugs. Nucleic Acids Res. 2016, 44, 6549–6563. [Google Scholar] [CrossRef] [PubMed]
- Davis, R.L.; Homer, V.M.; George, P.M.; Brennan, S.O. A deep intronic mutation in FGB creates a consensus exonic splicing enhancer motif that results in afibrinogenemia caused by aberrant mRNA splicing, which can be corrected in vitro with antisense oligonucleotide treatment. Hum. Mutat. 2009, 30, 221–227. [Google Scholar] [CrossRef] [PubMed]
- Nuzzo, F.; Radu, C.; Baralle, M.; Spiezia, L.; Hackeng, T.M.; Simioni, P.; Castoldi, E. Antisense-based RNA therapy of factor V deficiency: In vitro and ex vivo rescue of a F5 deep-intronic splicing mutation. Blood 2013, 122, 3825–3831. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Nuzzo, F.; Bulato, C.; Nielsen, B.I.; Lee, K.; Wielders, S.J.; Simioni, P.; Key, N.S.; Castoldi, E. Characterization of an apparently synonymous F5 mutation causing aberrant splicing and factor V deficiency. Haemophilia 2015, 21, 241–248. [Google Scholar] [CrossRef] [PubMed]
- Balestra, D.; Barbon, E.; Scalet, D.; Cavallari, N.; Perrone, D.; Zanibellato, S.; Bernardi, F.; Pinotti, M. Regulation of a strong F9 cryptic 5′ss by intrinsic elements and by combination of tailored U1snRNAs with antisense oligonucleotides. Hum. Mol. Genet. 2015, 24, 4809–4816. [Google Scholar] [CrossRef]
- Levin, A.A. Treating Disease at the RNA Level with Oligonucleotides. N. Engl. J. Med. 2019, 380, 57–70. [Google Scholar] [CrossRef] [PubMed]
- Crooke, S.T.; Liang, X.H.; Baker, B.F.; Crooke, R.M. Antisense technology: A review. J. Biol. Chem. 2021, 296, 100416. [Google Scholar] [CrossRef]
- Dahlbäck, B. Pro- and anticoagulant properties of factor V in pathogenesis of thrombosis and bleeding disorders. Int. J. Lab. Hematol. 2016, 38, 4–11. [Google Scholar] [CrossRef]
- Bos, M.H.A.; Camire, R.M. A Bipartite Autoinhibitory Region within the B-domain Suppresses Function in Factor V. J. Biol. Chem. 2012, 287, 26342–26351. [Google Scholar] [CrossRef]
- Rosing, J.; Tans, G.; Govers-Riemslag, J.W.; Zwaal, R.F.; Hemker, H.C. The role of phospholipids and factor Va in the prothrombinase complex. J. Biol. Chem. 1980, 255, 274–283. [Google Scholar] [CrossRef]
- Wood, J.P.; Ellery, P.E.R.; Maroney, S.A.; Mast, A.E. Biology of tissue factor pathway inhibitor. Blood 2014, 123, 2934–2943. [Google Scholar] [CrossRef]
- Wood, J.P.; Bunce, M.W.; Maroney, S.A.; Tracy, P.B.; Camire, R.M.; Mast, A.E. Tissue factor pathway inhibitor-alpha inhibits prothrombinase during the initiation of blood coagulation. Proc. Natl. Acad. Sci. USA 2013, 110, 17838–17843. [Google Scholar] [CrossRef]
- van Doorn, P.; Rosing, J.; Wielders, S.J.; Hackeng, T.M.; Castoldi, E. The C-terminus of tissue factor pathway inhibitor-alpha inhibits factor V activation by protecting the Arg1545 cleavage site. J. Thromb. Haemost. 2017, 15, 140–149. [Google Scholar] [CrossRef]
- Vincent, L.M.; Tran, S.; Livaja, R.; Bensend, T.A.; Milewicz, D.M.; Dahlbäck, B. Coagulation factor V(A2440G) causes east Texas bleeding disorder via TFPIα. J. Clin. Investig. 2013, 123, 3777–3787. [Google Scholar] [CrossRef] [PubMed]
- Petrillo, T.; Ayombil, F.; Van’t Veer, C.; Camire, R.M. Regulation of factor V and factor V-short by TFPIalpha: Relationship between B-domain proteolysis and binding. J. Biol. Chem. 2021, 296, 100234. [Google Scholar] [CrossRef]
- Duckers, C.; Simioni, P.; Spiezia, L.; Radu, C.; Gavasso, S.; Rosing, J.; Castoldi, E. Low plasma levels of tissue factor pathway inhibitor in patients with congenital factor V deficiency. Blood 2008, 112, 3615–3623. [Google Scholar] [CrossRef] [PubMed]
- Dahlbäck, B.; Guo, L.J.; Livaja-Koshiar, R.; Tran, S. Factor V-short and protein S as synergistic tissue factor pathway inhibitor (TFPIalpha) cofactors. Res. Pract. Thromb. Haemost. 2018, 2, 114–124. [Google Scholar] [CrossRef] [PubMed]
- Dahlbäck, B.; Tran, S. The preAR2 region (1458–1492) in Factor V-Short is crucial for the synergistic TFPIalpha-cofactor activity with protein S and the assembly of a trimolecular Factor Xa-inhibitory complex comprising FV-Short, protein S and TFPIalpha. J. Thromb. Haemost. 2021, in press. [Google Scholar] [CrossRef]
- Kuang, S.Q.; Hasham, S.; Phillips, M.D.; Wolf, D.; Wan, Y.; Thiagarajan, P.; Milewicz, D.M. Characterization of a novel autosomal dominant bleeding disorder in a large kindred from east Texas. Blood 2001, 97, 1549–1554. [Google Scholar] [CrossRef]
- Cunha, M.L.; Bakhtiari, K.; Peter, J.; Marquart, J.A.; Meijers, J.C.; Middeldorp, S. A novel mutation in the F5 gene (factor V Amsterdam) associated with bleeding independent of factor V procoagulant function. Blood 2015, 125, 1822–1825. [Google Scholar] [CrossRef]
- Zimowski, K.L.; Petrillo, T.; Ho, M.D.; Wechsler, J.; Shields, J.E.; Denning, G.; Jhita, N.; Rivera, A.A.; Escobar, M.A.; Kempton, C.L.; et al. F5-Atlanta: A novel mutation in F5 associated with enhanced East Texas splicing and FV-short productio. J. Thromb. Haemost. 2021, 19, 1653–1665. [Google Scholar] [CrossRef]
- Desmet, F.O.; Hamroun, D.; Lalande, M.; Collod-Beroud, G.; Claustres, M.; Beroud, C. Human Splicing Finder: An online bioinformatics tool to predict splicing signals. Nucleic Acids Res. 2009, 37, e67. [Google Scholar] [CrossRef]
- Summerton, J.; Weller, D. Morpholino antisense oligomers: Design, preparation, and properties. Antisense Nucleic Acid Drug Dev. 1997, 7, 187–195. [Google Scholar] [CrossRef]
- Lopez-Terrada, D.; Cheung, S.W.; Finegold, M.J.; Knowles, B.B. Hep G2 is a hepatoblastoma-derived cell line. Hum. Pathol. 2009, 40, 1512–1515. [Google Scholar] [CrossRef] [PubMed]
- Zhou, B.; Ho, S.S.; Greer, S.U.; Spies, N.; Bell, J.M.; Zhang, X.; Zhu, X.; Arthur, J.G.; Byeon, S.; Pattni, R.; et al. Haplotype-resolved and integrated genome analysis of the cancer cell line HepG2. Nucleic Acids Res. 2019, 47, 3846–3861. [Google Scholar] [CrossRef] [PubMed]
- Schouten, J.P.; McElgunn, C.J.; Waaijer, R.; Zwijnenburg, D.; Diepvens, F.; Pals, G. Relative quantification of 40 nucleic acid sequences by multiplex ligation-dependent probe amplification. Nucleic Acids Res. 2002, 30, e57. [Google Scholar] [CrossRef] [PubMed]
- Nuzzo, F.; Paraboschi, E.M.; Straniero, L.; Pavlova, A.; Duga, S.; Castoldi, E. Identification of a novel large deletion in a patient with severe factor V deficiency using an in-house F5 MLPA assay. Haemophilia 2015, 21, 140–147. [Google Scholar] [CrossRef]
- Ruijter, J.M.; Ramakers, C.; Hoogaars, W.M.; Karlen, Y.; Bakker, O.; van den Hoff, M.J.; Moorman, A.F. Amplification efficiency: Linking baseline and bias in the analysis of quantitative PCR data. Nucleic Acids Res. 2009, 37, e45. [Google Scholar] [CrossRef] [PubMed]
- Chowdary, P. Anti-tissue factor pathway inhibitor (TFPI) therapy: A novel approach to the treatment of haemophilia. Int. J. Hematol. 2020, 111, 42–50. [Google Scholar] [CrossRef] [PubMed]
- Sehgal, A.; Vaishnaw, A.; Fitzgerald, K. Liver as a target for oligonucleotide therapeutics. J. Hepatol. 2013, 59, 1354–1359. [Google Scholar] [CrossRef] [PubMed]
- Amantana, A.; Iversen, P.L. Pharmacokinetics and biodistribution of phosphorodiamidate morpholino antisense oligomers. Curr. Opin. Pharmacol. 2005, 5, 550–555. [Google Scholar] [CrossRef] [PubMed]
- Castoldi, E. F5-Atlanta: Factor V-short strikes again. J. Thromb. Haemost. 2021, 19, 1638–1640. [Google Scholar] [CrossRef] [PubMed]
- Shenasa, H.; Hertel, K.J. Combinatorial regulation of alternative splicing. Biochim. Biophys. Acta Gene Regul. Mech. 2019, 1862, 194392. [Google Scholar] [CrossRef]
- Chelle, P.; Montmartin, A.; Damien, P.; Piot, M.; Cournil, M.; Lienhart, A.; Genre-Volot, F.; Chambost, H.; Morin, C.; Tardy-Poncet, B. Tissue factor pathway inhibitor is the main determinant of thrombin generation in haemophilic patients. Haemophilia 2019, 25, 343–348. [Google Scholar] [CrossRef] [PubMed]
- MacDonald, S.; White, D.; Langdown, J.; Downes, K.; Thomas, W. Investigation of patients with unclassified bleeding disorder and abnormal thrombin generation for physiological coagulation inhibitors reveals multiple abnormalities and a subset of patients with increased tissue factor pathway inhibitor activity. Int. J. Lab. Hematol. 2020, 42, 246–255. [Google Scholar] [CrossRef] [PubMed]
- Mehic, D.; Tolios, A.; Hofer, S.; Ay, C.; Haslacher, H.; Rejto, J.; Ouwehand, W.H.; Downes, K.; Haimel, M.; Pabinger, I.; et al. Elevated levels of tissue factor pathway inhibitor in patients with mild to moderate bleeding tendency. Blood Adv. 2021, 5, 391–398. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Todaro, A.M.; Hackeng, T.M.; Castoldi, E. Antisense-Mediated Down-Regulation of Factor V-Short Splicing in a Liver Cell Line Model. Appl. Sci. 2021, 11, 9621. https://doi.org/10.3390/app11209621
Todaro AM, Hackeng TM, Castoldi E. Antisense-Mediated Down-Regulation of Factor V-Short Splicing in a Liver Cell Line Model. Applied Sciences. 2021; 11(20):9621. https://doi.org/10.3390/app11209621
Chicago/Turabian StyleTodaro, Alice M., Tilman M. Hackeng, and Elisabetta Castoldi. 2021. "Antisense-Mediated Down-Regulation of Factor V-Short Splicing in a Liver Cell Line Model" Applied Sciences 11, no. 20: 9621. https://doi.org/10.3390/app11209621
APA StyleTodaro, A. M., Hackeng, T. M., & Castoldi, E. (2021). Antisense-Mediated Down-Regulation of Factor V-Short Splicing in a Liver Cell Line Model. Applied Sciences, 11(20), 9621. https://doi.org/10.3390/app11209621




