Low-Rank Approximation of Difference between Correlation Matrices Using Inner Product
Abstract
1. Introduction
2. Methods
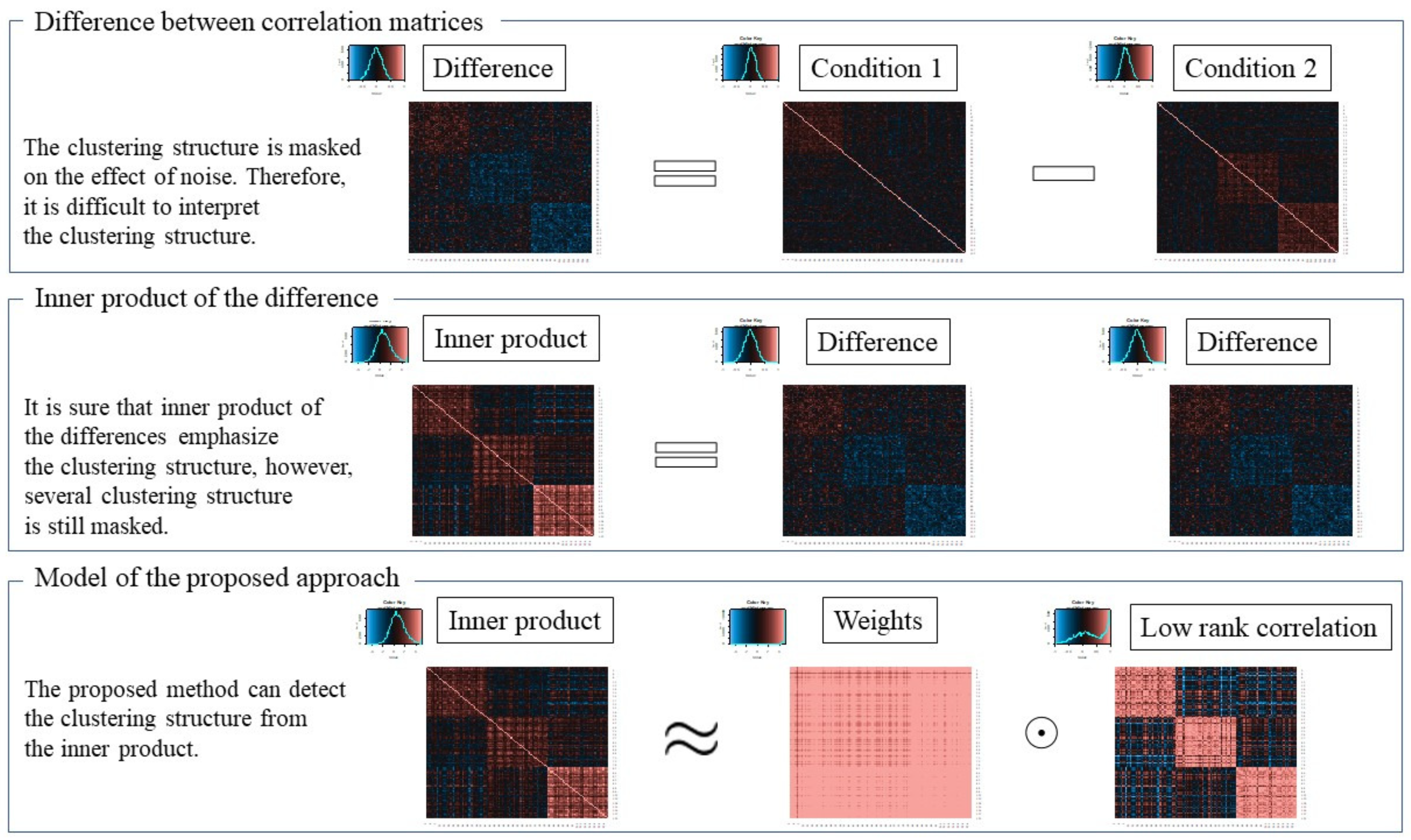
2.1. Model of Proposed Method
2.2. Formulation of Proposed Method
2.3. Algorithm for Estimating Low-Rank Correlation Matrix Based on MM Algorithm
| Algorithm 1 Estimating correlation matrix with rank d |
| Require: Inner product , rank d, initial vectors , and Ensure: coordinate matrix with rank d while do for do end for end while return |
2.4. Simulation Study
- Factor 1:
- Methods
- Factor 2:
- Rank
- Factor 3:
- Number of clusters
- Factor 4:
- Difference between true correlations
2.5. fMRI Data for Mental Arithmetic Task
2.5.1. Participants
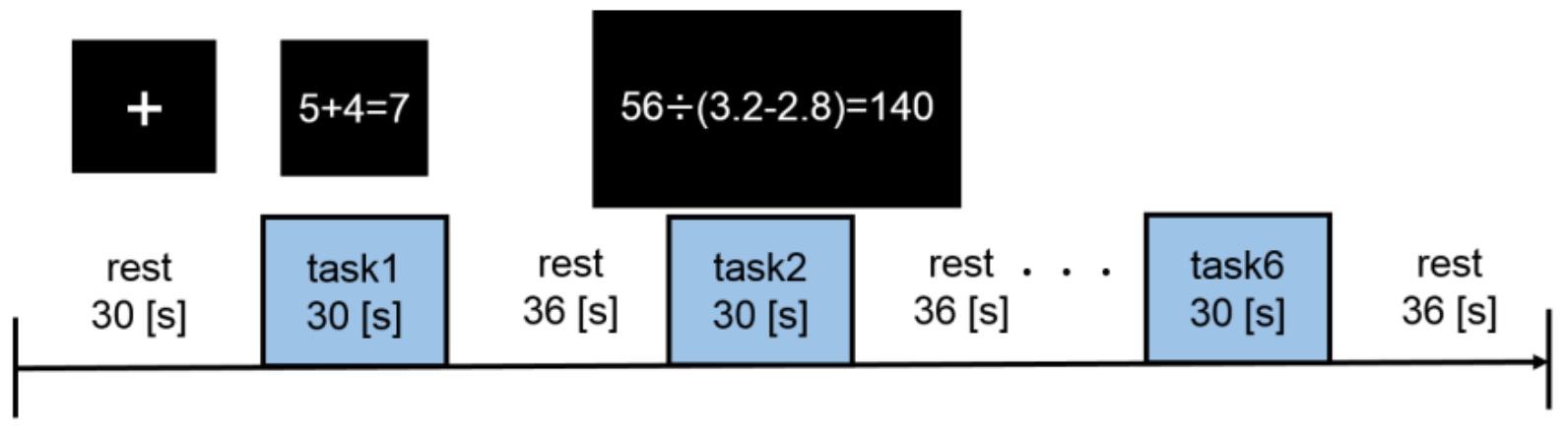
2.5.2. Experimental Design
2.5.3. Data Acquisition
2.5.4. Data Preprocessing
2.5.5. Functional Connectivity Analysis: Derivation of Correlation Matrices
3. Results
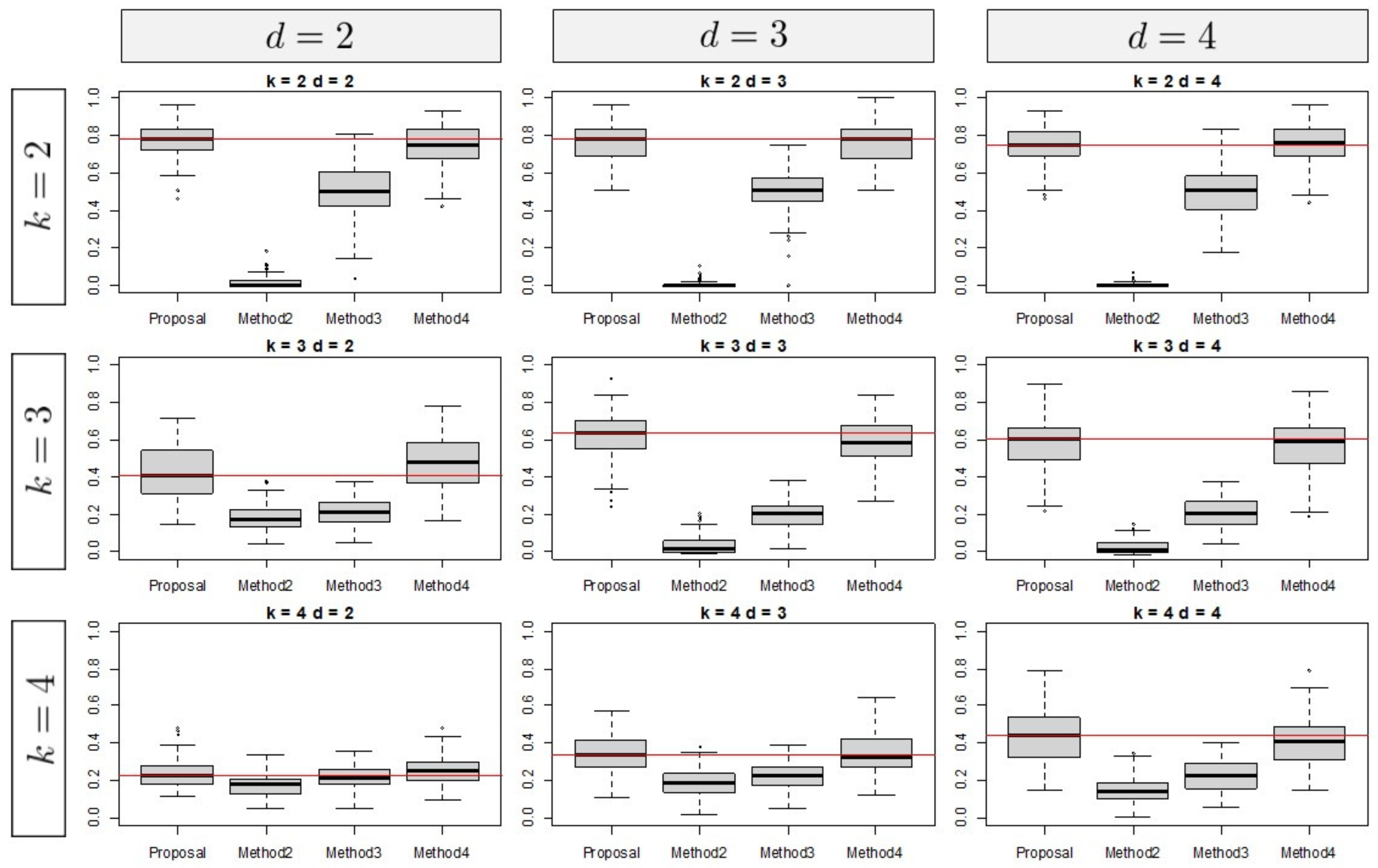
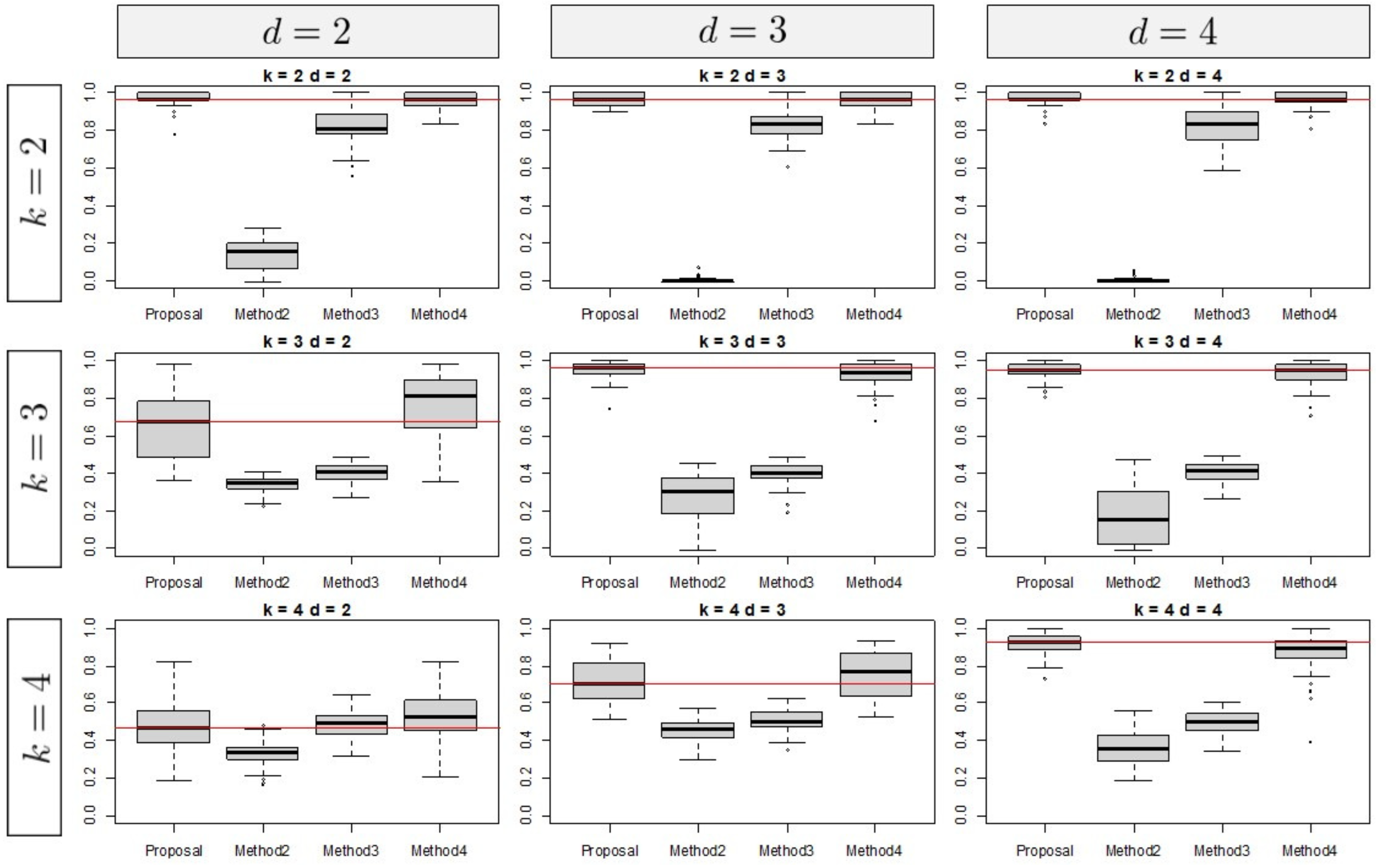
3.1. Simulation Results
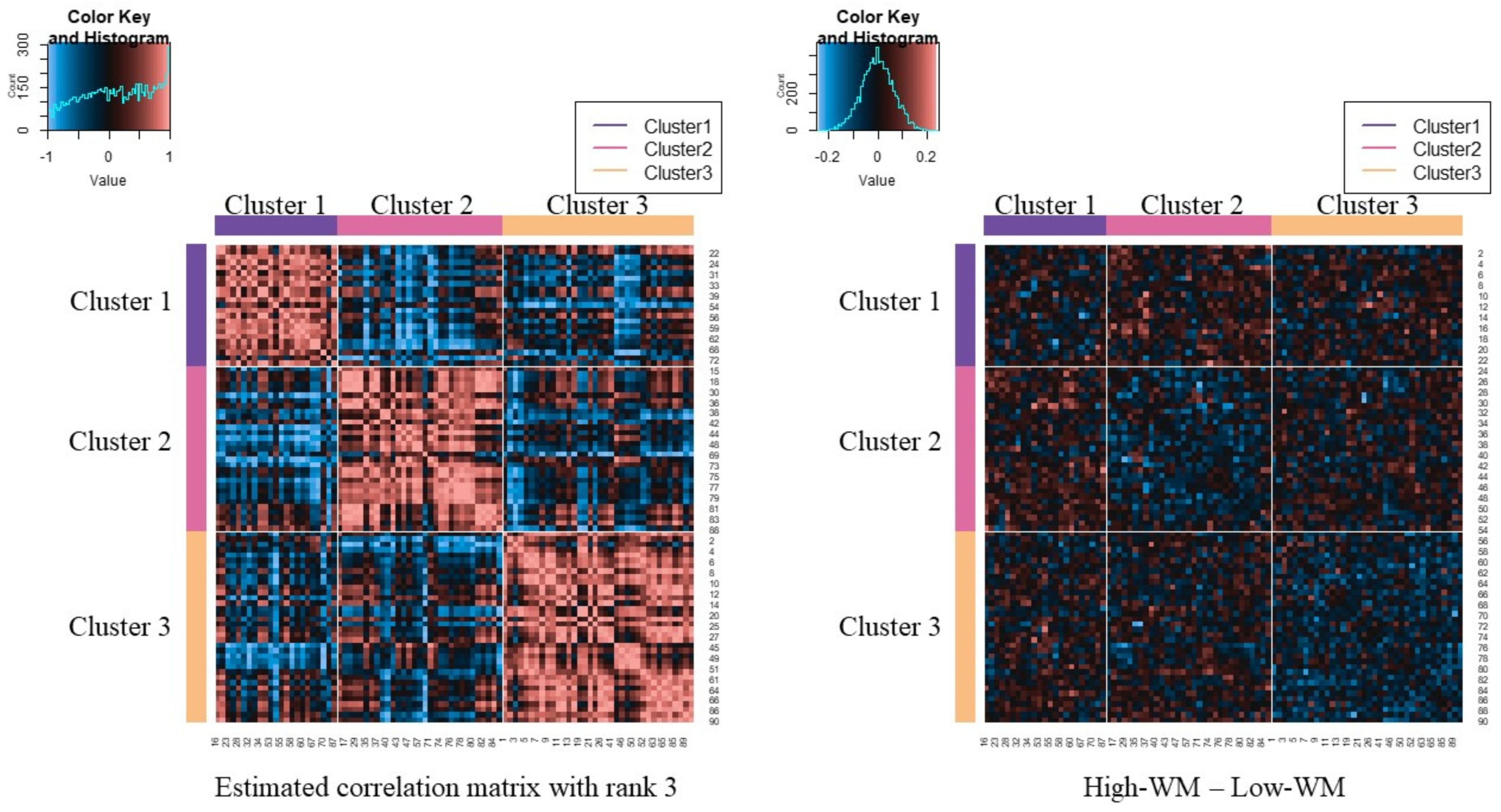
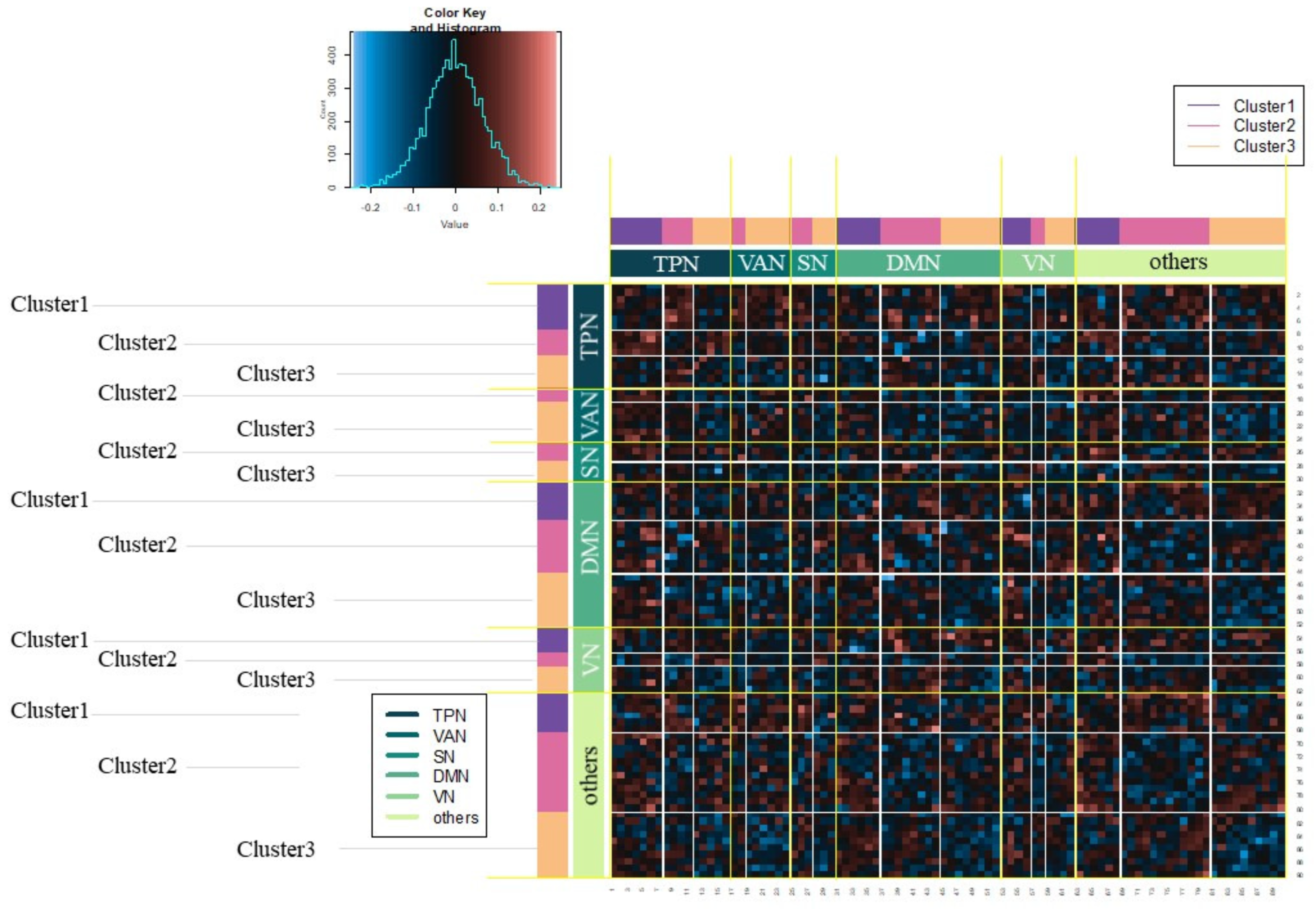
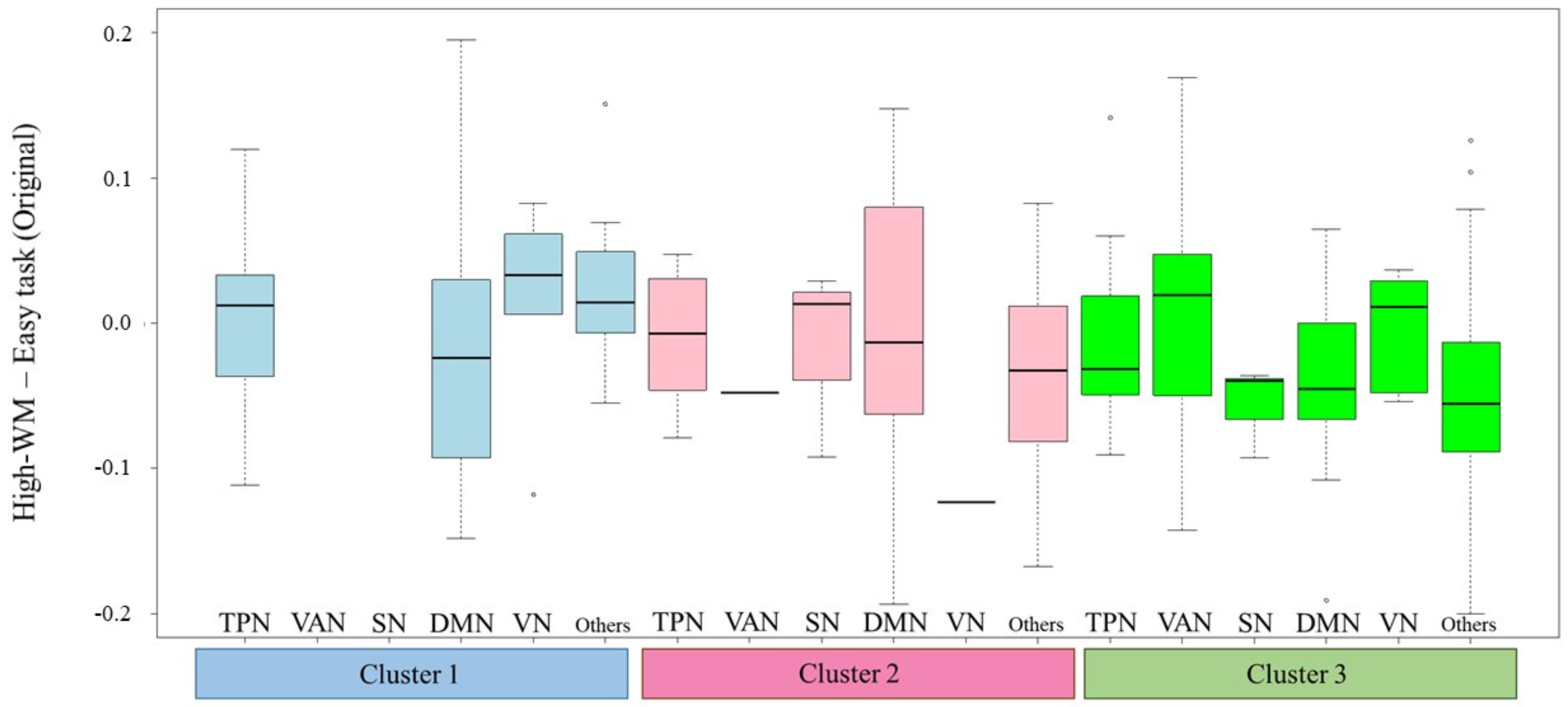
3.2. Results of fMRI Data Analysis
4. Conclusions and Remarks
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| AAL | automated anatomical labeling |
| ARI | adjusted Rand index |
| aCompCor | an anatomical component-based noise correction method |
| BOLD | blood-oxygen-level-dependent |
| CON | cingulo-opercular network |
| DAN | dorsal attention network |
| DMN | default mode network |
| EEG | electroencephalography |
| fMRI | functional magnetic resonance imaging |
| fNIRS | functional near-infrared spectroscopy |
| FPN | fronto-parietal network |
| IQRs | interquartile ranges |
| MNI | Montreal Neurological Institute |
| ROIs | regions of interest |
| SN | salience network |
| VN | visual network |
| TPN | task positive network |
| VAN | ventral attention network |
| WM | working memory |
References
- Fillipi, M. FMRI Techniques and Protocols; Springer Protocols, Humana Press: New York, NY, USA, 2009. [Google Scholar]
- Friston, K.; Ashburner, J.; Kiebel, S.; Nichols, T.; Penny, W. Statistical Parametric Mapping: The Analysis of Functional Brain Images; Academic Press: London, UK, 2007. [Google Scholar]
- Ferrari, M.; Quaresunama, V. A brief review on the history of human functional near-infrared spectroscopy (fnirs) development and fields of application. NeuroImage 2012, 63, 921–935. [Google Scholar] [CrossRef]
- Michel, C.M.; Murray, M.M.; Lantz, G.; Gonzalez, S.; Spinelli, L.; Grave de Peralta, R. Eeg source imaging. Clin. Neurophysiol. 2004, 114, 2195–2222. [Google Scholar] [CrossRef] [PubMed]
- Bullmore, E.; Sporns, O. Complex brain networks: Graph theoretical analysis of structural and functional systems. Nat. Rev. Neurosci. 2009, 10, 186–198. [Google Scholar] [CrossRef] [PubMed]
- Sporns, O. Structure and function of complex brain networks. Dialogues Clin. Neurosci. 2013, 15, 247–262. [Google Scholar]
- Varoquaux, G.; Craddock, R.C. Learning and comparing functional connectomes across subjects. NeuroImage 2013, 80, 405–415. [Google Scholar] [CrossRef] [PubMed]
- Lerman-Sinkoff, D.B.; Barch, D.M. Network community structure alterations in adult schizophrenia:identification and localization of alterations. Neuroimage Clin. 2016, 10, 96–106. [Google Scholar] [CrossRef]
- Cole, M.W.; Basset, D.S.; Power, J.D.; Braver, T.S.; Petersen, S.E. Intrinsic and task-evoked network architectures of the human brain. Neuron 2014, 83, 238–251. [Google Scholar] [CrossRef]
- Fränti, P.; Sieranoja, S. K-means properties on six clustering benchmark datasets. Appl. Intell. 2018, 48, 4743–4759. [Google Scholar] [CrossRef]
- Yang, J.; Rahardja, S.; Fränti, P. Mean-shift outlier detection and filtering. Pattern Recognit. 2021, 115, 107874. [Google Scholar] [CrossRef]
- Steinley, D. K-means clustering: A half-century synthesis. Br. J. Math. Stat. Psychol. 2006, 59, 1–34. [Google Scholar] [CrossRef]
- Terada, Y. Clustering for high-dimension, low-sample size data using distance vectors. arXiv 2013, arXiv:1312.3386. [Google Scholar]
- Forgy, E.W. Cluster analysis of multivariate data: Efficiency versus interpretability of classifications. Biometrics 1965, 21, 768–769. [Google Scholar]
- Knol, D.K.; Ten Berge, J.M.F. Least-squares approximation of an improper correlation matrix by a proper one. Psychometrika 1989, 54, 53–61. [Google Scholar] [CrossRef]
- Lurie, P.M.; Goldberg, M.S. An approximate method for sampling correlated random variables from partially-specified distributions. Manag. Sci. 1998, 44, 203–218. [Google Scholar] [CrossRef]
- Malick, J. A dual approach to solve semidefinite least squares problems. SIAM J. Matrix Anal. Appl. 2004, 26, 272–284. [Google Scholar] [CrossRef]
- Qi, H.D.; Sun, D. A quadratically convergent newton method for computing the nearest correlation matrix. SIAM J. Matrix Anal. Appl. 2006, 28, 360–385. [Google Scholar] [CrossRef]
- Borsdorf, R.; Higham, N.J. A preconditioned newton algorithm for the nearest correlation matrix. IMA J. Numer. Anal. 2010, 30, 94–107. [Google Scholar] [CrossRef]
- Pietersz, R.; Groenen, P. Rank reduction of correlation matrices by majorization. Quant. Financ. 2004, 4, 649–662. [Google Scholar] [CrossRef]
- Simon, D.; Abell, J. A majorization algorithm for constrained approximation. Linear Algebra Appl. 2010, 432, 1152–1164. [Google Scholar] [CrossRef][Green Version]
- Grubisic, I.; Pietersz, R. Efficient rank reduction of correlation matrices. Linear Algebra Appl. 2007, 422, 629–653. [Google Scholar] [CrossRef]
- Duan, X.F.; Bai, J.C.; Li, J.F.; Peng, J.J. On the low rank solution of the q-weighted nearest correlation matrix problem. Numer. Linear Algebra Appl. 2016, 23, 340–355. [Google Scholar] [CrossRef]
- Hunter, D.R.; Lange, K. A tutorial on mm algorithm. Am. Stat. 2004, 58, 30–37. [Google Scholar] [CrossRef]
- Borg, I.; Groenen, P. Modern Multidimensional Scaling; Springer: New York, NY, USA, 1997. [Google Scholar]
- Zhang, Z.; Dai, G. Optimal scoring for unsupervised learning. Neural Inf. Process. Syst. 2009, 23, 2241–2249. [Google Scholar]
- Wang, Y.; Fang, Y.; Wang, J. Sparse optimal discriminant clustering. Stat. Comput. 2016, 26, 629–639. [Google Scholar] [CrossRef]
- Hubert, L.; Arabie, P. Comparing partitions. J. Classif. 1985, 2, 193–218. [Google Scholar] [CrossRef]
- Arabie, P.; Hubert, L. Cluster analysis in marketing research. In Advanced Methods of Marketing Research; Bagozzi, R.P., Ed.; Blackwell: Oxford, UK.
- Baddeley, A.D.; Hitch, G. The psychology of learning and motivation. Work. Mem. 1974, 8, 47–89. [Google Scholar]
- Baddeley, A.D. The episodic buffer: A new component of working memory? Trends Cogn. Sci. 2000, 4, 417–423. [Google Scholar] [CrossRef]
- Susan, W.; Nieto-Castanon, A. Conn: A functional connectivity toolbox for correlated and anticorrelated brain networks. Brain Connect. 2012, 2, 125–141. [Google Scholar]
- Behzadi, Y.; Restom, K.; Liau, J.; Liu, T.T. A component based noise correction method (compcor) for bold and perfusion based fmri. Neuroimage 2007, 37, 90–101. [Google Scholar] [CrossRef] [PubMed]
- D’Esposito, M.; Postle, B.R. The cognitive neuroscience of working memory. Annu. Rev. Psychol. 2015, 66, 115–142. [Google Scholar] [CrossRef]
- Milligan, G.W.; Cooper, M.C. A study of standardization of variables in cluster analysis. J. Classif. 1988, 5, 181–204. [Google Scholar] [CrossRef]
- Cole, M.W.; Repovs, G.; Anticevic, A. The frontopaparietal control system: A central role in mental health. Neuroscientist 2014, 20, 652–664. [Google Scholar] [CrossRef]
- Wallis, G.; Stokes, M.; Cousijn, H.; Woolrich, M.; Nobre, A.C. Frontoparietal and cingulo-opercular networks play dissociable roles in control of working memory. J. Cogn. Neurosci. 2015, 27, 2019–2034. [Google Scholar] [CrossRef]
- Dosenbach, N.U.F.; Fair, D.A.; Miezin, F.M.; Cohen, A.L.; Wenger, K.K.; Dosenbach, R.A.T.; Fox, M.D.; Snyder, A.Z.; Vincent, J.L.; Raichle, M.E.; et al. Distinct brain networks for adaptive and stable task control in humans. Proc. Natl. Acad. Sci. USA 2007, 104, 11073–11078. [Google Scholar] [CrossRef] [PubMed]
- Ham, T.; de Boissezon, X.; Joffe, A.; Sharp, D.J. Cognitive control and the salience network: An investigation of error processing and effective connectivity. J. Neurosci. 2013, 33, 7091–7098. [Google Scholar] [CrossRef]
- Buckner, R.L.; Andrews-Hanna, J.R.; Schaacter, D.L. The brain’s default network: Anatomy, function, and relevance to disease. Ann. N. Y. Acad. Sci. 2008, 1124, 1–38. [Google Scholar] [CrossRef] [PubMed]
- Rousseeuw, P.J. Silhouettes: A graphical aid to the interpretation and validation of cluster analysis. J. Comput. Appl. Math. 1987, 20, 53–65. [Google Scholar] [CrossRef]
- Godwin, D.; Ji, A.; Kandala, S.; Mamah, D. Functional connectivity of cognitive brain networks in schizophrenia during a working memory task. Front. Psychiatry 2017, 8, 294. [Google Scholar] [CrossRef]
- Vossel, S.; Geng, J.J.; Fink, G.R. Dorsal and ventral attention systems: Distinct neural circuits but collaborative roles. Neuroscientist 2014, 20, 150–159. [Google Scholar] [CrossRef]
- Van der Hoef, H.; Warrens, M.J. Understanding information theoretic measures for comparing clustering. Behaviormetrika 2019, 46, 353–370. [Google Scholar] [CrossRef]
- Rezaei, M.; Fränti, P. Set matching measures for external cluster validity. IEEE Trans. Knowl. Data Eng. 2016, 28, 2173–2186. [Google Scholar] [CrossRef]
- Meilă, M. Comparing clusterings.an information based distance. J. Multivar. Anal. 2007, 98, 873–895. [Google Scholar] [CrossRef]
- Meilă, M. Criteria for comparing clustering. In Handbook of Cluster Analysis; Hennig, C., Meilă, M., Murtagh, F., Rocci, R., Eds.; Chapman and Hall: New York, NY, USA, 2015; pp. 619–636. [Google Scholar]
- Vinh, N.X.; Epps, J.; Bailey, J. Information theoretic measures for clustering comparison: Variants, properties, normalization and correction for chance. J. Mach. Learn. Res. 2010, 11, 2837–2854. [Google Scholar]
- De Souto, M.C.P.; Hielho, A.L.V.; Faceli, K.; Sakata, T.C.; Bonadia, V.; Costa, I.G. A comparison of external clustering evaluation indices in the context of imbalanced data sets. In Proceedings of the 2012 Brazilian Symposium on Neural Networks, Curitiba, Brazil, 20–25 October 2012; pp. 49–54. [Google Scholar]
- Meilă, M.; Heckerman, D. An experimental comparison of model based clustering methods. Mach. Learn. 2001, 41, 9–29. [Google Scholar] [CrossRef]
- Fränti, P.; Rezaei, M.; Zhao, Q. Centroid index:Cluster level similarity measure. Pattern Recognit. 2014, 47, 3034–3045. [Google Scholar] [CrossRef]
- Zhao, Q.; Fränti, P. Centroid ratio for a pairwise random swap clustering algorithm. IEEE Trans. Knowl. Data Eng. 2014, 26, 1090–1101. [Google Scholar] [CrossRef]
- Tibshirani, R.; Walther, G.; Hastie, T. Estimating the number of clusters in a data set via the gap statistic. J. R. Stat. Soc. Stat. Methodol. Ser. 2001, 63, 411–423. [Google Scholar] [CrossRef]
- Sugar, C.A.; James, G.M. Finding the Number of Clusters in a Dataset: An Information-Theoretic Approach. J. Am. Stat. Assoc. 2003, 98, 750–763. [Google Scholar] [CrossRef]
- Wang, J. Consistent selection of the number of clusters via crossvalidation. Biometrika 2010, 97, 893–904. [Google Scholar] [CrossRef]

| Names of Factors | Levels | Descriptions |
|---|---|---|
| Methods | 4 | proposal, Method 2, Method 3, and Method 4 |
| Rank | 3 | Rank and 4 |
| The number of clusters | 3 | , and 4 |
| The difference between true correlation | 2 | , and |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tanioka, K.; Hiwa, S. Low-Rank Approximation of Difference between Correlation Matrices Using Inner Product. Appl. Sci. 2021, 11, 4582. https://doi.org/10.3390/app11104582
Tanioka K, Hiwa S. Low-Rank Approximation of Difference between Correlation Matrices Using Inner Product. Applied Sciences. 2021; 11(10):4582. https://doi.org/10.3390/app11104582
Chicago/Turabian StyleTanioka, Kensuke, and Satoru Hiwa. 2021. "Low-Rank Approximation of Difference between Correlation Matrices Using Inner Product" Applied Sciences 11, no. 10: 4582. https://doi.org/10.3390/app11104582
APA StyleTanioka, K., & Hiwa, S. (2021). Low-Rank Approximation of Difference between Correlation Matrices Using Inner Product. Applied Sciences, 11(10), 4582. https://doi.org/10.3390/app11104582





