Hyperspectral Imaging for Minced Meat Classification Using Nonlinear Deep Features
Abstract
:1. Introduction
- (1)
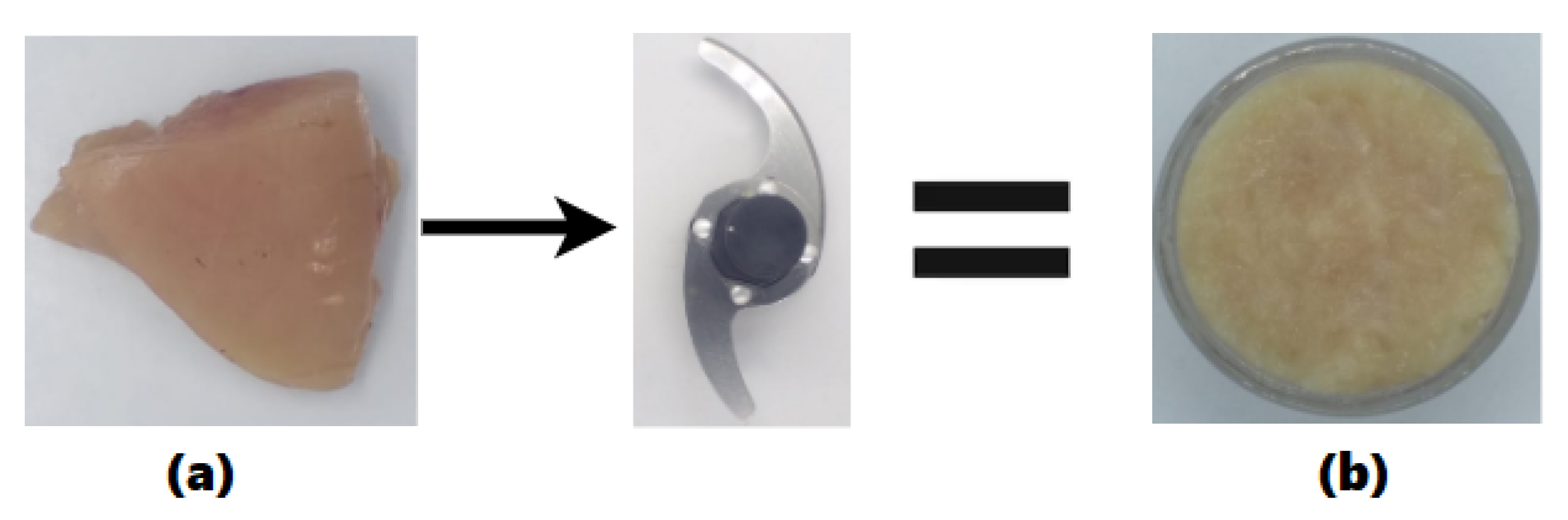
- Collection and mincing of meat types using two cross blades.
- (2)
- HSI system is used for image acquisition and image correction.
- (3)
- Reduction of dimensionality using Isos-bestic point in Mb pigments to retain maximum color information.
- (4)
- Classification of minced meat types using KNN, SVM, and 3D-CNN.
2. Material and Methods
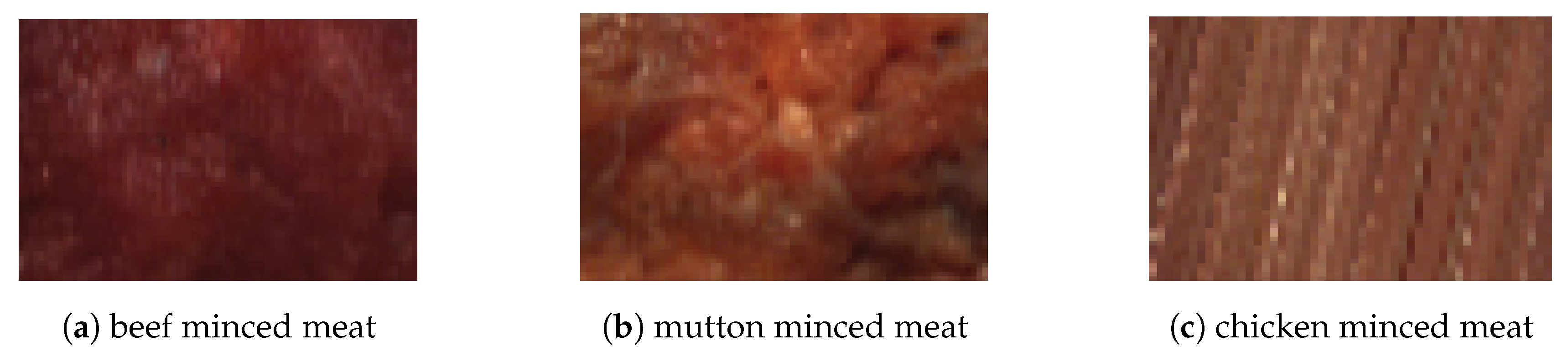
2.1. Preparation of Samples
2.2. Hyperspectral Imaging System (HSI)
2.2.1. Image Correction
2.2.2. Spatial Pre-Processing
2.2.3. Spectral Pre-Processing
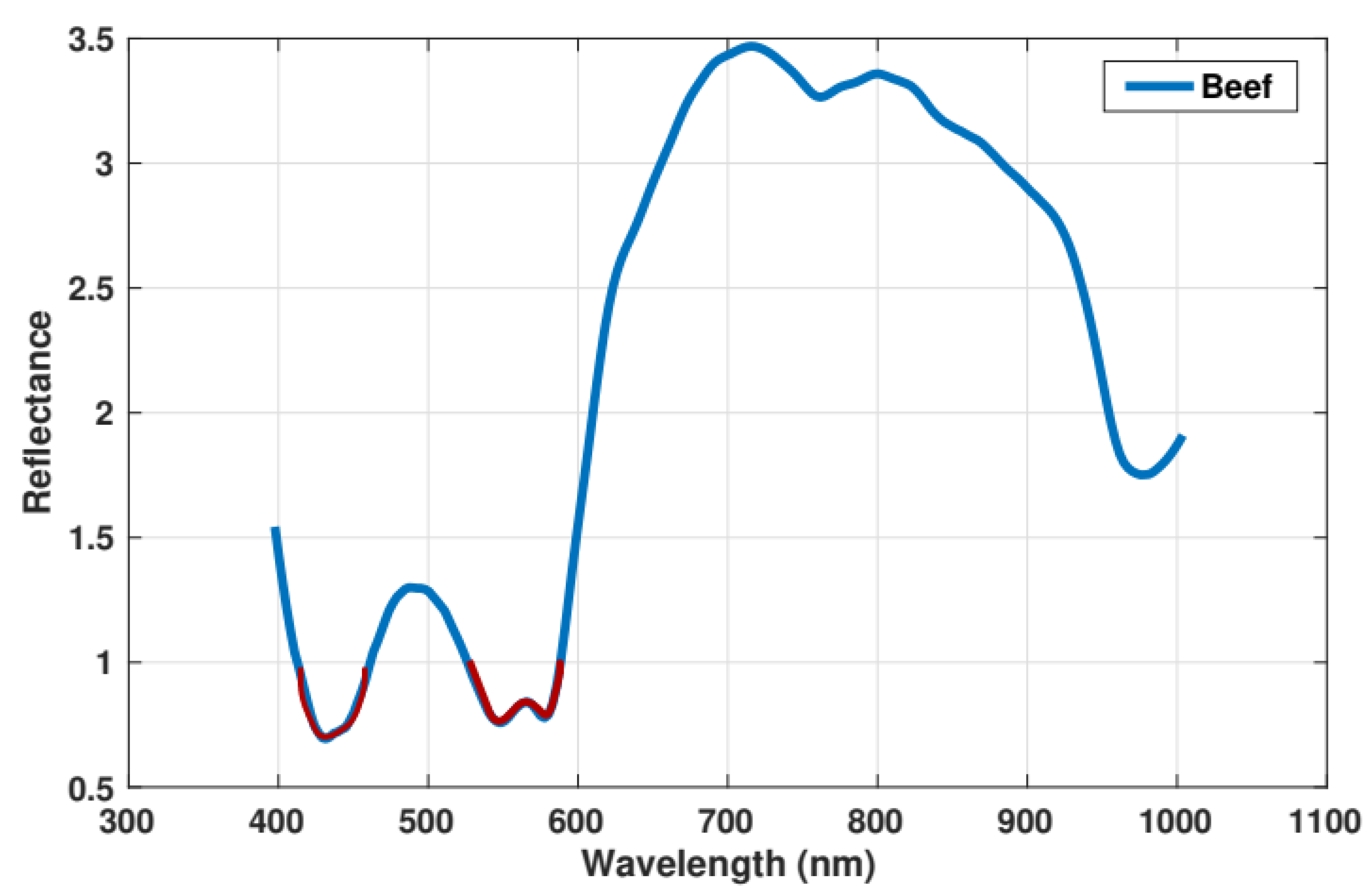
2.3. Isos-Bestic Point Reduction
2.4. Classification
2.4.1. Multivariate Data Analysis
SVM
KNN
2.4.2. Deep Convolution Neural Network
3. Results and Discussion
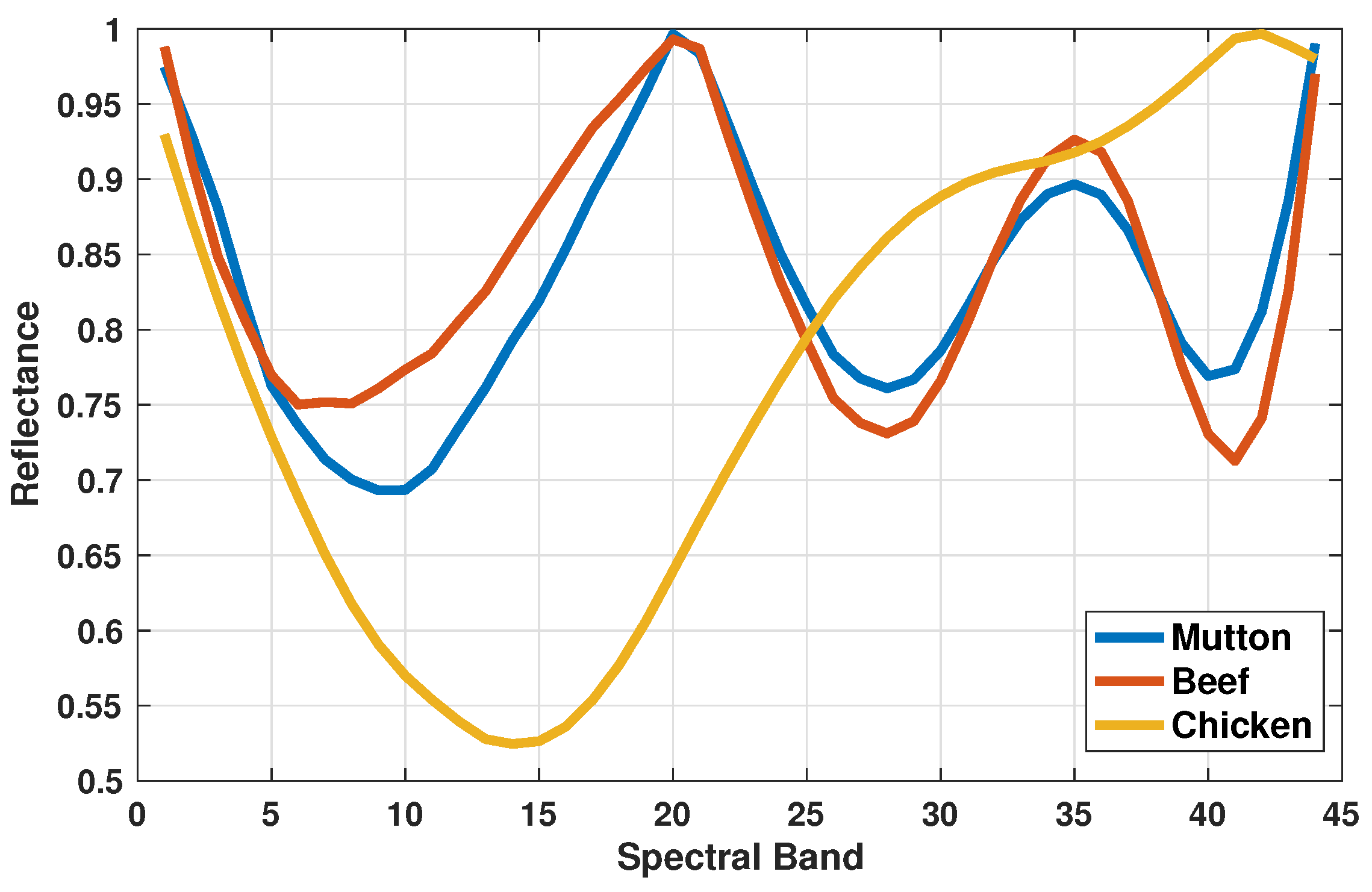
3.1. Minced Meat Spectral Analysis
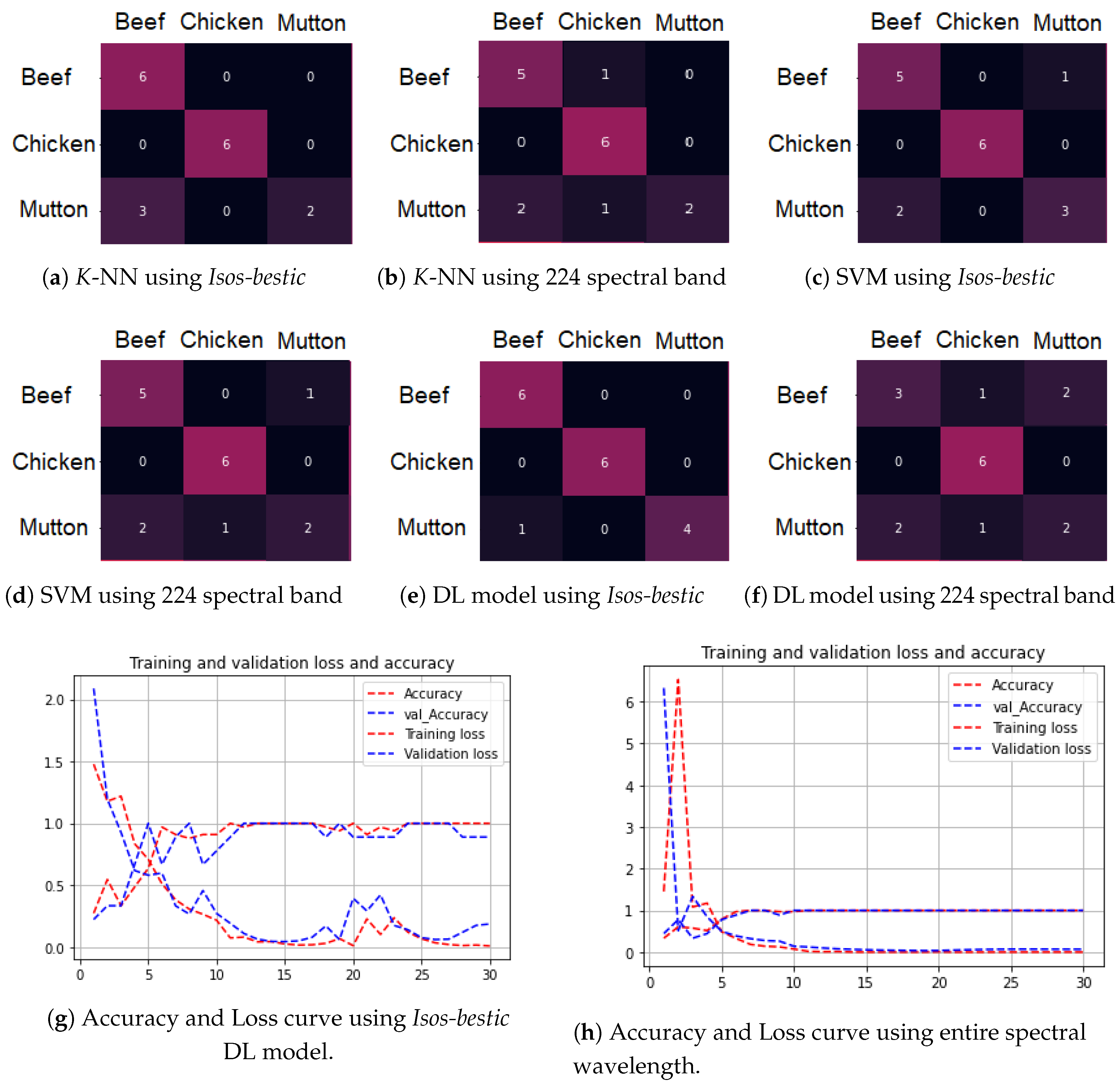
3.2. Classification of Minced Meat
3D-DL and Architecture
3.3. Comparison with SVM and KNN
3.3.1. Principle Component Analysis-Based Comparison
3.3.2. Model Evaluation
4. Conclusions
Author Contributions
Funding
Conflicts of Interest
Dataset and Code Availability:
References
- Ahmad, M.; Mazzara, M.; Raza, R.A.; Distefano, S.; Asif, M.; Sarfraz, M.S.; Khan, A.M.; Sohaib, A. Multiclass Non-Randomized Spectral–Spatial Active Learning for Hyperspectral Image Classification. Appl. Sci. 2020, 10, 4739. [Google Scholar] [CrossRef]
- Ahmad, M.; Khan, A.M.; Mazzara, M.; Distefano, S. Multi-layer Extreme Learning Machine-based Autoencoder for Hyperspectral Image Classification. In Proceedings of the VISIGRAPP (4: VISAPP), Prague, Czech Republic, 25–27 February 2019; pp. 75–82. [Google Scholar]
- Dixit, Y.; Casado-Gavalda, M.P.; Cama-Moncunill, R.; Cullen, P.; Sullivan, C. Challenges in model development for meat composition using multipoint NIR spectroscopy from at-line to in-line monitoring. J. Food Sci. 2017, 82, 1557–1562. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lu, G.; Fei, B. Medical hyperspectral imaging: A review. J. Biomed. Opt. 2014, 19, 010901. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Qiu, G.; Wang, J.; Zhou, X.; Shen, L.; Aleksic, M.; Vaddadi, S.; Zhuo, S. Multispectral Imaging System. U.S. Patent 9,692,991, 27 June 2017. [Google Scholar]
- Al-Sarayreh, M.; Reis, M.M.; Yan, W.Q.; Klette, R. Deep spectral-spatial features of snapshot hyperspectral images for red-meat classification. In Proceedings of the 2018 International Conference on Image and Vision Computing New Zealand (IVCNZ), Auckland, New Zealand, 19–21 November 2018; pp. 1–6. [Google Scholar]
- Ahmad, M.; Shabbir, S.; Oliva, D.; Mazzara, M.; Distefano, S. Spatial-prior generalized fuzziness extreme learning machine autoencoder-based active learning for hyperspectral image classification. Optik 2020, 206, 163712. [Google Scholar] [CrossRef]
- Geronimo, B.C.; Mastelini, S.M.; Carvalho, R.H.; Júnior, S.B.; Barbin, D.F.; Shimokomaki, M.; Ida, E.I. Computer vision system and near-infrared spectroscopy for identification and classification of chicken with wooden breast, and physicochemical and technological characterization. Infrared Phys. Technol. 2019, 96, 303–310. [Google Scholar] [CrossRef]
- Taheri-Garavand, A.; Fatahi, S.; Omid, M.; Makino, Y. Meat quality evaluation based on computer vision technique: A review. Meat Sci. 2019, 156, 183–195. [Google Scholar] [CrossRef]
- Zheng, Y.; Li, Y.; Satija, A.; Pan, A.; Sotos-Prieto, M.; Rimm, E.; Willett, W.C.; Hu, F.B. Association of changes in red meat consumption with total and cause specific mortality among US women and men: Two prospective cohort studies. BMJ 2019, 365. [Google Scholar] [CrossRef] [Green Version]
- Fuseini, A.; Wotton, S.B.; Knowles, T.G.; Hadley, P.J. Halal meat fraud and safety issues in the UK: A review in the context of the European Union. Food Ethics 2017, 1, 127–142. [Google Scholar] [CrossRef] [Green Version]
- López-Maestresalas, A.; Insausti, K.; Jarén, C.; Pérez-Roncal, C.; Urrutia, O.; Beriain, M.J.; Arazuri, S. Detection of minced lamb and beef fraud using NIR spectroscopy. Food Control 2019, 98, 465–473. [Google Scholar] [CrossRef]
- Ballin, N.Z. Authentication of meat and meat products. Meat Sci. 2010, 86, 577–587. [Google Scholar] [CrossRef]
- Ruslan, A.; Kamarulzaman, N.; Sanny, M. Muslim consumers’ awareness and perception of Halal food fraud. Int. Food Res. J. 2018, 25, S87–S96. [Google Scholar]
- Cheng, J.H.; Nicolai, B.; Sun, D.W. Hyperspectral imaging with multivariate analysis for technological parameters prediction and classification of muscle foods: A review. Meat Sci. 2017, 123, 182–191. [Google Scholar] [CrossRef] [PubMed]
- Sanz, J.A.; Fernandes, A.M.; Barrenechea, E.; Silva, S.; Santos, V.; Gonçalves, N.; Paternain, D.; Jurio, A.; Melo-Pinto, P. Lamb muscle discrimination using hyperspectral imaging: Comparison of various machine learning algorithms. J. Food Eng. 2016, 174, 92–100. [Google Scholar] [CrossRef]
- Xiong, Z.; Sun, D.W.; Zeng, X.A.; Xie, A. Recent developments of hyperspectral imaging systems and their applications in detecting quality attributes of red meats: A review. J. Food Eng. 2014, 132, 1–13. [Google Scholar] [CrossRef]
- ElMasry, G.; Barbin, D.F.; Sun, D.W.; Allen, P. Meat quality evaluation by hyperspectral imaging technique: An overview. Crit. Rev. Food Sci. Nutr. 2012, 52, 689–711. [Google Scholar] [CrossRef]
- Velásquez, L.; Cruz-Tirado, J.; Siche, R.; Quevedo, R. An application based on the decision tree to classify the marbling of beef by hyperspectral imaging. Meat Sci. 2017, 133, 43–50. [Google Scholar] [CrossRef]
- Barbon, S.; Costa Barbon, A.P.A.d.; Mantovani, R.G.; Barbin, D.F. Machine Learning Applied to Near-Infrared Spectra for Chicken Meat Classification. J. Spectrosc. 2018, 2018, 8949741. [Google Scholar] [CrossRef]
- Liu, Y.; Sun, D.W.; Cheng, J.H.; Han, Z. Hyperspectral imaging sensing of changes in moisture content and color of beef during microwave heating process. Food Anal. Methods 2018, 11, 2472–2484. [Google Scholar] [CrossRef]
- Liu, D.; Ma, J.; Sun, D.W.; Pu, H.; Gao, W.; Qu, J.; Zeng, X.A. Prediction of color and pH of salted porcine meats using visible and near-infrared hyperspectral imaging. Food Bioprocess Technol. 2014, 7, 3100–3108. [Google Scholar] [CrossRef]
- Xu, J.L.; Gowen, A.A. Spatial-spectral analysis method using texture features combined with PCA for information extraction in hyperspectral images. J. Chemom. 2020, 34, e3132. [Google Scholar] [CrossRef]
- Ayaz, H.; Ahmad, M.; Sohaib, A.; Yasir, M.N.; Zaidan, M.A.; Ali, M.; Khan, M.H.; Saleem, Z. Myoglobin-Based Classification of Minced Meat Using Hyperspectral Imaging. Appl. Sci. 2020, 10, 6862. [Google Scholar] [CrossRef]
- Xiong, Z.; Sun, D.W.; Pu, H.; Zhu, Z.; Luo, M. Combination of spectra and texture data of hyperspectral imaging for differentiating between free-range and broiler chicken meats. LWT-Food Sci. Technol. 2015, 60, 649–655. [Google Scholar] [CrossRef]
- Li, Y.; Zhang, H.; Shen, Q. Spectral–spatial classification of hyperspectral imagery with 3D convolutional neural network. Remote Sens. 2017, 9, 67. [Google Scholar] [CrossRef] [Green Version]
- Weng, S.; Guo, B.; Tang, P.; Yin, X.; Pan, F.; Zhao, J.; Huang, L.; Zhang, D. Rapid detection of adulteration of minced beef using Vis/NIR reflectance spectroscopy with multivariate methods. Spectrochim. Acta Part A Mol. Biomol. Spectrosc. 2020, 230, 118005. [Google Scholar] [CrossRef] [PubMed]
- Ahmad, M. A Fast 3D CNN for Hyperspectral Image Classification. arXiv 2020, arXiv:2004.14152. [Google Scholar]
- Al-Sarayreh, M.; M Reis, M.; Qi Yan, W.; Klette, R. Detection of red-meat adulteration by deep spectral–spatial features in hyperspectral images. J. Imaging 2018, 4, 63. [Google Scholar] [CrossRef] [Green Version]
- Farrand, W.H.; Singer, R.B.; Merényi, E. Retrieval of apparent surface reflectance from AVIRIS data: A comparison of empirical line, radiative transfer, and spectral mixture methods. Remote Sens. Environ. 1994, 47, 311–321. [Google Scholar] [CrossRef]
- Su, H.; Du, Q.; Du, P. Hyperspectral image visualization using band selection. IEEE J. Sel. Top. Appl. Earth Obs. Remote Sens. 2013, 7, 2647–2658. [Google Scholar] [CrossRef]
- Schafer, R.W. What is a Savitzky-Golay filter. IEEE Signal Process. Mag. 2011, 28, 111–117. [Google Scholar] [CrossRef]
- Cozzolino, D.; Murray, I. Identification of animal meat muscles by visible and near infrared reflectance spectroscopy. LWT-Food Sci. Technol. 2004, 37, 447–452. [Google Scholar] [CrossRef]
- American Meat Science Association. AMSA Meat Color Measurement Guidelines: AMSA; American Meat Science Association: Savoy, IL, USA, 2012. [Google Scholar]
- Huang, H.; Liu, L.; Ngadi, M.O. Recent developments in hyperspectral imaging for assessment of food quality and safety. Sensors 2014, 14, 7248–7276. [Google Scholar] [CrossRef] [Green Version]
- Akilan, T.; Wu, Q.J.; Zhang, H. Effect of fusing features from multiple DCNN architectures in image classification. IET Image Process. 2018, 12, 1102–1110. [Google Scholar] [CrossRef]
- Liang, H.; Li, Q. Hyperspectral imagery classification using sparse representations of convolutional neural network features. Remote Sens. 2016, 8, 99. [Google Scholar] [CrossRef] [Green Version]
- Carneiro, T.; Da Nóbrega, R.V.M.; Nepomuceno, T.; Bian, G.B.; De Albuquerque, V.H.C.; Reboucas Filho, P.P. Performance analysis of google colaboratory as a tool for accelerating deep learning applications. IEEE Access 2018, 6, 61677–61685. [Google Scholar] [CrossRef]
- Srivastava, N.; Hinton, G.; Krizhevsky, A.; Sutskever, I.; Salakhutdinov, R. Dropout: A simple way to prevent neural networks from overfitting. J. Mach. Learn. Res. 2014, 15, 1929–1958. [Google Scholar]
- Kingma, D.P.; Ba, J. Adam: A method for stochastic optimization. arXiv 2014, arXiv:1412.6980. [Google Scholar]




| Layers | Kernels | Filters | Activation | Output Size | Params |
|---|---|---|---|---|---|
| Input layer | — | — | — | (None, 60, 50, 44) | 0 |
| conv3d (Conv3D) | 8 | relu | (None, 58, 48, 42) | 224 | |
| conv3d (Conv3D) | 8 | relu | (None, 56, 46, 40) | 1736 | |
| max pooling | — | — | — | (None, 28, 23, 20) | 0 |
| flatten (Flatten) | — | — | — | (None, 51520) | 0 |
| dense (Dense) | — | — | relu | (None, 256) | 13,189,376 |
| dropout (0.4) | — | — | — | (None, 256) | 0 |
| dense (Dense) | — | — | — | (None, 3) | 771 |
| Total Trainable Parameters = | |||||
| Classifier | Beef | Chicken | Mutton |
|---|---|---|---|
| Precision | |||
| KNN | 0.67 | 0.99 | 0.99 |
| SVM | 0.71 | 0.99 | 0.75 |
| DL | 0.86 | 0.99 | 0.99 |
| Recall | |||
| KNN | 0.99 | 0.99 | 0.40 |
| SVM | 0.83 | 0.99 | 0.60 |
| DL | 0.99 | 0.99 | 0.80 |
| score | |||
| KNN | 0.80 | 0.99 | 0.57 |
| SVM | 0.77 | 0.99 | 0.67 |
| DL | 0.92 | 0.99 | 0.89 |
| Classifier | Beef | Chicken | Mutton | OA |
|---|---|---|---|---|
| KNN-Isos | 0.99 | 0.99 | 0.40 | 0.82 |
| KNN-PCA | 0.80 | 0.99 | 0.40 | 0.75 |
| KNN-224 | 0.83 | 0.99 | 0.40 | 0.75 |
| SVM-Isos | 0.83 | 0.99 | 0.60 | 0.82 |
| SVM-PCA | 0.40 | 0.99 | 0.40 | 0.62 |
| SVM-224 | 0.83 | 0.99 | 0.40 | 0.75 |
| DL-Isos | 0.99 | 0.99 | 0.80 | 0.94 |
| DL-PCA | 0.99 | 0.99 | 0.40 | 0.81 |
| DL-224 | 0.50 | 0.99 | 0.40 | 0.64 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ayaz, H.; Ahmad, M.; Mazzara, M.; Sohaib, A. Hyperspectral Imaging for Minced Meat Classification Using Nonlinear Deep Features. Appl. Sci. 2020, 10, 7783. https://doi.org/10.3390/app10217783
Ayaz H, Ahmad M, Mazzara M, Sohaib A. Hyperspectral Imaging for Minced Meat Classification Using Nonlinear Deep Features. Applied Sciences. 2020; 10(21):7783. https://doi.org/10.3390/app10217783
Chicago/Turabian StyleAyaz, Hamail, Muhammad Ahmad, Manuel Mazzara, and Ahmed Sohaib. 2020. "Hyperspectral Imaging for Minced Meat Classification Using Nonlinear Deep Features" Applied Sciences 10, no. 21: 7783. https://doi.org/10.3390/app10217783
APA StyleAyaz, H., Ahmad, M., Mazzara, M., & Sohaib, A. (2020). Hyperspectral Imaging for Minced Meat Classification Using Nonlinear Deep Features. Applied Sciences, 10(21), 7783. https://doi.org/10.3390/app10217783







