Healthcare Applications of Artificial Intelligence and Analytics: A Review and Proposed Framework
Abstract
1. Introduction
2. Methodology
- Found: articles matching our search strategy of AIA applications.
- Relevant: articles relevant as it presents one of the AIA applications we focused on.
- Cited: article cited in our review as it made significant advances complementary to others.
3. AI Applications
3.1. Data Mining
3.2. Ontologies and Semantic Reasoning
3.3. Clinical Decision Support Systems
- What factors are associated with decision-making complexity experienced by ID experts?
- What cognitive strategies do ID experts use to deal with complexity?
- overall clinical picture does not match the pattern,
- lack of comprehension of the situation, and
- social and emotional pressures.
3.4. Smart Homes
3.5. Medical Big Data
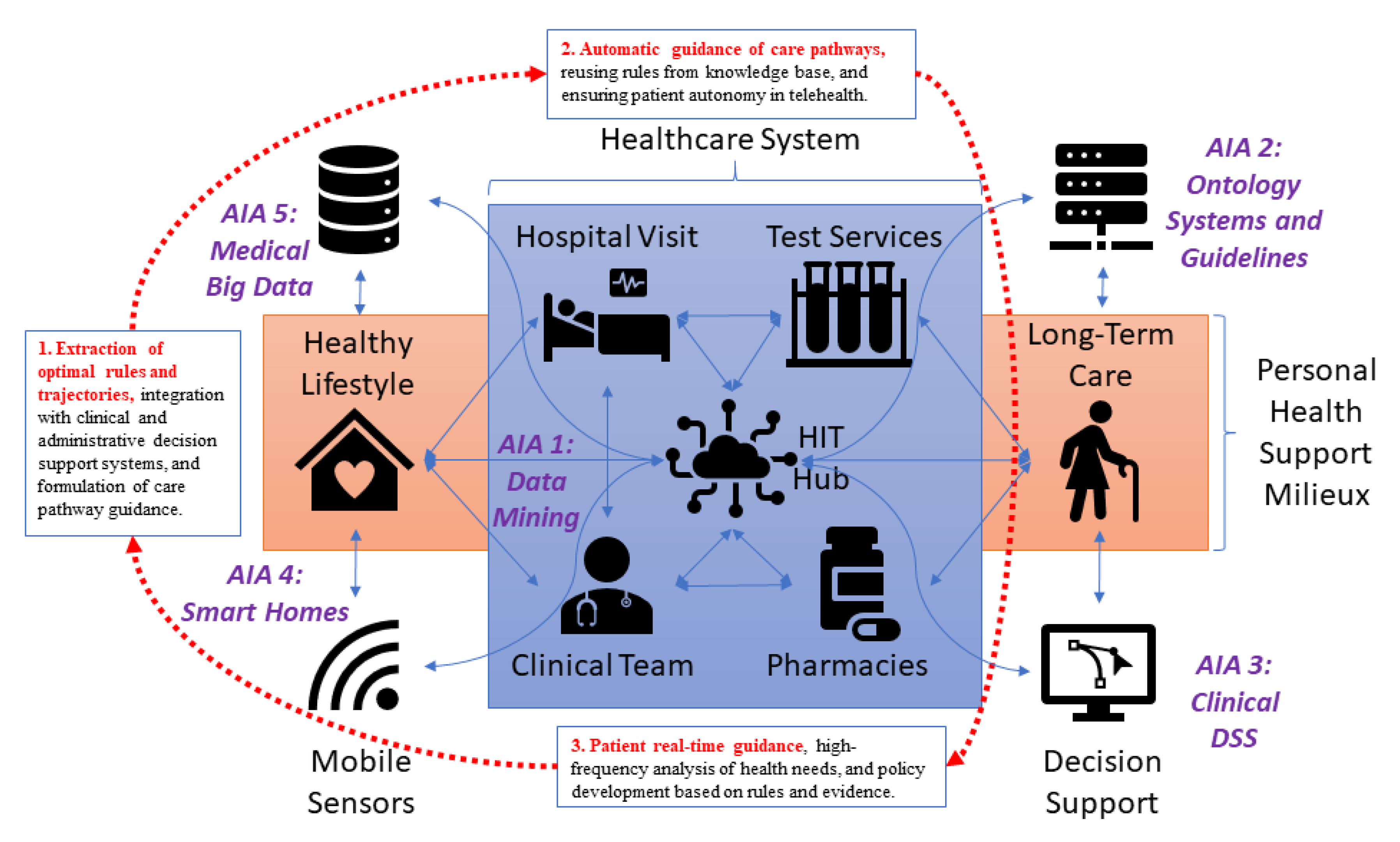
4. Discussion and Framework
- Extraction of optimal rules and trajectories, integration with clinical and administrative decision support systems, and formulation of care pathway guidance (AIA 1-4-5).
- Automatic guidance of care pathways, reusing rules from knowledge base, and ensuring patient autonomy in telehealth (AIA 1-2-3).
- Patient real-time guidance, high-frequency analysis of health needs, and policy development based on rules and evidence (AIA 2-3-4).
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Turing, A.M. Computing machinery and intelligence. In Parsing the Turing Test; Springer: Berlin/Heidelberg, Germany, 2009; pp. 23–65. [Google Scholar]
- Buchanan, B.G. A (very) brief history of artificial intelligence. Ai Mag. 2005, 26, 53. [Google Scholar]
- Moher, D.; Liberati, A.; Tetzlaff, J.; Altman, D.G.; The, P.G. Preferred Reporting Items for Systematic Reviews and Meta-Analyses: The PRISMA Statement. PLoS Med. 2009, 6, e1000097. [Google Scholar] [CrossRef] [PubMed]
- Lashari, S.A.; Ibrahim, R.; Senan, N.; Taujuddin, N. Application of data mining techniques for medical data classification: A review. In Proceedings of the MATEC Web of Conferences, Warsaw, Poland, 5–7 October 2018; p. 06003. [Google Scholar]
- Lakshmi, B.; Indumathi, T.; Ravi, N. A Study on C. 5 decision tree classification algorithm for risk predictions during pregnancy. Procedia Technol. 2016, 24, 1542–1549. [Google Scholar] [CrossRef]
- Mirza, S.; Mittal, S.; Zaman, M. Decision Support Predictive model for prognosis of diabetes using SMOTE and Decision tree. Int. J. Appl. Eng. Res. 2018, 13, 9277–9282. [Google Scholar]
- Xue, D.; Frisch, A.; He, D. Differential Diagnosis of Heart Disease in Emergency Departments Using Decision Tree and Medical Knowledge. In Heterogeneous Data Management, Polystores, and Analytics for Healthcare; Springer: Berlin/Heidelberg, Germany, 2019; pp. 225–236. [Google Scholar]
- Chaurasia, V.; Pal, S.; Tiwari, B. Chronic kidney disease: A predictive model using decision tree. Int. J. Eng. Res. Technol. 2018, 11, 1781–1794. [Google Scholar]
- AlZu’bi, S.; Shehab, M.; Al-Ayyoub, M.; Jararweh, Y.; Gupta, B. Parallel implementation for 3d medical volume fuzzy segmentation. Pattern Recognit. Lett. 2020, 130, 312–318. [Google Scholar] [CrossRef]
- Abdulaziz Mohsen, A.; Alsurori, M.; Aldobai, B.; Mohsen, G.A. New approach to medical diagnosis using artificial neural network and decision tree algorithm: Application to dental diseases. Int. J. Inf. Eng. Electron. Bus. 2019, 11, 52. [Google Scholar] [CrossRef]
- Le, T.-L.; Huynh, T.-T.; Lin, L.-Y.; Lin, C.-M.; Chao, F. A K-means Interval Type-2 Fuzzy Neural Network for Medical Diagnosis. Int. J. Fuzzy Syst. 2019, 21, 2258–2269. [Google Scholar] [CrossRef]
- Shinde, A.; Kale, S.; Samant, R.; Naik, A.; Ghorpade, S. Heart Disease Prediction System using Multilayered Feed Forward Neural Network and Back Propagation Neural Network. Int. J. Comput. Appl. 2017, 166, 32–36. [Google Scholar]
- Karayılan, T.; Kılıç, Ö. Prediction of heart disease using neural network. In Proceedings of the 2017 International Conference on Computer Science and Engineering (UBMK), Antalya, Turkey, 5–7 October 2017; pp. 719–723. [Google Scholar]
- Sundaram, N.M. An Improved Elman Neural Network Classifier for classification of Medical Data for Diagnosis of Diabetes. Int. J. Eng. Sci. 2018, 16317, 16321. [Google Scholar]
- Rajalingam, B.; Priya, R. Multimodal medical image fusion based on deep learning neural network for clinical treatment analysis. Int. J. Chemtech Res. 2018, 11, 160–176. [Google Scholar]
- Rojas-Domínguez, A.; Padierna, L.C.; Valadez, J.M.C.; Puga-Soberanes, H.J.; Fraire, H.J. Optimal hyper-parameter tuning of SVM classifiers with application to medical diagnosis. IEEE Access 2017, 6, 7164–7176. [Google Scholar] [CrossRef]
- Zhu, H.; Liu, X.; Lu, R.; Li, H. Efficient and privacy-preserving online medical prediagnosis framework using nonlinear SVM. IEEE J. Biomed. Health Inform. 2016, 21, 838–850. [Google Scholar] [CrossRef] [PubMed]
- Zeng, N.; Qiu, H.; Wang, Z.; Liu, W.; Zhang, H.; Li, Y. A new switching-delayed-PSO-based optimized SVM algorithm for diagnosis of Alzheimer’s disease. Neurocomputing 2018, 320, 195–202. [Google Scholar] [CrossRef]
- Vogado, L.H.; Veras, R.M.; Araujo, F.H.; Silva, R.R.; Aires, K.R. Leukemia diagnosis in blood slides using transfer learning in CNNs and SVM for classification. Eng. Appl. Artif. Intell. 2018, 72, 415–422. [Google Scholar] [CrossRef]
- Khanmohammadi, S.; Adibeig, N.; Shanehbandy, S. An improved overlapping k-means clustering method for medical applications. Expert Syst. Appl. 2017, 67, 12–18. [Google Scholar] [CrossRef]
- Shakeel, P.M.; Baskar, S.; Dhulipala, V.S.; Jaber, M.M. Cloud based framework for diagnosis of diabetes mellitus using K-means clustering. Health Inf. Sci. Syst. 2018, 6, 16. [Google Scholar] [CrossRef]
- Gürüler, H. A novel diagnosis system for Parkinson’s disease using complex-valued artificial neural network with k-means clustering feature weighting method. Neural Comput. Appl. 2017, 28, 1657–1666. [Google Scholar] [CrossRef]
- Wu, M.; Krishna, S.; Thornhill, R.E.; Flood, T.A.; McInnes, M.D.; Schieda, N. Transition zone prostate cancer: Logistic regression and machine-learning models of quantitative ADC, shape and texture features are highly accurate for diagnosis. J. Magn. Reson. Imaging 2019, 50, 940–950. [Google Scholar] [CrossRef]
- Enriko, I.K.A.; Suryanegara, M.; Gunawan, D. Heart Disease Diagnosis System with k-Nearest Neighbors Method Using Real Clinical Medical Records. In Proceedings of the 4th International Conference on Frontiers of Educational Technologies, Moscow, Russia, 25–27 June 2018; pp. 127–131. [Google Scholar]
- Khanna, D.; Sharma, A. Kernel-Based Naive Bayes Classifier for Medical Predictions. In Intelligent Engineering Informatics; Springer: Berlin/Heidelberg, Germany, 2018; pp. 91–101. [Google Scholar]
- Shen, Y.; Li, Y.; Zheng, H.-T.; Tang, B.; Yang, M. Enhancing ontology-driven diagnostic reasoning with a symptom-dependency-aware Naïve Bayes classifier. BMC Bioinform. 2019, 20, 330. [Google Scholar] [CrossRef]
- Orphanou, K.; Dagliati, A.; Sacchi, L.; Stassopoulou, A.; Keravnou, E.; Bellazzi, R. Combining naive bayes classifiers with temporal association rules for coronary heart disease diagnosis. In Proceedings of the 2016 IEEE International Conference on Healthcare Informatics (ICHI), Chicago, IL, USA, 4–7 October 2016; pp. 81–92. [Google Scholar]
- Pourhomayoun, M.; Shakibi, M. Predicting mortality risk in patients with COVID-19 using artificial intelligence to help medical decision-making. medRxiv 2020. [Google Scholar] [CrossRef]
- Wang, L.; Li, J.; Guo, S.; Xie, N.; Yao, L.; Cao, Y.; Day, S.W.; Howard, S.C.; Graff, J.C.; Gu, T. Real-time estimation and prediction of mortality caused by COVID-19 with patient information based algorithm. Sci. Total Environ. 2020, 727, 138394. [Google Scholar] [CrossRef] [PubMed]
- Narin, A.; Kaya, C.; Pamuk, Z. Automatic detection of coronavirus disease (covid-19) using X-ray images and deep convolutional neural networks. arXiv 2020, arXiv:2003.10849. [Google Scholar]
- Gruber, T.R. A translation approach to portable ontology specifications. Knowl. Acquis. 1993, 5, 199–220. [Google Scholar] [CrossRef]
- Rubin, D.L.; Lewis, S.E.; Mungall, C.J.; Misra, S.; Westerfield, M.; Ashburner, M.; Sim, I.; Chute, C.G.; Storey, M.-A.; Smith, B. National center for biomedical ontology: Advancing biomedicine through structured organization of scientific knowledge. Omics A J. Integr. Biol. 2006, 10, 185–198. [Google Scholar] [CrossRef]
- Babcock, S.; Cowell, L.G.; Beverley, J.; Smith, B. The Infectious Disease Ontology in the Age of COVID-19; OSF Preprints; Center for Open Science: Charlottesville, VA, USA, 2020. [Google Scholar]
- Hoehndorf, R.; Schofield, P.N.; Gkoutos, G.V. The role of ontologies in biological and biomedical research: A functional perspective. Brief. Bioinform. 2015, 16, 1069–1080. [Google Scholar] [CrossRef]
- Chien, H.-Y.; Chen, J.-F.; Chen, Y.-Y.; Lin, P.-S.; Chang, Y.-T.; Chen, R.-C. An Ontology-Based Herb Therapy Recommendation for Respiration System. In Proceedings of the 2017 International Conference on Intelligent Information Hiding and Multimedia Signal Processing, Matsue, Japan, 12–15 August 2017; pp. 74–81. [Google Scholar]
- Asad, S.; Saba, T.; Hussain, S.; Ahmed, M.; Akram, S.; Khan, A.; Anjum, A.; Shah, M.A.; Javaid, N. An ontology-based approach for detecting drug abuse epidemiology. J. Med. Imaging Health Inform. 2017, 7, 1324–1337. [Google Scholar] [CrossRef]
- Fareedi, A.A.; Ghazawneh, A. An Ontology Approach for Knowledge Acquisition and Development of Health Information System (HIS). In Proceedings of the 27th International Conference on Information Systems Development, Lund, Sweden, 21–24 August 2018. [Google Scholar]
- Mortensen, J.M.; Minty, E.P.; Januszyk, M.; Sweeney, T.E.; Rector, A.L.; Noy, N.F.; Musen, M.A. Using the wisdom of the crowds to find critical errors in biomedical ontologies: A study of SNOMED CT. J. Am. Med. Inform. Assoc. 2015, 22, 640–648. [Google Scholar] [CrossRef]
- Lester, R.J. Ground zero: Ontology, recognition, and the elusiveness of care in American eating disorders treatment. Transcult. Psychiatry 2018, 55, 516–533. [Google Scholar] [CrossRef]
- El-Sappagh, S.; Ali, F. DDO: A diabetes mellitus diagnosis ontology. In Applied Informatics; Springer: Berlin/Heidelberg, Germany, 2016; p. 5. [Google Scholar]
- El-Sappagh, S.; Kwak, D.; Ali, F.; Kwak, K.-S. DMTO: A realistic ontology for standard diabetes mellitus treatment. J. Biomed. Semant. 2018, 9, 8. [Google Scholar] [CrossRef]
- Chen, L.; Lu, D.; Zhu, M.; Muzammal, M.; Samuel, O.W.; Huang, G.; Li, W.; Wu, H. OMDP: An ontology-based model for diagnosis and treatment of diabetes patients in remote healthcare systems. Int. J. Distrib. Sens. Netw. 2019, 15, 1550147719847112. [Google Scholar] [CrossRef]
- Messaoudi, R.; Jaziri, F.; Mtibaa, A.; Grand-Brochier, M.; Ali, H.M.; Amouri, A.; Fourati, H.; Chabrot, P.; Gargouri, F.; Vacavant, A. Ontology-based approach for liver cancer diagnosis and treatment. J. Digit. Imaging 2019, 32, 116–130. [Google Scholar] [CrossRef] [PubMed]
- Hu, S.; Zuo, H.; Qi, J.; Hu, Y.; Yu, B. Analysis of effects of Schisandra in the treatment of myocardial infarction based on three-mode gene ontology network. Front. Pharmacol. 2019, 10, 232. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.; Qi, X.; Zhang, J.; Wang, Z. Classical Formula Ontology Construction and Application in the Diagnosis and Treatment of Dermatosis. In Proceedings of the 2018 IEEE/WIC/ACM International Conference on Web Intelligence (WI), Santiago, Chile, 3–6 December 2018; pp. 615–618. [Google Scholar]
- Jie, W.; Yan, P.; Xiaoxiao, R.; Yixuan, Q. An Expert System for Diagnosis and Treatment of Hypertension Based on Ontology. In Proceedings of the International Conference on Bio-Inspired Computing: Theories and Applications, Beijing, China, 2–4 November 2018; pp. 264–274. [Google Scholar]
- Joob, B.; Wiwanitkit, V. Feasibility of using melatonin as a new treatment agent for Ebola virus infection: A gene ontology study. Asian Pac. J. Trop. Biomed. 2016, 2, 179–180. [Google Scholar] [CrossRef]
- Ryerson, C.J.; Walsh, S.L.; Collard, H.R. Reply to Moodley: A Standardized Diagnostic Ontology for Fibrotic Interstitial Lung Disease. Am. J. Respir. Crit. Care Med. 2018, 197, 1366–1367. [Google Scholar] [CrossRef]
- Salem, Y.B.; Idoudi, R.; Ettabaa, K.S.; Hamrouni, K.; Solaiman, B. Ontology based possibilistic reasoning for breast cancer aided diagnosis. In Proceedings of the European, Mediterranean, and Middle Eastern Conference on Information Systems, Coimbra, Potugal, 7–8 September 2017; pp. 353–366. [Google Scholar]
- Ameur, S.T.B.; Cloppet, F.; Wendling, L.; Sellami, D. Ontology based decision system for breast cancer diagnosis. In Proceedings of the Tenth International Conference on Machine Vision (ICMV 2017), Vienna, Austria, 13–15 November 2017; p. 106962. [Google Scholar]
- Azzi, S.; Iglewski, M.; Nabelsi, V. Ontology design for pneumonia diagnostic. In Proceedings of the International Conference on Biomedical and Health Informatics (BHI) 2018 and the IEEE Conference on Body Sensor Networks (BSN), Las Vegas, NV, USA, 4–7 March 2018. [Google Scholar]
- Azzi, S.; Iglewski, M.; Nabelsi, V. Design of a Pneumonia Ontology for Diagnosis Decision Support System. Int. J. Med. Health Sci. 2018. [Google Scholar] [CrossRef]
- Azzi, S.; Iglewski, M.; Nabelsi, V. Competency questions for biomedical ontology reuse. Procedia Comput. Sci. 2019, 160, 362–368. [Google Scholar] [CrossRef]
- Shoaip, N.; Barakat, S.; Elmogy, M. Alzheimer’s Disease Integrated Ontology (ADIO). In Proceedings of the 2019 14th International Conference on Computer Engineering and Systems (ICCES), Cairo, Egypt, 15–16 December 2019; pp. 374–379. [Google Scholar]
- Filice, R.W.; Kahn, C.E. Integrating an ontology of radiology differential diagnosis with ICD-10-cm, RadLex, and SNOMED CT. J. Digit. Imaging 2019, 32, 206–210. [Google Scholar] [CrossRef]
- Gribova, V.; Okun, D.; Petryaeva, M.; Shalfeeva, E.; Tarasov, A. Ontology for Differential Diagnosis of Acute and Chronic Diseases. In Proceedings of the Russian Conference on Artificial Intelligence, Mosco, Russia, 24–27 September 2018; pp. 152–163. [Google Scholar]
- Köhler, S.; Øien, N.C.; Buske, O.J.; Groza, T.; Jacobsen, J.O.; McNamara, C.; Vasilevsky, N.; Carmody, L.C.; Gourdine, J.; Gargano, M. Encoding Clinical Data with the Human Phenotype Ontology for Computational Differential Diagnostics. Curr. Protoc. Hum. Genet. 2019, 103, e92. [Google Scholar] [CrossRef]
- Lin, Y.; Mehta, S.; Küçük-McGinty, H.; Turner, J.P.; Vidovic, D.; Forlin, M.; Koleti, A.; Nguyen, D.-T.; Jensen, L.J.; Guha, R. Drug target ontology to classify and integrate drug discovery data. J. Biomed. Semant. 2017, 8, 50. [Google Scholar] [CrossRef]
- Li, F.; Wang, M.; Pham, H.A.; Xiang, Y.; Tao, C.; Du, J.; Rao, G. Systematic Design of Drug Repurposing-Oriented Alzheimer’s Disease Ontology. In Proceedings of the 2019 IEEE International Conference on Healthcare Informatics (ICHI), Xi’an, China, 10–13 June 2019; pp. 1–5. [Google Scholar]
- Mutowo, P.; Bento, A.P.; Dedman, N.; Gaulton, A.; Hersey, A.; Lomax, J.; Overington, J.P. A drug target slim: Using gene ontology and gene ontology annotations to navigate protein-ligand target space in ChEMBL. J. Biomed. Semant. 2016, 7, 59. [Google Scholar] [CrossRef] [PubMed]
- Fox, J.; Gutenstein, M.; Khan, O.; South, M.; Thomson, R. OpenClinical. net: A platform for creating and sharing knowledge and promoting best practice in healthcare. Comput. Ind. 2015, 66, 63–72. [Google Scholar] [CrossRef]
- Kawamoto, K.; Del Fiol, G. Clinical decision support systems in healthcare. Health Inform.-E-Book 2016, 170, 203. [Google Scholar]
- Buntin, M.B.; Burke, M.F.; Hoaglin, M.C.; Blumenthal, D. The benefits of health information technology: A review of the recent literature shows predominantly positive results. Health Aff. 2011, 30, 464–471. [Google Scholar] [CrossRef] [PubMed]
- Lyman, J.A.; Cohn, W.F.; Bloomrosen, M.; Detmer, D.E. Clinical decision support: Progress and opportunities. J. Am. Med. Inform. Assoc. 2010, 17, 487–492. [Google Scholar] [CrossRef] [PubMed]
- Kawamoto, K.; Lobach, D.F. Proposal for fulfilling strategic objectives of the US roadmap for national action on decision support through a service-oriented architecture leveraging HL7 services. J. Am. Med. Inform. Assoc. 2007, 14, 146–155. [Google Scholar] [CrossRef]
- Chaudhry, B.; Wang, J.; Wu, S.; Maglione, M.; Mojica, W.; Roth, E.; Morton, S.C.; Shekelle, P.G. Systematic review: Impact of health information technology on quality, efficiency, and costs of medical care. Ann. Intern. Med. 2006, 144, 742–752. [Google Scholar] [CrossRef] [PubMed]
- Osheroff, J.A.; Teich, J.M.; Middleton, B.; Steen, E.B.; Wright, A.; Detmer, D.E. A roadmap for national action on clinical decision support. J. Am. Med. Inform. Assoc. 2007, 14, 141–145. [Google Scholar] [CrossRef]
- El-Sappagh, S.; Alonso, J.M.; Ali, F.; Ali, A.; Jang, J.-H.; Kwak, K.-S. An ontology-based interpretable fuzzy decision support system for diabetes diagnosis. IEEE Access 2018, 6, 37371–37394. [Google Scholar] [CrossRef]
- Ali, F.; Islam, S.R.; Kwak, D.; Khan, P.; Ullah, N.; Yoo, S.-j.; Kwak, K.S. Type-2 fuzzy ontology–aided recommendation systems for IoT–based healthcare. Comput. Commun. 2018, 119, 138–155. [Google Scholar] [CrossRef]
- Amland, R.C.; Hahn-Cover, K.E. Clinical decision support for early recognition of sepsis. Am. J. Med. Qual. 2016, 31, 103–110. [Google Scholar] [CrossRef] [PubMed]
- Middleton, B.; Sittig, D.; Wright, A. Clinical decision support: A 25 year retrospective and a 25 year vision. Yearb. Med. Inform. 2016, 25, S103–S116. [Google Scholar]
- Ekstrom, H.L.; Kharbanda, E.O.; Ballard, D.W.; Vinson, D.R.; Vazquez-Benitez, G.; Chettipally, U.K.; Dehmer, S.P.; Kunisetty, G.; Sharma, R.; Rauchwerger, A.S. Development of a Clinical Decision Support System for Pediatric Abdominal Pain in Emergency Department Settings Across Two Health Systems Within the HCSRN. EGEMS 2019, 7, 15. [Google Scholar] [CrossRef] [PubMed]
- Pota, M.; Esposito, M.; De Pietro, G. Fuzzy partitioning for clinical DSSs using statistical information transformed into possibility-based knowledge. Knowl.-Based Syst. 2014, 67, 1–15. [Google Scholar] [CrossRef]
- Jensen, J.D.; Durand, D.J. Partnering with your health system to select and implement clinical decision support for imaging. J. Am. Coll. Radiol. 2017, 14, 262–268. [Google Scholar] [CrossRef]
- Dayan, P.S.; Ballard, D.W.; Tham, E.; Hoffman, J.M.; Swietlik, M.; Deakyne, S.J.; Alessandrini, E.A.; Tzimenatos, L.; Bajaj, L.; Vinson, D.R. Use of traumatic brain injury prediction rules with clinical decision support. Pediatrics 2017, 139, e20162709. [Google Scholar] [CrossRef]
- Islam, R.; Weir, C.R.; Jones, M.; Del Fiol, G.; Samore, M.H. Understanding complex clinical reasoning in infectious diseases for improving clinical decision support design. BMC Med. Inform. Decis. Mak. 2015, 15, 101. [Google Scholar] [CrossRef]
- Denecke, K.; Deng, Y. Sentiment analysis in medical settings: New opportunities and challenges. Artif. Intell. Med. 2015, 64, 17–27. [Google Scholar] [CrossRef]
- Araque, O.; Zhu, G.; Iglesias, C.A. A semantic similarity-based perspective of affect lexicons for sentiment analysis. Knowl.-Based Syst. 2019, 165, 346–359. [Google Scholar] [CrossRef]
- Teng, Z.; Vo, D.-T.; Zhang, Y. Context-sensitive lexicon features for neural sentiment analysis. In Proceedings of the 2016 Conference on Empirical Methods in Natural Language Processing, Austin, TX, USA, 1–5 November 2016; pp. 1629–1638. [Google Scholar]
- Khoo, C.S.; Johnkhan, S.B. Lexicon-based sentiment analysis: Comparative evaluation of six sentiment lexicons. J. Inf. Sci. 2018, 44, 491–511. [Google Scholar] [CrossRef]
- Lohr, S. IBM is counting on its bet on Watson, and paying big money for it. NY Times 2016, 17, 2016. [Google Scholar]
- Katzman, J.L.; Shaham, U.; Cloninger, A.; Bates, J.; Jiang, T.; Kluger, Y. DeepSurv: Personalized treatment recommender system using a Cox proportional hazards deep neural network. BMC Med. Res. Methodol. 2018, 18, 24. [Google Scholar] [CrossRef] [PubMed]
- Ronao, C.A.; Cho, S.-B. Human activity recognition with smartphone sensors using deep learning neural networks. Expert Syst. Appl. 2016, 59, 235–244. [Google Scholar] [CrossRef]
- Ma, M.; Fan, H.; Kitani, K.M. Going deeper into first-person activity recognition. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Las Vegas, NV, USA, 27–30 June 2016; pp. 1894–1903. [Google Scholar]
- Hassan, M.M.; Uddin, M.Z.; Mohamed, A.; Almogren, A. A robust human activity recognition system using smartphone sensors and deep learning. Future Gener. Comput. Syst. 2018, 81, 307–313. [Google Scholar] [CrossRef]
- Ignatov, A. Real-time human activity recognition from accelerometer data using Convolutional Neural Networks. Appl. Soft Comput. 2018, 62, 915–922. [Google Scholar] [CrossRef]
- Cippitelli, E.; Gasparrini, S.; Gambi, E.; Spinsante, S. A human activity recognition system using skeleton data from RGBD sensors. Comput. Intell. Neurosci. 2016, 2016, 4351435. [Google Scholar] [CrossRef]
- Wang, A.; Chen, G.; Shang, C.; Zhang, M.; Liu, L. Human activity recognition in a smart home environment with stacked denoising autoencoders. In Proceedings of the 2016 International Conference on Web-Age Information Management, Nanchang, China, 3–5 June 2016; pp. 29–40. [Google Scholar]
- Cicirelli, F.; Fortino, G.; Giordano, A.; Guerrieri, A.; Spezzano, G.; Vinci, A. On the design of smart homes: A framework for activity recognition in home environment. J. Med. Syst. 2016, 40, 200. [Google Scholar] [CrossRef]
- Mehr, H.D.; Polat, H.; Cetin, A. Resident activity recognition in smart homes by using artificial neural networks. In Proceedings of the 2016 4th International Istanbul Smart Grid Congress and Fair (ICSG), Istanbul, Turkey, 20–21 April 2016; pp. 1–5. [Google Scholar]
- Fan, X.; Xie, Q.; Li, X.; Huang, H.; Wang, J.; Chen, S.; Xie, C.; Chen, J. Activity recognition as a service for smart home: Ambient assisted living application via sensing home. In Proceedings of the 2017 IEEE International Conference on AI & Mobile Services (AIMS), Honolulu, HI, USA, 15–30 June 2017; pp. 54–61. [Google Scholar]
- Gayathri, K.; Easwarakumar, K.; Elias, S. Activity Recognition and Dementia Care in Smart Home. In Digital India; Springer: Berlin/Heidelberg, Germany, 2018; pp. 33–47. [Google Scholar]
- Zou, H.; Zhou, Y.; Yang, J.; Jiang, H.; Xie, L.; Spanos, C.J. WiFi-enabled device-free gesture recognition for smart home automation. In Proceedings of the 2018 IEEE 14th International Conference on Control and Automation (ICCA), Anchorage, AK, USA, 12–15 June 2018; pp. 476–481. [Google Scholar]
- Singh, G.; Nelson, A.; Lu, S.; Robucci, R.; Patel, C.; Banerjee, N. Event-driven low-power gesture recognition using differential capacitance. IEEE Sens. J. 2016, 16, 4955–4967. [Google Scholar] [CrossRef]
- Kaur, M.; Kaur, G.; Sharma, P.K.; Jolfaei, A.; Singh, D. Binary cuckoo search metaheuristic-based supercomputing framework for human behavior analysis in smart home. J. Supercomput. 2020, 76, 2479–2502. [Google Scholar] [CrossRef]
- Pan, Z.; Pacheco, J.; Hariri, S.; Chen, Y.; Liu, B. Context Aware Anomaly Behavior Analysis for Smart Home Systems. Int. J. Inf. Commun. Eng. 2019, 13, 261–274. [Google Scholar]
- Chen, M.; Yang, J.; Zhu, X.; Wang, X.; Liu, M.; Song, J. Smart home 2.0: Innovative smart home system powered by botanical IoT and emotion detection. Mob. Netw. Appl. 2017, 22, 1159–1169. [Google Scholar] [CrossRef]
- Fu, C. Smart Home Control Method Based on Emotion Recognition and the System Thereof. U.S. Patent No. 10,013,977, 3 July 2018. [Google Scholar]
- Demir, E.; Köseoğlu, E.; Sokullu, R.; Şeker, B. Smart home assistant for ambient assisted living of elderly people with dementia. Procedia Comput. Sci. 2017, 113, 609–614. [Google Scholar] [CrossRef]
- Ghayvat, H.; Liu, J.; Mukhopadhyay, S.C.; Gui, X. Wellness sensor networks: A proposal and implementation for smart home for assisted living. IEEE Sens. J. 2015, 15, 7341–7348. [Google Scholar] [CrossRef]
- Stahl, C.; Laub, P. Maintaining multiple sclerosis patients’ quality of life: A case study on environment control assistance in a smart home. In Proceedings of the 10th International Conference on PErvasive Technologies Related to Assistive Environments, Rhodes, Greece, 21–23 June 2017; pp. 83–86. [Google Scholar]
- Aman, F.; Aubergé, V.; Vacher, M. Influence of expressive speech on ASR performances: Application to elderly assistance in smart home. In Proceedings of the International Conference on Text, Speech, and Dialogue, Brno, Czech Republic, 12–16 September 2016; pp. 522–530. [Google Scholar]
- Yassine, A.; Singh, S.; Hossain, M.S.; Muhammad, G. IoT big data analytics for smart homes with fog and cloud computing. Future Gener. Comput. Syst. 2019, 91, 563–573. [Google Scholar] [CrossRef]
- Sukor, A.S.A.; Zakaria, A.; Rahim, N.A.; Kamarudin, L.M.; Setchi, R.; Nishizaki, H. A hybrid approach of knowledge-driven and data-driven reasoning for activity recognition in smart homes. J. Intell. Fuzzy Syst. 2019, 36, 4177–4188. [Google Scholar] [CrossRef]
- Hudec, M.; Smutny, Z. RUDO: A home ambient intelligence system for blind people. Sensors 2017, 17, 1926. [Google Scholar] [CrossRef] [PubMed]
- Warren, D.A.; Marschalkowski, E. Smart-Home Device Telemetry Circuits for Detecting Electrical Anomalies in a Population of Devices. U.S. Patent No. 10,557,640, 11 February 2020. [Google Scholar]
- Boettcher, J.; Sloo, D.H.; Boyd, J.A.; Le Guen, S. Systems and Methods of Providing Status Information in a Smart Home Security Detection System. U.S. Patent No. 10,593,190, 17 March 2020. [Google Scholar]
- Hargreaves, T.; Wilson, C.; Hauxwell-Baldwin, R. Learning to live in a smart home. Build. Res. Inf. 2018, 46, 127–139. [Google Scholar] [CrossRef]
- Manogaran, G.; Thota, C.; Lopez, D.; Vijayakumar, V.; Abbas, K.M.; Sundarsekar, R. Big data knowledge system in healthcare. In Internet of Things and big Data Technologies for Next Generation Healthcare; Springer: Berlin/Heidelberg, Germany, 2017; pp. 133–157. [Google Scholar]
- Simonsen, L.; Gog, J.R.; Olson, D.; Viboud, C. Infectious disease surveillance in the big data era: Towards faster and locally relevant systems. J. Infect. Dis. 2016, 214, S380–S385. [Google Scholar] [CrossRef]
- Benbow, N.; Kirkpatrick, C.; Gupta, A.; Villamar, J.; Chernyshov, K.; Cramer, N.; Mena, L.; Mayer, R.; Young, S. An Iterative Process of Integrating and Developing Big Data Modeling and Visualization Tools in Collaboration With Public Health Officials; SAGE Publications Ltd.: New York, NY, USA, 2020. [Google Scholar]
- Salerno, J.; Knoppers, B.M.; Lee, L.M.; Hlaing, W.M.; Goodman, K.W. Ethics, big data and computing in epidemiology and public health. Ann. Epidemiol. 2017, 27, 297–301. [Google Scholar] [CrossRef]
- Chae, S.; Kwon, S.; Lee, D. Predicting infectious disease using deep learning and big data. Int. J. Environ. Res. Public Health 2018, 15, 1596. [Google Scholar] [CrossRef]
- Choirat, C.; Braun, D.; Kioumourtzoglou, M.-A. Data Science in Environmental Health Research. Curr. Epidemiol. Rep. 2019, 6, 291–299. [Google Scholar] [CrossRef] [PubMed]
- Ehrenstein, V.; Nielsen, H.; Pedersen, A.B.; Johnsen, S.P.; Pedersen, L. Clinical epidemiology in the era of big data: New opportunities, familiar challenges. Clin. Epidemiol. 2017, 9, 245. [Google Scholar] [CrossRef] [PubMed]
- Brody, J.A.; Morrison, A.C.; Bis, J.C.; O’Connell, J.R.; Brown, M.R.; Huffman, J.E.; Ames, D.C.; Carroll, A.; Conomos, M.P.; Gabriel, S. Analysis commons, a team approach to discovery in a big-data environment for genetic epidemiology. Nat. Genet. 2017, 49, 1560–1563. [Google Scholar] [CrossRef] [PubMed]
- Bansal, S.; Chowell, G.; Simonsen, L.; Vespignani, A.; Viboud, C. Big data for infectious disease surveillance and modeling. J. Infect. Dis. 2016, 214, S375–S379. [Google Scholar] [CrossRef]
- Bragazzi, N.L.; Guglielmi, O.; Garbarino, S. SleepOMICS: How big data can revolutionize sleep science. Int. J. Environ. Res. Public Health 2019, 16, 291. [Google Scholar] [CrossRef] [PubMed]
- Boone-Heinonen, J.; Tillotson, C.; O’Malley, J.; Cottrell, E.; Gaudino, J.; Amofah, A.; Rivo, M.; Brickman, A.; Mayer, K.; McBurnie, M. Characterizing a “big data” cohort of over 200,000 low-income US infants and children for obesity research: The Advance Early Life Cohort. Matern. Child Health J. 2017, 21, 421–431. [Google Scholar] [CrossRef]
- Lamminpää, R.; Gissler, M.; Vehviläinen-Julkunen, K. Utilization of large data sets in maternal health in Finland. J. Perinat. Neonatal Nurs. 2017, 31, 236–243. [Google Scholar] [CrossRef]
- Maugard, C.; Bosson-Rieutort, D.; François, O.; Bonneterre, V. 1154 Big Data and Occupational Health Surveillance: Use of French Medico-Administrative Databases for Hypothesis Generation Regarding Occupational Risks in Agriculture; BMJ Publishing Group Ltd.: London, UK, 2018. [Google Scholar]
- Maugard, C.; Rieutort, D.B.; Ozenfant, D.; François, O.; Bonneterre, V. Big-data and occupational health surveillance: Screening of occupational determinants of health among French agricultural workers, through data mining of medico-administrative databases. Rev. D’épidémiologie Et De St. Publique 2018, 66, S262–S263. [Google Scholar] [CrossRef]
- Phillips, C.A.; Pollock, B.H. Big Data for Nutrition Research in Pediatric Oncology: Current State and Framework for Advancement. Jnci Monogr. 2019, 2019, 127–131. [Google Scholar] [CrossRef]
- McDonald, D.; Glusman, G.; Price, N.D. Personalized nutrition through big data. Nat. Biotechnol. 2016, 34, 152. [Google Scholar] [CrossRef]
- Paul, P.; Aithal, P.; Bhuimali, A. Enhancing Cloud and Big Data Systems for healthy Food and Nutrition Information Systems Practice: A Conceptual Study. Int. J. Sci. Res. Biol. Sci. 2017, 4, 5. [Google Scholar] [CrossRef]
- Al-Ali, A.-R.; Zualkernan, I.A.; Rashid, M.; Gupta, R.; AliKarar, M. A smart home energy management system using IoT and big data analytics approach. IEEE Trans. Consum. Electron. 2017, 63, 426–434. [Google Scholar] [CrossRef]
- Yassine, A.; Singh, S.; Alamri, A. Mining human activity patterns from smart home big data for health care applications. IEEE Access 2017, 5, 13131–13141. [Google Scholar] [CrossRef]
- Kanevsky, J.; Corban, J.; Gaster, R.; Kanevsky, A.; Lin, S.; Gilardino, M. Big data and machine learning in plastic surgery: A new frontier in surgical innovation. Plast. Reconstr. Surg. 2016, 137, 890e–897e. [Google Scholar] [CrossRef] [PubMed]
- Lee, C.H.; Yoon, H.-J. Medical big data: Promise and challenges. Kidney Res. Clin. Pract. 2017, 36, 3. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.; Groeneveld, P.W. Big data, health informatics, and the future of cardiovascular medicine. J. Am. Coll. Cardiol. 2017, 69, 899–902. [Google Scholar] [CrossRef]
- Sanaei, A.; Sepehri, M.M. Quality Improvement through Intensive Care Registries: The Value of Big Data in Clinical Decision Making. Arch. Anesth. Crit. Care 2020. [Google Scholar] [CrossRef]
- Van den Heuvel, L.; Dorsey, R.R.; Prainsack, B.; Post, B.; Stiggelbout, A.M.; Meinders, M.J.; Bloem, B.R. Quadruple Decision Making for Parkinson’s Disease Patients: Combining Expert Opinion, Patient Preferences, Scientific Evidence, and Big Data Approaches to Reach Precision Medicine. J. Parkinson’s Dis. 2020, 10, 223–231. [Google Scholar] [CrossRef]
- Duan, Y.; Edwards, J.S.; Dwivedi, Y.K. Artificial intelligence for decision making in the era of Big Data–evolution, challenges and research agenda. Int. J. Inf. Manag. 2019, 48, 63–71. [Google Scholar] [CrossRef]
- Roosan, D.; Samore, M.; Jones, M.; Livnat, Y.; Clutter, J. Big-data based decision-support systems to improve clinicians’ cognition. In Proceedings of the 2016 IEEE International Conference on Healthcare Informatics (ICHI), Chicago, IL, USA, 4–7 October 2016; pp. 285–288. [Google Scholar]
- Saunders, G.H.; Christensen, J.H.; Gutenberg, J.; Pontoppidan, N.H.; Smith, A.; Spanoudakis, G.; Bamiou, D.-E. Application of Big Data to Support Evidence-Based Public Health Policy Decision-Making for Hearing. Ear Hear. 2020. [Google Scholar] [CrossRef]
- Kumar, Y.; Sood, K.; Kaul, S.; Vasuja, R. Big Data Analytics and Its Benefits in Healthcare. In Big Data Analytics in Healthcare; Springer: Berlin/Heidelberg, Germany, 2020; pp. 3–21. [Google Scholar]
- Price, W.N.; Cohen, I.G. Privacy in the age of medical big data. Nat. Med. 2019, 25, 37–43. [Google Scholar] [CrossRef] [PubMed]
- Kallianos, K.; Mongan, J.; Antani, S.; Henry, T.; Taylor, A.; Abuya, J.; Kohli, M. How far have we come? Artificial intelligence for chest radiograph interpretation. Clin. Radiol. 2019, 74, 338–345. [Google Scholar] [CrossRef] [PubMed]
- Barr, V.; Robinson, S.; Marin-Link, B.; Underhill, L.; Dotts, A.; Ravensdale, D.; Salivaras, S. The expanded chronic care model. Hosp. Q. 2003, 7, 73–82. [Google Scholar] [PubMed]
- Wagner, E.H.; Austin, B.T.; Davis, C.; Hindmarsh, M.; Schaefer, J.; Bonomi, A. Improving Chronic Illness Care: Translating Evidence Into Action. Health Aff. 2001, 20, 64–78. [Google Scholar] [CrossRef]
| Filtering | Search Query |
|---|---|
| Data mining | (“machine learning”[tiab] OR “data mining”[tiab] OR “text mining”[tiab] OR “mining”[tiab] OR “decision tree”[tiab] OR “neural network”[tiab] OR “SVM”[tiab] OR bayes*[tiab] OR “KNN”[tiab] OR “K-means”[tiab]) AND (med*[tiab] OR clinic*[tiab] OR health*[tiab]) |
| Ontologies and semantic reasoning | Ontology AND (med*[tiab] OR clinic*[tiab] OR health*[tiab] OR emergenc*[tiab] OR physician*[tiab] OR psycholog*[tiab] OR pain [tiab] OR therapy [tiab] OR treat*[tiab] OR drug*[tiab] OR disease*[tiab] OR diagnos*[tiab] OR emotion*[tiab]) OR (computerized clinical guideline*[tiab] OR computerized clinical practice guideline*[tiab])) |
| Clinical decision support systems | (monitor*[tiab] OR assist*[tiab] OR recommend*[tiab] OR “decision support”[tiab] OR “decision making”[tiab] OR “computer assisted”[tiab] OR “reminder system”[tiab] OR “support system”[tiab] OR expert system*[tiab]) AND (med*[tiab] OR clinic*[tiab] OR health*[tiab] OR emergenc*[tiab] OR physician*[tiab] OR psycholog*[tiab] OR pain [tiab] OR therapy [tiab] OR treat*[tiab] OR drug*[tiab] OR disease*[tiab] OR diagnos*[tiab] OR emotion*[tiab]) |
| Smart homes | (smart home*[tiab]) AND (activity recognition*[tiab] “expression recognition”[tiab] OR assist*[tiab] OR recogni*[tiab] OR detect*[tiab] OR guid*[tiab] OR algorithm*[tiab] OR “security”[tiab] OR web service*[tiab]OR “system”[tiab] OR “application”[tiab] OR “technology”[tiab] OR “engine”[tiab] OR “framework”[tiab] OR “behaviour analysis”[tiab]) |
| Medical big data | “Big data” [tiab] AND (med*[tiab] OR clinic*[tiab] OR health*[tiab]) |
| AI Applications | Found | Relevant | Cited |
|---|---|---|---|
| Data mining | 5987 | 1052 | 29 |
| Ontologies and semantic reasoning and clinical guidelines | 2912 | 675 | 34 |
| Clinical decision support systems | 3970 | 2198 | 21 |
| Smart homes | 6444 | 2932 | 30 |
| Medical big data | 5469 | 689 | 23 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Azzi, S.; Gagnon, S.; Ramirez, A.; Richards, G. Healthcare Applications of Artificial Intelligence and Analytics: A Review and Proposed Framework. Appl. Sci. 2020, 10, 6553. https://doi.org/10.3390/app10186553
Azzi S, Gagnon S, Ramirez A, Richards G. Healthcare Applications of Artificial Intelligence and Analytics: A Review and Proposed Framework. Applied Sciences. 2020; 10(18):6553. https://doi.org/10.3390/app10186553
Chicago/Turabian StyleAzzi, Sabrina, Stéphane Gagnon, Alex Ramirez, and Gregory Richards. 2020. "Healthcare Applications of Artificial Intelligence and Analytics: A Review and Proposed Framework" Applied Sciences 10, no. 18: 6553. https://doi.org/10.3390/app10186553
APA StyleAzzi, S., Gagnon, S., Ramirez, A., & Richards, G. (2020). Healthcare Applications of Artificial Intelligence and Analytics: A Review and Proposed Framework. Applied Sciences, 10(18), 6553. https://doi.org/10.3390/app10186553








