Machine Learning Applied to Diagnosis of Human Diseases: A Systematic Review
Abstract
1. Introduction
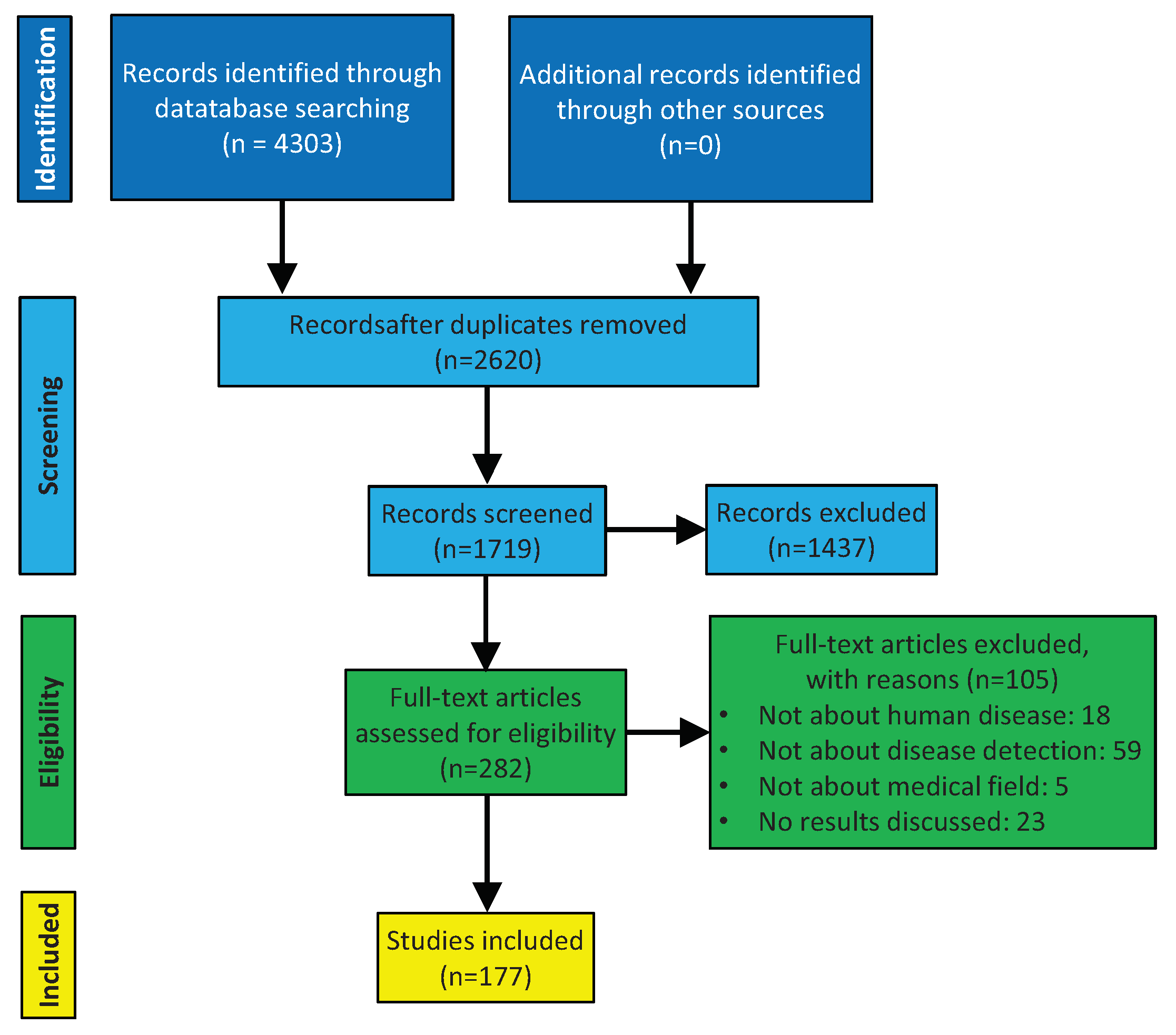
2. Methods
2.1. Data Sources
2.2. Data Extraction
2.3. Data Analyses
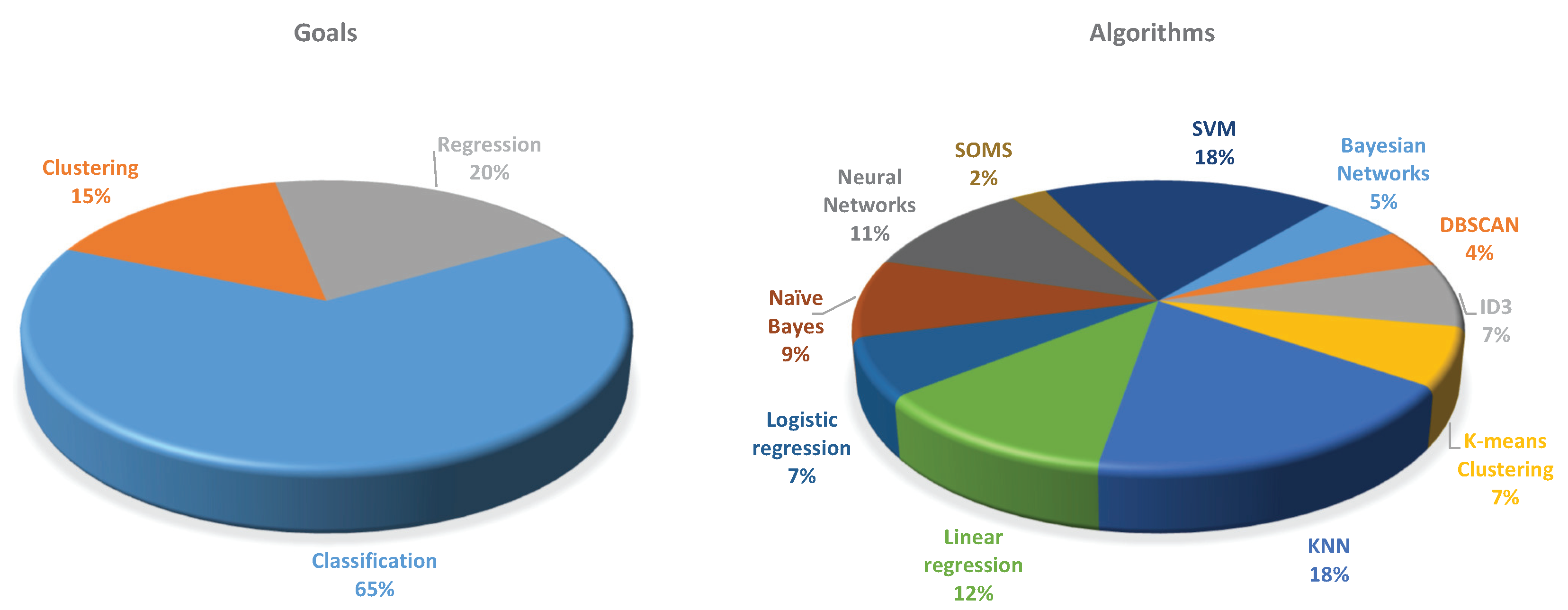
3. Results
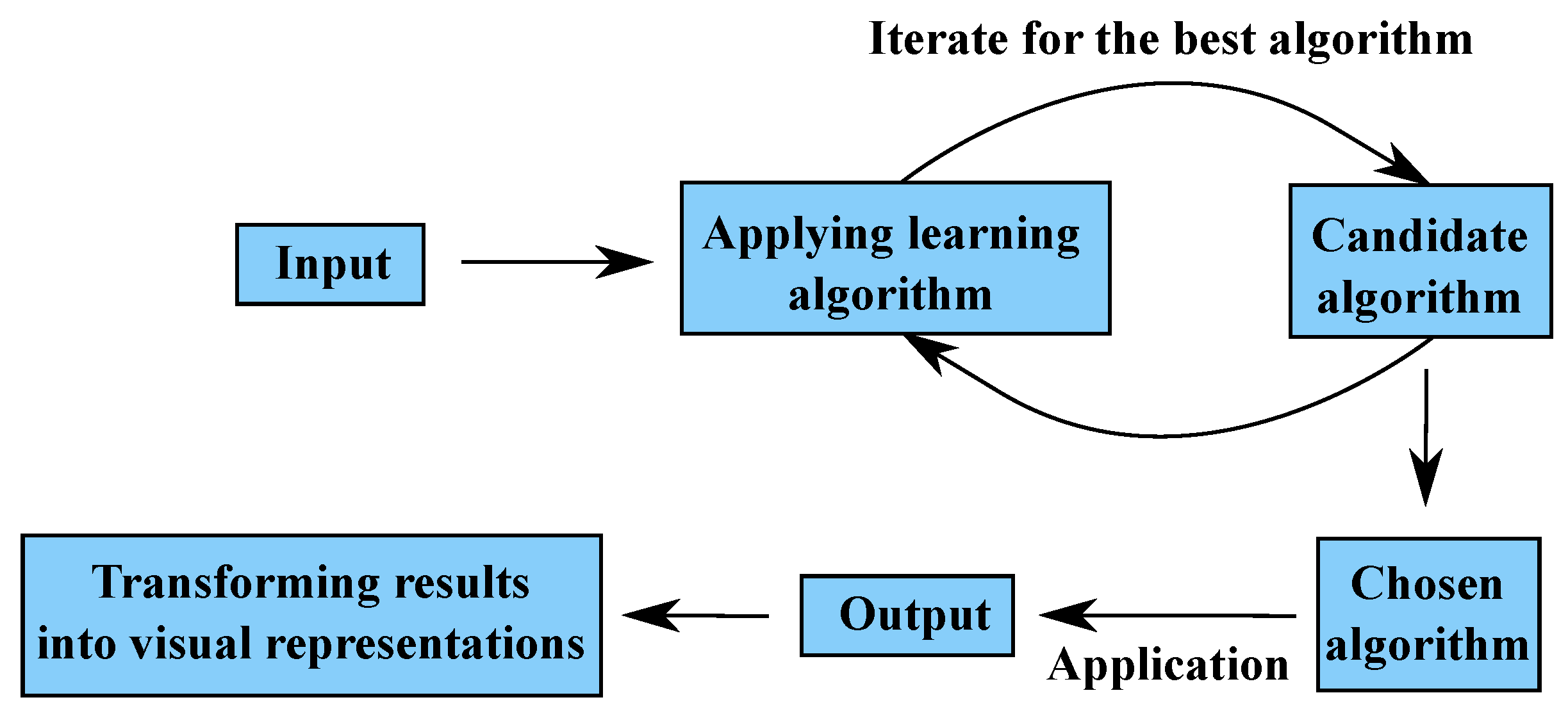
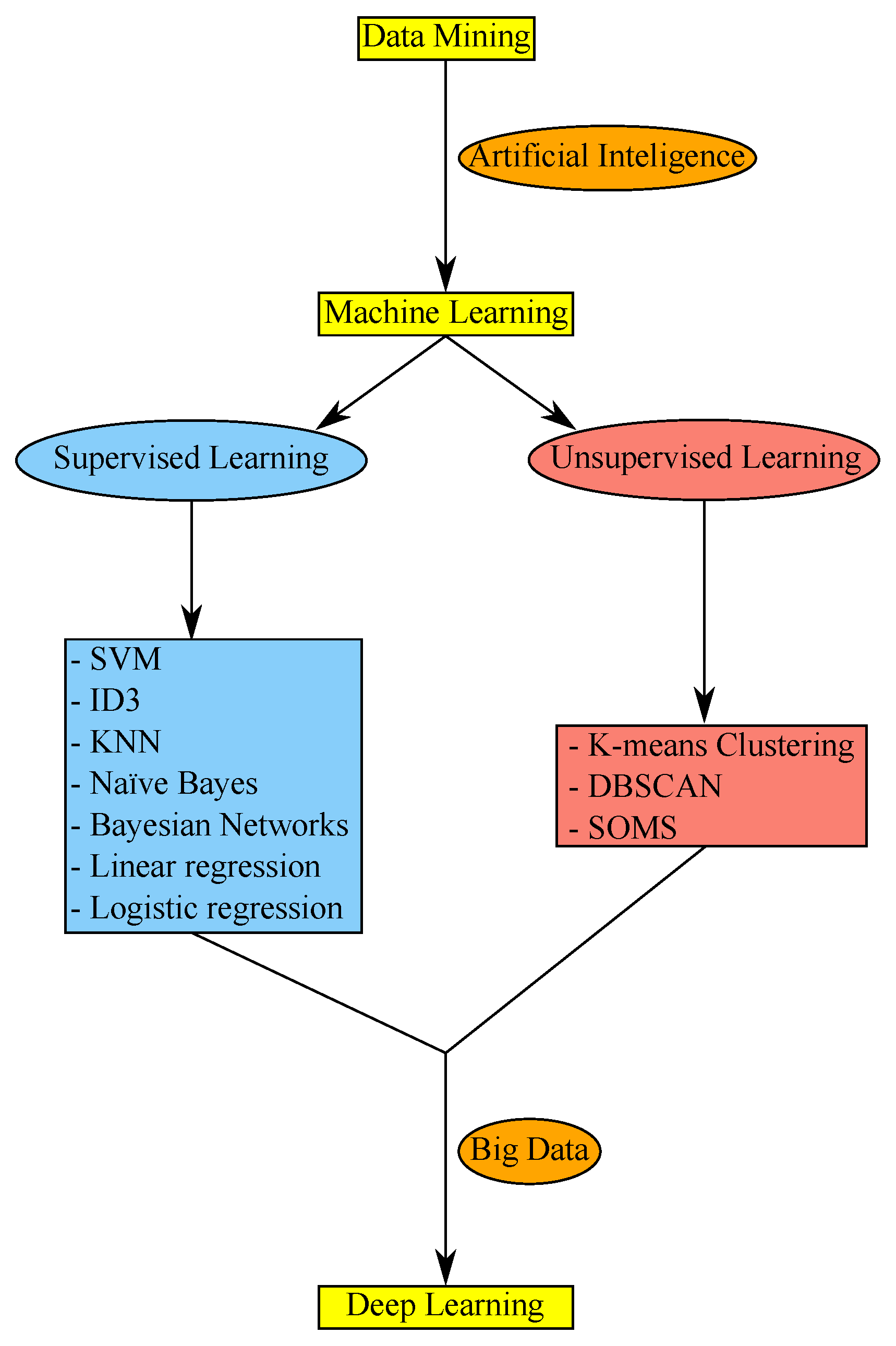
3.1. Machine Learning Principles
3.1.1. Input. Definition and Methods
3.1.2. Output. Learning Representation
- Decision tables are one type of information tables with a decision attribute giving the decision classes for all objects [38]. Table 1 shows a simple example of a decision table, where the last column represents the final decision. In decision tables, the way of inferring the output is to make the same as the input. Because of this, decision tables are one of the simplest way of representing the output from a ML classification model.
- Decision trees (DT) are predictive representations that can be used both for classification and regression models. Decision trees are a hierarchical way of partitioning the space, where the goal is to create a model that predicts the value of a target variable based on several input variables. A DT learns by splitting the source set into subsets that are based on an attribute value test. This process is repeated on each derived subset in a recursive way, called recursive partitioning. When a DT is used for classification tasks, it is more appropriately referred to as a classification tree. On the other hand, when it is used for regression tasks, it is called regression tree [39]. Breiman et al. [40] provided a simple example of a classification tree with medical data. In this example, it was predicted that the high or low risk of patients did not survive at least 30 days based on the initial 24 h. Serrano et al. [41] proposed an analysis through a regression tree for studying the successful weight loss from a commercial health app user for three distinct subgroups: the occasional users, the basic users, and the power users of the app. In both examples, a new sample can be inferred based on the corresponding representation.
- Regression lines are the most common representations for linear regression. The regression line is that one which is the best suited to the data point cloud. Yoon et al. [42] analysed the influence of the pulse pressure on the systolic blood pressure. A new sample can be inferred throughout the regression line known as the value of the pulse pressure.
- Hyper-plane diagrams are a specific type of representations of SVM algorithms. The basic idea is to find the maximum margin hyper-plane to separate different classes clearly and maximise the distance between them [43]. Tomar and Agarwal [44] applied the hyper-plane diagram for classifying diabetic patients basing on the blood glucose level and the body mass index. A new sample can be inferred throughout the known values of the blood glucose level and the body mass index.
- Clusters are a specific type of representations of clustering algorithms. In this case, the output takes the form of a diagram showing how the instances fall into clusters. In the simplest case, this involves associating a cluster number with each instance, which might be depicted by laying the instances out in two dimensional and partitioning the space [13]. Rawte and Anuradha [45] clustered patients, depending on whether they suffered hearth diseases, arthritis, or Parkinson’s disease. In this case, a new instance cannot be inferred, since clustering tasks are only designed for organising data.
3.1.3. Training and Testing
3.1.4. Credibility. Algorithm Evaluation
3.2. Deep Learning Principles
3.3. Applications
4. Discussion
4.1. Advantages and Disadvantages
4.2. Applicability of ML to Clinical Practice
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| AI | Artificial Intelligence |
| AL | Active Learning |
| BD | Big Data |
| CIMLR | Cancer Integration via Multikernel Learning |
| DBSCAN | Density-Based Spatial Clustering of Applications with Noise |
| DL | Deep Learning |
| DM | Data Mining |
| DMD | Dynamic Mode Decomposition |
| DMDC | DMD with Control |
| DT | Decision Trees |
| GloVe | Global Vectors |
| ID3 | Iterative Dichotomiser 3 |
| JCR | Journal Citation Reports |
| KNN | K-Nearest-Neighbour |
| LH-PCR | Length Heterogeneity Polymerase Chain Reaction |
| LOO | Leave-one-out |
| LPO | Leave-p-out |
| ML | Machine Learning |
| PCA | Principal Component Analysis |
| PINS | Perturbation Clustering for Data Integration and Disease Subtyping |
| POD | Proper Orthogonal Decomposition |
| RF | Random Forest |
| SHIMR | Sparse High-order Interaction Model with Rejection Option |
| SNF | Similarity Network Fusion |
| SOMS | Self-Organized Maps |
| SVM | Support Vector Machine |
References
- Bagga, P.; Hans, R. Applications of Mobile Agents in Healthcare Domain: A Literature Survey. Int. J. Grid Distrib. Comput. 2015, 8. [Google Scholar] [CrossRef]
- Grimson, J.; Stephens, G.; Jung, B.; Grimson, W.; Berry, D.; Pardon, S. Sharing health-care records over the Internet. IEEE Internet Comput. 2001, 5, 49–58. [Google Scholar] [CrossRef]
- Daniels, M.; Schroeder, S.A. Variation among physicians in use of laboratory tests II. Relation to clinical productivity and outcomes of care. Med. Care 1977, 482–487. [Google Scholar] [CrossRef] [PubMed]
- Wennberg, J.E. Dealing with medical practice variations: A proposal for action. Health Aff. 1984, 3, 6–32. [Google Scholar] [CrossRef]
- Smellie, W.S.A.; Galloway, M.J.; Chinn, D.; Gedling, P. Is clinical practice variability the major reason for differences in pathology requesting patterns in general practice? J. Clin. Pathol. 2002, 55, 312–314. [Google Scholar] [CrossRef]
- Stuart, P.J.; Crooks, S.; Porton, M. An interventional program for diagnostic testing in the emergency department. Med. J. Aust. 2002, 177, 131–134. [Google Scholar] [CrossRef]
- Pölsterl, S.; Conjeti, S.; Navab, N.; Katouzian, A. Survival analysis for high-dimensional, heterogeneous medical data: Exploring feature extraction as an alternative to feature selection. Artif. Intell. Med. 2016, 72, 1–11. [Google Scholar] [CrossRef]
- Dick, R.S.; Steen, E.B.; Detmer, D.E. The Computer-Based Patient Record: An Essential Technology for Health Care; National Academies Press: Washington, DC, USA, 1997. [Google Scholar]
- Zhuang, Z.Y.; Churilov, L.; Burstein, F.; Sikaris, K. Combining data mining and case-based reasoning for intelligent decision support for pathology ordering by general practitioners. Eur. J. Oper. Res. 2009, 195, 662–675. [Google Scholar] [CrossRef]
- Huang, M.J.; Chen, M.Y.; Lee, S.C. Integrating data mining with case-based reasoning for chronic diseases prognosis and diagnosis. Expert Syst. Appl. 2007, 32, 856–867. [Google Scholar] [CrossRef]
- Murdoch, T.B.; Detsky, A.S. The inevitable application of big data to health care. JAMA 2013, 309, 1351–1352. [Google Scholar] [CrossRef]
- Wu, X.; Zhu, X.; Wu, G.Q.; Ding, W. Data mining with big data. IEEE Trans. Knowl. Data Eng. 2014, 26, 97–107. [Google Scholar] [CrossRef]
- Witten, I.H.; Frank, E.; Hall, M.A.; Pal, C.J. Data Mining: Practical Machine Learning Tools and Techniques; Elsevier: Amsterdam, The Netherlands, 2016. [Google Scholar]
- Jordan, M.I.; Mitchell, T.M. Machine learning: Trends, perspectives, and prospects. Science 2015, 349, 255–260. [Google Scholar] [CrossRef] [PubMed]
- Kononenko, I. Machine learning for medical diagnosis: History, state of the art and perspective. Artif. Intell. Med. 2001, 23, 89–109. [Google Scholar] [CrossRef]
- Sriram, T.V.S.; Rao, M.V.; Narayana, G.V.S.; Kaladhar, D.S.V.G.K. A Comparison and Prediction Analysis for the Diagnosis of Parkinson Disease Using Data Mining Techniques on Voice Datasets. Int. J. Appl. Eng. Res. 2016, 11, 6355–6360. [Google Scholar]
- Goodfellow, I.; Bengio, Y.; Courville, A. Deep Learning; MIT Press: Cambridge, MA, USA, 2016. [Google Scholar]
- Dixon-Woods, M.; Bonas, S.; Booth, A.; Jones, D.R.; Miller, T.; Sutton, A.J.; Shaw, R.L.; Smith, J.A.; Young, B. How can systematic reviews incorporate qualitative research? A critical perspective. Qual. Res. 2006, 6, 27–44. [Google Scholar] [CrossRef]
- Kourou, K.; Exarchos, T.P.; Exarchos, K.P.; Karamouzis, M.V.; Fotiadis, D.I. Machine learning applications in cancer prognosis and prediction. Comput. Struct. Biotechnol. J. 2015, 13, 8–17. [Google Scholar] [CrossRef] [PubMed]
- Hartigan, J.A. Clustering Algorithms; Wiley: Hoboken, NJ, USA, 1975; Volume 209. [Google Scholar]
- Birant, D.; Kut, A. ST-DBSCAN: An algorithm for clustering spatial-temporal data. Data Knowl. Eng. 2007, 60, 208–221. [Google Scholar] [CrossRef]
- Kohonen, T. The self-organizing map. Neurocomputing 1998, 21, 1–6. [Google Scholar] [CrossRef]
- Dara, R.; Kremer, S.C.; Stacey, D.A. Clustering unlabeled data with SOMs improves classification of labeled real-world data. In Proceedings of the IEEE International Joint Conference on Neural Networks IJCNN’02, Honolulu, HI, USA, 12–17 May 2002; Volume 3, pp. 2237–2242. [Google Scholar] [CrossRef]
- Wang, B.; Mezlini, A.; Demir, F.; Fiume, M.; Tu, Z.; Brudno, M.; Haibe-Kains, B.; Goldenberg, A. Similarity network fusion for aggregating data types on a genomic scale. Nat. Methods 2014, 11. [Google Scholar] [CrossRef]
- Nguyen, T.; Tagett, R.; Diaz, D.; Draghici, S. A novel approach for data integration and disease subtyping. Genome Res. 2017, 27, 2025–2039. [Google Scholar] [CrossRef]
- Ramazzotti, D.; Lal, A.; Wang, B.; Batzoglou, S.; Sidow, A. Multi-omic tumor data reveal diversity of molecular mechanisms that correlate with survival. Nat. Commun. 2018, 9, 4453. [Google Scholar] [CrossRef] [PubMed]
- Nissim, N.; Boland, M.R.; Tatonetti, N.P.; Elovici, Y.; Hripcsak, G.; Shahar, Y.; Moskovitch, R. Improving condition severity classification with an efficient active learning based framework. J. Biomed. Informatics 2016, 61, 44–54. [Google Scholar] [CrossRef] [PubMed]
- Nissim, N.; Shahar, Y.; Elovici, Y.; Hripcsak, G.; Moskovitch, R. Inter-labeler and intra-labeler variability of condition severity classification models using active and passive learning methods. Artif. Intell. Med. 2017, 81, 12–32. [Google Scholar] [CrossRef] [PubMed]
- Cortes, C.; Vapnik, V. Support-vector networks. Mach. Learn. 1995, 20, 273–297. [Google Scholar] [CrossRef]
- Quinlan, J.R. Induction of decision trees. Mach. Learn. 1986, 1, 81–106. [Google Scholar] [CrossRef]
- Quinlan, J.R. C4.5: Programs for Machine Learning; Springer: Berlin, Germany, 1993. [Google Scholar]
- Fix, E.; Hodges, J.L. Discriminatory Analysis-Nonparametric Discrimination: Consistency Properties; Technical Report; DTIC Document; Defense Technical Information Center: Fort Belvoir, VA, USA, 1951.
- McCallum, A.; Nigam, K. A comparison of event models for naive bayes text classification. In Proceedings of the AAAI-98 Workshop on Learning for Text Categorization, Madison, WA, USA, 22–27 July 1998; Volume 752, pp. 41–48. [Google Scholar]
- Heckerman, D.; Horvitz, E.; Nathwani, B.N. Toward Normative Expert Systems: Part I. The Pathfinder project. Methods Inf. Med. 1992, 31, 90–105. [Google Scholar] [CrossRef]
- Heckerman, D.; Nathwani, B.N. Toward Normative Expert Systems: Part II. The Pathfinder project. Methods Inf. Med. 1992, 31, 106–116. [Google Scholar] [CrossRef]
- Lawson, C.L.; Hanson, R.J. Solving Least Squares Problems; Society for Industrial and Applied Mathematics: Philadelphia, PA, USA, 1995. [Google Scholar]
- Kleinbaum, D.G.; Klein, M. Analysis of matched data using logistic regression. In Logistic Regression; Springer: Berlin, Germany, 2010; pp. 389–428. [Google Scholar] [CrossRef]
- Miao, D.Q.; Zhao, Y.; Yao, Y.Y.; Li, H.X.; Xu, F.F. Relative reducts in consistent and inconsistent decision tables of the Pawlak rough set model. Inf. Sci. 2009, 179, 4140–4150. [Google Scholar] [CrossRef]
- Rokach, L.; Maimon, O. Data Mining with Decision Trees: Theory and Applications; World Scientific Publishing: Singapore, 2014. [Google Scholar]
- Breiman, L.; Friedman, J.; Stone, C.J.; Olshen, R.A. Classification and Regression Trees; Chapman and Hall/CRC: Boca Raton, FL, USA, 1984. [Google Scholar]
- Serrano, K.J.; Yu, M.; Coa, K.I.; Collins, L.M.; Atienza, A.A. Mining health app data to find more and less successful weight loss subgroups. J. Med. Internet Res. 2016, 18. [Google Scholar] [CrossRef]
- Yoon, Y.; Cho, J.H.; Yoon, G. Non-constrained blood pressure monitoring using ECG and PPG for personal healthcare. J. Med. Syst. 2009, 33, 261–266. [Google Scholar] [CrossRef]
- Yu, X.; Liu, J.; Zhou, Y.; Wan, W. Study of SVM decision-tree optimization algorithm based on genetic algorithm. In Proceedings of the IEEE International Conference on Audio Language and Image Processing (ICALIP), Shanghai, China, 23–25 November 2010; pp. 1079–1083. [Google Scholar] [CrossRef]
- Tomar, D.; Agarwal, S. A survey on Data Mining approaches for Healthcare. Int. J. Bio-Sci. Bio-Technol. 2013, 5, 241–266. [Google Scholar] [CrossRef]
- Rawte, V.; Anuradha, G. Fraud detection in health insurance using data mining techniques. In Proceedings of the IEEE International Conference on Communication, Information & Computing Technology (ICCICT), Mumbai, India, 15–17 January 2015; pp. 1–5. [Google Scholar] [CrossRef]
- Dietterich, T.G. Ensemble methods in machine learning. In Proceedings of the International Workshop on Multiple Classifier Systems, Cagliari, Italy, 21–23 June 2000; Springer: Berlin, Germany, 2000; pp. 1–15. [Google Scholar] [CrossRef]
- Ye, Q.; Zhang, Z.; Law, R. Sentiment classification of online reviews to travel destinations by supervised machine learning approaches. Expert Syst. Appl. 2009, 36, 6527–6535. [Google Scholar] [CrossRef]
- Veluswami, A.; Nakhla, M.S.; Zhang, Q.J. The application of neural networks to EM-based simulation and optimization of interconnects in high-speed VLSI circuits. IEEE Trans. Microw. Theory Tech. 1997, 45, 712–723. [Google Scholar] [CrossRef]
- Dreiseitl, S.; Ohno-Machado, L. Logistic regression and artificial neural network classification models: A methodology review. J. Biomed. Informatics 2002, 35, 352–359. [Google Scholar] [CrossRef]
- Crippa, A.; Salvatore, C.; Perego, P.; Forti, S.; Nobile, M.; Molteni, M.; Castiglioni, I. Use of machine learning to identify children with autism and their motor abnormalities. J. Autism Dev. Disord. 2015, 45, 2146–2156. [Google Scholar] [CrossRef] [PubMed]
- Samui, P. Handbook of Research on Advanced Computational Techniques for Simulation-Based Engineering; Advances in Computer and Electrical Engineering; IGI Global: Hershey, PA, USA, 2015. [Google Scholar]
- Wahbeh, A.H.; Al-Radaideh, Q.A.; Al-Kabi, M.N.; Al-Shawakfa, E.M. A comparison study between data mining tools over some classification methods. Int. J. Adv. Comput. Sci. Appl. 2011, 8, 18–26. [Google Scholar] [CrossRef]
- Khor, K.C.; Ting, C.Y.; Phon-Amnuaisuk, S. A cascaded classifier approach for improving detection rates on rare attack categories in network intrusion detection. Appl. Intell. 2012, 36, 320–329. [Google Scholar] [CrossRef]
- Kim, J.H. Estimating classification error rate: Repeated cross-validation, repeated hold-out and bootstrap. Comput. Stat. Data Anal. 2009, 53, 3735–3745. [Google Scholar] [CrossRef]
- Nguyen, T.T.T.; Armitage, G. A survey of techniques for internet traffic classification using machine learning. IEEE Commun. Surv. Tutorials 2008, 10, 56–76. [Google Scholar] [CrossRef]
- Kohavi, R. A study of cross-validation and bootstrap for accuracy estimation and model selection. In Proceedings of the International Joint Conference on Artificial Intelligence (IJCAI), Stanford, CA, USA, 20–25 August 1995; Volume 14, pp. 1137–1145. [Google Scholar]
- Arlot, S.; Celisse, A. A survey of cross-validation procedures for model selection. Stat. Surv. 2010, 4, 40–79. [Google Scholar] [CrossRef]
- Efron, B.; Tibshirani, R.J. Improvements on cross-validation: The 632+ bootstrap method. J. Am. Stat. Assoc. 1997, 92, 548–560. [Google Scholar] [CrossRef]
- Efron, B.; Tibshirani, R.J. An Introduction to the Bootstrap; Chapman & Hall/CRC: Boca Raton, FL, USA, 1994. [Google Scholar]
- Greenspan, H.; van Ginneken, B.; Summers, R.M. Guest editorial deep learning in medical imaging: Overview and future promise of an exciting new technique. IEEE Trans. Med. Imaging 2016, 35, 1153–1159. [Google Scholar] [CrossRef]
- Yuan, Z.; Lu, Y.; Wang, Z.; Xue, Y. Droid-Sec: Deep learning in android malware detection. ACM SIGCOMM Comput. Commun. Rev. 2014, 44, 371–372. [Google Scholar] [CrossRef]
- LeCun, Y.; Bengio, Y.; Hinton, G. Deep learning. Nature 2015, 521, 436–444. [Google Scholar] [CrossRef]
- Hecht-Nielsen, R. Theory of the backpropagation neural network. Neural Networks 1988, 1, 593–605. [Google Scholar] [CrossRef]
- Werbos, P.J. Beyond Regression: New Tools for Prediction and Analysis in Behavioral Sciences. Ph.D. Thesis, Harvard University, Cambridge, MA, USA, 1974. [Google Scholar]
- Rumenlhart, D.E.; McClelland, J.L. Parallel Distributed Processing; Volume 1. Explorations in the Microstructure of Cognition: Foundations; MIT Press: Cambridge, MA, USA, 1986. [Google Scholar]
- Schmidhuber, J. Deep learning in neural networks: An overview. Neural Networks 2015, 61, 85–117. [Google Scholar] [CrossRef]
- Erb, R.J. Introduction to backpropagation neural network computation. Pharm. Res. 1993, 10, 165–170. [Google Scholar] [CrossRef]
- Alimadadi, A.; Aryal, S.; Manandhar, I.; Munroe, P.B.; Joe, B.; Cheng, X. Artificial intelligence and machine learning to fight COVID-19. Physiol. Genom. 2020, 52, 200–202. [Google Scholar] [CrossRef]
- Arga, K.Y. COVID-19 and the Futures of Machine Learning. OMICS J. Integr. Biol. 2020. [Google Scholar] [CrossRef]
- Albahri, A.S.; Hamid, R.A.; Alwan, J.k.; Al-qays, Z.; Zaidan, A.A.; Zaidan, B.B.; Albahri, A.O.S.; AlAmoodi, A.H.; Khlaf, J.M.; Almahdi, E.M.; et al. Role of biological Data Mining and Machine Learning Techniques in Detecting and Diagnosing the Novel Coronavirus (COVID-19): A Systematic Review. J. Med. Syst. 2020, 44, 122. [Google Scholar] [CrossRef]
- Sujath, R.; Chatterjee, J.M.; Hassanien, A.E. A machine learning forecasting model for COVID-19 pandemic in India. Stoch. Environ. Res. Risk Assess. 2020, 34, 959–972. [Google Scholar] [CrossRef]
- Randhawa, G.S.; Soltysiak, M.P.M.; El Roz, H.; de Souza, C.P.E.; Hill, K.A.; Kari, L. Machine learning using intrinsic genomic signatures for rapid classification of novel pathogens: COVID-19 case study. PLoS ONE 2020, 15, 1–24. [Google Scholar] [CrossRef] [PubMed]
- Sear, R.F.; Velásquez, N.; Leahy, R.; Restrepo, N.J.; Oud, S.E.; Gabriel, N.; Lupu, Y.; Johnson, N.F. Quantifying COVID-19 Content in the Online Health Opinion War Using Machine Learning. IEEE Access 2020, 8, 91886–91893. [Google Scholar] [CrossRef]
- Maas, K.; Chan, S.; Parker, J.; Slater, A.; Moore, J.; Olsen, N.; Aune, T.M. Cutting edge: Molecular portrait of human autoimmune disease. J. Immunol. 2002, 169, 5–9. [Google Scholar] [CrossRef] [PubMed]
- Phillips, H.S.; Kharbanda, S.; Chen, R.; Forrest, W.F.; Soriano, R.H.; Wu, T.D.; Misra, A.; Nigro, J.M.; Colman, H.; Soroceanu, L. Molecular subclasses of high-grade glioma predict prognosis, delineate a pattern of disease progression, and resemble stages in neurogenesis. Cancer Cell 2006, 9, 157–173. [Google Scholar] [CrossRef]
- Ng, H.P.; Ong, S.H.; Foong, K.W.C.; Goh, P.S.; Nowinski, W.L. Medical image segmentation using k-means clustering and improved watershed algorithm. In Proceedings of the IEEE Southwest Symposium on Image Analysis and Interpretation, Denver, CO, USA, 26–28 March 2006; pp. 61–65. [Google Scholar] [CrossRef]
- Aarsland, D.; Brønnick, K.; Ehrt, U.; De Deyn, P.P.; Tekin, S.; Emre, M.; Cummings, J.L. Neuropsychiatric symptoms in patients with Parkinson’s disease and dementia: Frequency, profile and associated care giver stress. J. Neurol. Neurosurg. Psychiatry 2007, 78, 36–42. [Google Scholar] [CrossRef]
- Sun, G.; Hakozaki, Y.; Abe, S.; Vinh, N.Q.; Matsui, T. A novel infection screening method using a neural network and k-means clustering algorithm which can be applied for screening of unknown or unexpected infectious diseases. J. Infect. 2012, 65, 591–592. [Google Scholar] [CrossRef]
- Seok, J.; Warren, H.S.; Cuenca, A.G.; Mindrinos, M.N.; Baker, H.V.; Xu, W.; Richards, D.R.; McDonald-Smith, G.P.; Gao, H.; Hennessy, L.; et al. Genomic responses in mouse models poorly mimic human inflammatory diseases. Proc. Natl. Acad. Sci. USA 2013, 110, 3507–3512. [Google Scholar] [CrossRef]
- Manickavasagam, K.; Sutha, S.; Kamalanand, K. An automated system based on 2 d empirical mode decomposition and k-means clustering for classification of Plasmodium species in thin blood smear images. BMC Infect. Dis. 2014, 14, P13. [Google Scholar] [CrossRef]
- Antony, S.J.S.; Ravi, S. A new approach to determine the classification of mammographic image using K-means clustering algorithm. Int. J. Adv. Res. Technol. 2015, 4, 40–44. [Google Scholar]
- Sari, B.N. Identification of Tuberculosis Patient Characteristics Using K-Means Clustering. Sci. J. Informatics 2016, 3, 129–138. [Google Scholar] [CrossRef]
- Kumar, P.R.; Prasath, T.A.; Rajasekaran, M.P.; Vishnuvarthanan, G. Brain Subject Estimation Using PSO K-Means Clustering-An Automated Aid for the Assessment of Clinical Dementia. In Proceedings of the International Conference on Information and Communication Technology for Intelligent Systems, Ahmedabad, India, 15–16 March 2017; Springer: Berlin, Germany, 2017; pp. 482–489. [Google Scholar] [CrossRef]
- Chauhan, R.; Kaur, H.; Alam, M.A. Data clustering method for discovering clusters in spatial cancer databases. Int. J. Comput. Appl. 2010, 10, 9–14. [Google Scholar] [CrossRef]
- Çelik, M.; Dadaşer-Çelik, F.; Dokuz, A.Ş. Anomaly detection in temperature data using dbscan algorithm. In Proceedings of the IEEE International Symposium on Innovations in Intelligent Systems and Applications (INISTA), Istanbul, Turkey, 15–18 June 2011; pp. 91–95. [Google Scholar] [CrossRef]
- Radhakrishnan, M.; Kuttiannan, T. Comparative analysis of feature extraction methods for the classification of prostate cancer from TRUS medical images. Int. J. Comput. Sci. 2012, 9, 171–179. [Google Scholar]
- Antonelli, D.; Baralis, E.; Bruno, G.; Cerquitelli, T.; Chiusano, S.; Mahoto, N. Analysis of diabetic patients through their examination history. Expert Syst. Appl. 2013, 40, 4672–4678. [Google Scholar] [CrossRef]
- Sriram, T.V.S.; Rao, M.V.; Narayana, G.V.S.; Kaladhar, D.S.V.G.K. Diagnosis of Parkinson Disease Using Machine Learning and Data Mining Systems from Voice Dataset. In Proceedings of the 3rd International Conference on Frontiers of Intelligent Computing: Theory and Applications (FICTA), Durgapur, West Bengal, India, 16–18 November 2015; Springer: Berlin, Germany, 2015; pp. 151–157. [Google Scholar] [CrossRef]
- Plant, C.; Teipel, S.J.; Oswald, A.; Böhm, C.; Meindl, T.; Mourao-Miranda, J.; Bokde, A.W.; Hampel, H.; Ewers, M. Automated detection of brain atrophy patterns based on MRI for the prediction of Alzheimer’s disease. Neuroimage 2010, 50, 162–174. [Google Scholar] [CrossRef]
- Aidos, H.; Fred, A. Discrimination of Alzheimer’s Disease using longitudinal information. Data Min. Knowl. Discov. 2017, 31, 1006–1030. [Google Scholar] [CrossRef]
- Stebbins, G.T.; Goetz, C.G.; Burn, D.J.; Jankovic, J.; Khoo, T.K.; Tilley, B.C. How to identify tremor dominant and postural instability/gait difficulty groups with the movement disorder society unified Parkinson’s disease rating scale: Comparison with the unified Parkinson’s disease rating scale. Mov. Disord. 2013, 28, 668–670. [Google Scholar] [CrossRef]
- Lyketsos, C.G.; Sheppard, J.M.E.; Steinberg, M.; Tschanz, J.A.T.; Norton, M.C.; Steffens, D.C.; Breitner, J. Neuropsychiatric disturbance in Alzheimer’s disease clusters into three groups: The Cache County study. Int. J. Geriatr. Psychiatry 2001, 16, 1043–1053. [Google Scholar] [CrossRef]
- Harold, D.; Abraham, R.; Hollingworth, P.; Sims, R.; Gerrish, A.; Hamshere, M.L.; Pahwa, J.S.; Moskvina, V.; Dowzell, K.; Williams, A. Genome-wide association study identifies variants at CLU and PICALM associated with Alzheimer’s disease. Nat. Genet. 2009, 41, 1088–1093. [Google Scholar] [CrossRef]
- Lambert, J.C.; Ibrahim-Verbaas, C.A.; Harold, D.; Naj, A.C.; Sims, R.; Bellenguez, C.; Jun, G.; DeStefano, A.L.; Bis, J.C.; Beecham, G.W. Meta-analysis of 74,046 individuals identifies 11 new susceptibility loci for Alzheimer’s disease. Nat. Genet. 2013, 45, 1452–1458. [Google Scholar] [CrossRef]
- Huang, C.L.; Liao, H.C.; Chen, M.C. Prediction model building and feature selection with support vector machines in breast cancer diagnosis. Expert Syst. Appl. 2008, 34, 578–587. [Google Scholar] [CrossRef]
- Avci, E. A new intelligent diagnosis system for the heart valve diseases by using genetic-SVM classifier. Expert Syst. Appl. 2009, 36, 10618–10626. [Google Scholar] [CrossRef]
- Fei, S.W. Diagnostic study on arrhythmia cordis based on particle swarm optimization-based support vector machine. Expert Syst. Appl. 2010, 37, 6748–6752. [Google Scholar] [CrossRef]
- Yao, J.; Dwyer, A.; Summers, R.M.; Mollura, D.J. Computer-aided diagnosis of pulmonary infections using texture analysis and support vector machine classification. Acad. Radiol. 2011, 18, 306–314. [Google Scholar] [CrossRef]
- Sartakhti, J.S.; Zangooei, M.H.; Mozafari, K. Hepatitis disease diagnosis using a novel hybrid method based on support vector machine and simulated annealing (SVM-SA). Comput. Methods Programs Biomed. 2012, 108, 570–579. [Google Scholar] [CrossRef]
- Abdi, M.J.; Giveki, D. Automatic detection of erythemato-squamous diseases using PSO–SVM based on association rules. Eng. Appl. Artif. Intell. 2013, 26, 603–608. [Google Scholar] [CrossRef]
- Berna, A.Z.; McCarthy, J.S.; Wang, R.X.; Saliba, K.J.; Bravo, F.G.; Cassells, J.; Padovan, B.; Trowell, S.C. Analysis of breath specimens for biomarkers of Plasmodium falciparum infection. J. Infect. Dis. 2015, 212, 1120–1128. [Google Scholar] [CrossRef]
- Kesorn, K.; Ongruk, P.; Chompoosri, J.; Phumee, A.; Thavara, U.; Tawatsin, A.; Siriyasatien, P. Morbidity rate prediction of dengue hemorrhagic fever (DHF) using the support vector machine and the Aedes aegypti infection rate in similar climates and geographical areas. PLoS ONE 2015, 10, e0125049. [Google Scholar] [CrossRef]
- Khan, S.; Ullah, R.; Khan, A.; Wahab, N.; Bilal, M.; Ahmed, M. Analysis of dengue infection based on Raman spectroscopy and support vector machine (SVM). Biomed. Opt. Express 2016, 7, 2249–2256. [Google Scholar] [CrossRef]
- Meystre, S.; Gouripeddi, R.; Tieder, J.; Simmons, J.; Srivastava, R.; Shah, S. Enhancing Comparative Effectiveness Research with Automated Pediatric Pneumonia Detection in a Multi-Institutional Clinical Repository: A PHIS+ Pilot Study. J. Med. Internet Res. 2017, 19. [Google Scholar] [CrossRef]
- Hernandez, B.; Herrero, P.; Rawson, T.M.; Moore, L.S.P.; Evans, B.; Toumazou, C.; Holmes, A.H.; Georgiou, P. Supervised learning for infection risk inference using pathology data. BMC Med. Informatics Decis. Mak. 2017, 17, 168. [Google Scholar] [CrossRef] [PubMed]
- Forsström, J.; Nuutila, P.; Irjala, K. Using the ID3 algorithm to find discrepant diagnoses from laboratory databases of thyroid patients. Med. Decis. Mak. 1991, 11, 171–175. [Google Scholar] [CrossRef]
- Tanner, L.; Schreiber, M.; Low, J.G.H.; Ong, A.; Tolfvenstam, T.; Lai, Y.L.; Ng, L.C.; Leo, Y.S.; Puong, L.T.; Vasudevan, S.G.; et al. Decision tree algorithms predict the diagnosis and outcome of dengue fever in the early phase of illness. PLoS Neglected Trop. Dis. 2008, 2, e196. [Google Scholar] [CrossRef] [PubMed]
- Ture, M.; Tokatli, F.; Kurt, I. Using Kaplan-Meier analysis together with decision tree methods (C&RT, CHAID, QUEST, C4.5 and ID3) in determining recurrence-free survival of breast cancer patients. Expert Syst. Appl. 2009, 36, 2017–2026. [Google Scholar] [CrossRef]
- Bashir, S.; Qamar, U.; Khan, F.H.; Javed, M.Y. An Efficient Rule-Based Classification of Diabetes Using ID3, C4.5, & CART Ensembles. In Proceedings of the IEEE 12-th International Conference on Frontiers of Information Technology (FIT), Islamabad, Pakistan, 17–19 December 2014; pp. 226–231. [Google Scholar] [CrossRef]
- Thenmozhi, K.; Deepika, P. Heart disease prediction using classification with different decision tree techniques. Int. J. Eng. Res. Gen. Sci. 2014, 2, 6–11. [Google Scholar]
- Buczak, A.L.; Baugher, B.; Guven, E.; Ramac-Thomas, L.C.; Elbert, Y.; Babin, S.M.; Lewis, S.H. Fuzzy association rule mining and classification for the prediction of malaria in South Korea. BMC Med. Informatics Decis. Mak. 2015, 15, 47. [Google Scholar] [CrossRef]
- Subasi, A.; Alickovic, E.; Kevric, J. Diagnosis of Chronic Kidney Disease by Using Random Forest. In Proceedings of the International Conference on Medical and Biological Engineering, CMBEBIH 2017, Sarajevo, Bosnia and Herzegovina, 16–18 March 2017; pp. 589–594. [Google Scholar] [CrossRef]
- Abdar, M.; Zomorodi-Moghadam, M.; Das, R.; Ting, I.H. Performance analysis of classification algorithms on early detection of liver disease. Expert Syst. Appl. 2017, 67, 239–251. [Google Scholar] [CrossRef]
- Singh, G.; ManjotKaur, E. A Review Paper: Decision Tree Algorithms for diagnosis of Angioplasty and Stents for Heart Disease Treatment. Int. J. Eng. Sci. 2017, 7, 6643–6645. [Google Scholar]
- Jen, C.H.; Wang, C.C.; Jiang, B.C.; Chu, Y.H.; Chen, M.S. Application of classification techniques on development an early-warning system for chronic illnesses. Expert Syst. Appl. 2012, 39, 8852–8858. [Google Scholar] [CrossRef]
- Liu, D.Y.; Chen, H.L.; Yang, B.; Lv, X.E.; Li, L.N.; Liu, J. Design of an enhanced fuzzy k-nearest neighbor classifier based computer aided diagnostic system for thyroid disease. J. Med. Syst. 2012, 36, 3243–3254. [Google Scholar] [CrossRef]
- Zuo, W.L.; Wang, Z.Y.; Liu, T.; Chen, H.L. Effective detection of Parkinson’s disease using an adaptive fuzzy k-nearest neighbor approach. Biomed. Signal Process. Control. 2013, 8, 364–373. [Google Scholar] [CrossRef]
- Wisittipanit, N.; Rangwala, H.; Sikaroodi, M.; Keshavarzian, A.; Mutlu, E.A.; Gillevet, P. Classification methods for the analysis of LH–PCR data associated with inflammatory bowel disease patients. Int. J. Bioinform. Res. Appl. 2015, 11, 111–129. [Google Scholar] [CrossRef] [PubMed]
- Papakostas, G.A.; Savio, A.; Graña, M.; Kaburlasos, V.G. A lattice computing approach to Alzheimer’s disease computer assisted diagnosis based on MRI data. Neurocomputing 2015, 150, 37–42. [Google Scholar] [CrossRef]
- Chandel, K.; Kunwar, V.; Sabitha, S.; Choudhury, T.; Mukherjee, S. A comparative study on thyroid disease detection using K-nearest neighbor and Naive Bayes classification techniques. CSI Trans. ICT 2016, 4, 313–319. [Google Scholar] [CrossRef]
- Mahajan, P.; Kuppermann, N.; Mejias, A.; Suarez, N.; Chaussabel, D.; Casper, T.C.; Smith, B.; Alpern, E.R.; Anders, J.; Atabaki, S.M.; et al. Association of RNA biosignatures with bacterial infections in febrile infants aged 60 days or younger. JAMA 2016, 316, 846–857. [Google Scholar] [CrossRef]
- Biswas, S.; Acharyya, S. Identification of disease critical genes causing Duchenne muscular dystrophy (DMD) using computational intelligence. CSI Trans. ICT 2017, 5, 3–8. [Google Scholar] [CrossRef]
- Nelson, K.N.; Hui, Q.; Rimland, D.; Xu, K.; Freiberg, M.S.; Justice, A.C.; Marconi, V.C.; Sun, Y.V. Identification of HIV infection-related DNA methylation sites and advanced epigenetic aging in HIV-positive, treatment-naive US veterans. Aids 2017, 31, 571–575. [Google Scholar] [CrossRef]
- Vargas, J.C.B.; Cernea, A.; Fernández-Martínez, J.L. Improvements in Resampling Techniques for Phenotype Prediction: Applications to Neurodegenerative Diseases. In Computational Mathematics, Numerical Analysis and Applications; Springer: Berlin, Germany, 2017; Volume 13, pp. 245–248. [Google Scholar] [CrossRef]
- Mabrouk, R.; Chikhaoui, B.; Bentabet, L. Machine Learning Models Classification using Clinical and DaTSCAN SPECT Imaging features: A Study on SWEDD and Parkinson’s disease. IEEE Trans. Radiat. Plasma Med. Sci. 2018, 1. [Google Scholar] [CrossRef]
- Soni, J.; Ansari, U.; Sharma, D.; Soni, S. Predictive data mining for medical diagnosis: An overview of heart disease prediction. Int. J. Comput. Appl. 2011, 17, 43–48. [Google Scholar] [CrossRef]
- Pattekari, S.A.; Parveen, A. Prediction system for heart disease using Naïve Bayes. Int. J. Adv. Comput. Math. Sci. 2012, 3, 290–294. [Google Scholar]
- Chaurasia, V.; Pal, S. Data mining approach to detect heart diseases. Int. J. Adv. Comput. Sci. Inf. Technol. 2013, 2, 56–66. [Google Scholar]
- Vijayarani, S.; Dhayanand, S. Liver disease prediction using SVM and Naïve Bayes algorithms. Int. J. Sci. Eng. Technol. Res. 2015, 4, 816–820. [Google Scholar]
- Thangaraju, P.; Mehala, R. Novel Classification based approaches over Cancer Diseases. System 2015, 4, 294–297. [Google Scholar] [CrossRef]
- Vijayarani, S.; Dhayanand, S. Data mining classification algorithms for kidney disease prediction. Int. J. Cybern. Informatics 2015, 4, 13–25. [Google Scholar] [CrossRef]
- Zhou, X.; Wang, S.; Xu, W.; Ji, G.; Phillips, P.; Sun, P.; Zhang, Y. Detection of Pathological Brain in MRI Scanning Based on Wavelet-Entropy and Naive Bayes Classifier. In Proceedings of the International Conference on Bioinformatics and Biomedical Engineering IWBBIO, Granada, Spain, 15–17 April 2015; pp. 201–209. [Google Scholar]
- Trihartati, A.S.; Adi, C.K. An Identification of Tuberculosis (Tb) Disease in Humans using Naïve Bayesian Method. Sci. J. Informatics 2016, 3, 99–108. [Google Scholar] [CrossRef]
- Ferreira, F.L.; Cardoso, S.; Silva, D.; Guerreiro, M.; de Mendonça, A.; Madeira, S.C. Improving Prognostic Prediction from Mild Cognitive Impairment to Alzheimer’s Disease Using Genetic Algorithms. In Proceedings of the 11th International Conference on Practical Applications of Computational Biology & Bioinformatics, Porto, Portugal, 21–23 January 2017; pp. 180–188. [Google Scholar] [CrossRef]
- Stern, S.; Santos, R.; Marchetto, M.C.; Mendes, A.P.D.; Rouleau, G.A.; Biesmans, S.; Wang, Q.W.; Yao, J.; Charnay, P.; Bang, A.G.; et al. Neurons derived from patients with bipolar disorder divide into intrinsically different sub-populations of neurons, predicting the patients’ responsiveness to lithium. Mol. Psychiatry 2017. [Google Scholar] [CrossRef]
- Gevaert, O.; Smet, F.D.; Timmerman, D.; Moreau, Y.; Moor, B.D. Predicting the prognosis of breast cancer by integrating clinical and microarray data with Bayesian networks. Bioinformatics 2006, 22, e184–e190. [Google Scholar] [CrossRef]
- Vázquez-Castellanos, J.F.; Serrano-Villar, S.; Latorre, A.; Artacho, A.; Ferrus, M.L.; Madrid, N.; Vallejo, A.; Sainz, T.; Martínez-Botas, J.; Ferrando-Martínez, S. Altered metabolism of gut microbiota contributes to chronic immune activation in HIV-infected individuals. Mucosal Immunol. 2015, 8, 760–772. [Google Scholar] [CrossRef]
- Castro, A.; Pinheiro, P.; Pinheiro, M. A multicriteria model applied in the diagnosis of alzheimer’s disease. Rough Sets Knowl. Technol. 2008, 612–619. [Google Scholar] [CrossRef]
- Wu, X.; Li, R.; Fleisher, A.S.; Reiman, E.M.; Guan, X.; Zhang, Y.; Chen, K.; Yao, L. Altered default mode network connectivity in Alzheimer’s disease—A resting functional MRI and Bayesian network study. Hum. Brain Mapp. 2011, 32, 1868–1881. [Google Scholar] [CrossRef]
- Zhang, B.; Gaiteri, C.; Bodea, L.G.; Wang, Z.; McElwee, J.; Podtelezhnikov, A.A.; Zhang, C.; Xie, T.; Tran, L.; Dobrin, R. Integrated systems approach identifies genetic nodes and networks in late-onset Alzheimer’s disease. Cell 2013, 153, 707–720. [Google Scholar] [CrossRef] [PubMed]
- Rowe, J.B.; Hughes, L.E.; Barker, R.A.; Owen, A.M. Dynamic causal modelling of effective connectivity from fMRI: Are results reproducible and sensitive to Parkinson’s disease and its treatment? Neuroimage 2010, 52, 1015–1026. [Google Scholar] [CrossRef] [PubMed]
- Sciarretta, S.; Palano, F.; Tocci, G.; Baldini, R.; Volpe, M. Antihypertensive treatment and development of heart failure in hypertension: A Bayesian network meta-analysis of studies in patients with hypertension and high cardiovascular risk. Arch. Intern. Med. 2011, 171, 384–394. [Google Scholar] [CrossRef] [PubMed]
- Wu, H.Y.; Huang, J.W.; Lin, H.J.; Liao, W.C.; Peng, Y.S.; Hung, K.Y.; Wu, K.D.; Tu, Y.K.; Chien, K.L. Comparative effectiveness of renin-angiotensin system blockers and other antihypertensive drugs in patients with diabetes: Systematic review and bayesian network meta-analysis. BMJ 2013, 347, f6008. [Google Scholar] [CrossRef]
- Raggi, P.; Boulay, A.; Chasan-Taber, S.; Amin, N.; Dillon, M.; Burke, S.K.; Chertow, G.M. Cardiac calcification in adult hemodialysis patients: A link between end-stage renal disease and cardiovascular disease? J. Am. Coll. Cardiol. 2002, 39, 695–701. [Google Scholar] [CrossRef]
- Lanzkron, S.; Carroll, C.P.; Haywood, C., Jr. Mortality rates and age at death from sickle cell disease: US, 1979–2005. Public Health Rep. 2013, 128, 110–116. [Google Scholar] [CrossRef]
- Zhou, Z.G.; Liu, F.; Jiao, L.C.; Wang, Z.L.; Zhang, X.P.; Wang, X.D.; Luo, X.Z. An evidential reasoning based model for diagnosis of lymph node metastasis in gastric cancer. BMC Med. Informatics Decis. Mak. 2013, 13, 123. [Google Scholar] [CrossRef]
- Smith, K.F.; Goldberg, M.; Rosenthal, S.; Carlson, L.; Chen, J.; Chen, C.; Ramachandran, S. Global rise in human infectious disease outbreaks. J. R. Soc. Interface 2014, 11, 20140950. [Google Scholar] [CrossRef]
- Althoff, K.N.; McGinnis, K.A.; Wyatt, C.M.; Freiberg, M.S.; Gilbert, C.; Oursler, K.K.; Rimland, D.; Rodriguez-Barradas, M.C.; Dubrow, R.; Park, L.S.; et al. Comparison of risk and age at diagnosis of myocardial infarction, end-stage renal disease, and non-AIDS-defining cancer in HIV-infected versus uninfected adults. Clin. Infect. Dis. 2014, 60, 627–638. [Google Scholar] [CrossRef]
- Williams, J.P.; Hurst, J.; Stöhr, W.; Robinson, N.; Brown, H.; Fisher, M.; Kinloch, S.; Cooper, D.; Schechter, M.; Tambussi, G.; et al. HIV-1 DNA predicts disease progression and post-treatment virological control. Elife 2014, 3, e03821. [Google Scholar] [CrossRef]
- Nowak, P.; Troseid, M.; Avershina, E.; Barqasho, B.; Neogi, U.; Holm, K.; Hov, J.R.; Noyan, K.; Vesterbacka, J.; Svärd, J.; et al. Gut microbiota diversity predicts immune status in HIV-1 infection. Aids 2015, 29, 2409–2418. [Google Scholar] [CrossRef]
- Horvath, S.; Levine, A.J. HIV-1 infection accelerates age according to the epigenetic clock. J. Infect. Dis. 2015, 212, 1563–1573. [Google Scholar] [CrossRef] [PubMed]
- Gjoneska, E.; Pfenning, A.R.; Mathys, H.; Quon, G.; Kundaje, A.; Tsai, L.H.; Kellis, M. Conserved epigenomic signals in mice and humans reveal immune basis of Alzheimer’s disease. Nature 2015, 518, 365–369. [Google Scholar] [CrossRef]
- Fischer, R.; Judson, S.; Miazgowicz, K.; Bushmaker, T.; Prescott, J.; Munster, V.J. Ebola virus stability on surfaces and in fluids in simulated outbreak environments. Emerg. Infect. Dis. 2015, 21, 1243–1246. [Google Scholar] [CrossRef] [PubMed]
- Ly, K.N.; Hughes, E.M.; Jiles, R.B.; Holmberg, S.D. Rising mortality associated with hepatitis C virus in the United States, 2003–2013. Clin. Infect. Dis. 2016, 62, 1287–1288. [Google Scholar] [CrossRef]
- Chaplot, S.; Patnaik, L.M.; Jagannathan, N.R. Classification of magnetic resonance brain images using wavelets as input to support vector machine and neural network. Biomed. Signal Process. Control. 2006, 1, 86–92. [Google Scholar] [CrossRef]
- Saritha, M.; Joseph, K.P.; Mathew, A.T. Classification of MRI brain images using combined wavelet entropy based spider web plots and probabilistic neural network. Pattern Recognit. Lett. 2013, 34, 2151–2156. [Google Scholar] [CrossRef]
- El-Dahshan, E.S.A.; Hosny, T.; Salem, A.B.M. Hybrid intelligent techniques for MRI brain images classification. Digit. Signal Process. 2010, 20, 433–441. [Google Scholar] [CrossRef]
- Chen, H.L.; Huang, C.C.; Yu, X.G.; Xu, X.; Sun, X.; Wang, G.; Wang, S.J. An efficient diagnosis system for detection of Parkinson’s disease using fuzzy k-nearest neighbor approach. Expert Syst. Appl. 2013, 40, 263–271. [Google Scholar] [CrossRef]
- Hariharan, M.; Polat, K.; Sindhu, R. A new hybrid intelligent system for accurate detection of Parkinson’s disease. Comput. Methods Programs Biomed. 2014, 113, 904–913. [Google Scholar] [CrossRef] [PubMed]
- Fan, C.Y.; Chang, P.C.; Lin, J.J.; Hsieh, J.C. A hybrid model combining case-based reasoning and fuzzy decision tree for medical data classification. Appl. Soft Comput. 2011, 11, 632–644. [Google Scholar] [CrossRef]
- Onan, A. A fuzzy-rough nearest neighbor classifier combined with consistency-based subset evaluation and instance selection for automated diagnosis of breast cancer. Expert Syst. Appl. 2015, 42, 6844–6852. [Google Scholar] [CrossRef]
- Marateb, H.R.; Mansourian, M.; Faghihimani, E.; Amini, M.; Farina, D. A hybrid intelligent system for diagnosing microalbuminuria in type 2 diabetes patients without having to measure urinary albumin. Comput. Biol. Med. 2014, 45, 34–42. [Google Scholar] [CrossRef] [PubMed]
- Norouzi, J.; Yadollahpour, A.; Mirbagheri, S.A.; Mazdeh, M.M.; Hosseini, S.A. Predicting renal failure progression in chronic kidney disease using integrated intelligent fuzzy expert system. Comput. Math. Methods Med. 2016, 2016. [Google Scholar] [CrossRef] [PubMed]
- Das, A.; Rad, P.; Choo, K.K.R.; Nouhi, B.; Lish, J.; Martel, J. Distributed machine learning cloud teleophthalmology IoT for predicting AMD disease progression. Future Gener. Comput. Syst. 2018, 93. [Google Scholar] [CrossRef]
- Obolski, U.; Gori, A.; Lourençp, J.; Thompson, C.; Thompson, R.; French, N.; Heyderman, R.; Gupta, S. Identifying genes associated with invasive disease in S. pneumoniae by applying a machine learning approach to whole genome sequence typing data. Sci. Rep. 2019, 9, 4049. [Google Scholar] [CrossRef]
- Mcdonnell, M.; Harris, R.; Mills, T.; Downey, L.; Dharmasiri, S.; Felwick, R.; Borca, F.; Phan, H.; Cummings, J.R.F.; Gwiggner, M. P384 High incidence of hyperglycaemia in steroid treated hospitalised inflammatory bowel disease (IBD) patients and its risk factors identified by machine learning methods. J. Crohn’S Colitis 2019, 13, S299–S300. [Google Scholar] [CrossRef]
- Das, D.; Ito, J.; Kadowaki, T.; Tsuda, K. An interpretable machine learning model for diagnosis of Alzheimer’s disease. PeerJ 2019, 7. [Google Scholar] [CrossRef]
- Patrick, M.; Raja, K.; Miller, K.; Sotzen, J.; Gudjonsson, J.; Elder, J.; Tsoi, L. Drug Repurposing Prediction for Immune-Mediated Cutaneous Diseases using a Word-Embedding–Based Machine Learning Approach. J. Investig. Dermatol. 2018, 139. [Google Scholar] [CrossRef]
- Xi, J.; Zhao, W. Correlating exhaled aerosol images to small airway obstructive diseases: A study with dynamic mode decomposition and machine learning. PLoS ONE 2019, 14. [Google Scholar] [CrossRef]
- Konerman, M.A.; Beste, L.A.; Van, T.; Liu, B.; Zhang, X.; Zhu, J.; Saini, S.D.; Su, G.L.; Nallamothu, B.K.; Ioannou, G.N.; et al. Machine learning models to predict disease progression among veterans with hepatitis C virus. PLoS ONE 2019, 14. [Google Scholar] [CrossRef]
- Klein, J.; Baker, N.; Zorn, K.; Russo, D.; Rubio, A.; Clark, A.; Ekins, S. Data mining and machine learning for lysosomal disease drug discovery and beyond. Mol. Genet. Metab. 2019, 126, S86. [Google Scholar] [CrossRef]
- Rathore, S.; Niazi, T.; Iftikhar, M.A.; Chaddad, A. Glioma Grading via Analysis of Digital Pathology Images Using Machine Learning. Cancers 2020, 12, 578. [Google Scholar] [CrossRef] [PubMed]
- Vey, J.; Kapsner, L.A.; Fuchs, M.; Unberath, P.; Veronesi, G.; Kunz, M. A Toolbox for Functional Analysis and the Systematic Identification of Diagnostic and Prognostic Gene Expression Signatures Combining Meta-Analysis and Machine Learning. Cancers 2019, 11, 1606. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.Y.; Chen, C.H.; Shi, S.; Chung, C.R.; Wen, Y.H.; Wu, M.H.; Lebowitz, M.S.; Zhou, J.; Lu, J.J. Improving Multi-Tumor Biomarker Health Check-Up Tests with Machine Learning Algorithms. Cancers 2020, 12, 1442. [Google Scholar] [CrossRef] [PubMed]
- Gaidano, V.; Tenace, V.; Santoro, N.; Varvello, S.; Cignetti, A.; Prato, G.; Saglio, G.; De Rosa, G.; Geuna, M. A Clinically Applicable Approach to the Classification of B-Cell Non-Hodgkin Lymphomas with Flow Cytometry and Machine Learning. Cancers 2020, 12, 1684. [Google Scholar] [CrossRef]
- Zhu, W.; Xie, L.; Han, J.; Guo, X. The Application of Deep Learning in Cancer Prognosis Prediction. Cancers 2020, 12, 603. [Google Scholar] [CrossRef]
- Betrouni, N.; Delval, A.; Chaton, L.; Defebvre, L.; Duits, A.; Moonen, A.; Leentjens, A.; Dujardin, K. Electroencephalography-based machine learning for cognitive profiling in Parkinson’s disease: Preliminary results. Mov. Disord. 2018, 34. [Google Scholar] [CrossRef]
- Wan, K.R.; Maszczyk, T.; See, A.A.Q.; Dauwels, J.; King, N.K.K. A review on microelectrode recording selection of features for machine learning in deep brain stimulation surgery for Parkinson’s disease. Clin. Neurophysiol. 2019, 130, 145–154. [Google Scholar] [CrossRef]
- Kautzky, A.; Seiger, R.; Hahn, A.; Fischer, P.; Krampla, W.; Kasper, S.; Kovacs, G.G.; Lanzenberger, R. Prediction of Autopsy Verified Neuropathological Change of Alzheimer’s Disease Using Machine Learning and MRI. Front. Aging Neurosci. 2018, 10. [Google Scholar] [CrossRef]
- Dogan, M.V.; Beach, S.R.H.; Simons, R.L.; Lendasse, A.; Penaluna, B.; Philibert, R.A. Blood-Based Biomarkers for Predicting the Risk for Five-Year Incident Coronary Heart Disease in the Framingham Heart Study via Machine Learning. Genes 2018, 9, 641. [Google Scholar] [CrossRef] [PubMed]
- Kim, T.; Heo, J.; Jang, D.K.; Sunwoo, L.; Kim, J.; Lee, K.; Kang, S.H.; Park, S.J.; Kwon, O.K.; Oh, C. Machine learning for detecting moyamoya disease in plain skull radiography using a convolutional neural network. EBioMedicine 2018. [Google Scholar] [CrossRef]
- Dongping, L. Automatic Detection of Cardiovascular Disease Using Deep Kernel Extreme Learning Machine. Biomed. Eng. Appl. Basis Commun. 2018, 30, 1850038. [Google Scholar] [CrossRef]
- Kannan, R.; Vasanthi, V. Machine Learning Algorithms with ROC Curve for Predicting and Diagnosing the Heart Disease. In Soft Computing and Medical Bioinformatics; Springer: Berlin, Germany, 2019; pp. 63–72. [Google Scholar] [CrossRef]
- Dimopoulos, A.; Nikolaidou, M.; Caballero, F.F.; Engchuan, W.; Sanchez-Niubo, A.; Arndt, H.; Ayuso-Mateos, J.; Haro, J.M.; Chatterji, S.; Georgousopoulou, E.; et al. Machine learning methodologies versus cardiovascular risk scores, in predicting disease risk. BMC Med. Res. Methodol. 2018, 18. [Google Scholar] [CrossRef]
- Wu, C.C.; Yeh, W.C.; Hsu, W.D.; Islam, M.M.; Nguyen, P.A.; Poly, T.N.; Wang, Y.C.; Yang, H.C.; Li, Y.C. Prediction of fatty liver disease using machine learning algorithms. Comput. Methods Programs Biomed. 2019, 170, 23–29. [Google Scholar] [CrossRef] [PubMed]
- Canbay, A.; Kälsch, J.; Neumann, U.; Rau, M.; Hohenester, S.; Baba, H.; Rust, C.; Geier, A.; Heider, D.; Sowa, J.P. Non-invasive assessment of NAFLD as systemic disease—A machine learning perspective. PLoS ONE 2019, 14, e0214436. [Google Scholar] [CrossRef] [PubMed]
- Rahman, T.; Siddiqua, S.; Rabby, S.; Hasan, N.; Imam, M. Early Detection of Kidney Disease Using ECG Signals through Machine Learning Based Modelling. In Proceedings of the International Conference on Robotics, Electrical and Signal Processing Techniques (ICREST), Dhaka, Bangladesh, 10–12 January 2019. [Google Scholar] [CrossRef]
- Danter, W. DeepNEU: Cellular reprogramming comes of age—A machine learning platform with application to rare diseases research. Orphanet J. Rare Dis. 2019, 14. [Google Scholar] [CrossRef] [PubMed]
- Jia, J.; Wang, R.; An, Z.; Guo, Y.; Ni, X.L.; Shi, T. RDAD: A Machine Learning System to Support Phenotype-Based Rare Disease Diagnosis. Front. Genet. 2018, 9. [Google Scholar] [CrossRef]
- Olden, J.D.; Lawler, J.J.; Poff, N.L. Machine learning methods without tears: A primer for ecologists. Q. Rev. Biol. 2008, 83, 171–193. [Google Scholar] [CrossRef]
- Wang, H.; Raj, B. A survey: Time travel in deep learning space: An introduction to deep learning models and how deep learning models evolved from the initial ideas. arXiv 2015, arXiv:1510.04781. [Google Scholar]
- Eerman, J.; Mahanti, A.; Arlitt, M. Internet traffic identification using machine learning techniques. In Proceedings of the 49th IEEE Global Telecommunications Conference (GLOBECOM), San Francisco, CA, USA, 27 November–1 December 2006; pp. 1–6. [Google Scholar] [CrossRef]
- Tu, J.V. Advantages and disadvantages of using artificial neural networks versus logistic regression for predicting medical outcomes. J. Clin. Epidemiol. 1996, 49, 1225–1231. [Google Scholar] [CrossRef]
- Rajkomar, A.; Oren, E.; Chen, K.; Dai, A.; Hajaj, N.; Liu, P.; Liu, X.; Sun, M.; Sundberg, P.; Yee, H.; et al. Scalable and accurate deep learning for electronic health records. NPJ Digit. Med. 2018, 1. [Google Scholar] [CrossRef] [PubMed]
- Kelly, C.; Karthikesalingam, A.; Suleyman, M.; Corrado, G.; King, D. Key challenges for delivering clinical impact with artificial intelligence. BMC Med. 2019, 17. [Google Scholar] [CrossRef] [PubMed]
| Patient | Head Ache | Muscle Ache | Fever | Flu |
|---|---|---|---|---|
| P1 | Yes | No | Yes | Yes |
| P2 | No | Yes | Yes | Yes |
| P3 | Yes | No | Yes | No |
| P4 | Yes | Yes | Yes | Yes |
| P5 | No | Yes | Yes | Yes |
| P6 | No | Yes | No | No |
| Area | Author | Goal | Algorithm |
|---|---|---|---|
| Metabolic diseases | [74,79] | Clustering | K-means Clustering |
| [87] | Clustering | DBSCAN | |
| [171] | Regression | Random Forest | |
| [100] | Classification | SVM | |
| [106,109] | Classification | ID3 | |
| [115,116,118,120,122] | Classification | KNN | |
| [135] | Classification | Naïve Bayes | |
| [137,143] | Classification | Bayesian Networks | |
| [145] | Regression | Linear regression | |
| Cancer | [75,81] | Clustering | K-means Clustering |
| [84,86] | Clustering | DBSCAN | |
| [24] | Clustering | SNF | |
| [25] | Clustering | PINS | |
| [26] | Clustering | CIMLR | |
| [95,172] | Classification | SVM | |
| [108] | Classification | ID3 | |
| [130] | Classification | Naïve Bayes | |
| [136] | Classification | Bayesian Networks | |
| [148,173] | Regression | Linear regression | |
| [146,174] | Regression | Logistic regression | |
| [157] | Classification | Neural Networks + KNN | |
| [156] | Classification | Neural Networks + SVM | |
| [160] | Classification | Neural Networks + ID3 | |
| [161] | Classification | KNN | |
| [175] | Classification | DT | |
| [176] | Classification | DL | |
| Parkinson’s disease | [77,80] | Clustering | K-means Clustering |
| [91] | Clustering | SOMS | |
| [177] | Classification | KNN + SVM | |
| [117,124,125] | Classification | KNN | |
| [134] | Classification | Naïve Bayes | |
| [141] | Classification | Bayesian Networks | |
| [152] | Regression | Linear regression | |
| [125,152] | Regression | Logistic regression | |
| [155,159] | Classification | Neural Networks + SVM | |
| [125,158] | Classification | Neural Networks + KNN | |
| Alzheimer’s diseases | [83] | Clustering | K-means Clustering |
| [89,90] | Clustering | DBSCAN | |
| [92,93,94] | Clustering | SOMS | |
| [119,124] | Classification | KNN | |
| [132] | Classification | Naïve Bayes | |
| [138,139,140] | Classification | Bayesian Networks | |
| [155] | Classification | Neural Networks + SVM | |
| [157] | Classification | Neural Networks + KNN | |
| [167] | Classification | SHIMR | |
| [178] | Classification | Naïve Bayes + SVM + RF | |
| [179] | Classification | RF | |
| Heart and vascular diseases | [180] | Classification | RF |
| [96,97] | Classification | SVM | |
| [110,114] | Classification | ID3 | |
| [115] | Classification | KNN | |
| [126,127,128] | Classification | Naïve Bayes | |
| [142] | Classification | Bayesian Networks | |
| [148] | Regression | Linear regression | |
| [181,182] | Classification | DL | |
| [183] | Regression | Gradient boosting | |
| [184] | Classification | KNN + RF + DT | |
| Hepatic diseases | [99] | Classification | SVM |
| [113] | Classification | ID3 | |
| [185] | Regression | Linear regression | |
| [115] | Classification | KNN | |
| [129,185] | Classification | Naïve Bayes | |
| [186] | Classification | Ensemble Feature Selection | |
| [170] | Classification | Cross-sectional models | |
| Infectious diseases | [78,82] | Clustering | K-means Clustering |
| [85] | Clustering | DBSCAN | |
| [72,98,101,102,103,104,105] | Classification | SVM | |
| [107,111] | Classification | ID3 | |
| [72,121,123] | Classification | KNN | |
| [133] | Classification | Naïve Bayes | |
| [71,147,148,149,150,151,153,154] | Regression | Linear regression | |
| [151,154] | Regression | Logistic regression | |
| [71,156] | Classification | Neural Networks + SVM | |
| [165] | Classification | Random Forest | |
| Renal diseases | [112] | Classification | ID3 |
| [115] | Classification | KNN | |
| [129] | Classification | Naïve Bayes | |
| [144,148] | Regression | Linear regression | |
| [144] | Regression | Logistic regression | |
| [162] | Classification | Neural Networks + Naïve Bayes | |
| [162,163] | Classification | Neural Networks + SVM | |
| [187] | Classification | SVM | |
| Other diseases | |||
| Vision | [164] | Classification | Neural Networks |
| Digestive | [166] | Regression | RF |
| Cutaneous | [168] | Regression | GloVe |
| Respiratory | [169] | Classification | SVM + RF |
| Rare | [188,189] | Classification | KNN + RF + NB + DL |
| Method | Advantages | Disadvantages |
|---|---|---|
| ML | Algorithms are often easy to be implemented. Algorithms are flexible enough to handle complex problems with multiple interacting variables. Input and output are not necessarily fixed. | Complex relationships between dependent and independent variables are not identified easily in high-dimensional databases. High computational cost. |
| DL | Complex relationships between dependent and independent variables are identified easily in high-dimensional databases. Ability to handle databases with high noise. | Input and output are fixed. Overfitting problem possibility is high. Implementation is not so easy than in ML. Training process requires a higher computational cost than ML. |
| Method | Advantages | Disadvantages |
|---|---|---|
| Unsupervised learning | It does not require a training data to be labelled. The automatic labelling of the training data set saving the time spent in hand classification. Classification task is fast. | There are no notions of the output along the learning process. It does not allow to estimate or map the results of a new sample. Results vary considerably in the presence of outliers. It only performs classification tasks. |
| Supervised learning | It exists notions of the output along the learning process. It performs classification and regression tasks. It allows estimating or mapping the results to a new sample. | It requires a labelled data set. It requires a training process. |
| Method | Advantages | Disadvantages |
|---|---|---|
| K-means Clustering | Simple clustering approach. Efficient clustering method. Method is easy to be implemented. | Requires a number of clusters in advance. Handling categorical attributes cause problems. Results vary considerably in the presence of outliers. |
| DBSCAN/SOMS | Simple clustering approach. A number of clusters in advance is not required. Efficient clustering method. | Handling categorical attributes cause problems. Results vary considerably in the presence of outliers. |
| SVM | Better accuracy compared to other classifiers. Overfitting problem is not so great as in other methods. | High computational cost. Training process requires more time than other methods. |
| ID3 | There are no domain requirements. Exact value results are provided for various actions, minimising the ambiguity of complex decisions. High dimensional databases are processed easily. Classifier and output are easy to be interpreted. | Results are restricted to one output attribute. Only categorical output is generated. Classifier performance depends on the type of dataset, making it unstable. |
| KNN | Method is easy to be implemented. Training process requires low computational cost. | Large storage space is required. Sensitivity to databases with high noise. Testing process requires high computational cost. |
| Naïve Bayes Bayesian Networks | Method is easy to be implemented. Method is speeder and provide more accuracy in high dimensional databases than other methods. | Low accuracy is provided in cases where exists dependence between variables. |
| Linear regression | Better accuracy compared to other classifiers. Complex relationships between dependent and independent variables are identified easily. | Results vary considerably in the presence of outliers. Training process requires more time than other methods. Classifier performance depends on the type of dataset, making it unstable. Only numerical output is generated. |
| Logistic regression | Better accuracy compared to other classifiers. Complex relationships between dependent and independent variables are identified easily. | Results vary considerably in the presence of outliers. Training process requires more time than other methods. Classifier performance depends on the type of dataset, making it unstable. Only categorical output is generated. |
| Neural network | Complex relationships between dependent and independent variables are identified easily. Ability to handle databases with high noise. A previous feature extraction task is not required. | High possibility of local minima. High possibility of overfitting problem. Classifier is difficult to be interpreted. High computational time is required if there is a large number of layers. No explanation or justification of decisions can be given, i.e., a “black box” characteristic. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Caballé-Cervigón, N.; Castillo-Sequera, J.L.; Gómez-Pulido, J.A.; Gómez-Pulido, J.M.; Polo-Luque, M.L. Machine Learning Applied to Diagnosis of Human Diseases: A Systematic Review. Appl. Sci. 2020, 10, 5135. https://doi.org/10.3390/app10155135
Caballé-Cervigón N, Castillo-Sequera JL, Gómez-Pulido JA, Gómez-Pulido JM, Polo-Luque ML. Machine Learning Applied to Diagnosis of Human Diseases: A Systematic Review. Applied Sciences. 2020; 10(15):5135. https://doi.org/10.3390/app10155135
Chicago/Turabian StyleCaballé-Cervigón, Nuria, José L. Castillo-Sequera, Juan A. Gómez-Pulido, José M. Gómez-Pulido, and María L. Polo-Luque. 2020. "Machine Learning Applied to Diagnosis of Human Diseases: A Systematic Review" Applied Sciences 10, no. 15: 5135. https://doi.org/10.3390/app10155135
APA StyleCaballé-Cervigón, N., Castillo-Sequera, J. L., Gómez-Pulido, J. A., Gómez-Pulido, J. M., & Polo-Luque, M. L. (2020). Machine Learning Applied to Diagnosis of Human Diseases: A Systematic Review. Applied Sciences, 10(15), 5135. https://doi.org/10.3390/app10155135







