Deep Learning System for COVID-19 Diagnosis Aid Using X-ray Pulmonary Images
Abstract
Featured Application
Abstract
1. Introduction
2. Materials and Methods
2.1. Dataset
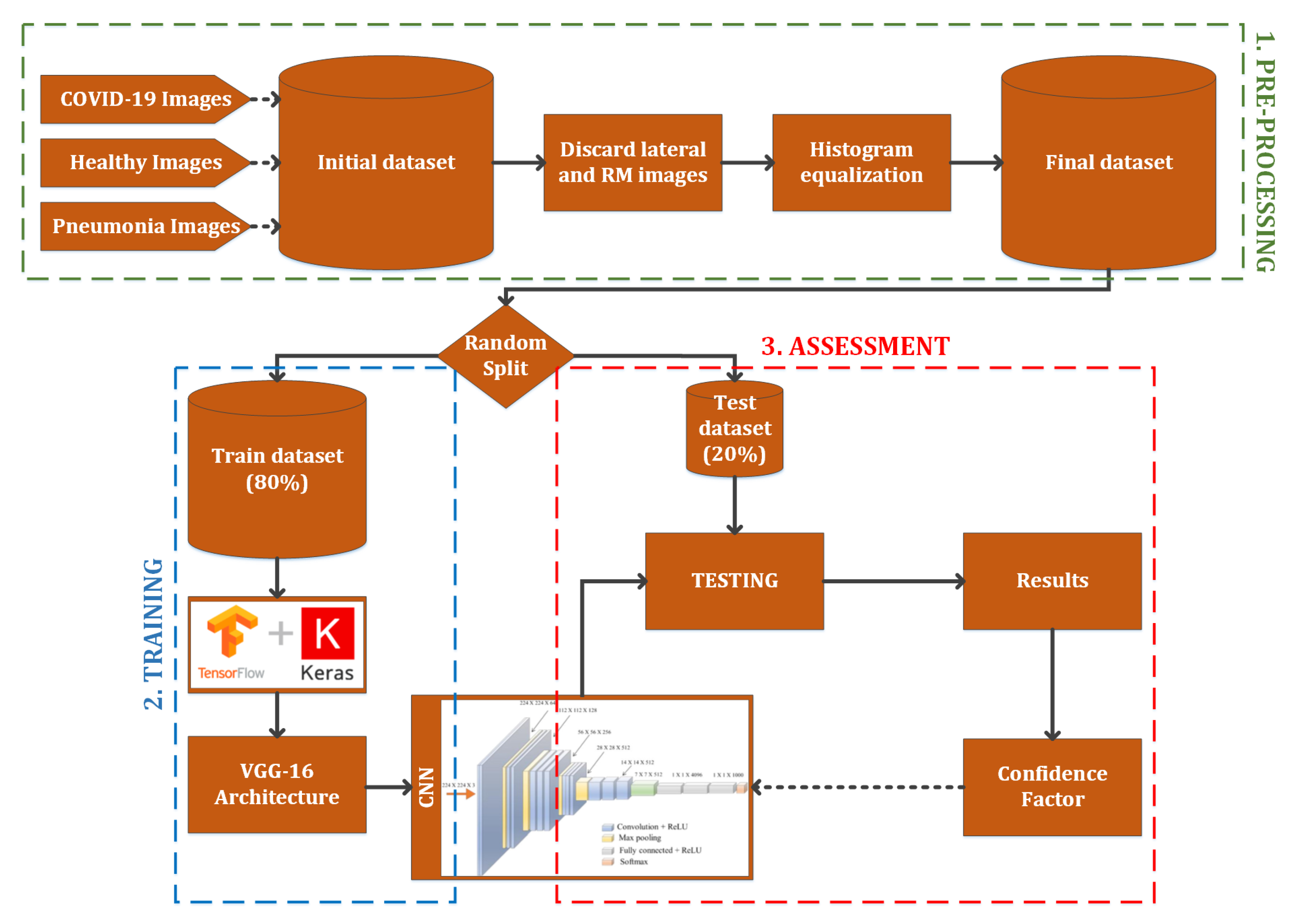
2.2. Processing Architecture
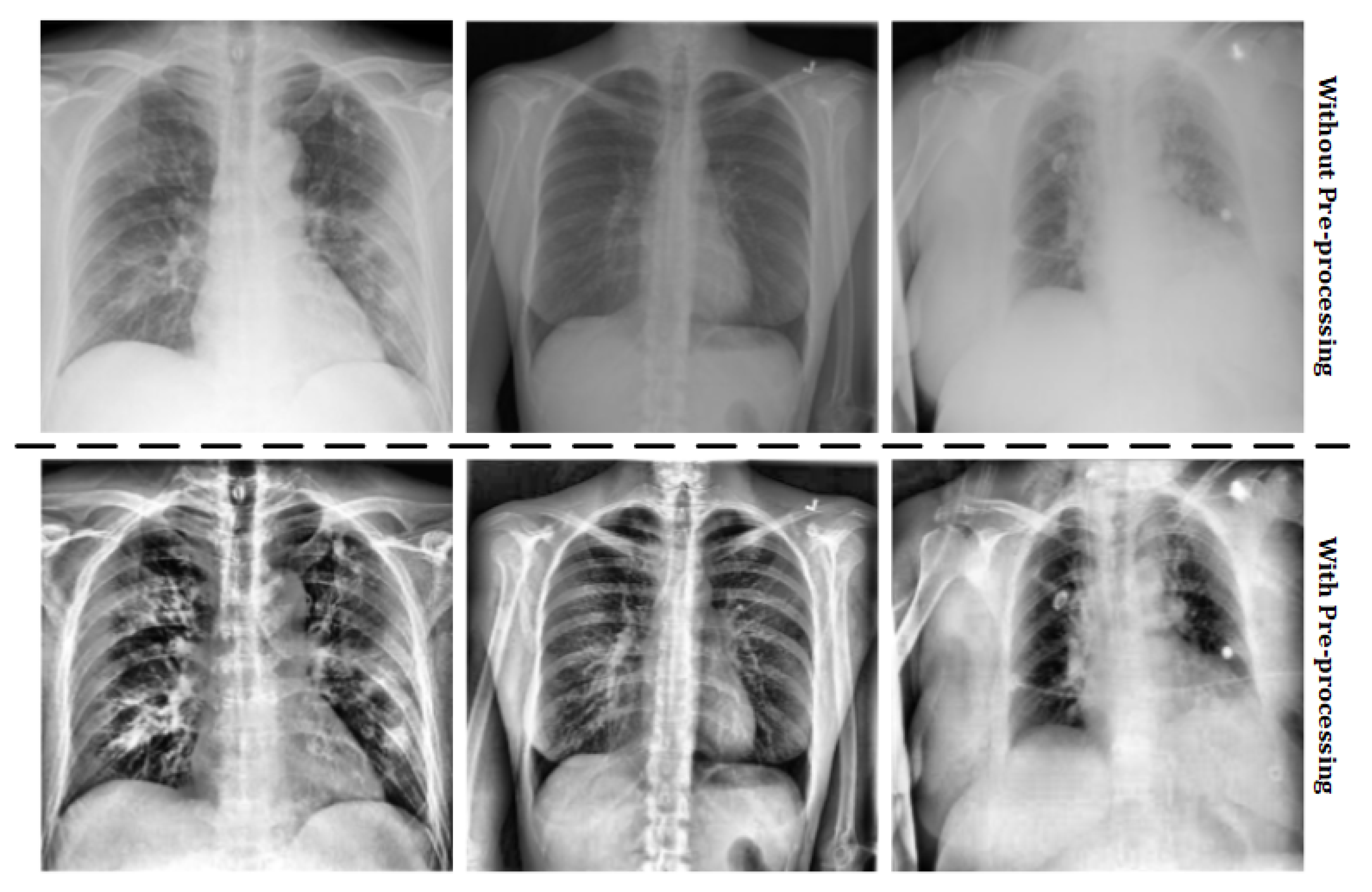
- Pre-processing: the images stored in the original dataset contains lung X-ray images of healthy patients, patients with pneumonia and COVID-19 positives. However, some images of the COVID-19 positive cases were not obtained with the same parameters as detailed above, so these images must not be taken into account. Moreover, in order to work with images of the same characteristics, an histogram equalization is applied. These two treatments compose the pre-processing stage. The results of the pre-processing step can be observed in Figure 2.
- Training: using TensorFlow framework with Keras, a VGG-16 architecture [22] is implemented and combined with a final inference layer to train a classification system with three classes (healthy, pneumonia and COVID-19). The output of this stage is the convolutional neural network model.
- Assessment: after the model is obtained, the testing dataset is used to evaluate the classification effectiveness, obtaining a confidence factor. This one is used to analyze the CNN performance in order to evaluate the usefulness as a diagnostic tool.
3. Results and Discussion
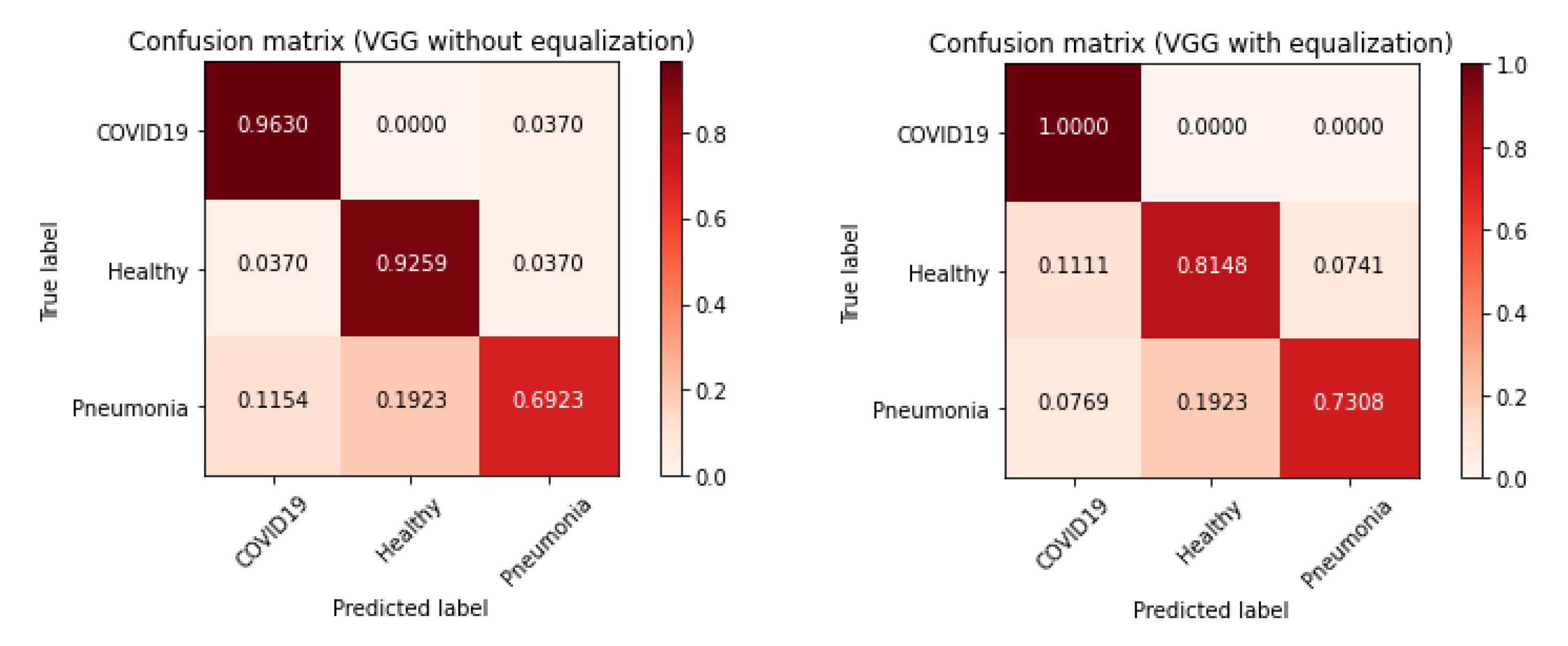
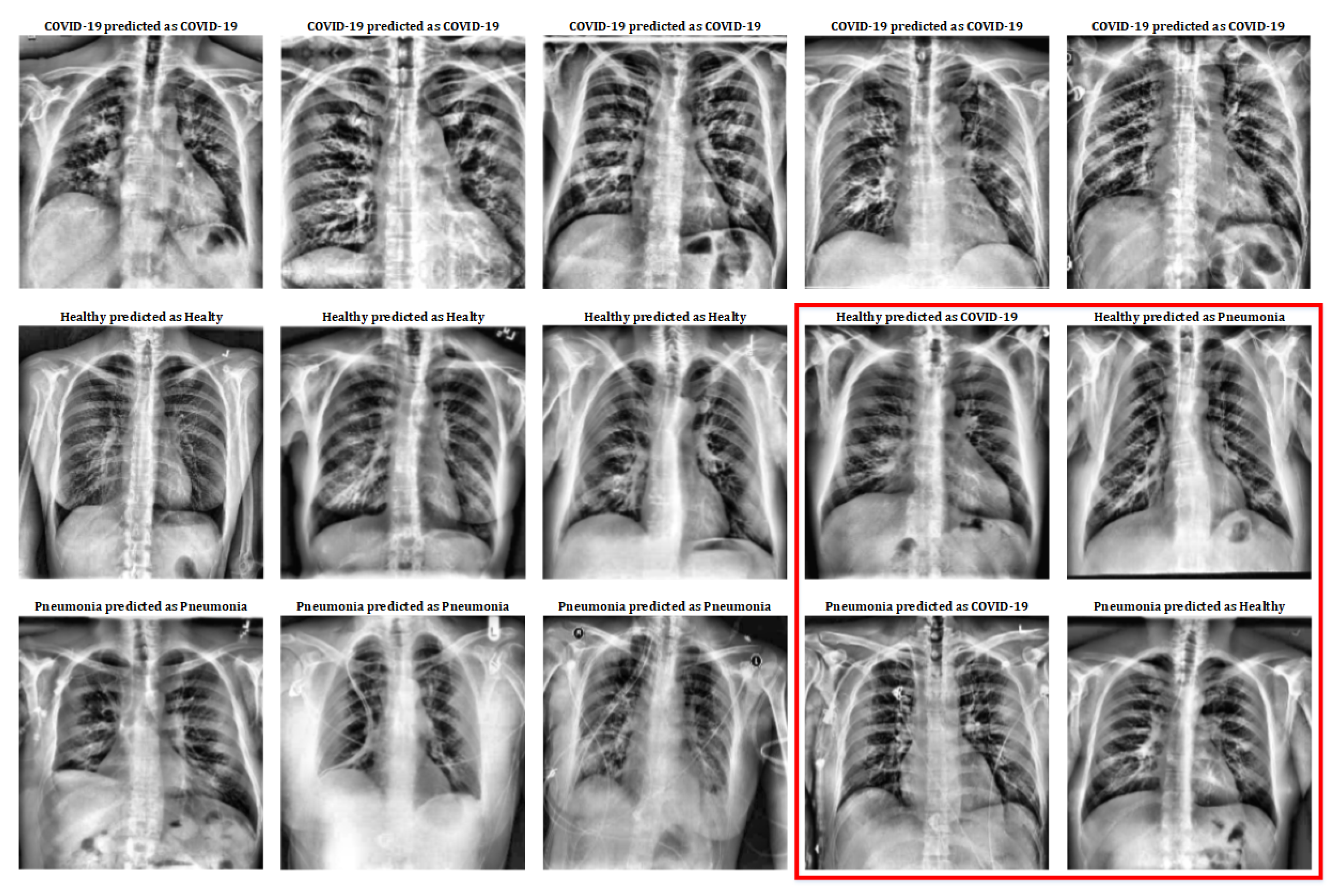
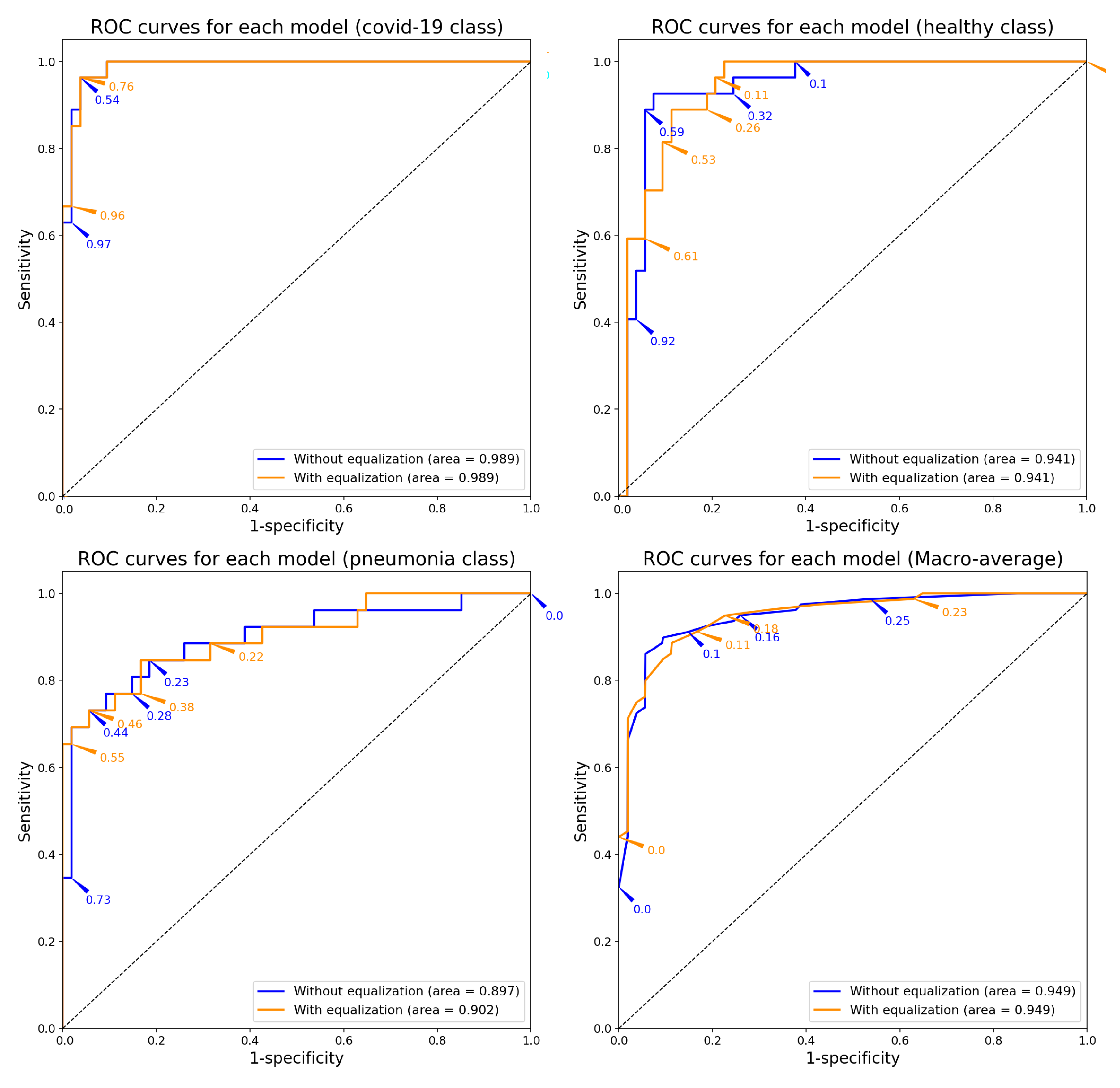
Effectiveness Results
4. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| SARS-CoV | Severe Acute Respiratory Syndrome Coronavirus |
| COVID-19 | Coronavirus Disease 2019 |
| RT-PCR | Reverse Transcription Polymerase Chain Reaction |
| BAL | Bronchoalveolar Lavage |
| MR | Magnetic Resonance |
| CT | Computerized Tomography |
| VGG-16 | Visual Geometry Group 16 |
| ROC | Receiver Operating Characteristic |
| AUC | Area Under Curve |
References
- Repici, A.; Maselli, R.; Colombo, M.; Gabbiadini, R.; Spadaccini, M.; Anderloni, A.; Carrara, S.; Fugazza, A.; Di Leo, M.; Galtieri, P.A.; et al. Coronavirus (COVID-19) outbreak: What the department of endoscopy should know. Gastrointest. Endosc. 2020, 92, 192–197. [Google Scholar] [CrossRef] [PubMed]
- Dong, E.; Du, H.; Gardner, L. An interactive web-based dashboard to track COVID-19 in real time. Lancet Infect. Dis. 2020, 20, P533–P534. [Google Scholar] [CrossRef]
- Sohrabi, C.; Alsafi, Z.; O’Neill, N.; Khan, M.; Kerwan, A.; Al-Jabir, A.; Iosifidis, C.; Agha, R. World Health Organization declares global emergency: A review of the 2019 novel coronavirus (COVID-19). Int. J. Surg. 2020, 76, 71–76. [Google Scholar] [CrossRef] [PubMed]
- Rothan, H.A.; Byrareddy, S.N. The epidemiology and pathogenesis of coronavirus disease (COVID-19) outbreak. J. Autoim. 2020, 109, 102433. [Google Scholar] [CrossRef] [PubMed]
- Li, Q.; Guan, X.; Wu, P.; Wang, X.; Zhou, L.; Tong, Y.; Ren, R.; Leung, K.S.; Lau, E.H.; Wong, J.Y.; et al. Early transmission dynamics in Wuhan, China, of novel coronavirus–infected pneumonia. N. Engl. J. Med. 2020, 382, 1199–1207. [Google Scholar] [CrossRef] [PubMed]
- Lauer, S.A.; Grantz, K.H.; Bi, Q.; Jones, F.K.; Zheng, Q.; Meredith, H.R.; Azman, A.S.; Reich, N.G.; Lessler, J. The incubation period of coronavirus disease 2019 (COVID-19) from publicly reported confirmed cases: Estimation and application. Ann. Intern. Med. 2020, 172, 577–582. [Google Scholar] [CrossRef] [PubMed]
- Cascella, M.; Rajnik, M.; Cuomo, A.; Dulebohn, S.C.; Di Napoli, R. Features, evaluation and treatment coronavirus (COVID-19). In Statpearls [Internet]; StatPearls Publishing: Tampa, FL, USA, 2020. [Google Scholar]
- World Health Organization. Modes of Transmission of Virus Causing COVID-19: Implications for IPC Precaution Recommendations; Scientific Brief, 27 March 2020; Technical Report; World Health Organization: Geneva, Switzerland, 2020. [Google Scholar]
- Xu, Z.; Shi, L.; Wang, Y.; Zhang, J.; Huang, L.; Zhang, C.; Liu, S.; Zhao, P.; Liu, H.; Zhu, L.; et al. Pathological findings of COVID-19 associated with acute respiratory distress syndrome. Lancet Respir. Med. 2020, 8, 420–422. [Google Scholar] [CrossRef]
- Lan, L.; Xu, D.; Ye, G.; Xia, C.; Wang, S.; Li, Y.; Xu, H. Positive RT-PCR test results in patients recovered from COVID-19. JAMA 2020, 323, 1502–1503. [Google Scholar] [CrossRef] [PubMed]
- Serebriakova, O.; Romanova, E.; Govorin, A.; Filev, A.; Shangin, A. Peculiarities of clinical and X-ray picture of pneumonia in patients with influenza A (H1N1). Klin. Med. 2012, 90, 70–72. [Google Scholar]
- Lin, Z.Q.; Xu, X.Q.; Zhang, K.B.; Zhuang, Z.G.; Liu, X.S.; Zhao, L.Q.; Lin, C.Y.; Li, Y.; Hua, X.L.; Zhao, H.L.; et al. Chest X-ray and CT findings of early H7N9 avian influenza cases. Acta Radiol. 2015, 56, 552–556. [Google Scholar] [CrossRef] [PubMed]
- Tse, G.M.; To, K.; Chan, P.K.; Lo, A.; Ng, K.; Wu, A.; Lee, N.; Wong, H.; Mak, S.; Chan, K.; et al. Pulmonary pathological features in coronavirus associated severe acute respiratory syndrome (SARS). J. Clin. Pathol. 2004, 57, 260–265. [Google Scholar] [CrossRef] [PubMed]
- Xie, X.; Li, X.; Wan, S.; Gong, Y. Mining X-ray Images of SARS Patients; Data Mining; Springer: Berlin, Germany, 2006; pp. 282–294. [Google Scholar]
- Ker, J.; Wang, L.; Rao, J.; Lim, T. Deep learning applications in medical image analysis. IEEE Access 2017, 6, 9375–9389. [Google Scholar] [CrossRef]
- Luna-Perejón, F.; Domínguez-Morales, M.J.; Civit-Balcells, A. Wearable fall detector using recurrent neural networks. Sensors 2019, 19, 4885. [Google Scholar] [CrossRef] [PubMed]
- Domínguez-Morales, M.J.; Luna-Perejón, F.; Miró-Amarante, L.; Hernández-Velázquez, M.; Sevillano-Ramos, J.L. Smart Footwear Insole for Recognition of Foot Pronation and Supination Using Neural Networks. Appl. Sci. 2019, 9, 3970. [Google Scholar] [CrossRef]
- Litjens, G.; Kooi, T.; Bejnordi, B.E.; Setio, A.A.A.; Ciompi, F.; Ghafoorian, M.; Van Der Laak, J.A.; Van Ginneken, B.; Sánchez, C.I. A survey on deep learning in medical image analysis. Med. Image Anal. 2017, 42, 60–88. [Google Scholar] [CrossRef] [PubMed]
- Asri, H.; Mousannif, H.; Al Moatassime, H.; Noel, T. Using machine learning algorithms for breast cancer risk prediction and diagnosis. Procedia Comp. Sci. 2016, 83, 1064–1069. [Google Scholar] [CrossRef]
- Jhuo, S.L.; Hsieh, M.T.; Weng, T.C.; Chen, M.J.; Yang, C.M.; Yeh, C.H. Trend prediction of influenza and the associated pneumonia in taiwan using machine learning. In 2019 International Symposium on Intelligent Signal Processing and Communication Systems (ISPACS); IEEE: Piscataway, NJ, USA, 2019; pp. 1–2. [Google Scholar]
- Civit-Masot, J.; Luna-Perejon, F.; Vicente-Diaz, S.; Corral, J.M.R.; Civit, A. TPU Cloud-Based Generalized U-Net for Eye Fundus Image Segmentation. IEEE Access 2019, 7, 142379–142387. [Google Scholar] [CrossRef]
- Simonyan, K.; Zisserman, A. Very deep convolutional networks for large-scale image recognition. arXiv 2014, arXiv:1409.1556. [Google Scholar]
- Jaeger, S.; Antani, S.; Thoma, G. Tuberculosis screening of chest radiographs. SPIE Newsroom 2011, 3, 89–99. [Google Scholar] [CrossRef]
| Subset | COVID-19 | Healthy | Pneumonia | Total |
|---|---|---|---|---|
| Total | 132 | 132 | 132 | 396 |
| Training | 105 | 105 | 106 | 316 |
| Test | 27 | 27 | 26 | 80 |
| Model | Accuracy | Precision | F1-Score | Specificity | Sensitivity |
|---|---|---|---|---|---|
| Original | 0.86 | 0.86 | 0.86 | 0.93 | 0.86 |
| Equalized | 0.85 | 0.85 | 0.85 | 0.92 | 0.85 |
| Class | Precision | Recall | F1-Score |
|---|---|---|---|
| COVID-19 | 0.87 | 0.96 | 0.91 |
| Healthy | 0.83 | 0.93 | 0.88 |
| Pneumonia | 0.90 | 0.69 | 0.78 |
| Class | Precision | Recall | F1-Score |
|---|---|---|---|
| COVID-19 | 0.84 | 1.00 | 0.92 |
| Healthy | 0.81 | 0.81 | 0.81 |
| Pneumonia | 0.90 | 0.73 | 0.81 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Civit-Masot, J.; Luna-Perejón, F.; Domínguez Morales, M.; Civit, A. Deep Learning System for COVID-19 Diagnosis Aid Using X-ray Pulmonary Images. Appl. Sci. 2020, 10, 4640. https://doi.org/10.3390/app10134640
Civit-Masot J, Luna-Perejón F, Domínguez Morales M, Civit A. Deep Learning System for COVID-19 Diagnosis Aid Using X-ray Pulmonary Images. Applied Sciences. 2020; 10(13):4640. https://doi.org/10.3390/app10134640
Chicago/Turabian StyleCivit-Masot, Javier, Francisco Luna-Perejón, Manuel Domínguez Morales, and Anton Civit. 2020. "Deep Learning System for COVID-19 Diagnosis Aid Using X-ray Pulmonary Images" Applied Sciences 10, no. 13: 4640. https://doi.org/10.3390/app10134640
APA StyleCivit-Masot, J., Luna-Perejón, F., Domínguez Morales, M., & Civit, A. (2020). Deep Learning System for COVID-19 Diagnosis Aid Using X-ray Pulmonary Images. Applied Sciences, 10(13), 4640. https://doi.org/10.3390/app10134640








