1. Introduction
There are many factors that can caused by voice pathologies. Some of include infections of voice tissue, tiredness, environmental changes, muscular dystrophy, face soreness and others [
1]. The voice pathology has a negative impact on vibration regularity and voice functionality, which leads to an increase in vocal noise. The normal voice turned to be tense, weak and hoarse [
2] that affects the quality of voice [
3]. To date, the current vocal pathology detection methods have a biased evaluation based on subjective matters [
4]. An example of the subjective evaluation is auditory-perceptual assessment in hospitals, which is widely applied by visual laryngostroboscopy assessment [
5]. Several clinical examinations are applied for auditory-perceptual parameters to scale the rate of severity diagnosis [
6]. However, those evaluation methods are subject to parameter sensitivity and are also time consuming and laborious [
7]. In addition, these methods require a physical patient examination in the clinic which could be difficult for patients with severe conditions. An example of objective evaluation is using a computer-aided tool to identify and analyse vocal signals without any surgical intervention. The automatic detection can also recognize inaudible sounds [
1].
These evaluation methods are not subjective as they do not depend on a human decision. Besides, they are easy to apply since the voice recordings can be made available remotely via different internet recoding applications. Therefore, some studies such as [
8] have developed vocal processing methods to determine the vocal pathology aspects can be effectively combined with a machine learning method to detect the voice pathology automatically in one framework to accurately distinguish healthy people from people with voice pathologies. In the literature, various voice pathology databases have been widely applied for the objective evaluation of voice pathology. The most common voice pathology databases are the Saarbruecken Voice Database (SVD) [
9], Arabic Voice Pathology Database (AVPD) [
10] and the Massachusetts Eye and Ear Infirmary Database (MEEI) [
11]. The vocalization of the vowel /a/. [
1,
11] is available in many language databases [
2], therefore, it is commonly analysed by researchers. Other combinations of vowels are also analyzed by researchers [
1,
12]. Notably, the majority of the researchers in the voice pathologies community have limited the datasets to specific pathologies sets [
12].
Usually, the clinical interpretation of vocal features is conducted before the process of pathology detection [
13]. Examples of vocal features are glottal-to-noise excitation ratio (GNE) [
14], Mel frequency cepstral coefficients (MFCC) [
15], multidimensional voice program parameters (MDVP) [
16] and many others. See [
17] for more details about the settings of speech pathologies. Once the vocal features are extracted, many conventional classification methods are applied for voice pathology detection. For detection purposes, most studies have used Random Forests (RF), Artificial Neural Networks (ANN) [
18], Gaussian Mixture Models (GMM), Support Vector Machines (SVM) and other classifiers [
19,
20]. It is observed that the study results show notable differences. Because of the different set selection, a sample of voice pathology, a vocal feature and classification method are applied in the experiments. This drives us to the following conclusions: most works analyse a single speech task, mainly the sustained phonation of the vowel /a/ (language-independent speech task)
The majority of studies focus only on analyzing a single voice segment. In particular the vocalization of the vowel /a/ in the independent language speech task.
The most analysis is conducted on limited acoustic pathologies from the SVD, AVPD and MEEI databases.
The conventional dysphonic feature is the feature most extracted to determine the voice aspect for a particular voice pathology.
Artificial Neural Network (ANN), Random Forest (RF) and Support Vector Machine (SVM) are the most conventional machine learning methods employed for vocal-pathologies detection.
In this paper, we attempt to make a comparative analysis with published results on the speech recordings of the vocalization of the vowel /a/. In spite of other studies, the comparison analysis will cover a bigger segment of SVD [
21]. In order to widen the problem scope and maximize the generality of application of the proposed method, we will not limit the vocal pathologies in the database to the popular subtest that is commonly used in the literature. Therefore, a big number of voice pathologies with minimum voice recordings will be included in our dataset for this study. As far as we are concerned, no study presented in the literature [
22], is based on a deep learning method for detection of vocal pathologies. In this study, a conventional voice pathology detection method is used and combined with a vocal feature selection method. We also will employ a gradient boosting method as a classifier. An investigation of anomaly detection methods usage is also conducted in this study to manage the wide distribution of vocal pathologies that are associated with a limited number of voice phytology recordings. In this study, we propose automatic rapid voice pathology detection based on a deep learning classifier, namely a DNN system for voice pathology detection. Our proposed methods are applied through four primary phases, including preparation of dataset following by learning process phase then a training and validation phase and finally an inference processing phase.
This paper is organized as follows: An overview of currents studies and some related works are presented in
Section 2. In
Section 3, we describe our proposed voice pathology detection based on deep learning. We present the experimental results in
Section 4. Finally, we present our conclusions and directions of future research in
Section 5.
2. Related Work
The utilization of machine learning (ML) can be useful in many applications such as medical diagnosis [
23], cancer detection [
24], smart building applications [
25], and others [
1,
11,
26]. Machine learning methods are valuable for discriminatory detection and classification tasks [
27,
28]. These methods have been used in diverse speech identification uses, where one of these uses is pathological voice investigation [
29]. The identification and the classification of voice pathology techniques are still one of the difficult domains within the investigation of speech detection. Besides, these basic techniques are expensive and need more time and many sorts of gear [
30]. Many researchers focus on the Saarbrücken voice database (SVD) in their studies. The researchers that utilized SVD extracted different features from voice records prior to pathology identification. The features that are frequently extracted are entropy, energy, time, contained Mel-frequency cepstral coefficients (MFCC), cepstral domains, frequency, harmonics-to-noise ratio, short-term cepstral parameters, normalized noise energy, and others [
2,
31,
32,
33]. After this stage the classification task will begin. Many binary and multi-classification methods have been used such K-means clustering, Support Vector Machine, and so on. To our best knowledge, our study is the first study to present voice pathology detection and classification using a convolutional neural network (CNN).
The outcomes of the published studies vary greatly because of the variances among the datasets used in the experimental results. According to Martınez et al. [
34], the accuracy achieved utilizing 200 records of sustained vowel /a/ represent a high value and it’s very close to our study. Other studies utilized the combination of vowels /a/, /i/ and /u/ to get high accuracy and do not focus on the pathology causes. In the studies by Souissi et al. in [
35] they achieved high accuracy of 87.82% utilizing subset involving four kinds of voice pathologies that include 71 types. Also, Al-Nasheri et al. [
16,
36] achieved an accuracy of 99.68% due to their use of a subset involving a few of the pathologies to conduct a test on information that was moreover displayed in other accessible datasets, such as Arabic Voice Pathology Database (AVPD), and Massachusetts Eye and Ear Infirmary Database (MEEI). Another study conducted by Muhammad et al. [
13] utilized a subset involving three kinds of voice pathologies that achieved an accuracy of 93.20%. In addition, they utilized a combination of voice records as an electroglottograph signal to increase the accuracy to 99.98%. However, in another study conducted by Hemmerling et al. [
37] they achieved a high accuracy of 100% in the detection issue by their method to separate male and female speakers.
The study of Hammami et al. [
38] assessed the execution of the proposed high order statistic feature highlights extricated from wavelet space to segregate between normal voices and pathological ones. Traditional features such as Cruel Wavelet Esteem, Cruel Wavelet Vitality and Cruel Wavelet Entropy were used in the experiments. These highlights, combined with a SVM classifier, reach the most elevated correctness of 99.26% within the location step and 100% when classifying the information. In order to include concrete logical included values a clinical evaluation was performed on information collected from subjects from a healing center in Tunez. The results were acceptable and the precisions were 94.82% and 94.44% for the location and classification, respectively. Fonseca et al. [
39] worked on the discovery of co-existent laryngeal issues for which the major phonic side effect is the same, creating features with noteworthy inter-class coverage. Based on the combination of SE, ZCR and SH, all utilized for extraction, related with DPM, particularly received for classification, the proposed approach was effectively concluded, productively dealing within definitions and inconsistencies with an estimated precision of 95%. The ongoing challenge of dysphonia voice research is the small size of the database produced by Rueda and Krishnan [
40]. It is very complicated to use more advanced deep learning methods without underfitting or overfitting. They proposed an adaptive method utilized to break down a signal into its components employing a Fourier-based sychrosqueezing change (FSST) for information enlargement and change. The 2D TF representation output becomes the input to CNN.
It is clear that each voice disorder produces distinctive frequencies depending on the sort of voice disorder and its area on the vocal folds, as we observed. Thus, monitoring the frequency groups is exceptionally vital to evaluate which one contributes more to the discovery and classification of voice afflictions. For example, Pouchoulin et al. [
41] stated that lower frequencies (3000 Hz) are more reasonable for recognizing dysphonic voices than higher frequencies. Furthermore, Fraile et al. [
42] demonstrated that the control of dysphonic voice flags is altogether less steady within the recurrence area between 2000 and 6400 Hz than the other recurrence regions. To discuss the outcomes of a comparative literature review, they analysed voice records of maintained phonation of the vowel /a/ as well. However, in contrast to past studies, we analyze a bigger database collected from SVD [
1]. Moreover, to propose systems capable of powerful voice pathology detection and classification, we do not confine the database as if it were a subset of popular voice pathologies. In this study, the database includes an expansive number of pathologies with small recordings. As we observed in the related works, in spite of past work [
1], no other studies have utilized deep learning methods for voice pathology identification. In the following sections we utilize a robust voice pathology identification model based on the acoustic feature extraction strategy. We use voice pathology detection and identification utilizing a CNN approach. We utilize the transfer learning method for using the current powerful CNN models. Particularly, the ResNet34 models were used. To handle the issue of inadequate distribution of an assortment of voice pathologies with few recordings in the datasets, we also explore the utilization of abnormality detection methods.
3. Proposed Methodology
3.1. Dataset Used
As already stated, we have opted to use continuous vowel /a/ phonation as the base for our experiments. A speaker is asked to maintain vowel phoning during this specific speech task, to maintain the amplitude and frequency at a realistic rate [
21]. The benefit of this speech task is that it is free of articulative and other linguistic confusions compared with other language standard tasks such as reading or speaking activities. This uniqueness makes it an ideal alternative for this mission for building the large database required for supervised deep learning models [
43]. Thus, the only speech task used in this process is sustained /a/ vowel phoning.
The Saarbruecken Voice Database (SVD) is built based on 2000 speakers [
10] and voice and electroglottography (EGG) signal sets are included in this dataset. It comprises records of 687 healthy individuals (259 men and428 women) and 1356 individuals (629 men and 727 women) with different pathologies. The recording procedure involves: (a) vowels /i, a, u/ formed in normal speech, (b) high and low pitches; vowels /i, a, u/ with rising-falling pitch; and (c) the German sentence “Guten Morgen, wiegeht es Ihnen?” (“Good morning, how are you?”). Each recorded SVD voice was sampled with a resolution of 16-bit at 50 kHz. This dataset is fairly recent and has therefore been used by very few studies in the field of voice pathology. Following the three diseases criteria, we downloaded files from the website listed in [
44] and selected only the continuous vowel /a/ samples generated at normal pitch.
3.2. Proposed Method
The key aim of the study is to extract features that enhance the accuracy for detection and classification of voice pathology and to investigate the impact on the detection and classification processes of different frequency regions (bands). Before feeding to a convolutionary neural network (CNN), the voice signals are processed. To use existing stable CNN models, we use a transfer-learning platform. The paper explores, in particular, the ResNet34 models. The block diagram of the proposed solution is shown in
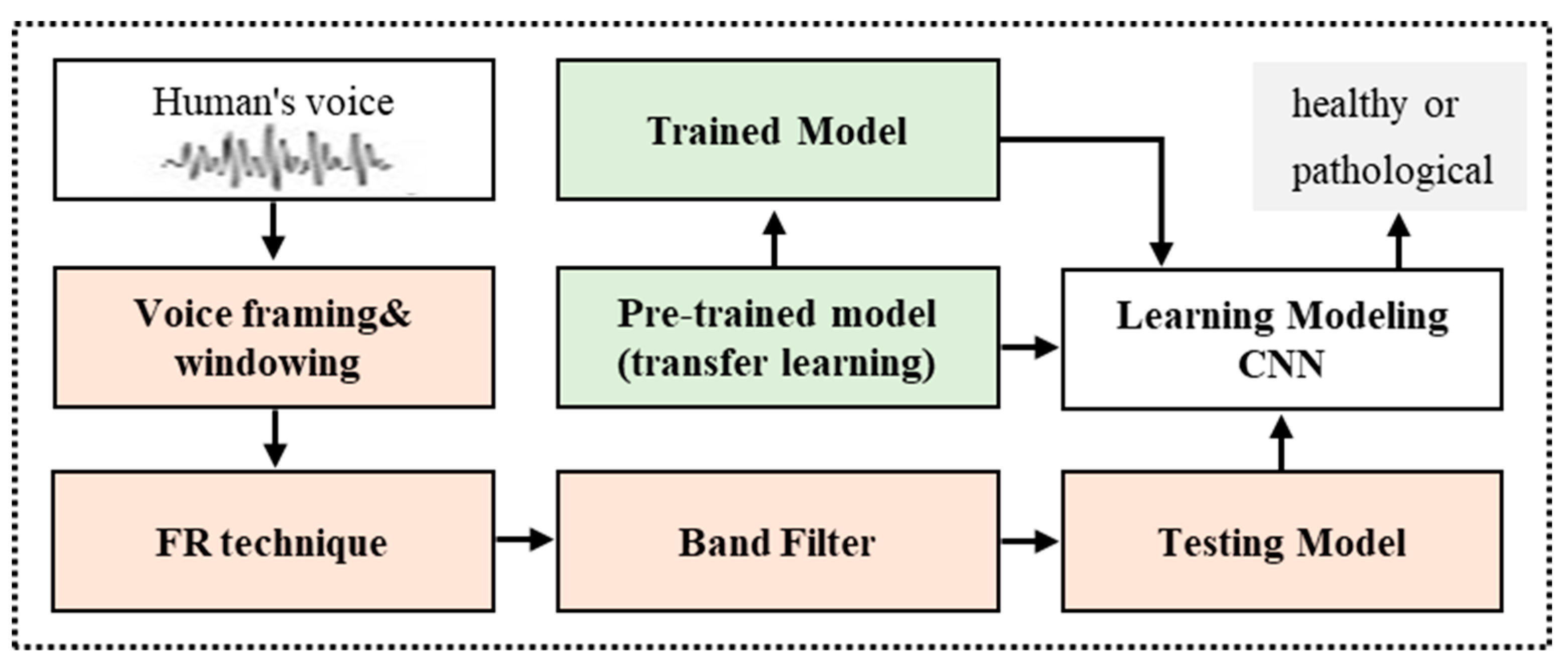
Figure 1.
Figure 1 demonstrates the proposed deep learning approach to the detection of voice pathology. The system is fed a patient’s voice, and the output determines whether the patient’s voice is normal or pathological. The signal of the voice is 1 s. If an input reaches 1 s, then a signal of 1 s is cut from the centre. The signal is split in 40 ms frames, of which the gap is 20 ms. The 40 ms frame duration is a well-equilibrated pitch capture and voice breaks smoothing option. If this is very long, then the voice breaks or some sounds cause the vocal folds to be irregularly opened and closed. The continuation effect and pitch duration are lost if the frame length is short. The framed signal is transformed by a fast Fourier transform to a frequency-domain signal. We get a spectrogram after concatenating all frequency-domains of the frames. The spectrogram could be viewed as an image. The spectrogram includes a minimum of 20 filters for the band pass. The filters are based on the octave. In the area of voice pathology detection, the octave scale typically functions better than the Mel scale [
34]. Time derivatives of the first and second order for the octave spectrum output are used. After this method, we get three image-like patterns: the octave and its derivatives of first and second order. The input of the CNN models is made up of three image patterns. We tested ResNet34 in the proposed method.
In this paper, for several reasons the transfer learning strategy for the CNN training is applied: (i) to overcome the lack of adequate voice pathologic attributes, in particular voice diseases derived from patients with reported infections, (ii) reducing the learning duration needed to acquire the final learned typical, and (iii) increasing the classification precision of the voice pathology identification. The technique of transfer learning is intended to boost neural network output in realistic applications bypassing learning from another task [
45]. For example, the training of a CNN to classify the case into two groups (e.g., pathological or healthy) may help to classify cases of different disease types. In this case, we used an effective ResNet34 pertained model. Residual Network (ResNet) is one of the highest-profile CNNs and the recipient of the 2015 ILSVRC ImageNet classification award [
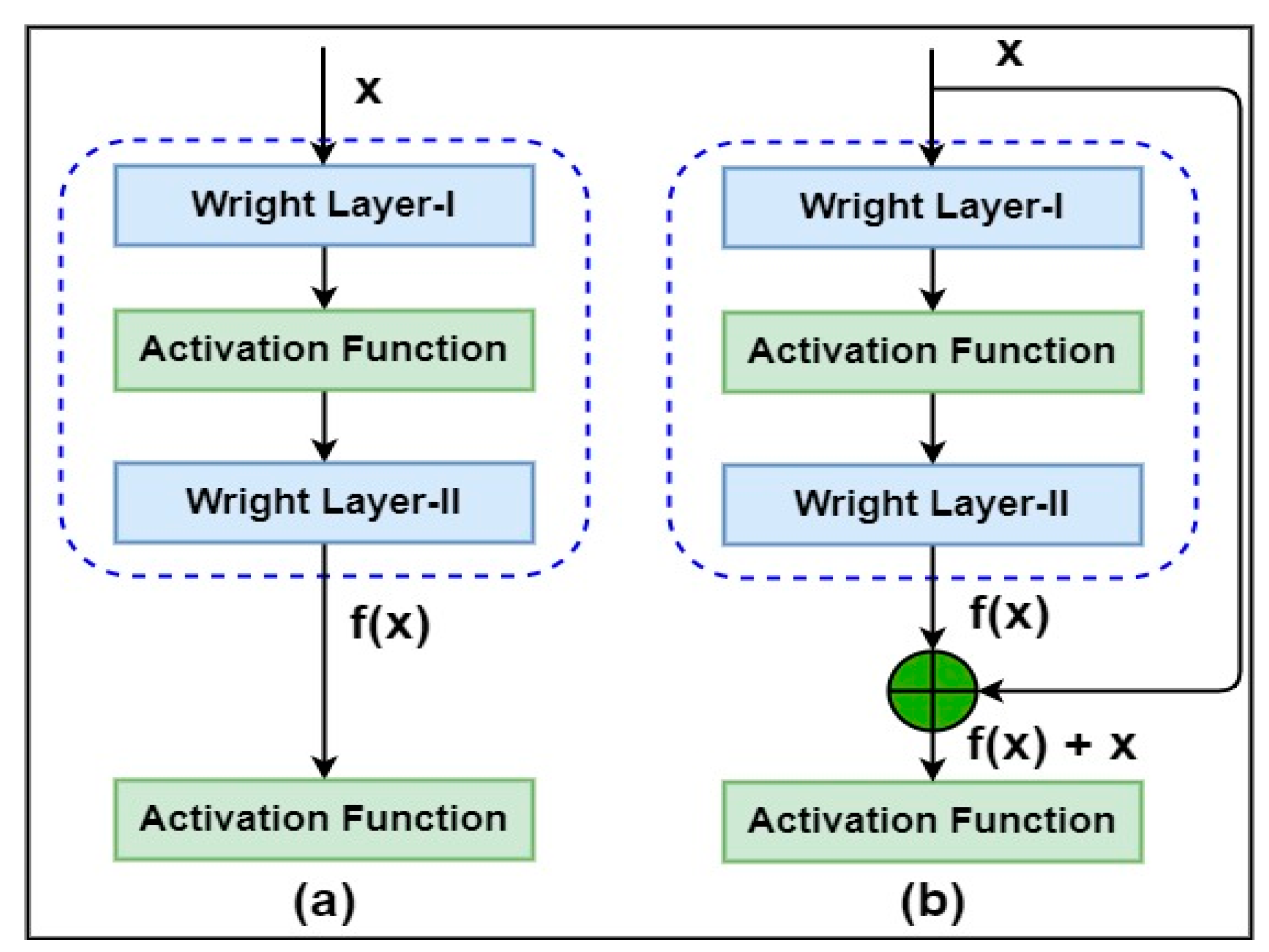
46]. ResNet is much like the other CNNs, which are sequentially packed with convolutionary, pooling, activation maps and fully interconnected layers. The only big difference between ResNet and other CNNs is the connection identity from the input layer to the end of the residual block (as shown in
Figure 2b). The architecture of ResNet34 begins with a convolutionary operation and max-pooling of the use of size kernels (5*5) pixels and (2*2) pixels, respectively. Thereafter, four stages with a different number with residual blocks are introduced, using size kernels (2*2) pixels to perform the convolutionary operation. When one passes from one point to the next, the depth of the channel is doubled, and the size of the input sample is halved. In this study the ResNet34 has an average pooling layer with two neurons (for example, positive pathological and normal case as healthy) followed by a completely connected layer.
Table 1 illustrates the main details of the ResNet34 architecture. Following the proposed training methodology, experiments were performed using k-folds cross validation samples of SVD voice pathology as a ResNet34 training sample, although the reset samples were used for the testing. In the course of the training, 20 percent of the training set was chosen randomly and used as a validator set to test the model’s general capacity and store the configuration of weights that gave the validation set the minimum error rate. The best model (based on hyper-parameters) used in the proposed Voice Pathology Detection System can be found in
Table 2. To summarize, the principal steps in the proposed approach for training are as follows:
- (1)
Divide the SVD dataset into three separate sets: training, test and validation set.
- (2)
Select initial hyper parameter values (e.g., learning rate, dynamic, and so on).
- (3)
Train the ResNet34 utilizing the training group and the hyper-parameters group in step 2.
- (4)
The validation set is used to evaluate ResNet34 performance during the training phase.
- (5)
Reiterate phases 3 to 4 for 12 epochs.
- (6)
Choose the best candidate typical with minimum validation error rate.
- (7)
Through utilizing the test group, the best-trained model is identified.
3.3. Training and Validation Stage
To train and validate the deep learning model, the dataset have been split into different sets as mentioned before. Subsequently, 10-fold cross-validation indices were generated for each set in the training and validation phases so that for each experiment we can use the same data sets. For the final evaluation of the models, the test set was left. Next, the testing and validation sets are stratified to the age and gender classes, by medical status (Healthy—H, Pathology—P). The long recordings were divided into several chunks which were necessary to prevent leakages into the test or validation set. These chunks have been carefully removed from the set. The other chunks were included in the training set. In each point of the validation confusion matrix, we used specific number samples that were taken from 150 healthy—H, pathological—P samples. To detect a pathology using the CNN model, we used 874 pathological and 200 healthy samples for testing the confusion matrix. We separated all the dataset into training, validation, and testing sets and made sure that the number of healthy and pathological samples was equivalent in each of the training and validation sets. The remainder has been added to the test set. In sum, 960 (480 healthy, 480 pathological) samples have been used for the training, 300 (150 healthy, 150 pathologic) samples have been applied for validation, and 874 (200 healthy and 674 pathological) samples have been used for testing. In the training phase, the distribution of the samples is unequal. We responded to this by adjusting the weights of samples, which are used for the minority groups during training to compensate. A 3-part weight product is the weight of the final sample. The number of subgroups in the group chosen (e.g., ratio of normal as well as pathological) is quantified by increasing partial weight. To this end, we presented a class weight α, gender weight β, as well as a group of gender-age weight γ that led to a final sample weight ω that is calculated as ω = α·β·γ. Furthermore, weights can be determined for a given sample in subgroup αi in group α, βi in group β, and γi in group γ. We have chosen the best hyperparameters for the cross-validation configuration as an output measurement. After tuning the hyperparameters, we have retrofitted and then evaluated the deep learning algorithms with the unsurpassed hyper- parameters over the whole set of training. In terms of a classification report (CR) and a confusion matrix (CM), the final results are presented. Formulation 1, 2 and 3 define how the CR tables calculate the recall, precision, and F1 score (weighting the average accuracy and recall). These three measurements are determined as follows:
The
Precision metric is used for measuring the proportion of the subjects that are of great importance. With this metric, the classifier’s ability to reject unimportant subjects is measured. The following is an expression of the metric:
The
F1 score is described as the weighted average of the precision and recall, the best value of F1 score is reached at 1, while the worst score is at 0. The precision and recall make an equal relative contribution to the F1 score. The F1 formula is given as follows:
The
Recall metric is used for the evaluation of the proportion of important subjects that are identified. With this metric, the classifier’s ability to provide all subjects that are of importance are measured. The following is an expression of the recall metric
4. Experimental Results
In this work, we propose a deep learning Convolutional Neural Network (CNN) model to perform pathology detection based on numerical analysis of voice signals. It is implemented in a Voice Pathology Detection DeepNet system using Python programming language. It is trained and tested by using a Google Colaboratory server. The testing computer has a 69 K GPU graphics card and 8 GB of RAM. It runs the Windows 10.1 operating system. The model architecture includes four fully connected layers. The CNN model of the Voice Pathology Detection DeepNet system is tested using the Saarbruecken Voice Database (SVD) dataset [
1]. The dataset contains recordings of 71 types of voice signals in which the signals were split into 64 ms long Hamming windowed segments with 30 ms overlap. It is arranged as a sequence of time-based vectors. Each vector as labelled as healthy or pathological class.
Table 3 presents the training confusion matrix of the SVD dataset. The table shows that the 300 training samples are divided equally to 150 healthy and 150 pathological classes to ensure a balanced training process.
The CNN model is able to achieve an average prediction score of 93.72% accuracy, 94.11% sensitivity or recall and 95.41% specificity.
Table 4 shows the results of precision, f1-score and recall for healthy and pathological classes in the training phase.
Table 5 presents the testing confusion matrix to the SVD dataset. The table shows that the 1074 testing instances are divided into 200 healthy and 874 pathological classes to insure robust testing process.
In the testing phase, the CNN model is able to achieve an average prediction score of 96.11% accuracy, 95.38% sensitivity or recall and 95.97% specificity.
Table 6 shows the results of precision, f1-score and recall for healthy and pathological classes in the testing phase.
Based on the training methodology that is mentioned early, the setting of the prediction model includes 0.02 learning rate, 2 batch size, 10 number of epochs, and 0.93 momentum as presented in
Table 2.
Table 7 shows the training Train Loss and testing Valid Loss for each epoch along with the accuracy and time per epoch results. From the observation of the results, during the progress of the Epoch, the Train Loss and testing Valid Loss are slightly increased, the accuracy result is slightly increased and the time per epoch is almost constant at 02:54 s. This result can be attributed to the ResNet34 training algorithm’s ability to rapidly adapt to the discriminative features of the SVD dataset and provide significant generalization.
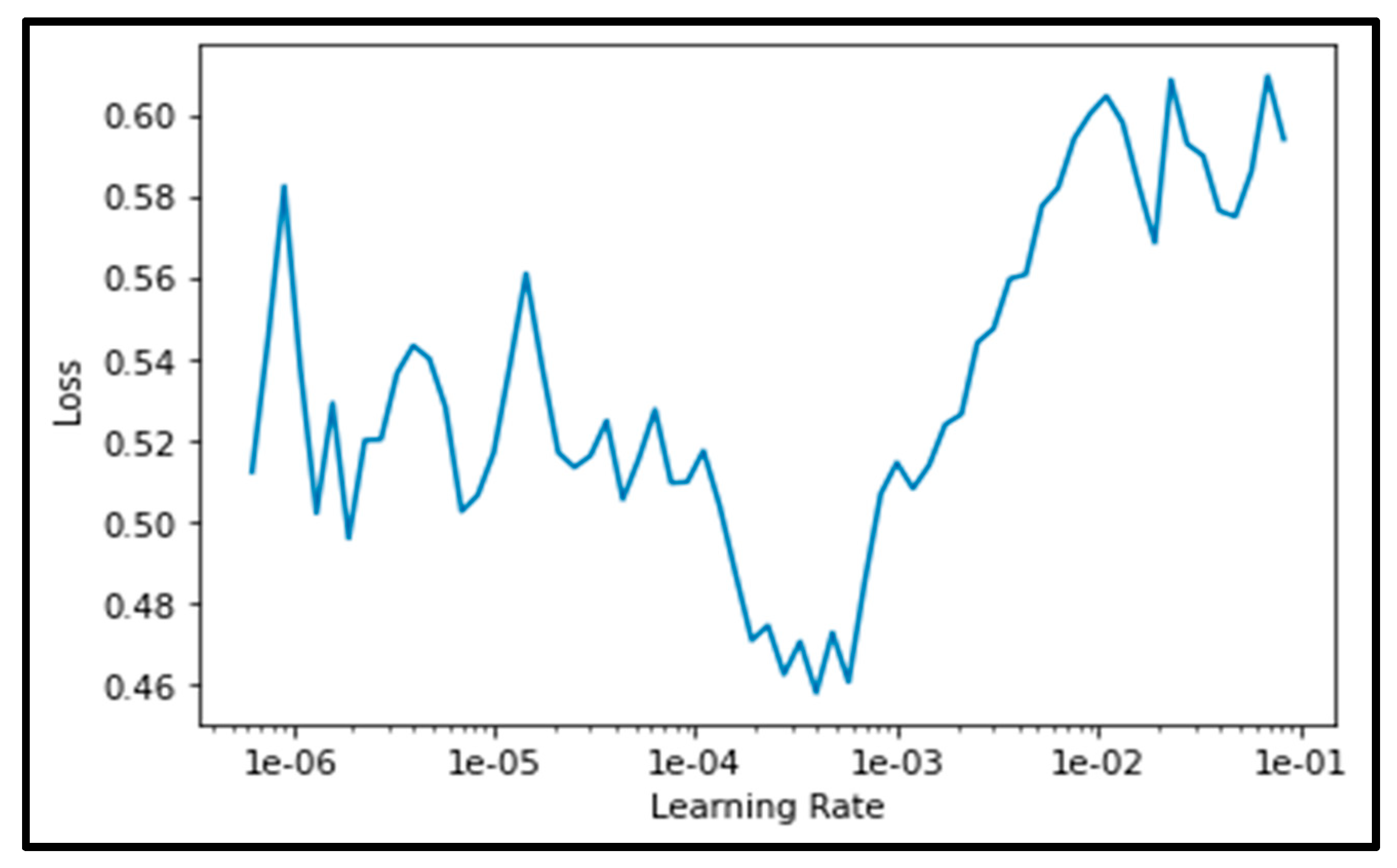
To further analyze the behaviour of the model, we investigated the relationship between the loss and learning rate. The analysis result shows that the best order of magnetite falls in the middle of the learning rate when the loss have values between 0.46 and 0.48 (i.e., between log scales le-04 and le-03) as shown in
Figure 3. Hence, the centroid of this area is selected to set the learning rate of the ResNet34 algorithm.
Some of the SVD voice pathology features can clinically identify pathological cases without the need for advanced analytical systems. However, voice pathology features changes with the changes in the stages of the disease development which affects the accuracy of the voice pathology detection results. In the early stages of the disease, the voice of the patient will not be significantly affected while when the disease becomes severe there will be a high irregularity in the voice of the patient. Hence, this work proposes a Voice Pathology Detection system that integrates a CNN model to perform voice pathology detection. The deep learning CNN of the Voice Pathology Detection system achieves high prediction accuracy results of up to 96.28% in distinguishing healthy from pathological cases. The main contribution of this work lays on modelling and tuning a deep learning CNN with ResNet34 layers to provide accurate voice pathology detection. Additionally, most of the related works use conventional dysphonic voice pattern analysis features which are easy to predict and even clinically interpretable while this dataset considered complex and challenging [
1,
12]. On the other hand, the limitations of this work are mainly due to the SVD testing dataset used and can be summarized as: (i) the small size of the tested cases, (ii) the absence of gender separation in the cases and (iii) ignoring the severity of the pathology in the features.