PySpark-Based Optimization of Microwave Image Reconstruction Algorithm for Head Imaging Big Data on High-Performance Computing and Google Cloud Platform
Abstract
1. Introduction
- The parallelism of the microwave image reconstruction (MIR) algorithm is first identified. Then, the computational model of Apache Spark is studied, and the parallel version of the algorithm is presented.
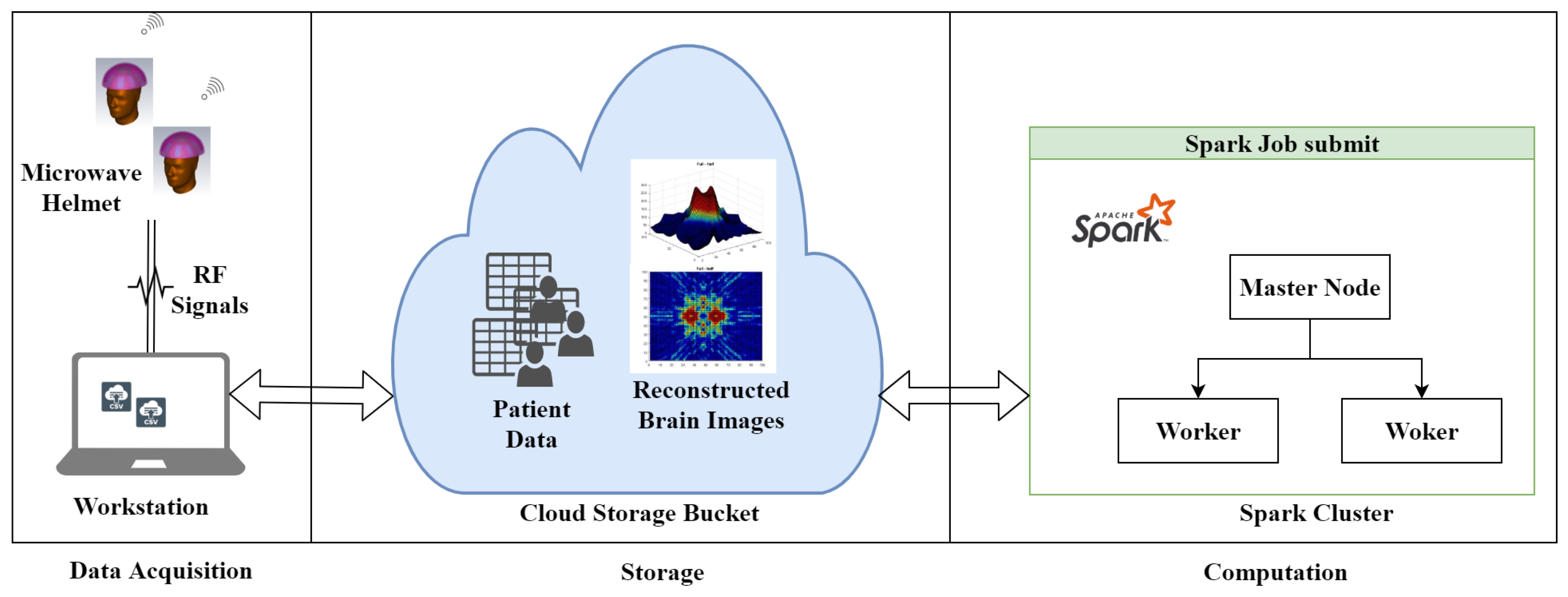
- A novel distributed approach, which adopts both data and algorithm in parallel, for efficient image reconstruction of head imaging is proposed. The imaging algorithm is optimized using PySpark on HPC clusters to improve the processing speed and make it real-time. The imaging system retrieves input data generated through radio frequency (RF) sensors and store in Eddie. An integrated imaging algorithm optimized through PySpark creates and saves images back to Eddie Storage.
2. Related Work
3. Image Reconstruction and the MIR Algorithm
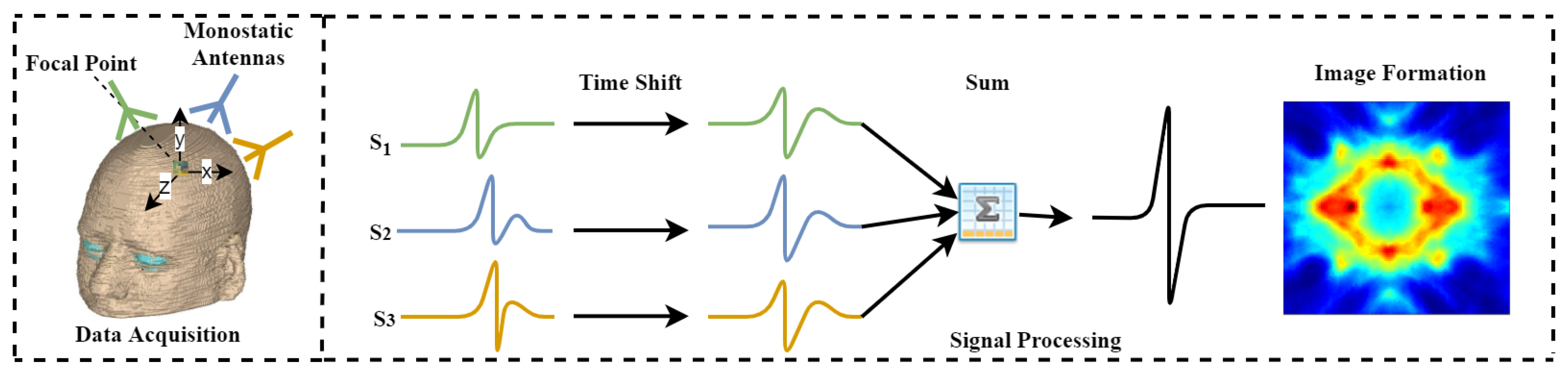
3.1. Principle of MIR Algorithm
| Algorithm 1 MIR Algorithm | |
| Input: Radio Frequency Sensors Data | |
| Output: Brain Images | |
| 1: | for← 1 to do |
| 2: | for x, y ← 1 to do |
| 3: | Compute using Equation (1) |
| 4: | end for |
| 5: | end for |
| 6: | for u, v ← 1 to do |
| 7: | = 0 |
| 8: | for ∀ S do |
| 9: | = 0 |
| 10: | for ← 1 to |
| 11: | if in range then |
| 12: | = + + as described in Equation (2) |
| 13: | end if |
| 14: | end for |
| 15: | = + ( * ) |
| 16: | end for |
| 17: | end for |
| = round () | |
3.2. Apache Spark Framework
4. Data Acquisition and Parallel Design and Implementation of MIR Algorithm
4.1. Identifying the Parallelism of MIR Algorithm
4.2. Design and Implementation of PMIR Algorithm
| Algorithm 2 PMIR algorithm | |
| Input: Radio Frequency Sensors Data | |
| Output: Brain Images | |
| 1: | Read data from Eddie distributed file system |
| 2: | Copy the RDD for delay and Antennas location to each worker node |
| 3: | Set the variables z, distance, and energy to zero |
| 4: | While < range do |
| 5: | Calculate the subset of pixel points on each worker node in parallel based on Equation (2) using Map operation |
| 6: | Update the master node concurrently with a subset of pixel values using Reduce operation |
| 7: | End While |
| 8: | Reconstruct the image from the matrix of pixel values on master node |
| 9: | Save the image to the Distributed file system on Eddie |
5. Experimental Evaluation
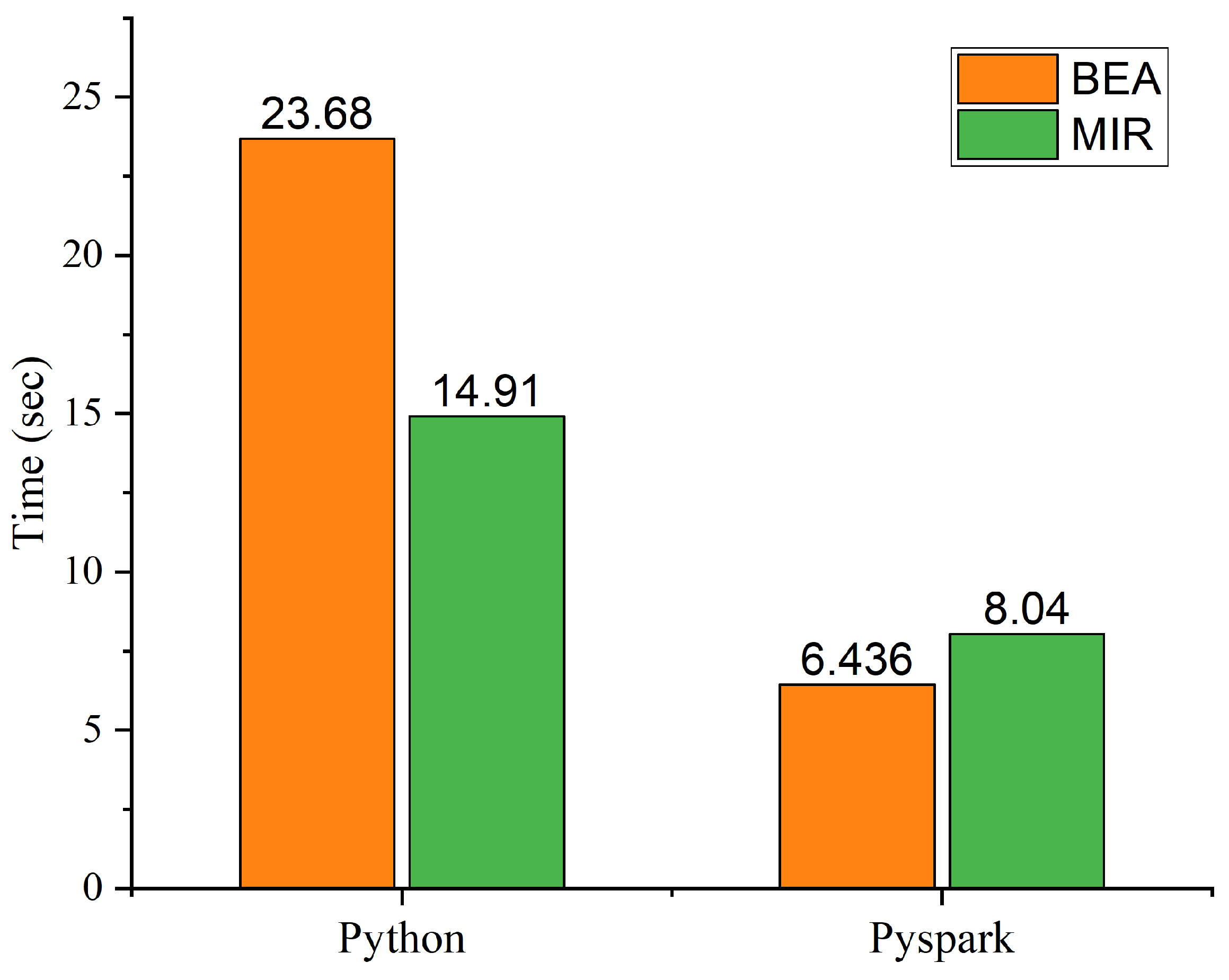
Performance Analysis of MIR and PMIR
6. Conclusions and Future Work
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| HPC | High Performance Computing |
| GCP | Google Cloud Platform |
| RF | Radio Frequency |
| Signal of Antenna A at Location n | |
| Number of Input Points | |
| Propagation time of signal n | |
| Signal Samples | |
| Energy for Pixel i | |
| RDD | Resilient Distributed Dataset |
References
- Manogaran, G.; Varatharajan, R.; Lopez, D.; Kumar, P.M.; Sundarasekar, R.; Thota, C. A new architecture of Internet of Things and big data ecosystem for secured smart healthcare monitoring and alerting system. Future Gener. Comput. Syst. 2018, 82, 375–387. [Google Scholar] [CrossRef]
- Makkie, M.; Li, X.; Quinn, S.; Lin, B.; Ye, J.; Mon, G.; Liu, T. A Distributed Computing Platform for fMRI Big Data Analytics. IEEE Trans. Big Data 2018, 5, 109–119. [Google Scholar] [CrossRef] [PubMed]
- Dhayne, H.; Haque, R.; Kilany, R.; Taher, Y. In Search of Big Medical Data Integration Solutions—A Comprehensive Survey. IEEE Access 2019, 7, 91265–91290. [Google Scholar] [CrossRef]
- Karadima, O.; Rahman, M.; Sotiriou, I.; Ghavami, N.; Lu, P.; Ahsan, S.; Kosmas, P. Experimental Validation of Microwave Tomography with the DBIM-TwIST Algorithm for Brain Stroke Detection and Classification. Sensors 2020, 20, 840. [Google Scholar] [CrossRef]
- Makarov, S.N.; Noetscher, G.M.; Arum, S.; Rabiner, R.; Nazarian, A. Concept of a Radiofrequency Device for osteopenia/osteoporosis Screening. Sci. Rep. 2020, 10, 3540. [Google Scholar] [CrossRef]
- Guo, B.; Li, J.; Zmuda, H.; Sheplak, M. Multifrequency microwave-induced thermal acoustic imaging for breast cancer detection. IEEE Trans. Biomed. Eng. 2007, 54, 2000–2010. [Google Scholar] [CrossRef]
- Nasiriavanaki, M.; Xia, J.; Wan, H.; Bauer, A.Q.; Culver, J.P.; Wang, L.V. High-resolution photoacoustic tomography of resting-state functional connectivity in the mouse brain. Proc. Natl. Acad. Sci. USA 2014, 111, 21–26. [Google Scholar] [CrossRef] [PubMed]
- Mozaffarzadeh, M.; Mahloojifar, A.; Orooji, M.; Adabi, S.; Nasiriavanaki, M. Double-stage delay multiply and sum beamforming algorithm: Application to linear-array photoacoustic imaging. IEEE Trans. Biomed. Eng. 2017, 65, 31–42. [Google Scholar] [CrossRef]
- Islam, M.; Mahmud, M.; Islam, M.T.; Kibria, S.; Samsuzzaman, M. A Low Cost and Portable Microwave Imaging System for Breast Tumor Detection Using UWB Directional Antenna array. Sci. Rep. 2019, 9, 15491. [Google Scholar] [CrossRef]
- Chandra, R.; Zhou, H.; Balasingham, I.; Narayanan, R.M. On the opportunities and challenges in microwave medical sensing and imaging. IEEE Trans. Biomed. Eng. 2015, 62, 1667–1682. [Google Scholar] [CrossRef]
- Stancombe, A.E.; Bialkowski, K.S.; Abbosh, A.M. Portable microwave head imaging system using software-defined radio and switching network. IEEE J. Electromagn. RF Microwaves Med. Biol. 2019, 3, 284–291. [Google Scholar] [CrossRef]
- Bolomey, J.C. Crossed viewpoints on microwave-based imaging for medical diagnosis: From genesis to earliest clinical outcomes. In The World of Applied Electromagnetics; Springer: Berlin/Heidelberg, Germany, 2018; pp. 369–414. [Google Scholar]
- O’Loughlin, D.; O’Halloran, M.; Moloney, B.M.; Glavin, M.; Jones, E.; Elahi, M.A. Microwave breast imaging: Clinical advances and remaining challenges. IEEE Trans. Biomed. Eng. 2018, 65, 2580–2590. [Google Scholar] [CrossRef] [PubMed]
- Wang, F.; Arslan, T.; Wang, G. Breast cancer detection with microwave imaging system using wearable conformal antenna arrays. In Proceedings of the 2017 IEEE International Conference on Imaging Systems and Techniques (IST), Beijing, China, 20 October 2017; pp. 1–6. [Google Scholar]
- Vasquez, J.T.; Turvani, G.; Dassano, G.; Casu, M.; Vipiana, F.; Joachimowicz, N.; Scapaticci, R.; Crocco, L.; Duchêne, B. Ongoing developments towards the realization of a microwave device for brain stroke monitoring. In Proceedings of the 2018 IEEE International Symposium on Antennas and Propagation & USNC/URSI National Radio Science Meeting, Boston, MA, USA, 13 July 2018; pp. 1139–1140. [Google Scholar]
- Mobashsher, A.T.; Abbosh, A. On-site rapid diagnosis of intracranial hematoma using portable multi-slice microwave imaging system. Sci. Rep. 2016, 6, 37620. [Google Scholar] [CrossRef] [PubMed]
- Saied, I.; Chandran, S.; Arslan, T. Integrated Flexible Hybrid Silicone-Textile Dual-Resonant Sensors and Switching Circuit for Wearable Neurodegeneration Monitoring Systems. IEEE Trans. Biomed. Circuits Syst. 2019, 13, 1304–1312. [Google Scholar] [CrossRef] [PubMed]
- Bao, S.; Parvarthaneni, P.; Huo, Y.; Barve, Y.; Plassard, A.J.; Yao, Y.; Sun, H.; Lyu, I.; Zald, D.H.; Landman, B.A.; et al. Technology Enablers for Big Data, Multi-Stage Analysis in Medical Image Processing. In Proceedings of the 2018 IEEE International Conference on Big Data, Seattle, WA, USA, 10 Decembr 2019; pp. 1337–1346. [Google Scholar] [CrossRef]
- Etminan, A.; Moghaddam, M. A Novel Global Optimization Technique for Microwave Imaging Based on the Simulated Annealing and Multi -Directional Search. In Proceedings of the 2018 IEEE International Symposium on Antennas and Propagation & USNC/URSI National Radio Science Meeting, Boston, MA, USA, 13 July 2018; pp. 1791–1792. [Google Scholar] [CrossRef]
- Chen, D.; Hu, Y.; Cai, C.; Zeng, K.; Li, X. Brain big data processing with massively parallel computing technology: Challenges and opportunities. Softw. Pract. Exp. 2017, 47, 405–420. [Google Scholar] [CrossRef]
- Siddiqui, F.; Amiri, S.; Minhas, U.I.; Deng, T.; Woods, R.; Rafferty, K.; Crookes, D. FPGA-based processor acceleration for image processing applications. J. Imaging 2019, 5, 16. [Google Scholar] [CrossRef]
- Wong, K.K.L.; Fong, S.; Wang, D.; Kian, K.; Wong, L. Impact of advanced parallel or cloud computing technologies for image guided diagnosis and therapy. J. X-ray Sci. Technol. 2017, 25, 187–192. [Google Scholar] [CrossRef]
- Ianni, M.; Masciari, E.; Mazzeo, G.M.; Mezzanzanica, M.; Zaniolo, C. Fast and effective Big Data exploration by clustering. Future Gener. Comput. Syst. 2019, 102, 84–94. [Google Scholar] [CrossRef]
- Basha, S.A.K.; Basha, S.M.; Vincent, D.R.; Rajput, D.S. Challenges in Storing and Processing Big Data Using Hadoop and Spark. In Deep Learning and Parallel Computing Environment for Bioengineering Systems; Elsevier: Amsterdam, The Netherlands, 2019; pp. 179–187. [Google Scholar] [CrossRef]
- Fu, G.S.; Levin-Schwartz, Y.; Lin, Q.H.; Zhang, D. Machine Learning for Medical Imaging. J. Healthc. Eng. 2019. [Google Scholar] [CrossRef]
- Essa, Y.M.; Hemdan, E.E.D.; El-Mahalawy, A.; Attiya, G.; El-Sayed, A. IFHDS: Intelligent Framework for Securing Healthcare BigData. J. Med. Syst. 2019, 43, 124. [Google Scholar] [CrossRef]
- Edinburgh Compute and Data Facility Website. University of Edinburgh. Available online: http://www.ecdf.ed.ac.uk/ (accessed on 23 January 2020).
- Li, Y.; Liu, P.; Li, Y.; Fan, H.; Su, P.; Peng, S.L.; Park, D.C.; Rodrigue, K.M.; Jiang, H.; Faria, A.V.; et al. ASL-MRICloud: An online tool for the processing of ASL MRI data. NMR Biomed. 2019, 32, e4051. [Google Scholar] [CrossRef] [PubMed]
- Fahmi, F.; Nasution, T.H.; Anggreiny, A. Smart cloud system with image processing server in diagnosing brain diseases dedicated for hospitals with limited resources. Technol. Health Care 2017, 25, 607–610. [Google Scholar] [CrossRef] [PubMed]
- Sarraf, S.; Ostadhashem, M. Big data application in functional magnetic resonance imaging using apache Spark. In Proceedings of the 2016 Future Technologies Conference (FTC), San Francisco, CA, USA, 7 December 2016; pp. 281–284. [Google Scholar]
- Liu, B.; He, S.; He, D.; Zhang, Y.; Guizani, M. A Spark-Based Parallel Fuzzy c -Means Segmentation Algorithm for Agricultural Image Big Data. IEEE Access 2019, 7, 42169–42180. [Google Scholar] [CrossRef]
- Cui, L.; Feng, J.; Zhang, Z.; Yang, L. High throughput automatic muscle image segmentation using parallel framework. BMC Bioinform. 2019, 20, 158. [Google Scholar] [CrossRef] [PubMed]
- Munro, I.; GarcÍA, E.; Yan, M.; Guldbrand, S.; Kumar, S.; Kwakwa, K.; Dunsby, C.; Neil, M.; French, P. Accelerating single molecule localization microscopy through parallel processing on a high-performance computing cluster. J. Microsc. 2019, 273, 148–160. [Google Scholar] [CrossRef] [PubMed]
- Qin, Y.; Wu, J.; Hu, Q.; Ghista, D.N.; Wong, K.K. Computational evaluation of smoothed particle hydrodynamics for implementing blood flow modelling through CT reconstructed arteries. J. X-ray Sci. Technol. 2017, 25, 213–232. [Google Scholar] [CrossRef]
- Cai, G.; Wang, J.; Mei, X.; Zhang, W.; Luan, G.; Liu, X. Electroclinical semiology of the bilateral asymmetric tonic seizures observed in patients with supplementary sensorimotor area epilepsy confirmed by pre- and post-operative MRI. J. X-ray Sci. Technol. 2017, 25, 247–259. [Google Scholar] [CrossRef]
- Liu, L.; Chen, W.; Nie, M.; Zhang, F.; Wang, Y.; He, A.; Wang, X.; Yan, G. iMAGE cloud: Medical image processing as a service for regional healthcare in a hybrid cloud environment. Environ. Health Prev. Med. 2016, 21, 563–571. [Google Scholar] [CrossRef]
- Chard, R.; Madduri, R.; Karonis, N.T.; Chard, K.; Duffin, K.L.; Ordoñez, C.E.; Uram, T.D.; Fleischauer, J.; Foster, I.T.; Papka, M.E.; et al. Scalable pCT Image Reconstruction Delivered as a Cloud Service. IEEE Trans. Cloud Comput. 2018, 6, 182–195. [Google Scholar] [CrossRef]
- Roychowdhury, S.; Hage, P.; Vasquez, J. Azure-Based Smart Monitoring System for Anemia-Like Pallor. Future Internet 2017, 9, 39. [Google Scholar] [CrossRef]
- Serrano, E.; Garcia-Blas, J.; Carretero, J. A Cloud Environment for Ubiquitous Medical Image Reconstruction. In Proceedings of the 2018 IEEE Intl Conf on Parallel & Distributed Processing with Applications, Ubiquitous Computing & Communications, Big Data & Cloud Computing, Social Computing & Networking, Sustainable Computing & Communications (ISPA/IUCC/BDCloud/SocialCom/SustainCom), Melbourne, Australia, 13 December 2018; pp. 1048–1055. [Google Scholar] [CrossRef]
- Meng, B.; Pratx, G.; Xing, L. Ultrafast and scalable cone-beam CT reconstruction using MapReduce in a cloud computing environment. Med. Phys. 2011, 38, 6603–6609. [Google Scholar] [CrossRef] [PubMed]
- Bao, S.; Damon, S.M.; Landman, B.A.; Gokhale, A. Performance Management of High Performance Computing for Medical Image Processing in Amazon Web Services; Zhang, J., Cook, T.S., Eds.; SPIE: Bellingham, WA, USA, 2016; Volume 9789, p. 97890Q. [Google Scholar] [CrossRef]
- Tchagna Kouanou, A.; Tchiotsop, D.; Kengne, R.; Zephirin, D.T.; Adele Armele, N.M.; Tchinda, R. An optimal big data workflow for biomedical image analysis. Inf. Med. Unlocked 2018, 11, 68–74. [Google Scholar] [CrossRef]
- Bond, E.J.; Li, X.; Hagness, S.C.; Van Veen, B.D. Microwave imaging via space-time beamforming for early detection of breast cancer. IEEE Trans. Antennas Propag. 2003, 51, 1690–1705. [Google Scholar] [CrossRef]
- Kibria, S.; Samsuzzaman, M.; Islam, M.T.; Mahmud, M.Z.; Misran, N.; Islam, M.T. Breast phantom imaging using iteratively corrected coherence factor delay and sum. IEEE Access 2019, 7, 40822–40832. [Google Scholar] [CrossRef]
- Been Lim, H.; Thi Tuyet Nhung, N.; Li, E.P.; Duc Thang, N. Confocal microwave imaging for breast cancer detection: Delay-multiply-and-sum image reconstruction algorithm. IEEE Trans. Biomed. Eng. 2008, 55, 1697–1704. [Google Scholar] [CrossRef]
- Elahi, M.A.; O’Loughlin, D.; Lavoie, B.R.; Glavin, M.; Jones, E.; Fear, E.C.; O’Halloran, M. Evaluation of image reconstruction algorithms for confocal microwave imaging: Application to patient data. Sensors 2018, 18, 1678. [Google Scholar] [CrossRef]
- Saied, I.; Arslan, T. Microwave Imaging Algorithm for Detecting Brain Disorders. In Proceedings of the 29th International Conference Radioelektronika (RADIOELEKTRONIKA), Kosice, Slovakia, Czech Republic, 16 April 2019; pp. 1–5. [Google Scholar]
- Apache Hadoop YARN. Available online: https://hadoop.apache.org/docs/current/hadoop-yarn/hadoop-yarn-site/YARN.html (accessed on 4 March 2020).
- Apache Mesos. Available online: https://http://mesos.apache.org/ (accessed on 4 March 2020).
- Zaharia, M.; Chowdhury, M.; Franklin, M.J.; Shenker, S. Spark: Cluster Computing with Working Sets. Technical Report. 2010. Available online: https://www.usenix.org/legacy/event/hotcloud10/tech/full_papers/Zaharia.pdf (accessed on 2 February 2020).
- Apache Spark. Available online: https://Spark.apache.org/docs/latest/index.html (accessed on 18 September 2019).
- Saied, I.; Arslan, T. Non-Invasive Wearable RF Device towards Monitoring Brain Atrophy and Lateral Ventricle Enlargement. IEEE J. Electromagn. RF Microw. Med. Biol. 2019. [Google Scholar] [CrossRef]
- Chew, K.M.; Yong, C.Y.; Sudirman, R.; Wei, S.T.C. Human brain modeling tumor detection in 2D and 3D representation using microwave signal analysis. In Proceedings of the ISCAIE 2018—IEEE Symposium on Computer Applications and Industrial Electronics, Penang, Malaysia, 29 April 2018; pp. 310–316. [Google Scholar] [CrossRef]
- Chew, K.M.; Sudirman, R.; Mahmood, N.H.; Seman, N.; Yong, C.Y. Human Brain Microwave Imaging Signal Processing: Frequency Domain (S-parameters) to Time Domain Conversion. Engineering 2013, 5, 31–36. [Google Scholar] [CrossRef]





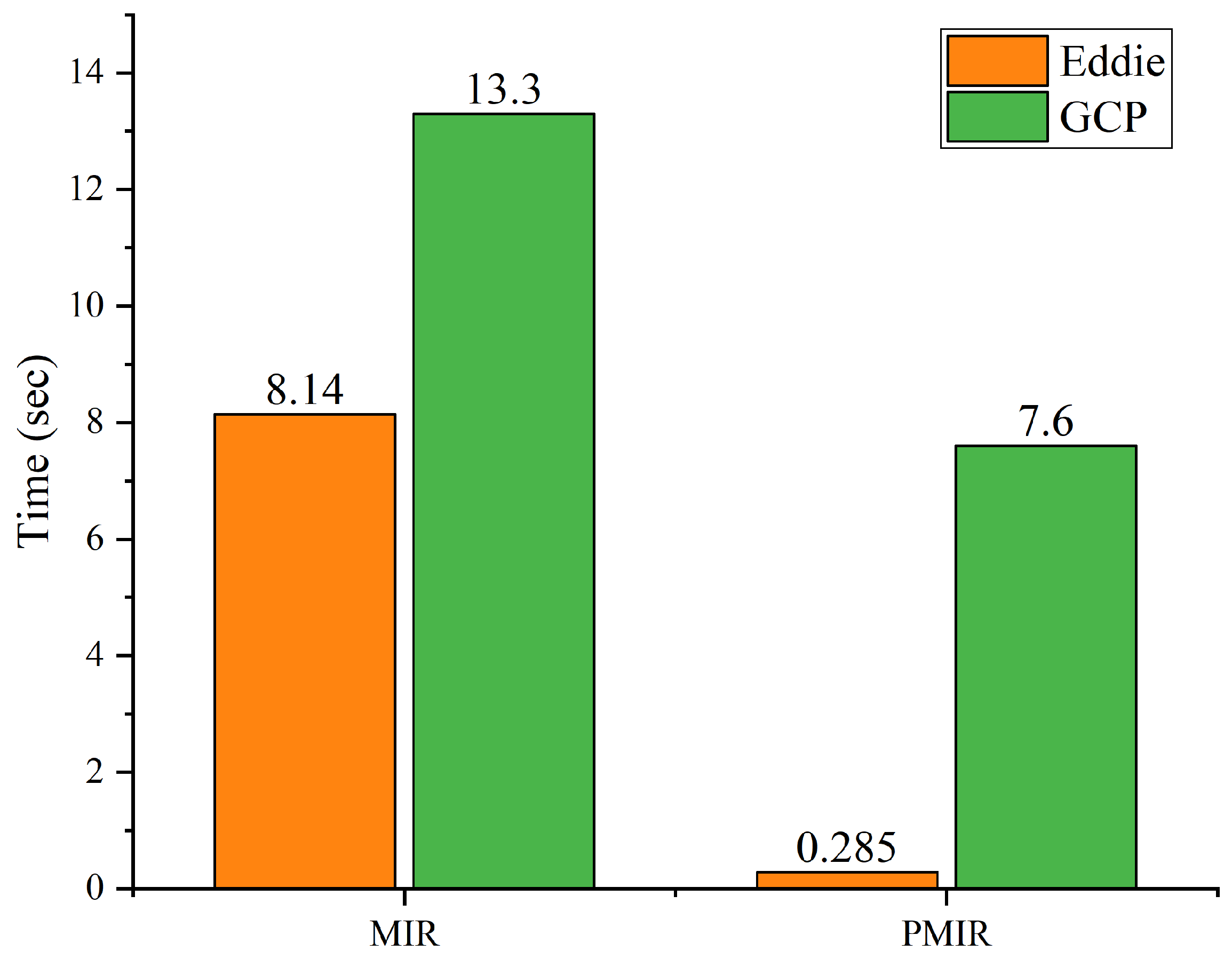
| Proposed Technique | Advantages | Limitations | Ref. |
|---|---|---|---|
| ASL-magnetic resonance imaging (MRI) Cloud | Cloud-based tool to process MRI data, image segmentation, better performance due to scalability | No parallel techniques implemented | [28] |
| Smart Cloud System | Cloud-based system for processing Neuroimaging data, image compression | No parallel techniques implemented, network congestion | [29] |
| cone-beam computerized tomography (CT) reconstruction | Faster reconstruction of image using Hadoop | No in-memory computation, no evaluation on computing cluster | [30] |
| Parallel Fuzzy c-Means Segmentation Algorithm | 12.54 time speed improvement, in-memory computation | No generality evaluation | [31] |
| Muscle image segmentation algorithm | 10 time speed improvement, in-memory computation, both data and model parallelism | No generality evaluation | [32] |
| Parallel computing approach to accelerate microscopy data | High throughput | Does not considered big data frameworks | [33] |
| Item | Value |
|---|---|
| CPU | Intel® Xeon® Processor E5-2630 v3 (2.4 GHz) |
| Memory | 16 G |
| Operating system | Scientific Linux 7 |
| PySpark version | 2.4.1 |
| JDK version | 1.8.0 |
| Hadoop version | 3.1.1 |
| Python version | 3.4.3 |
| Algorithm | Stand-Alone | Eddie | GCP |
|---|---|---|---|
| MIR | 14.91 | 8.14 | 13.3 |
| PMIR | 8.04 | 0.285 | 7.6 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ullah, R.; Arslan, T. PySpark-Based Optimization of Microwave Image Reconstruction Algorithm for Head Imaging Big Data on High-Performance Computing and Google Cloud Platform. Appl. Sci. 2020, 10, 3382. https://doi.org/10.3390/app10103382
Ullah R, Arslan T. PySpark-Based Optimization of Microwave Image Reconstruction Algorithm for Head Imaging Big Data on High-Performance Computing and Google Cloud Platform. Applied Sciences. 2020; 10(10):3382. https://doi.org/10.3390/app10103382
Chicago/Turabian StyleUllah, Rahmat, and Tughrul Arslan. 2020. "PySpark-Based Optimization of Microwave Image Reconstruction Algorithm for Head Imaging Big Data on High-Performance Computing and Google Cloud Platform" Applied Sciences 10, no. 10: 3382. https://doi.org/10.3390/app10103382
APA StyleUllah, R., & Arslan, T. (2020). PySpark-Based Optimization of Microwave Image Reconstruction Algorithm for Head Imaging Big Data on High-Performance Computing and Google Cloud Platform. Applied Sciences, 10(10), 3382. https://doi.org/10.3390/app10103382






