1. Introduction
Cyclospora cayetanensis is a food- and water-borne protozoan pathogen [
1]. Infected individuals shed immature oocysts into the environment where they mature given satisfactory environmental conditions [
2]. The oocysts take between 7 and 10 days to mature into sporulated oocysts after which they can excyst and become infective [
3]. During the environmental phase, oocysts are influenced by the conditions of the environment and can be transported and deposited in new areas [
4].
Humans can become infected after drinking contaminated water or after using the water to prepare food. Irrigation water can also contaminate food products in the field because it is sprayed directly onto produce and the oocysts can become trapped in small spaces. Similarly, produce can be contaminated if pesticides are diluted with contaminated water prior to field application [
5].
Areas with advanced sanitation practices are generally considered non-endemic for
Cyclospora [
6,
7]. Most people living in these regions acquire infections during travel to endemic regions or by consuming fresh produce imported from endemic areas [
8,
9]. However, some studies have reported detection of oocysts in US wastewater treatment plant effluent, even after undergoing chlorine disinfection [
10], showing some risk of exposure even in areas with advanced sanitation practices.
While
Cyclospora is a pathogen of emerging concern, it has historically been underdiagnosed. From 2017 to 2018, there was a dramatic increase in the number of reported cases of cyclosporiasis [
8,
11]. The increase in diagnosed cases is attributed to several factors, including the number of separate outbreaks and the use of new multiplex PCR techniques [
11]. However, there is some preliminary evidence that contact with animals, both domestic and wild, are associated with human infection [
12] and could be a route of exposure as urbanization encourages more interaction between humans and animals.
Soil is a potential route of transmission. Several studies have indicated a connection between contact with soil and rates of infection [
4,
9,
13]. Chacín-Bonilla et al. [
9] determined that contact with fecal-contaminated soil was associated with cyclosporiasis on San Carlos Island, Venezuela.
At this time, no natural animal hosts for
C. cayetanensis have been identified [
14]. However, Chu et al. [
15] detected
C. cayetanensis DNA in fecal samples of various animal species in Nepal. The DNA was found in fecal samples obtained from two dogs, a monkey, and a chicken [
15], indicating the presence of the pathogen, but not whether it was infective or not.
Humans are currently the only known host of
C. cayetanensis [
13]. Therefore, untreated human waste contaminated with
C. cayetanensis oocysts must enter the environment for environmental contamination to occur. In areas with advanced sanitation, there are fewer opportunities for environmental contamination. However, the opportunity arises during large rainfall events in communities that utilize combined sewage and stormwater management systems. During large or intense rain events, combined systems are unable to contain the excess volume of water, and stormwater combined with untreated sewage is discharged into streams at combined sewer outfalls. The extent to which combined sewer outfall (CSO) areas are contaminated with
C. cayetanensis is unknown.
If CSO systems discharge C. cayetanensis into the environment, the oocysts can move to new locations where they can be harmful to humans. While not infected, wildlife may still be able to transport C. cayetanensis oocysts from site to site. For example, if an animal ingests C. cayetanensis oocysts from a contaminated CSO site, and subsequently travels to a food production site where it defecates in a field, the C. cayetanensis can enter the food system. To help prevent cases of cyclosporiasis, potential sources of contamination need to be understood and managed. Water may be contaminated directly by CSO discharge. Wildlife may drink the water and defecate on soil or in other water sources, spreading oocysts further afield.
Several studies have examined the environmental presence of
C. cayetanensis. A study in Italy found
Cyclospora in tap water in bathrooms on national rail trains [
16]. Also in Italy, Giangaspero et al. [
4] identified contaminated water, food, and soil contaminated with
Cyclospora oocysts. These studies looked at specific locations at discrete time points and there is very little currently known about the distribution of
Cyclospora cayetanensis in the environment [
17]. The objective of this study was to determine the presence of
C. cayetanensis oocysts in water, soil, and wildlife scat in close proximity to CSO systems following discharge events in the Chicago metropolitan area. If
C. cayetanensis is found in certain locations and types of samples, risk management strategies can be developed to minimize the risk of human infection.
2. Materials and Methods
2.1. Sample Collection
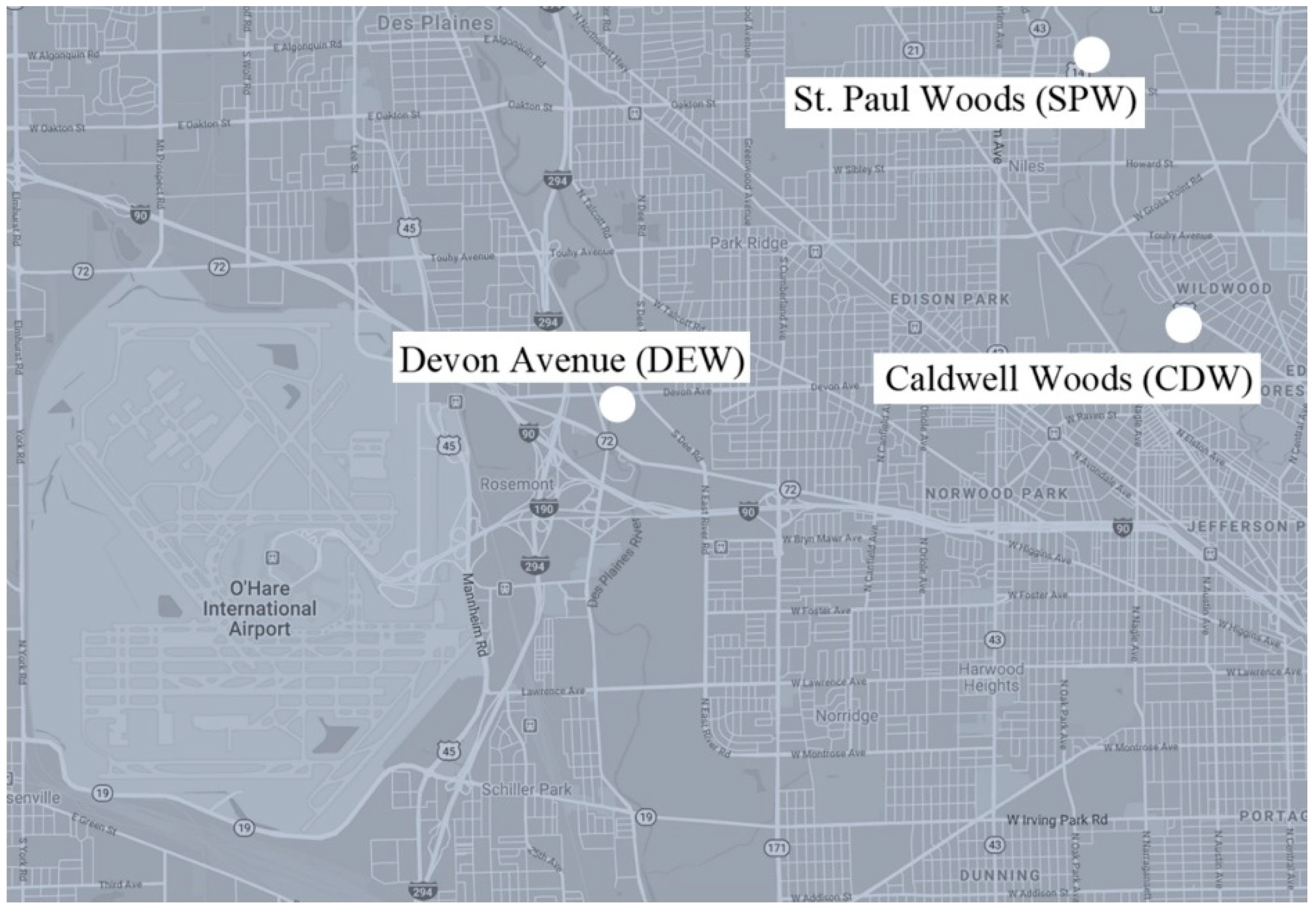
The Metropolitan Water Reclamation District of Greater Chicago monitors CSO activity and notifies the public when the CSO systems actively discharge sewage. CSO systems are located along waterways throughout the Greater Chicago Area. All samples were collected within one week after active discharge events. The following three CSO sites (
Figure 1) were selected because of their accessibility via public parks.
St. Paul Woods: Morton Grove, IL, along North Branch of the Chicago River
Caldwell Woods: Chicago, IL, along North Branch of the Chicago River
Devon Avenue: Forest Preserve of Cook County, at Devon Ave and S. Dee Rd, along the Des Plaines River
Each site was divided into four sampling points, one upstream of the CSO and three downstream. Point Z was 30 m upstream of the CSO, and A, B, and C were downstream of the CSO at 30 m increments. Soil was collected at each sampling point, both at the edge of the water and in the upland area. Soil was collected from within 6.1 m (20 ft.) of the water’s edge in a place that would cause minimal disruption to the vegetation. A visual inspection was used to find a representative soil sample from the sampling point. If there were multiple types of soil, then all types were collected from the top 2.54 cm (1 in.) of soil. Since the soil samples were collected directly adjacent to the stream, soils were primarily clay and clay loam soil types. Water samples were collected from each sampling point as well, within 1.52 m (5 ft.) of the bank for the safety of the researchers. All observed wildlife feces found between the extreme upstream and downstream sampling points was also collected.
Using a new disposable spoon for each new sample, soil or feces was collected to fill a 50 mL conical tube. After sampling and during transport to the lab, collection tubes were stored in a lunchbox cooler containing an ice pack.
U.S. Environmental Protection Agency (EPA) method 1623 [
18] describes options to concentrate
Cryptosporidium and
Giardia. The method was modified to collect and concentrate
Cyclospora because of the similarity in morphology to
Cryptosporidium. Some of the steps require agents that bind to the surface of the organisms, and there are no alternatives for
Cyclospora. Thus, a modified approach was used where approximately 30.28–37.85 L (8–10 gal.) of water were collected using a transfer pump. Later the water was transported back to the lab for mechanical concentration. The inlet hose was continually moved from the surface to the middle of the stream flow and back to the surface to achieve a depth-integrated water sample, approximately 0.91 m (3 ft.) from the bank, until the 5-gallon (18.93 L) collection bucket was full. During transport, the 5-gallon buckets of water were wrapped in foil thermal blankets and packed with ice. A total of 61 samples were collected from the Chicago, IL, USA, metropolitan area; 16 samples from Devon Ave, 23 from Caldwell Woods, and 22 from St. Paul Woods.
2.2. Sample Preparation
Feces and soil samples were stored at –80 °C for three or more days to render any
Cyclospora oocysts non-infectious [
2]. Keeping oocysts viable was not required for this study, and to improve the safety in the lab, oocysts were inactivated to minimize any chance of infection. Sathyanarayanan and Ortega [
2] found that there was no sporulation when
Cyclospora oocysts (in water or basil samples) were stored at −70 °C for one or more hours, so the environmental samples were stored at −80 °C and were left until they were processed. It was not possible to freeze the 5-gallon water buckets due to their size, so the water samples were concentrated before storing at −80 °C. Large particulates were filtered out of the water by pouring the water through cheesecloth. Then, the water was poured into six 250 mL plastic Nalgene centrifuge bottles and centrifuged at 2125 ×
g for 30 min. Water was removed from the top and discarded, leaving approximately 10 mL of water with a film of fine particulates at the bottom of each. For each Nalgene bottle, the particles were resuspended in the 10 mL solution by pipette mixing.
Then, additional water from the same 5-gallon sample was filtered with a cheesecloth and added to the 10 mL solution. Again, the sample was centrifuged, resuspended, and supernatant was removed. This process was repeated until all the particulates in the 5-gallon water sample were concentrated down into six bottles of 10 mL.
After the last of the 5-gallon water sample was concentrated into the six 10 mL bottles, it was pipette mixed and transferred into 50 mL conical tubes. The third round of centrifugation used 50 mL conical tubes. All the supernatant was removed except for the pellet and a thin layer of water to ensure that the pellet remained undisturbed. The pellet in each conical tube was resuspended in the fluid remaining in the tube by pipette mixing. The solutions from each conical tube were then consolidated into one 50 mL conical tube, which was stored at −80 °C to inactivate any C. cayetanensis oocysts.
2.3. DNA Extraction and PCR
DNA was extracted from each sample using the PowerSoil DNA Isolation Kit (MoBio) following the manufacturer’s instructions with a few exceptions. To mechanically break open the oocysts to release the DNA, a manufacturer representative suggested using a tissuelyzer for 5 min at 30 Hz rather than vortexing the samples for 10 min at maximum speed. All centrifugation steps were reduced from 10,000 × g to 9,500 × g due to centrifuge limitations. Additionally, to increase the genomic DNA concentration, the elution step was completed using 75 µL of Solution C6. DNA was stored at −20 °C until PCR was performed.
New primers were designed to specifically target
C. cayetanensis. This study utilized KX618190.1 as the template sequence and newly designed primers using PrimerQuest (IDT, Coralville, IA, USA). The resulting primers (
Table 1) were designed to anneal at positions where the ribosomal 18S gene sequences differed between
C. cayetanensis and
Eimera spp. A specificity check using Primer-BLAST indicated no cross amplification was expected for the primer pair. Concentrated
Eimeria acervulina was processed following the same procedure as the other samples and used to optimize the CYCLO18S primers. Annealing temperatures varied from 55–65 °C to identify the optimal temperature for the primers to anneal to
C. cayetanensis and not to
E. acervulina samples. We found 60 °C to be the optimal temperature (data not shown).
The 20 μL PCR reaction volume contained 7.8 μL of nuclease-free water, 0.1 μL of each CYCLO18S 100 μM of forward and reverse primer, 10.0 μL of GoTaq G2 Hot Start Green Master Mix (Promega, Madison, WI, USA), and 2.0 μL of template DNA. Amplification parameters began with 2 min at 95 °C, followed by 20 cycles of 15 s at 94 °C, 1 min at 60 °C, and final extension at 72 °C for 5 s and 10 °C for 5 min. For the second round of PCR, the same procedure was used, including use of the same primer pair. We used the product from the first round of PCR as the DNA template and increased the reaction to 35 cycles rather than 20.
The novel primers were designed to produce a short length fragment (100 bp) following the first two rounds of standard PCR. One round of PCR was insufficient to amplify low concentrations of C. cayetanensis DNA, therefore we added the second round of PCR to amplify low concentrations of DNA and allow us to correctly identify positive samples. Functionally, reducing the first round of PCR to 20 cycles ensured that the DNA concentrations did not plateau, that sufficient quantities of Cyclospora were amplified if DNA was in fact in the sample, and that non-target DNA was not amplified to the same extent.
We visualized amplified fragments after separating them by gel electrophoresis in a 2% agarose-0.5X TBE gel containing ethidium bromide. We loaded 10 µl of sample gel and HyperLadder™ 100 bp (Bioline; molecular weight marker) into gels.
3. Results and Discussion
The number and types of samples collected from each site are shown in
Table 2.
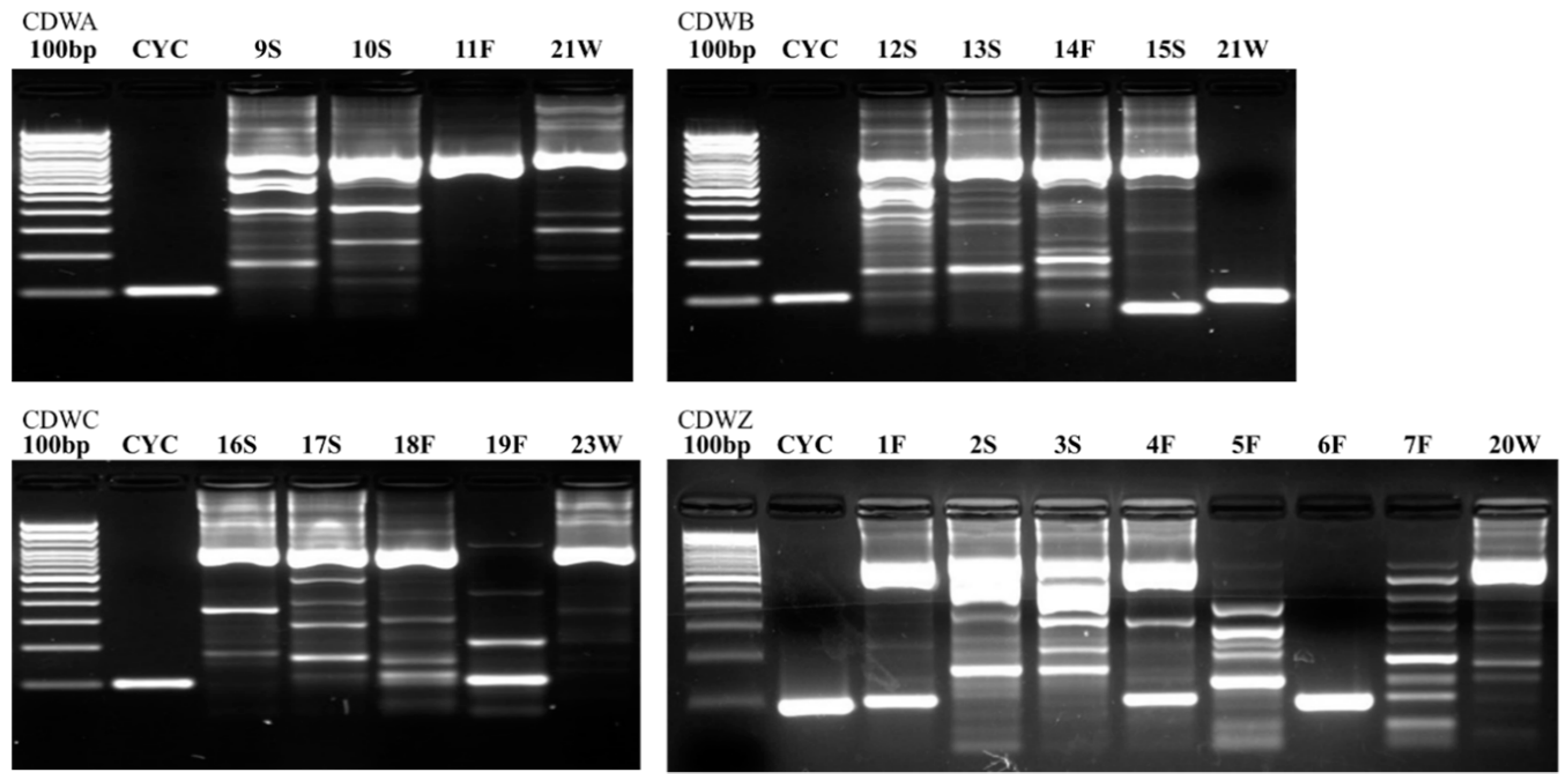
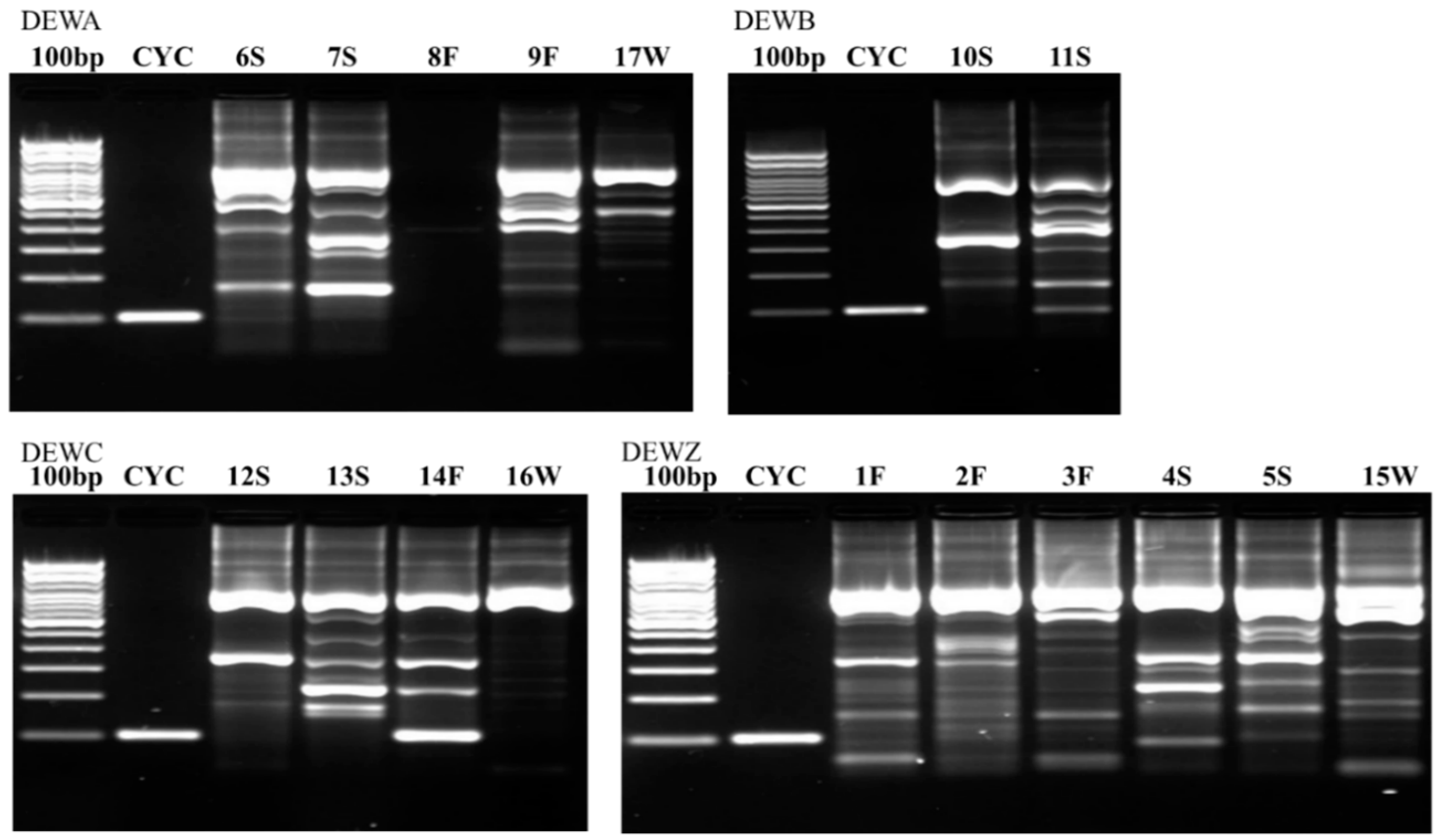
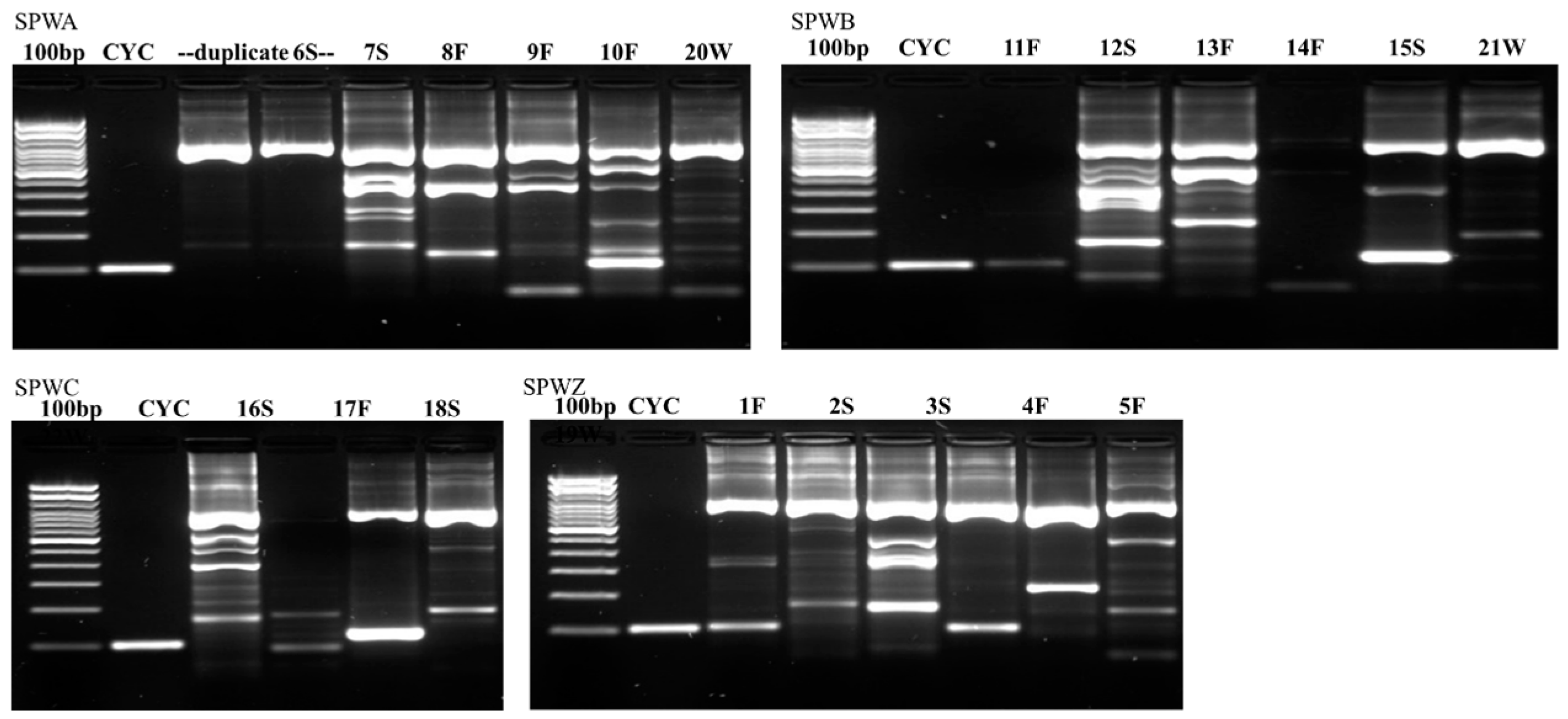
Figure 2,
Figure 3 and
Figure 4 illustrate the results from the three sites. Most samples produced many non-specific bands, indicating that the PCR technique may benefit from further optimization, but amplification of the target DNA was evident regardless of extraneous bands. Samples were designated positive if there was a discrete band at 100 bp (e.g., see CDWB21W in
Figure 2). If the smear of fragmented DNA ran through the ≤200 bp range with no distinct band in the 100 bp region, the sample was deemed negative (see CDWZ3S in
Figure 2).
Many samples had artifacts around 100 bp in length. With many artifacts, the smaller size bands can move more slowly through the agarose, and so may appear to have a slightly larger size than their true value. Bands that appeared to be at 100 bp or just slightly larger (in the presence of artifacts) were considered positive. A good example of this difference in position can be found in sample CDWZ1F beside CYC (
Figure 2).
The results from examining the gels are consolidated in
Table 3,
Table 4 and
Table 5, below. The “-” indicates there is no band at the target DNA size, meaning the primers did not amplify any DNA in the expected size range for
C. cayetanensis. The “+” indicates that there was a band of the appropriate size, which could be
C. cayetanensis.
As shown in
Table 3,
Table 4 and
Table 5, there did not appear to be much clustering of positive samples. All but one of the sampling points at each site had at least one positive band. As expected, none of the water samples were positive for
C. cayetanensis upstream of the CSO (transect Z). Interestingly, we recovered
C. cayetanensis DNA from fecal samples that were collected upstream of the CSO in both Caldwell Woods and St. Paul Woods, indicating wildlife may be consuming contaminated water downstream and then grazing in upstream areas. In addition, one upstream soil sample was positive at Devon Avenue. The pattern of positive fecal and soil samples in areas where the water was negative may indicate that wildlife transport of oocysts upstream, and possibly to areas away from the CSO location.
The highest proportion of positive samples was found at Caldwell Woods (11 positive samples). At all locations, most of the positive samples were wildlife feces (n = 13), and a few were soil (n = 7). There was one positive water sample. There were likely fewer positive samples found in the water compared to soil and feces because sampling occurred up to one week after the CSO discharge events. This study investigated the persistence of oocysts, in addition to being present in the CSO discharge, so allowing some time between the discharge event and sampling increased the opportunity for animals to ingest the water (and potentially oocysts as well) and then defecate. By the sampling time, much of the discharged Cyclospora, if present, may have been carried downstream, significantly diluted, or settled out of the water column. However, the data suggest that wildlife may often carry oocysts in their feces from one location to another. The fate and transport of Cyclospora should be studied more extensively especially in areas utilizing CSO systems.
While limited data on the environmental presence of
Cyclospora oocysts exist [
4,
9,
13,
16,
17], this study provides additional evidence that
Cyclospora oocysts are present in the environment, at least in close proximity to CSO systems.
4. Conclusions
To investigate the presence of
C. cayetanensis oocysts in water, soil, and wildlife scat near CSO locations following discharge events in the Chicago metropolitan area, this study evaluated 61 environmental samples during three discrete CSO discharge events. There were 21 positive samples; 13 from wildlife feces, 7 from soil, and 1 from water. Therefore, this case study indicates the likely presence of
C. cayetanensis in the environment. While small, this study does support past research [
10] indicating that
C. cayetanensis may be more common in North America than previously thought. Future work should include an in-depth study looking at temporal and spatial variability. Additional work should confirm where the organism is found in the environment and identify risks to the public based on the location and concentration. This study details methods to collect and detect
Cyclospora and provides new primers to selectively amplify
C. cayetanensis. Furthermore, it provides preliminary indication that
C. cayetanensis is present in the environment and may be transported by wildlife. Given the emerging threat of cyclosporiasis, additional studies are needed to confirm and expand these findings.