The Expanding Genetic Architecture of Arteriopathies: From Canonical TAAD Genes to Emerging Connective Tissue and Signaling Pathways
Abstract
1. Introduction
2. Classical Monogenic Syndromes and TAAD
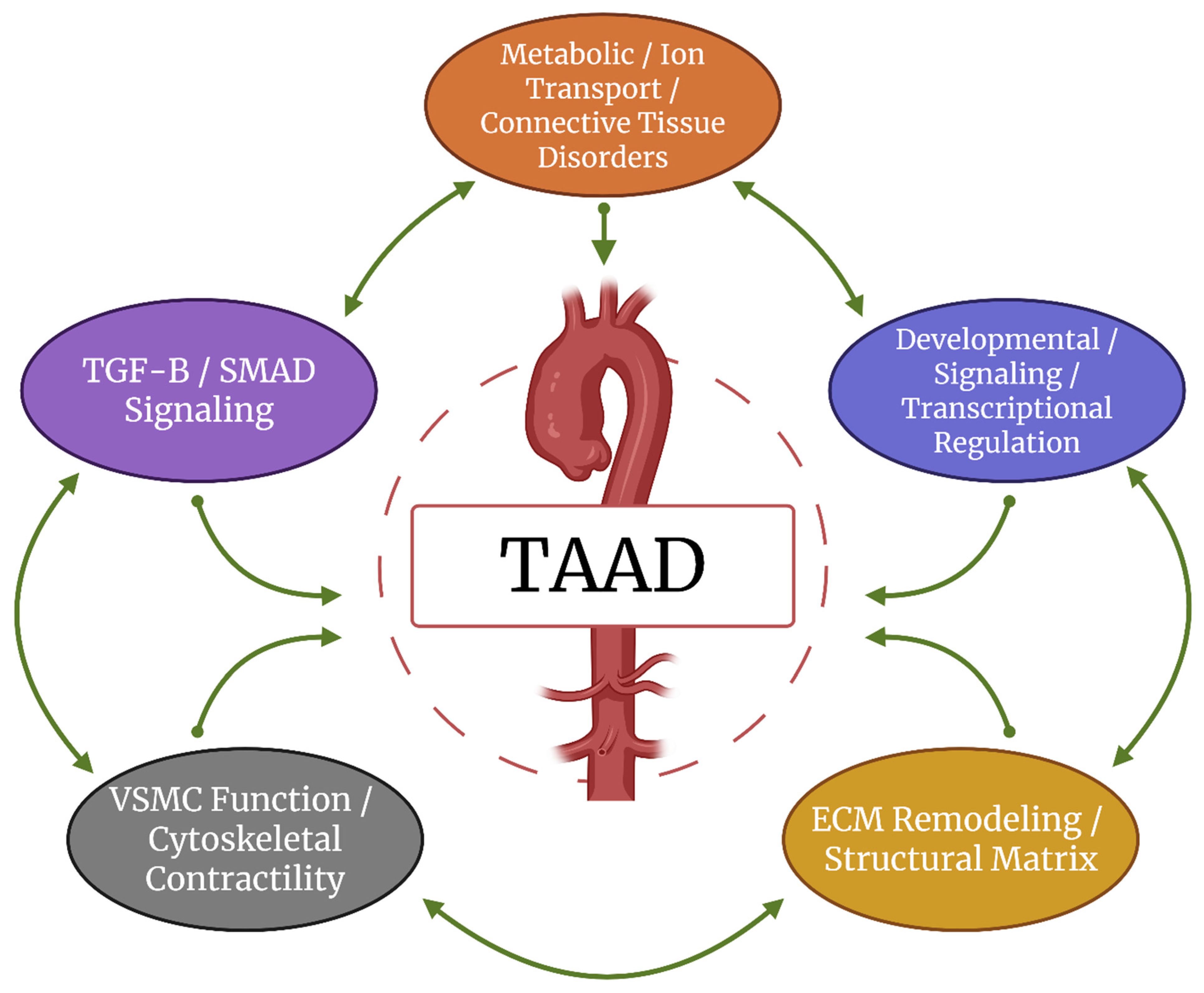
3. Functional Framework for TAAD Genes
3.1. TGF-β and SMAD Signaling
3.1.1. Core Pathway Overview
| Gene | Protein | Function | Location | Pathway Association | sTAAD | nsFTAAD | Phenotype | AA Size (cm) for SI | MoA | SoA | Category (A–D) | OMIM | Sources |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| ABCC6 | ATP-binding cassette, subfamily C, member 6 | ATP-binding cassette, serves as an export pump of unknown molecules that inhibit pathological calcification | 16p13.11 | Metabolic and Ion Transport | No | No | Pseudoxanthoma elasticum (PXE); arterial calcification, ECM fragmentation, increased stiffness | N/A | AD and AR | N/A | N/A | 603234 | [20,21,22,23,24] |
| ACTA2 | Smooth muscle alpha-actin | Contractile protein in vascular smooth muscle cells | 10q23.31 | Smooth Muscle Contraction/Mechanotransduction | Yes | Yes, most common cause | Aortic aneurysm and dissection (ascending), premature CAD, BAV, stroke | 4.5–5.0 cm | AD | Definitive | A2 | 611788 613834 614042 | [20,21,23,24,25] |
| ACTN2 | Alpha-actinin-2 | Crosslinks actin filaments at the Z-disc, modulates myofibrillar tension | 1q43 | Smooth Muscle Contraction/Mechanotransduction | No | Possibly | Left-dominant arrhythmogenic cardiomyopathy; suggested vascular overlap | N/A | AD | N/A | N/A | 102573 | [20,21,23,24,26] |
| ADAMTS2 | Procollagen 1 N-proteinase | Removes propeptides to allow for fibril self-assembly | 5q55.3 | Extracellular and Structural Matrix | Yes—EDS dermatosparaxis type (EDS VIIC) | No | Severe skin fragility, joint hypermobility, connective tissue disorder | N/A | AR | N/A | N/A | 604539 | [20,21,23,24,27] |
| ATP6V0A2 | ATPase, H+ transporting, lysosomal, V0 subunit A2 | Subunit of a vacuolar-type proton pump that acidifies endosomes, lysosomes, Golgi apparatus | 12q24.31 | Extracellular and Structural Matrix | Yes—cutis laxa and wrinkly skin syndrome (hypothesized to affect vasculature) | No | Connective tissue laxity, skin wrinkling, theoretical vascular remodeling impairment | N/A | AR | N/A | N/A | 611716 | [20,21,23,24,28] |
| ATP7A | ATPase, Cu2+-transporting alpha polypeptide | Transmembrane copper-transporting P-type ATPase supports lysyl oxidase function to crosslink collagen/elastin | Xq21.1 | Metabolic and Ion Transport | Yes—Menkes disease, occipital horn syndrome | No | Arterial tortuosity, dissection-prone vasculature, connective tissue fragility | N/A | XLR | N/A | N/A | 300011 | [20,21,23,24,29] |
| B3GALT6 | Beta-1,3-galactosyltransferase 6 | Facilitates synthesis of heparan sulfate and chondroitin sulfate (glycosaminoglycan matrix proteins) | 1p36.33 | Extracellular and Structural Matrix | Yes—spondylodysplastic EDS (spEDS) | No | Short stature, joint hypermobility, skin fragility, occasional vascular involvement | N/A | AR | N/A | N/A | 615291 | [20,21,23,24,30] |
| B4GALT7 | Beta-1,4-galactosyltransferase 7 | Facilitates synthesis of heparan sulfate and chondroitin sulfate (glycosaminoglycan matrix proteins) | 5q35.3 | Extracellular and Structural Matrix | Yes—spondylodysplastic EDS type 1 | No | Joint dislocations, short stature, limb anomalies, possible vascular fragility | N/A | AR | N/A | N/A | 604327 | [20,21,23,24,31] |
| BGN | Biglycan | Small leucine-rich proteoglycan, which binds to collagen fibrils and modulates ECM stiffness and profibrotic signaling | Xq28 | Overlap/Dual-Phenotype/Emerging Candidates | Yes, Meester–Loeys syndrome (X-linked, with vascular features) | No | Aortic dilation, reduced collagen content, increased pSMAD2, occasional aortic rupture | N/A | XL | N/A | N/A | 301870 | [20,21,23,24,32] |
| CBS | Cystathionine beta-synthase | Catalyzes the first irreversible step of transsulfuration, leading to the production of homocysteine | 21q22.3 | Metabolic and Ion Transport | Yes—homocystinuria | No | Endothelial dysfunction, thrombosis, ECM degradation due to hyperhomocysteinemia | N/A | AR | Limited | C | 613381 | [20,21,23,24,33] |
| CDH2 | Cadherin 2 | Neuronal-cadherin is also involved in cell–cell adhesion between smooth muscle cells | 18q12.1 | Cell–Cell Junction/Adhesion | No | Possibly | arrhythmogenic right ventricular cardiomyopathy (ARVC); ocular and genital defects; cardiac anomalies | N/A | AD | N/A | N/A | 114020 | [20,21,23,24,34] |
| CHST14 | Carbohydrate sulfotransferase 14 | Catalyzes 4-O-sulfation of dermatan sulfate (stereoisomer of chondroitin sulfate) | 15q15.1 | Extracellular and Structural Matrix | Yes—musculocontractural EDS (EDSMC1) | No | Arterial dilation, aneurysm, or dissection reported in EDSMC1 | N/A | AR | N/A | N/A | 608429 | [20,21,23,24,35] |
| COL12A1 | Type XII collagen | Fibril-associated collagen with interrupted triple helices (FACIT) tethers fibrils to surrounding structures | 6q13-q14.1 | Extracellular and Structural Matrix | No | No | myopathic EDS, skeletal/connective abnormalities; no aortic findings | N/A | AD | N/A | N/A | 120320 | [20,21,23,24,36] |
| COL1A1 | Type I collagen, alpha-1 chain | Alpha-1 subunit of type 1 collagen | 17q21.33 | Extracellular and Structural Matrix | Yes—arthrochalasia-type EDS and classic EDS (rare vascular involvement) | No | Hyperextensible skin, joint laxity, early aortic valve disease, possible borderline aortic dilation | N/A | AD | No Evidence | D | 120150 | [14,20,21,23,24,37] |
| COL1A2 | Type I collagen, alpha-2 chain | Alpha-2 subunit of type 1 collagen | 7q21.3 | Extracellular and Structural Matrix | Yes—cardiac valvular EDS | No | Mitral/aortic valve disease, borderline root dilation, joint hypermobility | 5.0 cm | AD and AR | No Evidence | D | 120160 | [14,20,21,23,24,37] |
| COL3A1 | Type III collagen, alpha-1 chain | Dominant type 3 fibrillar collagen | 2q32.2 | Extracellular and Structural Matrix | Yes—vascular Ehlers–Danlos syndrome (vEDS) | No | Arterial rupture/dissection, thin translucent skin, easy bruising, uterine/intestinal rupture | 5.0 cm | AD | Definitive | A1 | 120180 | [14,20,21,23,24] |
| COL5A1 | Type V collagen, alpha-1 chain | Regulates collagen fibril assembly and ECM integrity; modulates the diameter of collagen I fibrils | 9q34.3 | Extracellular and Structural Matrix | Yes—classic EDS | No | Joint hypermobility, atrophic scarring, fragile skin; potential mild aortic dilation | Standard | AD | No Evidence | D | 120215 | [14,20,21,23,24,37] |
| COL5A2 | Type V collagen, alpha-2 chain | Alpha-2 subunit of type 5 collagen | 2q32.2 | Extracellular and Structural Matrix | Yes—Ehlers–Danlos syndrome, classic type 2 | No | Classic EDS (joint hypermobility, skin hyperextensibility), theoretical aortic impact | Standard | AD | No Evidence | D | 120190 | [14,20,21,23,24,37] |
| COLGALT1 | Collagen beta (1-O) galactosyltransferase 1 | Glycoyslates hydroxylysine residues, stabilizing triple-helix formation | 19p13.11 | Extracellular and Structural Matrix | No | No direct evidence; potential role in ECM disorder | Defects in collagen IV trafficking; no confirmed aortic phenotype | N/A | AR | N/A | N/A | 617531 | [20,21,23,24,38] |
| CTNNA1 | Catenin alpha 1 | Mediates adherens junction linkage to the actin cytoskeleton | 5q31.2 | Overlap/Dual-Phenotype/Emerging Candidates | No | Possibly | Germline inactivating variants predispose to early-onset diffuse gastric cancer, with some carriers also developing invasive lobular breast cancer; loss impairs cell–cell adhesion, alters focal adhesion dynamics, increases cell migration, and affects epithelial tissue integrity | N/A | AD | N/A | N/A | 116805 | [20,21,23,24,39,40] |
| CTNNB1 | Catenin beta 1 | Mediates adherens junction linkage to the actin cytoskeleton | 3p22.1 | Overlap/Dual-Phenotype/Emerging Candidates | No | Possibly | Global developmental delay, intellectual disability (often moderate–severe), speech impairment, microcephaly, hypotonia, spastic diplegia; implicated in heart disease (e.g., dilated cardiomyopathy) and cardiac development | N/A | AD | N/A | N/A | 116806 | [20,21,23,24,39,40] |
| DCN | Decorin | Small leucine-rich proteoglycan that regulates collagen fibrillogenesis by controlling fibril diameter and spacing | 12q21.33 | Overlap/Dual-Phenotype/Emerging Candidates | No | Possibly | Aortic aneurysm due to extracellular matrix disruption (based on role in proteoglycan regulation) | N/A | AD | N/A | N/A | 125255 | [20,21,23,24,41] |
| DSE | Dermatan sulfate epimerase | Converts D to L-glucuronic acid during dermatan sulfate synthesis | 6q22.1 | Extracellular and Structural Matrix | Yes—musculocontractural Ehlers–Danlos syndrome (mcEDS) with potential vascular features in severe cases | No | Ehlers–Danlos syndrome, musculocontractural type 2 | N/A | AR | N/A | N/A | 605942 | [20,21,23,24,42] |
| DSP | Desmoplakin | Core desmosome component, tethering intermediate filaments between adjacent cells to maintain cohesion during cyclic motion | 6p24.3 | Cell–Cell Junction/Adhesion | Yes, carvajal/naxos-like syndromes | No | Dilated cardiomyopathy, left ventricular noncompaction, arrhythmogenic cardiomyopathy with variable vascular involvement | N/A | AD and AR | N/A | N/A | 125647 | [20,21,23,24,43] |
| ELN | Elastin | Core component of the elastic fiber | 7q11.23 | Extracellular and Structural Matrix | Yes—Williams–Beuren syndrome and cutis laxa | Yes—reported in isolated SVAS and aortic aneurysms | Supravalvular aortic stenosis (SVAS), cutis laxa, arterial tortuosity, aneurysms, elastic fiber fragmentation | Standard | AD | Limited | B2 | 130160 | [20,21,23,24,44] |
| EMILIN1 | Emilin 1 | Elastic fiber-associated glycoprotein regulating TGF-β precursor maturation | 2p23.3 | TGF-β/SMAD Signaling | No | Proposed only | In mouse models, EMILIN1 deficiency results in aortic root aneurysms, likely driven by dysregulated TGF-β signaling | N/A | AR | N/A | N/A | 130660 | [20,21,23,24,45] |
| ENPP1 | Ectonucleotide pyrophosphatase/phosphodiesterase 1 | Generate inorganic pyrophosphate, which inhibits vascular calcification | 6q23.2 | Metabolic and Ion Transport | Yes—generalized arterial calcification of infancy (GACI1) | No | Arterial calcification, periarticular and aortic mineralization, PXE-like features | N/A | AR | N/A | N/A | 173335 | [20,21,23,24,46] |
| FBLN4 (EFEMP2) | EGF-containing fibulin-like extracellular matrix protein 2 | Promote elastic fiber crosslinking and stability between fibrillin, elastin, and lysyl oxidases | 11q13.1 | Extracellular and Structural Matrix | Yes—AR cutis laxa type IB | No | Severe cutis laxa, ascending aortic aneurysm, arterial tortuosity, pulmonary artery hypoplasia | Standard | AR | Moderate | B1 | 604633 | [20,21,23,24,47] |
| FBLN5 | Fibulin5 | Promote elastic fiber crosslinking and stability between fibrillin, elastin, and lysyl oxidases | 14q32.12 | Extracellular and Structural Matrix | Proposed—cutis laxa (AR or AD forms), but no confirmed aortic aneurysms in human cases | No | Elastic fiber defects, tortuosity, emphysema, pelvic organ prolapse (in mice); no clear human aortic aneurysm phenotype | N/A | AD and AR | N/A | N/A | 604580 | [20,21,23,24,47] |
| FBN1 | Fibrillin 1 | Central microfibril component, scaffolds elastic fibers, and sequesters latent TGF-β complexes | 15q21.1 | Extracellular and Structural Matrix | Yes—Marfan syndrome | Yes | Aortic root aneurysm, ectopia lentis, long limbs, scoliosis, pectus deformity, joint laxity, skin striae | 5.0 cm | AD | Definitive | A1 | 134797 | [20,21,23,24,48] |
| FBN2 | Fibrillin 2 | Fetally expressed microfibril scaffold for elastin | 5q23.3 | Extracellular and Structural Matrix | Yes—congenital contractural arachnodactyly (CCA) | No | Joint contractures, arachnodactyly, scoliosis, “crumpled” ears; mild aortic dilation occasionally noted | Standard | AD | N/A | N/A | 612570 | [16,20,21,23,24] |
| FLCN | Folliculin | Tumor suppressor, influencing mTOR signaling and SMC turnover behavior | 17p11.2 | Transcriptional/Nuclear Envelope Regulation | No | No | Hypothesized role in ECM remodeling and SMC behavior | N/A | AD | N/A | N/A | 607273 | [20,21,23,24,49] |
| FLNA | Filamin A | Actin-binding protein critical for cytoskeleton structure and signaling | Xq28 | Smooth Muscle Contraction/Mechanotransduction | Yes, periventricular nodular heterotopia syndrome with aortic dilation | No | Aortic aneurysm with connective tissue and neurologic features (PNH) | Standard | X-linked Dominant | Limited | B1 | 300017 | [20,21,23,24,50] |
| FLNC | Filamin C | Actin binding and crosslinking in muscle cells, cytoskeletal structure reshaping | 7q32.1 | Smooth Muscle Contraction/Mechanotransduction | No | Possible, based on altered expression | Skeletal and cardiac myopathies; possible VSMC cytoskeletal effects | N/A | AD | N/A | N/A | 102565 | [20,21,23,24,51] |
| GJA4 | Gap junction protein, alpha-4 (connexin 37) | Facilitate ionic and metabolic exchange between endothelial cells and endothelium/SMCs | 1p34.3 | Overlap/Dual-Phenotype/Emerging Candidates | No | Possibly | Disruption in GJA5 may impair vascular tone regulation and promote ECM remodeling. It has also been associated with essential hypertension and vessel wall dysfunction, features that could plausibly contribute to aneurysmal susceptibility, though not directly proven | N/A | N/A | N/A | N/A | 121012 | [20,21,23,24,40,52] |
| GJA5 | Gap junction protein, alpha-5 (connexin 40) | Facilitate ionic and metabolic exchange between endothelial cells and endothelium/SMCs | 1q21.2 | Overlap/Dual-Phenotype/Emerging Candidates | No | Possibly | Connexin-40 dysregulation; may influence vascular tone, but no direct evidence of TAAD | N/A | AD | N/A | N/A | 121013 | [20,21,23,24,40,52] |
| HEY2 | HES-related family bHLH transcription factor with YRPW Motif 2 | Helix–loop–helix transcription factor involved in early embryonic vessel formation and downstream Notch signaling | 6q22.31 | Transcriptional/Nuclear Envelope Regulation | No | Yes | Associated with congenital heart disease and thoracic aneurysms | N/A | Mixed | N/A | N/A | 604674 | [20,21,23,24,53] |
| HNRNPK | Heterogeneous nuclear ribonucleoprotein K | DNA- and RNA-binding protein involved in chromatin remodeling, transcription, mRNA splicing and stability, translation | 9q21.32 | Transcriptional/Nuclear Envelope Regulation | No | No | possibly influences SMC contractility and response to strain | N/A | AD | N/A | N/A | 600712 | [20,21,23,24,54,55] |
| IPO8 | Importin-8 | Nuclear transport protein involved in TGF-β signaling regulation | 12p11.21 | TGF-β/SMAD Signaling | Yes (associated with syndromic features including arterial tortuosity and aneurysm) | No | Aneurysms, arterial tortuosity, possibly other syndromic traits | N/A | AR | N/A | N/A | 605600 | [20,21,23,24,56,57] |
| JAG1 | Jagged1 | Notch receptor ligand, facilitating signaling leading to embryonic vascular development and vascular smooth muscle cell differentiation | 20p12.2 | Cell–Cell Junction/Adhesion | Yes, Alagille syndrome | Yes, reported in isolated cases | Congenital heart disease, possible isolated aortic aneurysm | N/A | AD | No Evidence | D | 601920 | [20,21,23,24,58] |
| KCNN1 | Potassium channel, calcium-activated, intermediate/small conductance, subfamily N, member 1 | Voltage-independent calcium-activated potassium channel, contributing to the depolarization of smooth muscle and endothelial cells | 19p13.11 | Metabolic and Ion Transport | No | Yes (based on proposed mechanistic link to SMC contractility and adaptation) | Vascular tone impairment, arterial stiffness, possible aneurysm predisposition | N/A | N/A | N/A | N/A | 602982 | [20,21,23,24,59] |
| LMNA | Lamin A/C | Structural nuclear envelope protein; involved in mechanotransduction | 1q22 | Smooth Muscle Contraction/Mechanotransduction | No | Yes | Aortic aneurysm associated with cardiomyopathy and conduction defects | N/A | AD | N/A | N/A | 150330 | [20,21,23,24,60] |
| LOX | Lysyl oxidase | Initiates crosslinking of collagens and elastin | 5q23.1 | Extracellular and Structural Matrix | No | Yes—FTAAD type 10 | Aortic aneurysm, dissection, elastic lamellae defects, variable penetrance | Standard | AD | Strong | A2 | 153455 | [20,21,23,24,61,62] |
| LTBP1 | Latent transforming growth factor beta-binding protein 1 | Regulates the availability of TGF-β ligands in the extracellular matrix | 2p22.3 | TGF-β/SMAD Signaling | Proposed | No | Connective tissue phenotype with possible vascular features | N/A | AR | N/A | N/A | 150390 | [20,21,23,24,63] |
| LTBP2 | Latent transforming growth factor-beta-binding protein 2 | Anchors latent growth factor complexes to microfibrils | 14q24.3 | Extracellular and Structural Matrix | No clear evidence | No | Ocular phenotypes, glaucoma-related connective tissue dysfunction | N/A | AR | N/A | N/A | 602091 | [20,21,23,24,63] |
| LTBP3 | Latent transforming growth factor beta-binding protein 3 | Binds TGF-β and modulates ECM and TGF-β bioavailability | 11q13.1 | TGF-β/SMAD Signaling | Yes—described in a syndromic context with aortic dilation | No | Aortic aneurysm (often syndromic), variable skeletal/connective features | N/A | AR | N/A | N/A | 602090 | [20,21,23,24,63] |
| LUM | Lumican | Small leucine-rich proteoglycan | 12q21.33 | Overlap/Dual-Phenotype/Emerging Candidates | No | Possibly | Aortic wall fragility and aneurysm susceptibility due to impaired ECM assembly | N/A | N/A | N/A | N/A | 600616 | [20,21,23,24,64] |
| MAT2A | Methionine adenosyltransferase II, alpha | Extrahepatic methionine adenosyltransferase, adds methyl groups for DNA, RNA, protein, and lipid modifications—specifically DNA methylation for epigenetic expressivity of vascular gene expression | 2p11.2 | Metabolic and Ion Transport | No | Yes | Aberrant methylation profiles; possible smooth muscle dysfunction and matrix turnover abnormalities | Standard | AD | N/A | N/A | 601468 | [20,21,23,24,65] |
| MED12 | Mediator complex subunit 12 | Bridges transcription factors to RNA polymerase II, influencing vascular SMC function | Xq13.1 | Transcriptional/Nuclear Envelope Regulation | Yes, in the context of Hardikar syndrome | No | Impaired SMC contractility and matrix remodeling; rare X-linked disorder with cardiac anomalies | N/A | X-linked | N/A | N/A | 300188 | [20,21,23,24,66] |
| MFAP5 | Microfibrillar-associated protein 5 | Integrates microfibrils into regular elastic recoil | 12p13.31 | Extracellular and Structural Matrix | No | Yes | Familial thoracic aortic aneurysm and dissection due to impaired microfibril integrity and TGF-β dysregulation | Standard | AD | Moderate | Uncertain | 601103 | [20,21,23,24,67] |
| MYH11 | Smooth muscle myosin heavy chain 11 | Major contractile protein in vascular smooth muscle | 16p13.11 | Smooth Muscle Contraction/Mechanotransduction | No | Yes | TAAD with or without PDA (patent ductus arteriosus), CAD, carotid IA | 4.5–5.0 cm | AD | Definitive | A2 | 160745 | [20,21,23,24,68,69] |
| MYLK | Myosin light chain kinase | Regulates smooth muscle contraction via phosphorylation of myosin | 3q21.1 | Smooth Muscle Contraction/Mechanotransduction | No | Yes | Aortic dissection (with minimal dilation), intrafamilial variability | 4.5–5.0 cm | AD | Definitive | A2 | 600922 | [20,21,23,24,70,71] |
| NOTCH1 | Notch receptor 1 | Primary receptor for Jagged-1, transcriptional regulator for cell fate of smooth muscle/ECM remodeling | 9q34.3 | Cell–Cell Junction/Adhesion | No | Yes, associated with BAV and aortic aneurysm | BAV, calcific valve disease, ascending aneurysm | Standard | AD | Limited | B2 | 190198 | [20,21,23,24,72,73] |
| PDLIM3 | PDZ and LIM domain protein 3 (a.k.a. ALP) | Actin-associated protein involved in cytoskeletal signaling | 4q35.1 | Smooth Muscle Contraction/Mechanotransduction | No | Possibly | SCAD, thoracic aortic aneurysm/dissection (rare), cardiomyopathy | N/A | Likely AD | N/A | N/A | 605889 | [20,21,23,24,74] |
| PKP2 | Plakophilin 2 | Core desmosome component, tethering intermediate filaments between adjacent cells to maintain cohesion during cyclic motion | 12p11.21 | Cell–Cell Junction/Adhesion | No | Possibly | Arrhythmogenic right ventricular cardiomyopathy (ARVC), the most common cause, with fibrofatty myocardial replacement, ventricular arrhythmias, and sudden death; Brugada syndrome; idiopathic ventricular fibrillation; biventricular and left-dominant cardiomyopathy variants | N/A | AD | N/A | N/A | 602861 | [20,21,23,24,75] |
| PLOD1 | Procollagen-Lysine, 2-oxoglutrate 5-dioxygenase 1 | Lysyl hydroxylase that forms hydroxylysine in collagens, allowing for later glycosylation/crosslinking | 1p36.22 | Extracellular and Structural Matrix | Yes—Ehlers–Danlos syndrome, kyphoscoliotic type (EDSKSCL1) | No | Progressive kyphoscoliosis, joint/skin hyperextensibility, hypotonia; indirect vessel involvement | N/A | AR | N/A | N/A | 153454 | [14,20,21,23,24,76] |
| PLOD3 | Procollagen-Lysine, 2-oxoglutrate 5-dioxygenase 3 | Lysyl hydroxylase that forms hydroxylysine in collagens, allowing for later glycosylation/crosslinking—additional galactosyltransferase and glucosyltransferase activities | 7q22.1 | Extracellular and Structural Matrix | Yes—BCARD syndrome with vascular features (connective tissue + arteriopathy) | No | Craniofacial dysmorphism, scoliosis, osteopenia, vascular fragility (BCARD syndrome) | N/A | AR | No Evidence | D | 603066 | [14,20,21,23,24,76] |
| PMEPA1 | Prostate transmembrane protein, androgen induced 1 | Transmembrane protein induced by TGF-β, promoting the degradation of activated SMADs | 20q13.31 | Transcriptional/Nuclear Envelope Regulation | No | No | Modulation of fibrosis and ECM remodeling; role in vascular remodeling is theoretical | N/A | N/A | N/A | N/A | 606564 | [20,21,23,24,77] |
| PPP1R12A | Protein phosphatase 1, regulatory subunit 12A | Regulatory subunit of myosin light chain phosphatase | 12q21.2-q21.31 | Smooth Muscle Contraction/Mechanotransduction | No | Proposed disease modifier | SMC relaxation and cytoskeletal remodeling; potential influence on aortic integrity | N/A | AD | N/A | N/A | 602021 | [20,21,23,24,62] |
| PRDM5 | PR domain-containing protein 5 | DNA-binding zinc finger transcription factor binds collagen and proteoglycan-expressing regions | 4q27 | Transcriptional/Nuclear Envelope Regulation | Yes, brittle cornea syndrome 2 | No | ECM disruption in BCS2, corneal fragility, possible vascular contributions | N/A | AR | N/A | N/A | 614161 | [20,21,23,24,78] |
| PRKG1 | cGMP-dependent protein kinase 1 | Regulates smooth muscle contraction via calcium desensitization | 10q11.23 | Smooth Muscle Contraction/Mechanotransduction | No | Yes | Familial aortic dissection with minimal dilation; early-onset dissection | 4.5–5.0 cm | AD | Definitive | A2 | 176894 | [8,20,21,23,24,79] |
| ROBO4 | Roundabout guidance receptor 4 | Endothelial receptor that suppresses pathological angiogenesis and vascular permeability via stabilization of the endothelial layer | 11q24.2 | Extracellular and Structural Matrix | No | Yes | Aortic valve disease, thoracic aneurysm, endothelial barrier dysfunction | Standard | AD | N/A | N/A | 607528 | [20,21,23,24,80] |
| ROCK2 | Rho-associated coiled coil containing protein kinase 2 (ROCK2) | Serine/threonine kinase involved in actin cytoskeleton dynamics | 2p25.1 | Smooth Muscle Contraction/Mechanotransduction | No | Proposed disease modifier | SMC tone dysregulation, ECM remodeling; proposed role in medial degeneration | N/A | AD | N/A | N/A | 604002 | [20,21,23,24,81] |
| SECISBP2 | Secis-binding protein 2 | Selenocysteine insertion sequence-binding protein that leads to synthesis of selenoproteins that protect cells from oxidative damage | 9q22.2 | Transcriptional/Nuclear Envelope Regulation | No | No | Not directly TAAD-associated; hypothesized to impact ECM via redox imbalance and inflammation | N/A | AR | N/A | N/A | 607693 | [20,21,23,24,82] |
| SELENOT | Selenoprotein T | Participates in redox homeostasis in mitochondria | 3q25.1 | Overlap/Dual-Phenotype/Emerging Candidates | No | Possibly | glucose intolerance due to impaired insulin response | N/A | N/A | N/A | N/A | 607912 | [20,21,23,24,82] |
| SKI | SKI proto-oncogene | Transcriptional repressor in the TGF-β signaling pathway | 1p36.33 | TGF-β/SMAD Signaling | Yes—associated with Shprintzen–Goldberg syndrome | No | Aneurysms, tortuosity, craniofacial, and skeletal anomalies | Standard | AD | Limited | B | 164780 | [20,21,23,24,83] |
| SLC2A10 (GLUT10) | Solute carrier family 2 (facilitated glucose transporter), member 10 | Facilitative glucose transporter family protein, which plays a role in the intracellular redox environment | 20q13.12 | Extracellular and Structural Matrix | Yes—arterial tortuosity syndrome | No | Arterial tortuosity, aneurysm, oxidative stress | Standard | AR | Limited | B2 | 606145 | [20,21,23,24,84] |
| SLC39A13 (ZIP13) | Solute carrier family 39 (zinc transporter), member 13 (also known as glucose transporter 10) | Transports zinc, regulating intracellular levels, which affects lysyl hydroxylase activity/GAG chain elongation | 11p11.2 | Extracellular and Structural Matrix | No | No | Congenital disorder of glycosylation with systemic findings (no clear TAAD link) | N/A | AR | N/A | N/A | 608735 | [20,21,23,24,85] |
| SMAD2 | SMAD family member 2 | Intracellular mediator of TGF-β signaling; regulates gene transcription | 18q21.1 | TGF-β/SMAD Signaling | Yes—associated with connective tissue disorder, Loeys–Dietz syndrome type 6 | No | Arterial aneurysms and dissections (ascending, vertebral, carotid) | Standard | AD | N/A | N/A | 601366 | [11,20,21,23,24] |
| SMAD3 | SMAD family member 3 | Intracellular mediator of TGF-β signaling; regulates gene transcription | 15q22.33 | TGF-β/SMAD Signaling | Yes—Loeys–Dietz syndrome type 3 | Yes | Arterial aneurysms and dissections, tortuosity, skeletal findings (osteoarthritis) | 4.0–4.2 cm | AD | Definitive | A1 | 603109 | [13,20,21,23,24] |
| SMAD6 | SMAD family member 6 | Inhibitory SMAD that dampens TGF-β and BMP signaling | 15q22.31 | TGF-β/SMAD Signaling | No | Yes—associated with BAV/TAAD overlap | Aortic valve disease, BAV-associated aortic aneurysm | Standard | AD | No Evidence | D | 602931 | [20,21,23,24,86] |
| SPARC | Secreted protein, acidic, cysteine-rich (osteonectin) | Influences the synthesis and interaction of proteoglycans and collagens | 5q33.1 | Extracellular and Structural Matrix | No | No | Associated with Osteogenesis Imperfecta XVII; no clear vascular phenotype | N/A | AR | N/A | N/A | [20,21,23,24,87] | |
| TBX20 | T-box transcription factor 20 | Binds the DNA-binding domain T-box 20, influences cardiomorphogenesis | 7p14.2 | Transcriptional/Nuclear Envelope Regulation | No | Yes | Associated with BAV and aortic dilation through disrupted valvulogenesis and aortic patterning | N/A | AD | N/A | N/A | 606061 | [20,21,23,24,88,89] |
| TGFB2 | Transforming growth factor beta-2 | Ligand in the TGF-β pathway; regulates vascular development and remodeling | 1q41 | TGF-β/SMAD Signaling | Yes—Loeys–Dietz syndrome type 4 | Yes | Aneurysms, arterial tortuosity, other vascular anomalies, BAV | 4.5–5.0 cm | AD | Definitive | A1 | 190220 | [20,21,23,24,90] |
| TGFB3 | Transforming growth factor beta-3 | TGF-β family cytokine; modulates ECM and cellular proliferation | 14q24.3 | TGF-β/SMAD Signaling | Yes—Loeys–Dietz syndrome type 5 | No | Aneurysms, dissections, intracranial aneurysms | Standard | AD | N/A | N/A | 190230 | [20,21,23,24,91] |
| TGFBR1 | Transforming growth factor-beta receptor type I | Serine/threonine kinase receptor for TGF-β; initiates SMAD signaling cascade | 9q22.33 | TGF-β/SMAD Signaling | Yes—Loeys–Dietz syndrome type 1 | Rare, but possible | Aneurysms, arterial tortuosity, craniofacial features (bifid uvula, hypertelorism), skeletal dysplasia | 4.0–4.5 cm | AD | Definitive | A1 | 190181 | [11,12,20,21,23,24] |
| TGFBR2 | Transforming growth factor-beta receptor type II | Serine/threonine kinase that binds TGF-β ligands and activates TGFBR1 | 3p24.1 | TGF-β/SMAD Signaling | Yes—Loeys–Dietz syndrome type 2 | Rare, but possible | Aneurysms, widespread arterial dissection, craniofacial and skeletal features | 4.0–4.5 cm | AD | Definitive | A1 | 190182 | [11,12,20,21,23,24] |
| THBS2 | Thrombospondin-2 | Suppresses metalloproteinase activity, limiting extracellular matrix degradation | 6q27 | Extracellular and Structural Matrix | Yes—classic-like EDS | No | ECM fragility, connective tissue disease with vascular involvement suspected | N/A | AD | N/A | N/A | 188061 | [20,21,23,24,92] |
| THSD4 (ADAMTSL6) | Thrombospondin type 1 domain-containing protein 4 | Promotes the assembly of microfibrils through fibrillin-1 assembly and suppression of TGF-B | 15q23 | Extracellular and Structural Matrix | Yes | Yes—familial thoracic aortic aneurysm with variable expressivity | Aortic dilation, familial TAAD (AD pattern with variable penetrance) | N/A | AD | N/A | N/A | 614476 | [20,21,23,24,93] |
| TLN-1 | Talin-1 | Cytoskeletal adaptor protein that links integrins to the actin cytoskeleton | 9p13.3 | Smooth Muscle Contraction/Mechanotransduction | No | Yes | SCAD, thoracic aortic aneurysm, arrhythmogenic cardiomyopathy features (in some cases) | N/A | AD | N/A | N/A | 186745 | [20,21,23,24,26,94,95] |
| TNXB | Tenascin-XB | Collagen-organizing glycoprotein, which modulates fibril spacing via intercalation | 6p21.33-p21.32 | Extracellular and Structural Matrix | Yes—EDS classic-like (EDSCLL) | No | Skin hyperextensibility, easy bruising, connective tissue fragility | N/A | AR | N/A | N/A | 600985 | [20,21,23,24,89] |
| VCL | Vinculin | Actin-binding protein is important in cell–matrix adhesion and mechanical force transmission | 10q22.2 | Smooth Muscle Contraction/Mechanotransduction | No | Yes | Thoracic aortic aneurysm (rare), SCAD, dilated/arrhythmogenic cardiomyopathy | N/A | AD | N/A | N/A | 193065 | [20,21,23,24,96] |
| ZBTB20 | Zinc finger and BTB-containing protein 20 | Influences PI3K and MAPK signaling pathways and has been linked to vascular remodeling and SMC phenotypic modulation | 3q13.31 | Overlap/Dual-Phenotype/Emerging Candidates | No | Possibly | Disrupted dendritic and synaptic structure; proposed developmental influence | N/A | AD | N/A | N/A | 606025 | [20,21,23,24,97] |
| ZFHX3 | Zinc finger homeobox 3 | Zinc finger homeobox gene expressed in vascular tissues and may regulate cytoskeletal gene expression and inflammatory pathways relevant to medial degeneration | 16q22.2-q22.3 | Overlap/Dual-Phenotype/Emerging Candidates | Possibly | No | Syndromic intellectual disability with vascular anomalies | N/A | AD | N/A | N/A | 104155 | [20,21,23,24,98] |
| ZNF469 | Zinc finger protein 469 | Zinc finger transcription factor that regulates ECM gene expression, including collagen and proteoglycans | 16q24.2 | Overlap/Dual-Phenotype/Emerging Candidates | No | Possibly | ZNF469 participates in extracellular matrix regulation. It downregulates several collagen and adhesion molecules. While it has no confirmed clinical role in TAAD, it may function as a modifier gene. It is known for causing brittle cornea syndrome and could influence vascular fragility due to its ECM effect | N/A | AR | N/A | N/A | 612078 | [20,21,23,24,78] |


3.1.2. Ligand and Receptor Variants
3.1.3. Intracellular Mediators and Transcriptional Regulation
3.1.4. Negative Feedback and Nuclear Transport
3.1.5. Extracellular Matrix Modulators
3.1.6. Paradoxical Hyperactivation and Therapeutic Implications
3.1.7. Clinical Significance
3.2. Smooth Muscle Cell Contractility and Mechanotransduction
3.2.1. Core Mechanotransduction Overview
3.2.2. Actomyosin Machinery
3.2.3. Kinase Regulation of Contractility
3.2.4. Cytoskeletal–Nuclear and ECM Linkers
3.2.5. Focal Adhesion Proteins
3.2.6. RhoA-ROCK Signaling Axis
3.2.7. Sarcomere-Associated Genes
3.2.8. Summary and Clinical Implications
3.3. Extracellular Matrix and Structural Matrix Genes
3.3.1. Overview of ECM Function in TAAD
3.3.2. Elastic Fiber Network
3.3.3. Collagen Biosynthesis and Organization
3.3.4. Post-translational Crosslinking Enzymes
3.3.5. Organizational Glycoproteins
3.3.6. ECM-TGF-β Modulators
3.3.7. Proteoglycan and GAG Metabolism
3.3.8. Golgi/Ion Transport in ECM Processing
3.3.9. Additional ECM-Interacting Proteins
3.3.10. Summary and Clinical Implication
3.4. Cell–Cell Junctions and Aortic Wall Cohesion
3.4.1. Overview of Cell–Cell Junctions in Aortic Wall Integrity
3.4.2. Notch Pathway Regulation of VSMC Differentiation
3.4.3. Adherens Junctions
3.4.4. Desmosomal Proteins
3.4.5. ECM–Cell Anchoring Proteins
3.4.6. Focal Adhesion Mechanosensors
3.4.7. Summary and Clinical Implications
3.5. Transcriptional Regulators and Nuclear Envelope Genes
3.5.1. Overview of Nuclear and Transcriptional Regulation in TAAD
3.5.2. Nuclear Envelope and Mechanotransduction
3.5.3. Transcription Factors in Aortic Wall Remodeling
3.5.4. SMAD Nuclear Transport
3.5.5. Post-Transcriptional Regulators
3.5.6. Redox Regulation of Transcriptional Control
3.5.7. Mediator Complex and mTOR-Linked Regulators
3.5.8. Summary and Clinical Implications
3.6. Metabolic and Ion Transport
3.6.1. Overview of Metabolic and Ion Transport in TAAD
3.6.2. Mineral Homeostasis and ECM Calcification
3.6.3. Sulfur Amino Acid Metabolism and Epigenetic Regulation
3.6.4. Ion Channel Regulation of VSMC Tone
3.6.5. Solute Carrier Family and Vesicular Transport
3.6.6. Summary and Therapeutic Perspectives
3.7. Developmental and Cardiac Transcription Factors
3.7.1. Overview of Developmental and Cardiac Transcription Factors in TAAD
3.7.2. Notch Pathway Regulators
3.7.3. TGF-β-Linked Transcriptional Repressors
3.7.4. SMC Differentiation Factors
3.7.5. Chromatin-Modifying and Cytoskeletal Regulators
3.7.6. Negative Feedback and Mediator Complex Components
3.7.7. Metabolic and Mechanical Signal Integration (FLCN)
3.7.8. Summary and Developmental–Vascular Intersection
3.8. Proteoglycans, Glycosylation, and Linkeropathies
3.8.1. Overview of Proteoglycans in the Aortic ECM
3.8.2. Linker Region Synthesis
3.8.3. Dermatan Sulfate Chain Elongation and Sulfation
3.8.4. Small Leucine-Rich Proteoglycans
3.8.5. Post-Translational Modifications in Proteoglycan Biology
3.8.6. Transcriptional Regulators of ECM and Proteoglycan Genes
3.8.7. Summary and Clinical Implications
3.9. Emerging Genes and Hypothesis-Generating Candidates
3.9.1. Overview of Emerging Gene Candidates in TAAD
3.9.2. Proteoglycan Remodeling and ECM Regulation
3.9.3. TGF-β Pathway Modulators
3.9.4. Cytoskeletal and Cell–Cell Adhesion Proteins
3.9.5. Focal Adhesion Integrity
3.9.6. Desmosomal Proteins with Vascular Roles
3.9.7. Gap Junction Communication in Aortic Wall Cohesion
3.9.8. Transcriptional Regulators with Vascular Implications
3.9.9. Summary and Clinical Implications
3.10. Overlap Syndromes and Dual-Phenotype Genes
3.10.1. Developmental Regulators Linking Cardiac Morphogenesis and Aortic Integrity
3.10.2. Transcription Factors Bridging CHD and Arteriopathy
3.10.3. Nuclear Transport and Chromatin Regulators
3.10.4. Cytoskeletal and Signaling Proteins
3.10.5. Mitochondrial and Redox Pathway Genes
3.10.6. Summary and Clinical Implications
4. Clinical Translation and Future Directions
4.1. Diagnostic Yield and Gene-Specific Patterns
4.2. Gene-Informed Surgical and Surveillance Strategies
4.3. From Variant Function to Therapeutic Targeting
4.4. Expanded Phenotyping for Comprehensive Risk Assessment
4.5. Translational Parallels in Peripheral Artery Disease (PAD)
4.6. Genetic Differential Diagnosis and Overlap with Hypertrophic Cardiomyopathy
4.7. Family-Based Screening and Gene Panel Curation
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Isselbacher, E.M. Thoracic and abdominal aortic aneurysms. Circulation 2005, 111, 816–828. [Google Scholar] [CrossRef] [PubMed]
- Milewicz, D.M.; Cecchi, A.C. Heritable Thoracic Aortic Disease Overview. In GeneReviews; Adam, M.P., Feldman, J., Mirzaa, G.M., Pagon, R.A., Wallace, S.E., Amemiya, A., Eds.; University of Washington: Seattle, WA, USA, 2003; Updated 4 May 2023. Available online: https://www.ncbi.nlm.nih.gov/books/NBK1120/ (accessed on 2 July 2025).
- Dreher, L. TAAD Pathway Integration Network. 2025. Available online: https://BioRender.com/owqlfp6 (accessed on 2 July 2025).
- Klarin, D.; Devineni, P.; Sendamarai, A.K.; Angueira, A.R.; Graham, S.E.; Shen, Y.H.; Levin, M.G.; Pirruccello, J.P.; Surakka, I.; Karnam, P.R.; et al. Genome-wide association study of thoracic aortic aneurysm and dissection in the Million Veteran Program. Nat. Genet. 2023, 55, 1106–1115. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Chen, M.H.; Deng, E.S.; Yamada, J.M.; Choudhury, S.; Scotellaro, J.; Kelley, L.; Isselbacher, E.; Lindsay, M.E.; Walsh, C.A.; Doan, R.N. Contributions of Germline and Somatic Mosaic Genetics to Thoracic Aortic Aneurysms in Nonsyndromic Individuals. J. Am. Heart Assoc. 2024, 13, e033232. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Chen, Y.; Wang, L.; Xu, X.; Li, K.; Sun, Y.; Wang, Y.; Wang, D.W. Genetic architecture of thoracic aortic dissection in the female population. Gene 2023, 887, 147727. [Google Scholar] [CrossRef] [PubMed]
- Magouliotis, D.E.; Sicouri, S.; Sicouri, N.; Baudo, M.; Cabrucci, F.; Yamashita, Y.; Ramlawi, B. Epigenetic Biomarkers in Thoracic Aortic Aneurysm, Dissection, and Bicuspid Aortopathy: A Comprehensive Review. Biomolecules 2025, 15, 568. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Takeda, N.; Komuro, I. Genetic basis of hereditary thoracic aortic aneurysms and dissections. J. Cardiol. 2019, 74, 136–143. [Google Scholar] [CrossRef] [PubMed]
- Rodrigues Bento, J.; Meester, J.; Luyckx, I.; Peeters, S.; Verstraeten, A.; Loeys, B. The Genetics and Typical Traits of Thoracic Aortic Aneurysm and Dissection. Annu. Rev. Genomics Hum. Genet. 2022, 23, 223–253. [Google Scholar] [CrossRef] [PubMed]
- Ostberg, N.P.; Zafar, M.A.; Ziganshin, B.A.; Elefteriades, J.A. The Genetics of Thoracic Aortic Aneurysms and Dissection: A Clinical Perspective. Biomolecules 2020, 10, 182. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- MacCarrick, G.; Black, J.H., 3rd; Bowdin, S.; El-Hamamsy, I.; Frischmeyer-Guerrerio, P.A.; Guerrerio, A.L.; Sponseller, P.D.; Loeys, B.; Dietz, H.C., 3rd. Loeys-Dietz syndrome: A primer for diagnosis and management. Genet. Med. 2014, 16, 576–587. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Loeys, B.L.; Chen, J.; Neptune, E.R.; Judge, D.P.; Podowski, M.; Holm, T.; Meyers, J.; Leitch, C.C.; Katsanis, N.; Sharifi, N.; et al. A syndrome of altered cardiovascular, craniofacial, neurocognitive and skeletal development caused by mutations in TGFBR1 or TGFBR2. Nat. Genet. 2005, 37, 275–281. [Google Scholar] [CrossRef] [PubMed]
- van de Laar, I.M.; Oldenburg, R.A.; Pals, G.; Roos-Hesselink, J.W.; de Graaf, B.M.; Verhagen, J.M.; Hoedemaekers, Y.M.; Willemsen, R.; Severijnen, L.A.; Venselaar, H.; et al. Mutations in SMAD3 cause a syndromic form of aortic aneurysms and dissections with early-onset osteoarthritis. Nat. Genet. 2011, 43, 121–126. [Google Scholar] [CrossRef] [PubMed]
- Malfait, F.; Francomano, C.; Byers, P.; Belmont, J.; Berglund, B.; Black, J.; Bloom, L.; Bowen, J.M.; Brady, A.F.; Burrows, N.P.; et al. The 2017 international classification of the Ehlers-Danlos syndromes. Am. J. Med. Genet. C Semin. Med. Genet. 2017, 175, 8–26. [Google Scholar] [CrossRef] [PubMed]
- Venable, E.; Knight, D.R.T.; Thoreson, E.K.; Baudhuin, L.M. COL1A1 and COL1A2 variants in Ehlers-Danlos syndrome phenotypes and COL1-related overlap disorder. Am. J. Med. Genet. C Semin. Med. Genet. 2023, 193, 147–159. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.; Fang, X.; Ma, L.; Zhang, J.; Wang, C.; Gao, T.; Peng, D. FBN2 pathogenic mutation in congenital contractural arachnodactyly with severe skeletal manifestations. Mol. Genet. Metab. Rep. 2025, 42, 101193. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Dagoneau, N.; Benoist-Lasselin, C.; Huber, C.; Faivre, L.; Mégarbané, A.; Alswaid, A.; Dollfus, H.; Alembik, Y.; Munnich, A.; Legeai-Mallet, L.; et al. ADAMTS10 mutations in autosomal recessive Weill-Marchesani syndrome. Am. J. Hum. Genet. 2004, 75, 801–806. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Hucthagowder, V.; Sausgruber, N.; Kim, K.H.; Angle, B.; Marmorstein, L.Y.; Urban, Z. Fibulin-4: A novel gene for an autosomal recessive cutis laxa syndrome. Am. J. Hum. Genet. 2006, 78, 1075–1080. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Isselbacher, E.M.; Preventza, O.; Black, J.H., 3rd; Augoustides, J.G.; Beck, A.W.; Bolen, M.A.; Braverman, A.C.; Bray, B.E.; Brown-Zimmerman, M.M.; Chen, E.P.; et al. 2022 ACC/AHA Guideline for the Diagnosis and Management of Aortic Disease: A Report of the American Heart Association/American College of Cardiology Joint Committee on Clinical Practice Guidelines. Circulation 2022, 146, e334–e482. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Faggion Vinholo, T.; Brownstein, A.J.; Ziganshin, B.A.; Zafar, M.A.; Kuivaniemi, H.; Body, S.C.; Bale, A.E.; Elefteriades, J.A. Genes Associated with Thoracic Aortic Aneurysm and Dissection: 2019 Update and Clinical Implications. Aorta 2019, 7, 99–107. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Lin, C.J.; Lin, C.Y.; Stitziel, N.O. Genetics of the extracellular matrix in aortic aneurysmal diseases. Matrix Biol. 2018, 71–72, 128–143. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Moitra, K.; Garcia, S.; Jaldin, M.; Etoundi, C.; Cooper, D.; Roland, A.; Dixon, P.; Reyes, S.; Turan, S.; Terry, S.; et al. ABCC6 and Pseudoxanthoma Elasticum: The Face of a Rare Disease from Genetics to Advocacy. Int. J. Mol. Sci. 2017, 18, 1488. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Renard, M.; Francis, C.; Ghosh, R.; Scott, A.F.; Witmer, P.D.; Adès, L.C.; Andelfinger, G.U.; Arnaud, P.; Boileau, C.; Callewaert, B.L.; et al. Clinical Validity of Genes for Heritable Thoracic Aortic Aneurysm and Dissection. J. Am. Coll. Cardiol. 2018, 72, 605–615. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Online Mendelian Inheritance in Man (OMIM). Available online: https://www.ncbi.nlm.nih.gov/omim (accessed on 16 June 2025).
- Guo, D.C.; Papke, C.L.; Tran-Fadulu, V.; Regalado, E.S.; Avidan, N.; Johnson, R.J.; Kim, D.H.; Pannu, H.; Willing, M.C.; Sparks, E.; et al. Mutations in smooth muscle alpha-actin (ACTA2) cause coronary artery disease, stroke, and Moyamoya disease, along with thoracic aortic disease. Am. J. Hum. Genet. 2009, 84, 617–627. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Wei, X.; Sun, Y.; Wu, Y.; Zhu, J.; Gao, B.; Yan, H.; Zhao, Z.; Zhou, J.; Jing, Z. Downregulation of Talin-1 expression associates with increased proliferation and migration of vascular smooth muscle cells in aortic dissection. BMC Cardiovasc Disord 2017, 17, 162. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Minotti, G.; Menna, P.; Camilli, M.; Salvatorelli, E.; Levi, R. Beyond hypertension: Diastolic dysfunction associated with cancer treatment in the era of cardio-oncology. Adv. Pharmacol. 2022, 94, 365–409. [Google Scholar] [CrossRef] [PubMed]
- Van Damme, T.; Gardeitchik, T.; Mohamed, M.; Guerrero-Castillo, S.; Freisinger, P.; Guillemyn, B.; Kariminejad, A.; Dalloyaux, D.; van Kraaij, S.; Lefeber, D.J.; et al. Mutations in ATP6V1E1 or ATP6V1A Cause Autosomal-Recessive Cutis Laxa. Am. J. Hum. Genet. 2017, 100, 216–227. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Sudhahar, V.; Das, A.; Horimatsu, T.; Ash, D.; Leanhart, S.; Antipova, O.; Vogt, S.; Singla, B.; Csanyi, G.; White, J.; et al. Copper Transporter ATP7A (Copper-Transporting P-Type ATPase/Menkes ATPase) Limits Vascular Inflammation and Aortic Aneurysm Development: Role of MicroRNA-125b. Arterioscler. Thromb. Vasc. Biol. 2019, 39, 2320–2337. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Nakajima, M.; Mizumoto, S.; Miyake, N.; Kogawa, R.; Iida, A.; Ito, H.; Kitoh, H.; Hirayama, A.; Mitsubuchi, H.; Miyazaki, O.; et al. Mutations in B3GALT6, which encodes a glycosaminoglycan linker region enzyme, cause a spectrum of skeletal and connective tissue disorders. Am. J. Hum. Genet. 2013, 92, 927–934. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Ritelli, M.; Dordoni, C.; Cinquina, V.; Venturini, M.; Calzavara-Pinton, P.; Colombi, M. Expanding the clinical and mutational spectrum of B4GALT7-spondylodysplastic Ehlers-Danlos syndrome. Orphanet J. Rare Dis. 2017, 12, 153. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Meester, J.A.; Vandeweyer, G.; Pintelon, I.; Lammens, M.; Van Hoorick, L.; De Belder, S.; Waitzman, K.; Young, L.; Markham, L.W.; Vogt, J.; et al. Loss-of-function mutations in the X-linked biglycan gene cause a severe syndromic form of thoracic aortic aneurysms and dissections. Genet. Med. 2017, 19, 386–395. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Ganguly, P.; Alam, S.F. Role of homocysteine in the development of cardiovascular disease. Nutr. J. 2015, 14, 6. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Mayosi, B.M.; Fish, M.; Shaboodien, G.; Mastantuono, E.; Kraus, S.; Wieland, T.; Kotta, M.C.; Chin, A.; Laing, N.; Ntusi, N.B.; et al. Identification of Cadherin 2 (CDH2) Mutations in Arrhythmogenic Right Ventricular Cardiomyopathy. Circ. Cardiovasc. Genet. 2017, 10, e001605. [Google Scholar] [CrossRef] [PubMed]
- Janecke, A.R.; Li, B.; Boehm, M.; Krabichler, B.; Rohrbach, M.; Müller, T.; Fuchs, I.; Golas, G.; Katagiri, Y.; Ziegler, S.G.; et al. The phenotype of the musculocontractural type of Ehlers-Danlos syndrome due to CHST14 mutations. Am. J. Med. Genet. A 2016, 170A, 103–115. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Chiquet, M.; Birk, D.E.; Bönnemann, C.G.; Koch, M. Collagen XII: Protecting bone and muscle integrity by organizing collagen fibrils. Int. J. Biochem. Cell Biol. 2014, 53, 51–54. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Wenstrup, R.J.; Florer, J.B.; Brunskill, E.W.; Bell, S.M.; Chervoneva, I.; Birk, D.E. Type V collagen controls the initiation of collagen fibril assembly. J. Biol. Chem. 2004, 279, 53331–53337. [Google Scholar] [CrossRef] [PubMed]
- Schegg, B.; Hülsmeier, A.J.; Rutschmann, C.; Maag, C.; Hennet, T. Core glycosylation of collagen is initiated by two beta(1-O)galactosyltransferases. Mol. Cell Biol. 2009, 29, 943–952. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Puertas-Umbert, L.; Varona, S.; Ballester-Servera, C.; Alonso, J.; Aguiló, S.; Orriols, M.; Martínez-Martínez, E.; Rodríguez-Sinovas, A.; Martínez-González, J.; Rodríguez, C. Activation of Wnt/β-catenin signaling in abdominal aortic aneurysm: A potential therapeutic opportunity? Genes Dis. 2022, 10, 639–642. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Meens, M.J.; Pfenniger, A.; Kwak, B.R. Risky communication in atherosclerosis and thrombus formation. Swiss Med. Wkly. 2012, 142, w13553. [Google Scholar] [CrossRef] [PubMed]
- Shen, Y.; Russo, V.; Zeglinski, M.R.; Sellers, S.L.; Wu, Z.; Oram, C.; Santacruz, S.; Merkulova, Y.; Turner, C.; Tauh, K.; et al. Recombinant Decorin Fusion Protein Attenuates Murine Abdominal Aortic Aneurysm Formation and Rupture. Sci. Rep. 2017, 7, 15857. [Google Scholar] [CrossRef]
- Schirwani, S.; Metcalfe, K.; Wagner, B.; Berry, I.; Sobey, G.; Jewell, R. DSE associated musculocontractural EDS, a milder phenotype or phenotypic variability. Eur. J. Med. Genet. 2020, 63, 103798. [Google Scholar] [CrossRef] [PubMed]
- Brandão, M.; Bariani, R.; Rigato, I.; Bauce, B. Desmoplakin Cardiomyopathy: Comprehensive Review of an Increasingly Recognized Entity. J. Clin. Med. 2023, 12, 2660. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Min, S.; Kinnear, C.; D’Alessandro, L.C.A.; Bouwmeester, J.; Yao, R.; Chiasson, D.; Keeley, F.; Mital, S. Genetic Diagnosis and the Severity of Cardiovascular Phenotype in Patients With Elastin Arteriopathy. Circ. Genom. Precis. Med. 2020, 13, e002971. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Munjal, C.; Opoka, A.M.; Osinska, H.; James, J.F.; Bressan, G.M.; Hinton, R.B. TGF-β mediates early angiogenesis and latent fibrosis in an Emilin1-deficient mouse model of aortic valve disease. Dis. Model. Mech. 2014, 7, 987–996. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Ferreira, C.R.; Carpenter, T.O.; Braddock, D.T. ENPP1 in Blood and Bone: Skeletal and Soft Tissue Diseases Induced by ENPP1 Deficiency. Annu. Rev. Pathol. 2024, 19, 507–540. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Papke, C.L.; Yanagisawa, H. Fibulin-4 and fibulin-5 in elastogenesis and beyond: Insights from mouse and human studies. Matrix Biol. 2014, 37, 142–149. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Regalado, E.S.; Guo, D.C.; Santos-Cortez, R.L.; Hostetler, E.; Bensend, T.A.; Pannu, H.; Estrera, A.; Safi, H.; Mitchell, A.L.; Evans, J.P.; et al. Pathogenic FBN1 variants in familial thoracic aortic aneurysms and dissections. Clin. Genet. 2016, 89, 719–723. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Chen, M.H.; Choudhury, S.; Hirata, M.; Khalsa, S.; Chang, B.; Walsh, C.A. Thoracic aortic aneurysm in patients with loss of function Filamin A mutations: Clinical characterization, genetics, and recommendations. Am. J. Med. Genet. A 2018, 176, 337–350. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Zhou, W.D.; Liu, C.H.; Yin, X.M.; Zeng, Q.Y. A Novel Mutation of Filamin A Gene in a Chinese Family with Periventricular Nodular Heterotopia. Chin. Med. J. 2016, 129, 2262–2263. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Song, S.; Shi, A.; Lian, H.; Hu, S.; Nie, Y. Filamin C in cardiomyopathy: From physiological roles to DNA variants. Heart Fail. Rev. 2022, 27, 1373–1385. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Wang, X.C.; Gu, Z.H.; Ren, G.W.; Zhao, X.H.; Qu, X.K.; Xu, Y.J.; Yang, Y.Q. A novel GJA5 variant associated with increased risk of essential hypertension. Am. J. Transl. Res. 2023, 15, 1259–1270. [Google Scholar] [PubMed] [PubMed Central]
- van Walree, E.S.; Dombrowsky, G.; Jansen, I.E.; Mirkov, M.U.; Zwart, R.; Ilgun, A.; Guo, D.; Clur, S.B.; Amin, A.S.; Savage, J.E.; et al. Germline variants in HEY2 functional domains lead to congenital heart defects and thoracic aortic aneurysms. Genet. Med. 2021, 23, 103–110, Erratum in Genet. Med. 2022, 24, 965. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Bomsztyk, K.; Denisenko, O.; Ostrowski, J. hnRNP K: One protein multiple processes. Bioessays 2004, 26, 629–638. [Google Scholar] [CrossRef] [PubMed]
- Makeyev, A.V.; Liebhaber, S.A. The poly(C)-binding proteins: A multiplicity of functions and a search for mechanisms. RNA 2002, 8, 265–278. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Van Gucht, I.; Meester, J.A.N.; Bento, J.R.; Bastiaansen, M.; Bastianen, J.; Luyckx, I.; Van Den Heuvel, L.; Neutel, C.H.G.; Guns, P.J.; Vermont, M.; et al. A human importin-β-related disorder: Syndromic thoracic aortic aneurysm caused by bi-allelic loss-of-function variants in IPO8. Am. J. Hum. Genet. 2021, 108, 1115–1125. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Ziegler, A.; Duclaux-Loras, R.; Revenu, C.; Charbit-Henrion, F.; Begue, B.; Duroure, K.; Grimaud, L.; Guihot, A.L.; Desquiret-Dumas, V.; Zarhrate, M.; et al. Bi-allelic variants in IPO8 cause a connective tissue disorder associated with cardiovascular defects, skeletal abnormalities, and immune dysregulation. Am. J. Hum. Genet. 2021, 108, 1126–1137. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Rodrigues Bento, J.; Krebsová, A.; Van Gucht, I.; Valdivia Callejon, I.; Van Berendoncks, A.; Votypka, P.; Luyckx, I.; Peldova, P.; Laga, S.; Havelka, M.; et al. Isolated aneurysmal disease as an underestimated finding in individuals with JAG1 pathogenic variants. Hum. Mutat. 2022, 43, 1824–1828. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Tykocki, N.R.; Boerman, E.M.; Jackson, W.F. Smooth Muscle Ion Channels and Regulation of Vascular Tone in Resistance Arteries and Arterioles. Compr. Physiol. 2017, 7, 485–581. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Crasto, S.; My, I.; Di Pasquale, E. The Broad Spectrum of LMNA Cardiac Diseases: From Molecular Mechanisms to Clinical Phenotype. Front. Physiol. 2020, 11, 761. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Brownstein, A.J.; Kostiuk, V.; Ziganshin, B.A.; Zafar, M.A.; Kuivaniemi, H.; Body, S.C.; Bale, A.E.; Elefteriades, J.A. Genes Associated with Thoracic Aortic Aneurysm and Dissection: 2018 Update and Clinical Implications. Aorta 2018, 6, 13–20. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Ganizada, B.H.; Veltrop, R.J.A.; Akbulut, A.C.; Koenen, R.R.; Accord, R.; Lorusso, R.; Maessen, J.G.; Reesink, K.; Bidar, E.; Schurgers, L.J. Unveiling cellular and molecular aspects of ascending thoracic aortic aneurysms and dissections. Basic. Res. Cardiol. 2024, 119, 371–395. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Rifkin, D.; Sachan, N.; Singh, K.; Sauber, E.; Tellides, G.; Ramirez, F. The role of LTBPs in TGF beta signaling. Dev. Dyn. 2022, 251, 95–104. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.W.; Chou, S.H.; Tung, Y.C.; Hsiao, F.C.; Ho, C.T.; Chan, Y.H.; Wu, V.C.; Chou, A.H.; Hsu, M.E.; Lin, P.J.; et al. Expression and role of lumican in acute aortic dissection: A human and mouse study. PLoS ONE 2021, 16, e0255238. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Guo, D.C.; Gong, L.; Regalado, E.S.; Santos-Cortez, R.L.; Zhao, R.; Cai, B.; Veeraraghavan, S.; Prakash, S.K.; Johnson, R.J.; Muilenburg, A.; et al. MAT2A mutations predispose individuals to thoracic aortic aneurysms. Am. J. Hum. Genet. 2015, 96, 170–177. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Zhou, Y.; Zha, L.; Wu, J.; Wang, M.; Zhou, M.; Wu, G.; Cheng, X.; Huang, Z.; Xie, Q.; Tu, X. MED12 Regulates Smooth Muscle Cell Functions and Participates in the Development of Aortic Dissection. Genes. 2022, 13, 692. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Cheng, B.; Zhong, J.P.; Fu, W.J.; Chen, H.J.; Fang, L.; Li, G.L.; Li, J.W.; Wen, M.H.; Lv, Y.B.; Wang, H.B. Microfiber-associated protein 5 (MFAP5): A promising approach to discover new biomarkers for heart failure and cardiac remodeling. Int. J. Cardiol. 2022, 366, 68–69. [Google Scholar] [CrossRef] [PubMed]
- Zhu, L.; Vranckx, R.; Khau Van Kien, P.; Lalande, A.; Boisset, N.; Mathieu, F.; Wegman, M.; Glancy, L.; Gasc, J.M.; Brunotte, F.; et al. Mutations in myosin heavy chain 11 cause a syndrome associating thoracic aortic aneurysm/aortic dissection and patent ductus arteriosus. Nat. Genet. 2006, 38, 343–349. [Google Scholar] [CrossRef] [PubMed]
- Renard, M.; Callewaert, B.; Baetens, M.; Campens, L.; MacDermot, K.; Fryns, J.P.; Bonduelle, M.; Dietz, H.C.; Gaspar, I.M.; Cavaco, D.; et al. Novel MYH11 and ACTA2 mutations reveal a role for enhanced TGFβ signaling in FTAAD. Int. J. Cardiol. 2013, 165, 314–321. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Rombouts, K.B.; van Merrienboer, T.A.R.; Ket, J.C.F.; Bogunovic, N.; van der Velden, J.; Yeung, K.K. The role of vascular smooth muscle cells in the development of aortic aneurysms and dissections. Eur. J. Clin. Investig. 2022, 52, e13697. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Spin, J.M. Gene mutations and familial thoracic aortic aneurysms: A walk on the mild side. Circ. Cardiovasc. Genet. 2011, 4, 4–6. [Google Scholar] [CrossRef] [PubMed]
- MacGrogan, D.; Nus, M.; de la Pompa, J.L. Notch signaling in cardiac development and disease. Curr. Top. Dev. Biol. 2010, 92, 333–365. [Google Scholar] [CrossRef] [PubMed]
- Garg, V.; Muth, A.N.; Ransom, J.F.; Schluterman, M.K.; Barnes, R.; King, I.N.; Grossfeld, P.D.; Srivastava, D. Mutations in NOTCH1 cause aortic valve disease. Nature 2005, 437, 270–274. [Google Scholar] [CrossRef] [PubMed]
- Wang, D.; Fang, J.; Lv, J.; Pan, Z.; Yin, X.; Cheng, H.; Guo, X. Novel polymorphisms in PDLIM3 and PDLIM5 gene encoding Z-line proteins increase risk of idiopathic dilated cardiomyopathy. J. Cell Mol. Med. 2019, 23, 7054–7062. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Dalal, D.; Molin, L.H.; Piccini, J.; Tichnell, C.; James, C.; Bomma, C.; Prakasa, K.; Towbin, J.A.; Marcus, F.I.; Spevak, P.J.; et al. Clinical features of arrhythmogenic right ventricular dysplasia/cardiomyopathy associated with mutations in plakophilin-2. Circulation 2006, 113, 1641–1649. [Google Scholar] [CrossRef] [PubMed]
- Ewans, L.J.; Colley, A.; Gaston-Massuet, C.; Gualtieri, A.; Cowley, M.J.; McCabe, M.J.; Anand, D.; Lachke, S.A.; Scietti, L.; Forneris, F.; et al. Pathogenic variants in PLOD3 result in a Stickler syndrome-like connective tissue disorder with vascular complications. J. Med. Genet. 2019, 56, 629–638. [Google Scholar] [CrossRef] [PubMed]
- Sharad, S.; Dobi, A.; Srivastava, S.; Srinivasan, A.; Li, H. PMEPA1 Gene Isoforms: A Potential Biomarker and Therapeutic Target in Prostate Cancer. Biomolecules 2020, 10, 1221. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Moore, P.; Wolf, A.; Sathyamoorthy, M. An Eye into the Aorta: The Role of Extracellular Matrix Regulatory Genes ZNF469 and PRDM5, from Their Previous Association with Brittle Cornea Syndrome to Their Novel Association with Aortic and Arterial Aneurysmal Diseases. Int. J. Mol. Sci. 2024, 25, 5848. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Guo, D.C.; Regalado, E.; Casteel, D.E.; Santos-Cortez, R.L.; Gong, L.; Kim, J.J.; Dyack, S.; Horne, S.G.; Chang, G.; Jondeau, G.; et al. Recurrent gain-of-function mutation in PRKG1 causes thoracic aortic aneurysms and acute aortic dissections. Am. J. Hum. Genet. 2013, 93, 398–404. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Jones, C.A.; London, N.R.; Chen, H.; Park, K.W.; Sauvaget, D.; Stockton, R.A.; Wythe, J.D.; Suh, W.; Larrieu-Lahargue, F.; Mukouyama, Y.S.; et al. Robo4 stabilizes the vascular network by inhibiting pathologic angiogenesis and endothelial hyperpermeability. Nat. Med. 2008, 14, 448–453, Erratum in Nat. Med. 2008, 14, 585. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Shimizu, T.; Narang, N.; Chen, P.; Yu, B.; Knapp, M.; Janardanan, J.; Blair, J.; Liao, J.K. Fibroblast deletion of ROCK2 attenuates cardiac hypertrophy, fibrosis, and diastolic dysfunction. JCI Insight 2017, 2, e93187. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Schoenmakers, E.; Marelli, F.; Jørgensen, H.F.; Visser, W.E.; Moran, C.; Groeneweg, S.; Avalos, C.; Jurgens, S.J.; Figg, N.; Finigan, A.; et al. Selenoprotein deficiency disorder predisposes to aortic aneurysm formation. Nat. Commun. 2023, 14, 7994. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Doyle, A.J.; Doyle, J.J.; Bessling, S.L.; Maragh, S.; Lindsay, M.E.; Schepers, D.; Gillis, E.; Mortier, G.; Homfray, T.; Sauls, K.; et al. Mutations in the TGF-β repressor SKI cause Shprintzen-Goldberg syndrome with aortic aneurysm. Nat. Genet. 2012, 44, 1249–1254. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Coucke, P.J.; Willaert, A.; Wessels, M.W.; Callewaert, B.; Zoppi, N.; De Backer, J.; Fox, J.E.; Mancini, G.M.; Kambouris, M.; Gardella, R.; et al. Mutations in the facilitative glucose transporter GLUT10 alter angiogenesis and cause arterial tortuosity syndrome. Nat. Genet. 2006, 38, 452–457. [Google Scholar] [CrossRef] [PubMed]
- Liu, M.J.; Bao, S.; Gálvez-Peralta, M.; Pyle, C.J.; Rudawsky, A.C.; Pavlovicz, R.E.; Killilea, D.W.; Li, C.; Nebert, D.W.; Wewers, M.D.; et al. ZIP8 regulates host defense through zinc-mediated inhibition of NF-κB. Cell Rep. 2013, 3, 386–400. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Park, J.E.; Park, J.S.; Jang, S.Y.; Park, S.H.; Kim, J.W.; Ki, C.S.; Kim, D.K. A novel SMAD6 variant in a patient with severely calcified bicuspid aortic valve and thoracic aortic aneurysm. Mol. Genet. Genom. Med. 2019, 7, e620. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Veith, C.; Vartürk-Özcan, I.; Wujak, M.; Hadzic, S.; Wu, C.Y.; Knoepp, F.; Kraut, S.; Petrovic, A.; Gredic, M.; Pak, O.; et al. SPARC, a Novel Regulator of Vascular Cell Function in Pulmonary Hypertension. Circulation 2022, 145, 916–933. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.J.; Zou, S.; Bian, Y.Z.; Liu, X.Y.; Yang, C.X.; Li, L.; Qiu, X.B.; Xu, Y.J.; Yang, Y.Q.; Huang, R.T. Chromosomal Location and Identification of TBX20 as a New Gene Responsible for Familial Bicuspid Aortic Valve. Diagnostics 2025, 15, 600. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Matsumoto, K.I.; Aoki, H. The Roles of Tenascins in Cardiovascular, Inflammatory, and Heritable Connective Tissue Diseases. Front. Immunol. 2020, 11, 609752. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Boileau, C.; Guo, D.C.; Hanna, N.; Regalado, E.S.; Detaint, D.; Gong, L.; Varret, M.; Prakash, S.K.; Li, A.H.; d’Indy, H.; et al. TGFB2 mutations cause familial thoracic aortic aneurysms and dissections associated with mild systemic features of Marfan syndrome. Nat. Genet. 2012, 44, 916–921. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Bertoli-Avella, A.M.; Gillis, E.; Morisaki, H.; Verhagen, J.M.A.; de Graaf, B.M.; van de Beek, G.; Gallo, E.; Kruithof, B.P.T.; Venselaar, H.; Myers, L.A.; et al. Mutations in a TGF-β ligand, TGFB3, cause syndromic aortic aneurysms and dissections. J. Am. Coll. Cardiol. 2015, 65, 1324–1336. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Yang, Z.; Kyriakides, T.R.; Bornstein, P. Matricellular proteins as modulators of cell-matrix interactions: Adhesive defect in thrombospondin 2-null fibroblasts is a consequence of increased levels of matrix metalloproteinase-2. Mol. Biol. Cell 2000, 11, 3353–3364. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Elbitar, S.; Renard, M.; Arnaud, P.; Hanna, N.; Jacob, M.P.; Guo, D.C.; Tsutsui, K.; Gross, M.S.; Kessler, K.; Tosolini, L.; et al. Pathogenic variants in THSD4, encoding the ADAMTS-like 6 protein, predispose to inherited thoracic aortic aneurysm. Genet. Med. 2021, 23, 111–122. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Guo, D.C.; Regalado, E.S.; Gong, L.; Duan, X.; Santos-Cortez, R.L.; Arnaud, P.; Ren, Z.; Cai, B.; Hostetler, E.M.; Moran, R.; et al. LOX Mutations Predispose to Thoracic Aortic Aneurysms and Dissections. Circ. Res. 2016, 118, 928–934. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Turley, T.N.; Theis, J.L.; Sundsbak, R.S.; Evans, J.M.; O’Byrne, M.M.; Gulati, R.; Tweet, M.S.; Hayes, S.N.; Olson, T.M. Rare Missense Variants in TLN1 Are Associated With Familial and Sporadic Spontaneous Coronary Artery Dissection. Circ. Genom. Precis. Med. 2019, 12, e002437. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Wang, H.Q.; Yang, H.; Tang, Q.; Gong, Y.C.; Fu, Y.H.; Wan, F.; Yang, B.; Guo, R.; Zhong, Y.L.; Zhu, J.M.; et al. Identification of Vinculin as a Potential Diagnostic Biomarker for Acute Aortic Dissection Using Label-Free Proteomics. Biomed. Res. Int. 2020, 2020, 7806409. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Jones, K.A.; Luo, Y.; Dukes-Rimsky, L.; Srivastava, D.P.; Koul-Tewari, R.; Russell, T.A.; Shapiro, L.P.; Srivastava, A.K.; Penzes, P. Neurodevelopmental disorder-associated ZBTB20 gene variants affect dendritic and synaptic structure. PLoS ONE 2018, 13, e0203760. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Pérez Baca, M.D.R.; Jacobs, E.Z.; Vantomme, L.; Leblanc, P.; Bogaert, E.; Dheedene, A.; De Cock, L.; Haghshenas, S.; Foroutan, A.; Levy, M.A.; et al. Haploinsufficiency of ZFHX3, encoding a key player in neuronal development, causes syndromic intellectual disability. Am. J. Hum. Genet. 2024, 111, 509–528. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Dreher, L. Central Illustration: Integrated Molecular Pathway Underlying Thoracic Aortic Aneurysm and Dissection. 2025. Available online: https://BioRender.com/1hkoar3 (accessed on 5 June 2025).
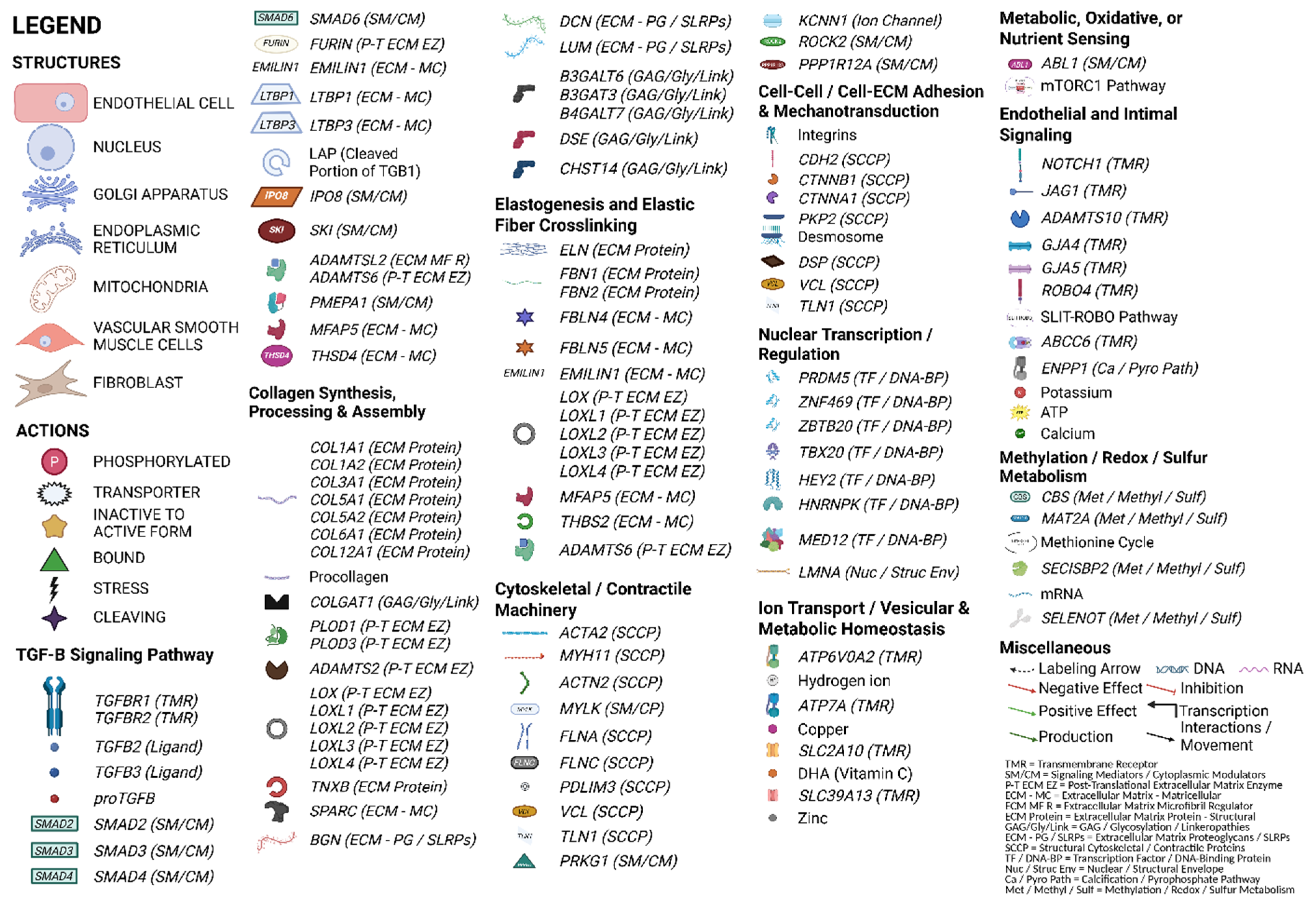
- Dreher, L. Legend. 2025. Available online: https://BioRender.com/ua3tzq9 (accessed on 5 June 2025).
- Pereira, L.; Andrikopoulos, K.; Tian, J.; Lee, S.Y.; Keene, D.R.; Ono, R.; Reinhardt, D.P.; Sakai, L.Y.; Biery, N.J.; Bunton, T.; et al. Targetting of the gene encoding fibrillin-1 recapitulates the vascular aspect of Marfan syndrome. Nat. Genet. 1997, 17, 218–222. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; Feng, H.; Tang, Y.; Wang, Z.; Qiu, P.; Huang, X.; Chang, L.; Zhang, J.; Chen, Y.E.; Mizrak, D.; et al. Bioengineered vascular grafts with a pathogenic TGFBR1 variant model aneurysm formation in vivo and reveal underlying collagen defects. Sci. Transl. Med. 2024, 16, eadg6298. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Good, J.M.; Fellmann, F.; Bhuiyan, Z.A.; Rotman, S.; Pruvot, E.; Schläpfer, J. ACTN2 variant associated with a cardiac phenotype suggestive of left-dominant arrhythmogenic cardiomyopathy. HeartRhythm Case Rep. 2019, 6, 15–19. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Dreher, L. Functional Overlap of Genes Implicated in Heritable TAAD. 2025. Available online: https://BioRender.com/jwulob5 (accessed on 1 July 2025).
- Tang, D.D.; Gerlach, B.D. The roles and regulation of the actin cytoskeleton, intermediate filaments and microtubules in smooth muscle cell migration. Respir. Res. 2017, 18, 54. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Ritelli, M.; Cinquina, V.; Giacopuzzi, E.; Venturini, M.; Chiarelli, N.; Colombi, M. Further Defining the Phenotypic Spectrum of B3GAT3 Mutations and Literature Review on Linkeropathy Syndromes. Genes 2019, 10, 631. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Ji, J.; Ding, K.; Luo, T.; Xu, R.; Zhang, X.; Huang, B.; Chen, A.; Zhang, D.; Miletic, H.; Bjerkvig, R.; et al. PMEPA1 isoform a drives progression of glioblastoma by promoting protein degradation of the Hippo pathway kinase LATS1. Oncogene 2020, 39, 1125–1139. [Google Scholar] [CrossRef]
- Huang, J.; Wang, H.; Xu, Y.; Li, C.; Lv, X.; Han, X.; Chen, X.; Chen, Y.; Yu, Z. The Role of CTNNA1 in Malignancies: An Updated Review. J. Cancer 2023, 14, 219–230. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Groenink, M.; den Hartog, A.W.; Franken, R.; Radonic, T.; de Waard, V.; Timmermans, J.; Scholte, A.J.; van den Berg, M.P.; Spijkerboer, A.M.; Marquering, H.A.; et al. Losartan reduces aortic dilatation rate in adults with Marfan syndrome: A randomized controlled trial. Eur. Heart J. 2013, 34, 3491–3500. [Google Scholar] [CrossRef] [PubMed]
- Mariscalco, G.; Debiec, R.; Elefteriades, J.A.; Samani, N.J.; Murphy, G.J. Systematic Review of Studies That Have Evaluated Screening Tests in Relatives of Patients Affected by Nonsyndromic Thoracic Aortic Disease. J. Am. Heart Assoc. 2018, 7, e009302. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Benrashid, E.; Ohman, J.W. Current management of the vascular subtype of Ehlers-Danlos syndrome. Curr. Opin. Cardiol. 2020, 35, 603–609. [Google Scholar] [CrossRef] [PubMed]
- Habashi, J.P.; Judge, D.P.; Holm, T.M.; Cohn, R.D.; Loeys, B.L.; Cooper, T.K.; Myers, L.; Klein, E.C.; Liu, G.; Calvi, C.; et al. Losartan, an AT1 antagonist, prevents aortic aneurysm in a mouse model of Marfan syndrome. Science 2006, 312, 117–121. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Chioncel, V.; Brezeanu, R.; Sinescu, C. New Directions in the Management of Peripheral Artery Disease. Am. J. Ther. 2019, 26, e284–e293. [Google Scholar] [CrossRef] [PubMed]
- Yousefzai, R.; Agarwal, A.; Jan, M.F.; Cho, C.; Anigbogu, M.; Shetabi, K.; Singh, M.; Bush, M.; Treiber, S.; Port, S.; et al. Hypertrophic cardiomyopathy with aortic dilation: A novel observation. Eur. Heart J. Cardiovasc. Imaging 2017, 18, 1398–1403. [Google Scholar] [CrossRef]
- Geske, J.B.; Nordhues, B.D.; Orme, N.M.; Tajik, A.J.; Spittell, P.C.; Ommen, S.R. Prevalence and Clinical Correlates of Aortic Dilation in Hypertrophic Cardiomyopathy. J. Am. Soc. Echocardiogr. 2021, 34, 279–285. [Google Scholar] [CrossRef] [PubMed]
- Mascia, G.; Olivotto, I.; Brugada, J.; Arbelo, E.; Di Donna, P.; Della Bona, R.; Canepa, M.; Porto, I. Sport practice in hypertrophic cardiomyopathy: Running to stand still? Int. J. Cardiol. 2021, 345, 77–82. [Google Scholar] [CrossRef] [PubMed]

Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Dreher, L.; Abdul Nabi, H.; Vandolah, H.; Brennan, S.; Bcharah, G.; Bcharah, H.; Osundiji, M.A.; Baudhuin, L.M.; Shamoun, F.E. The Expanding Genetic Architecture of Arteriopathies: From Canonical TAAD Genes to Emerging Connective Tissue and Signaling Pathways. Med. Sci. 2025, 13, 155. https://doi.org/10.3390/medsci13030155
Dreher L, Abdul Nabi H, Vandolah H, Brennan S, Bcharah G, Bcharah H, Osundiji MA, Baudhuin LM, Shamoun FE. The Expanding Genetic Architecture of Arteriopathies: From Canonical TAAD Genes to Emerging Connective Tissue and Signaling Pathways. Medical Sciences. 2025; 13(3):155. https://doi.org/10.3390/medsci13030155
Chicago/Turabian StyleDreher, Luke, Hussein Abdul Nabi, Hunter Vandolah, Stephen Brennan, George Bcharah, Hend Bcharah, Mayowa A. Osundiji, Linnea M. Baudhuin, and Fadi E. Shamoun. 2025. "The Expanding Genetic Architecture of Arteriopathies: From Canonical TAAD Genes to Emerging Connective Tissue and Signaling Pathways" Medical Sciences 13, no. 3: 155. https://doi.org/10.3390/medsci13030155
APA StyleDreher, L., Abdul Nabi, H., Vandolah, H., Brennan, S., Bcharah, G., Bcharah, H., Osundiji, M. A., Baudhuin, L. M., & Shamoun, F. E. (2025). The Expanding Genetic Architecture of Arteriopathies: From Canonical TAAD Genes to Emerging Connective Tissue and Signaling Pathways. Medical Sciences, 13(3), 155. https://doi.org/10.3390/medsci13030155






