Endolithic Microbial Habitats Hosted in Carbonate Nodules Currently Forming within Sediment at a High Methane Flux Site in the Sea of Japan
Abstract
1. Introduction
2. Materials and Methods
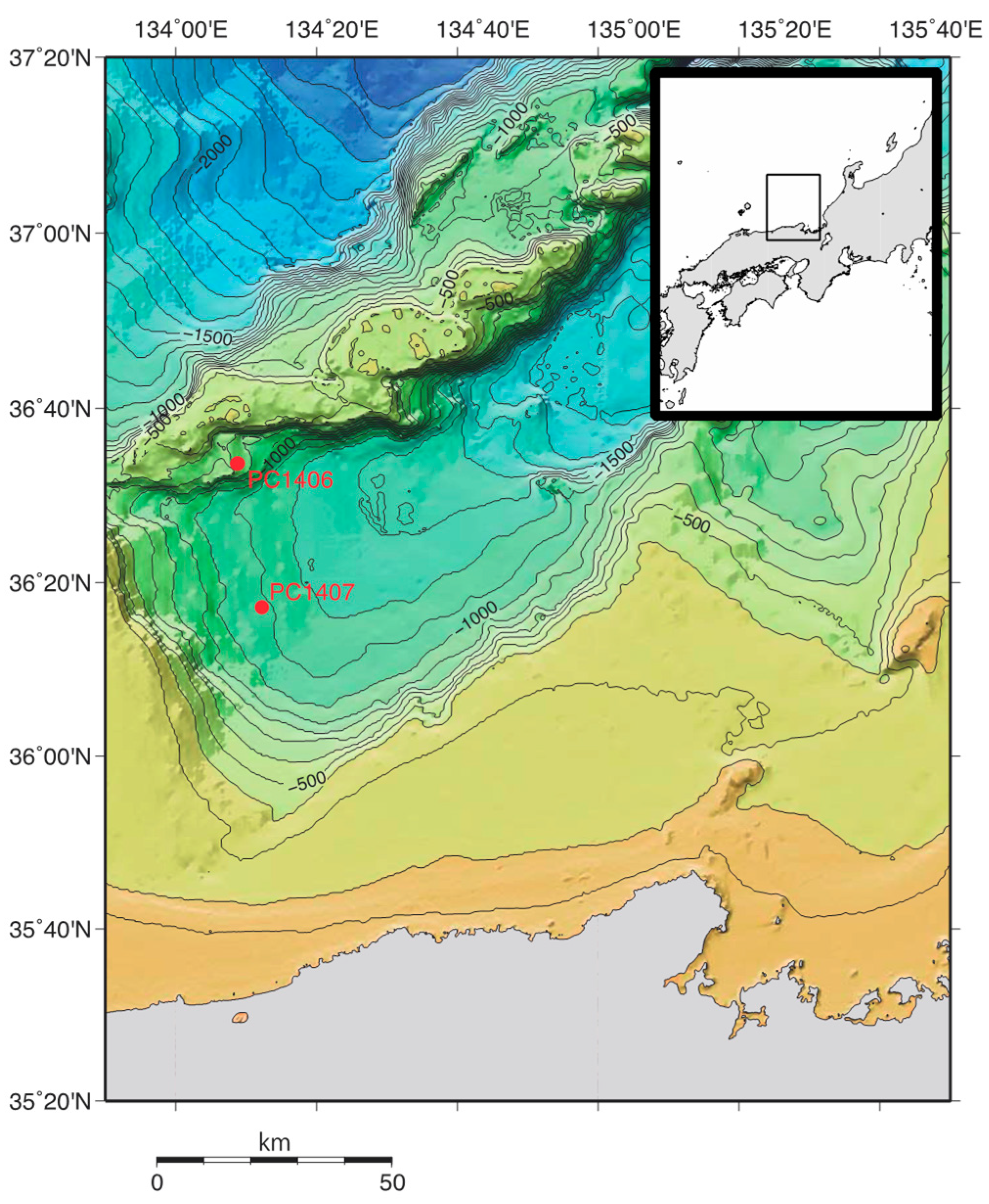
2.1. Sampling Sites and Sample Collection
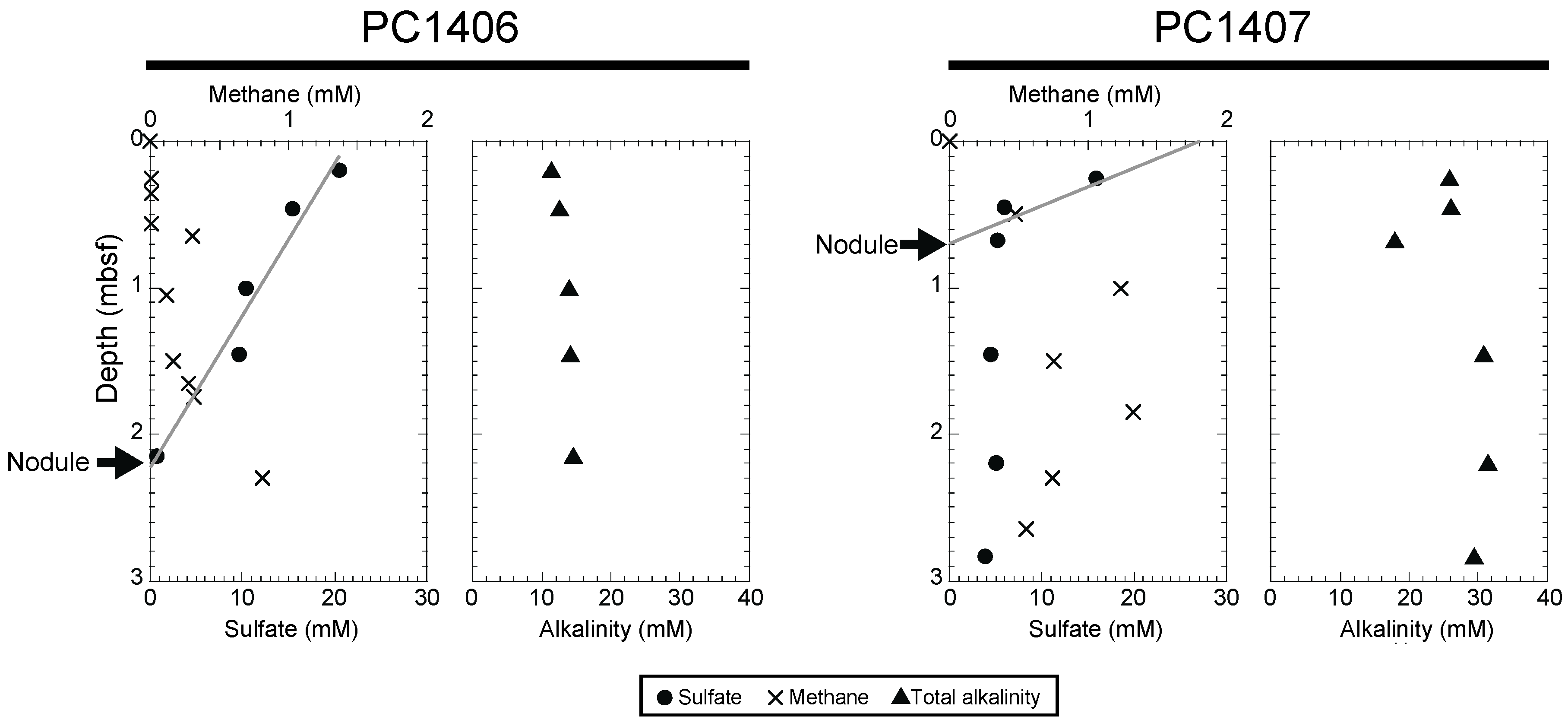
2.2. Pore Water Geochemistry
2.3. Mineral and Stable Isotopic Composition of Carbonate Nodules
2.4. Cell Count
2.5. DNA Extraction
2.6. Quantitative Real-time PCR (Q-PCR)
2.7. McrA Gene Clone Library Analysis
2.8. 16S rRNA Gene Phylotype Composition Analysis
3. Results and Discussion
3.1. Geochemical Characteristics of the Nodule-Bearing Sediment
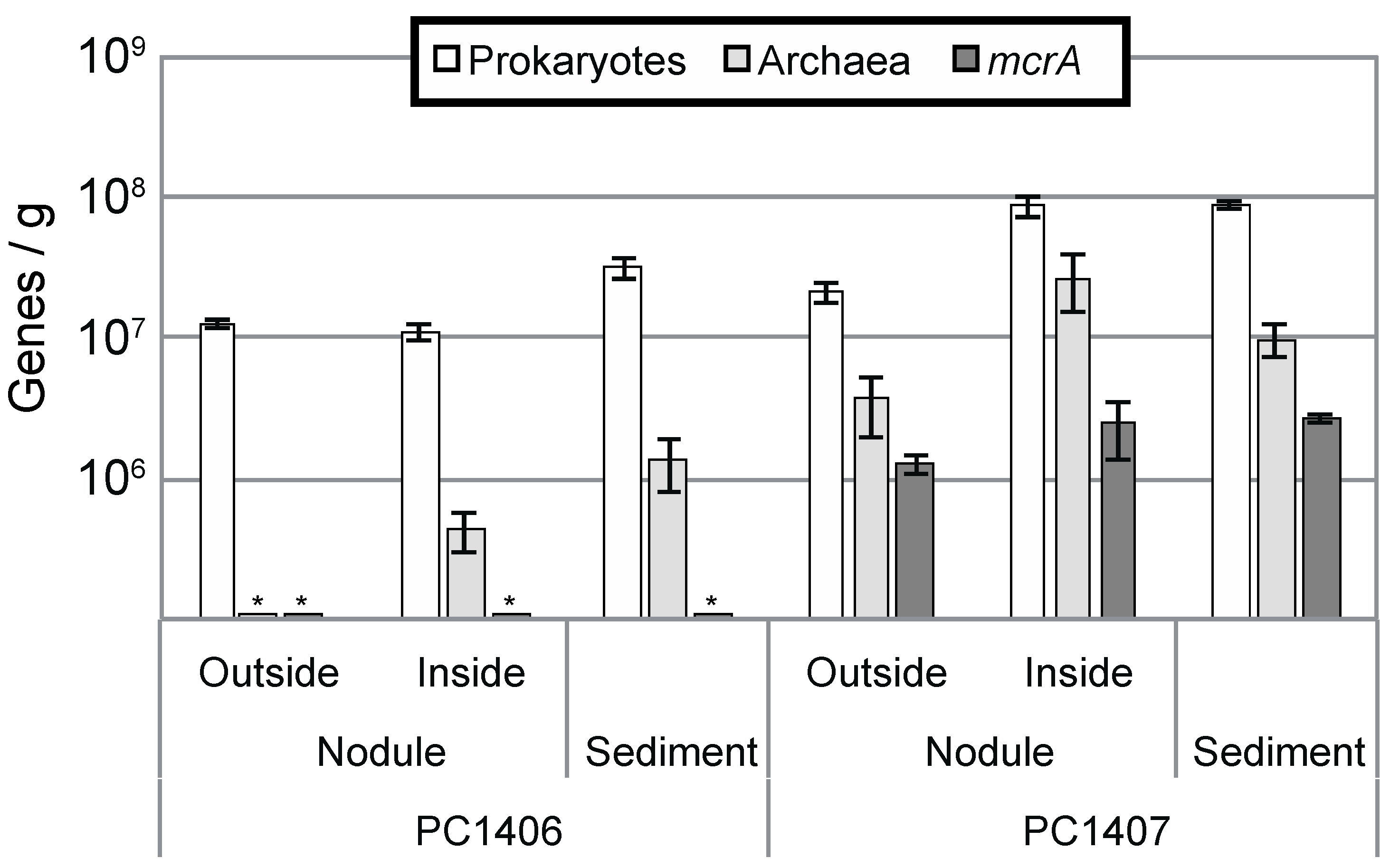
3.2. Microbial Abundance in the Nodule
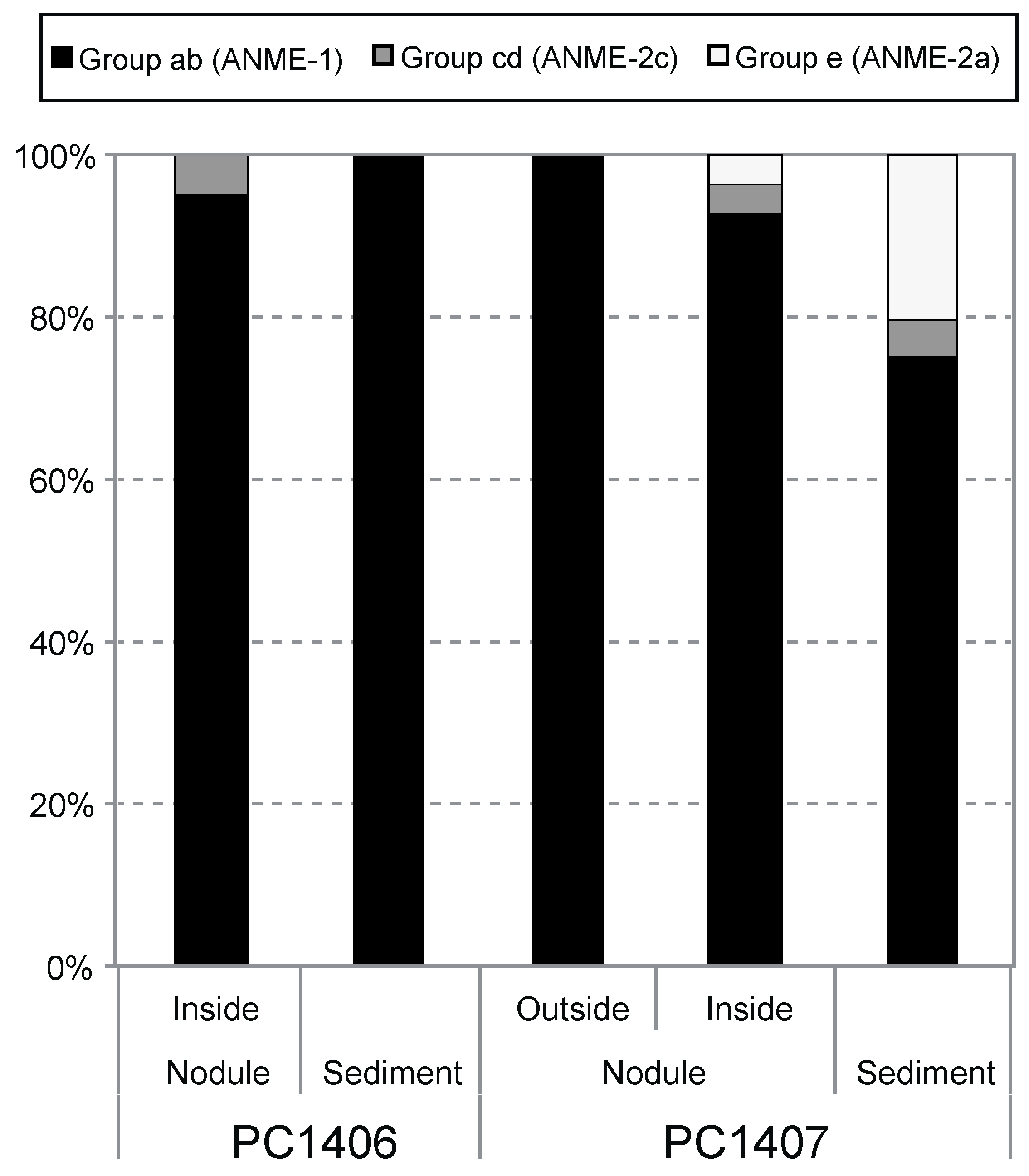
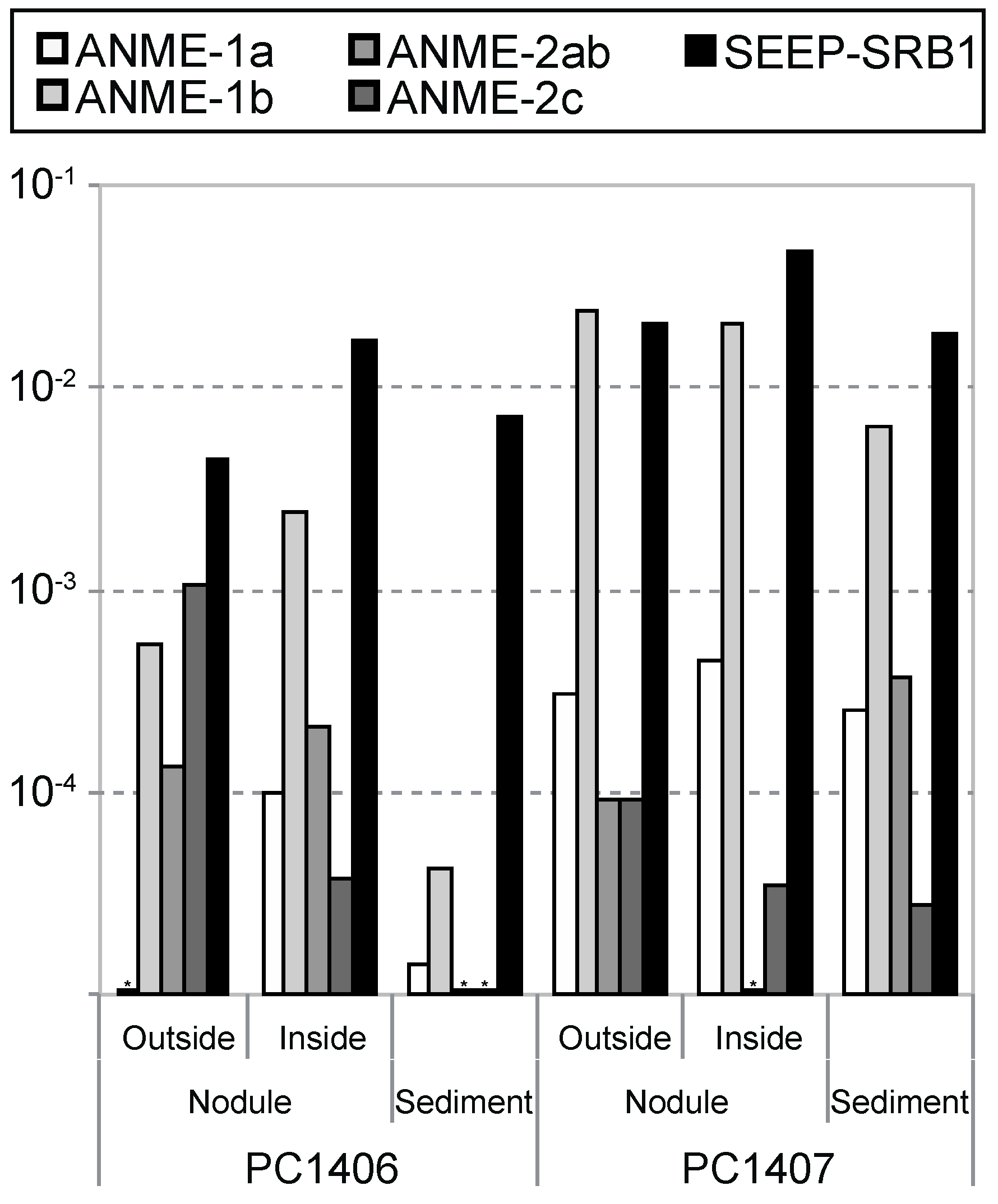
3.3. McrA Phylotype Composition in the Nodule Habitat
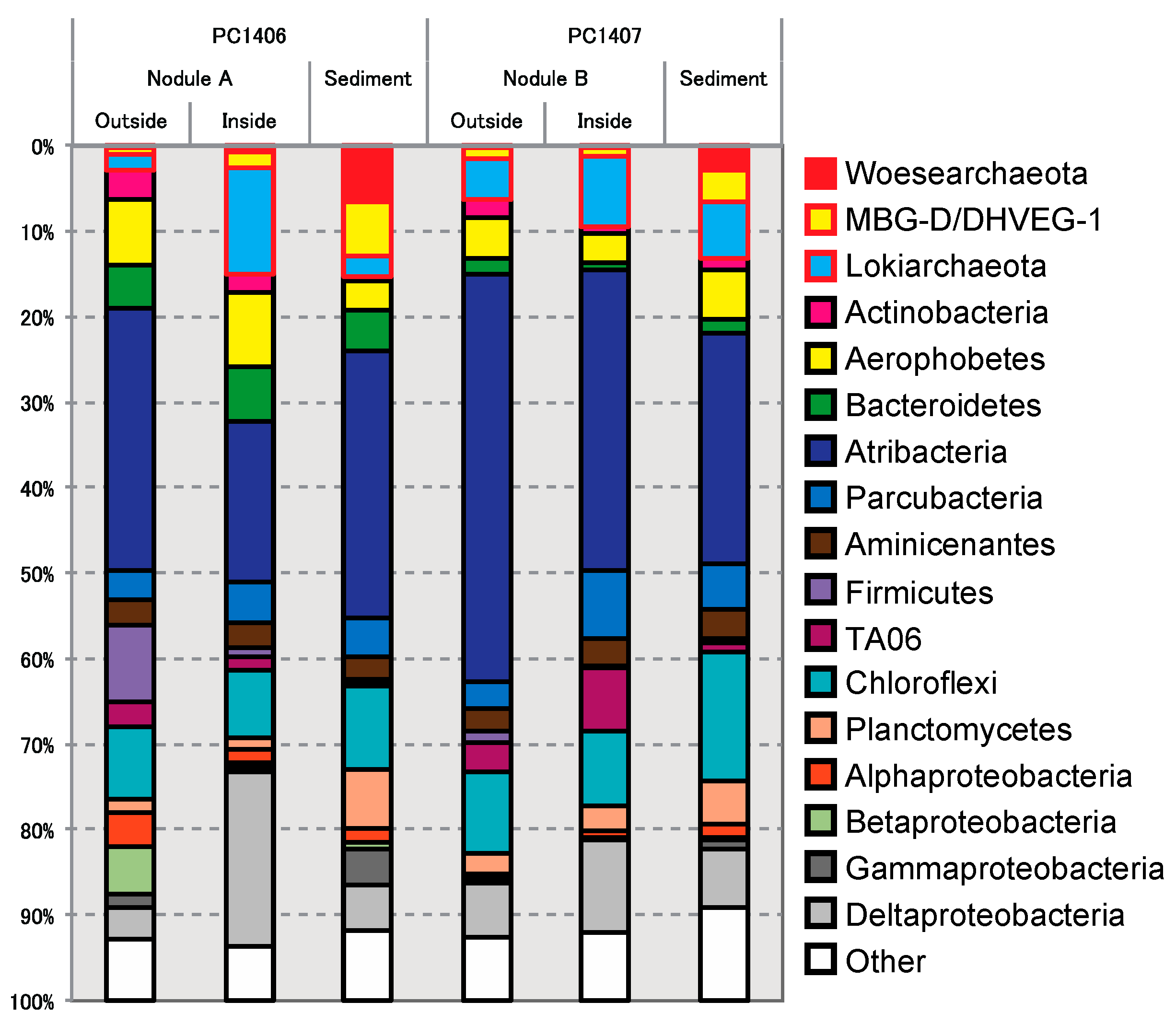
3.4. Nodule-Hosted Microbial Community Compositions
3.5. AOM-Responsible Microbial Life within the Nodules at the High Methane Flux Site
4. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Orphan, V.J.; Ussler, W.; Naehr, T.H.; House, C.H.; Hinrichs, K.U.; Paull, C.K. Geological, geochemical, and microbiological heterogeneity of the seafloor around methane vents in the Eel River Basin, offshore California. Chem. Geol. 2004, 205, 265–289. [Google Scholar] [CrossRef]
- Watanabe, Y.; Nakai, S.; Hiruta, A.; Matsumoto, R.; Yoshida, K. U-Th dating of carbonate nodules from methane seeps off Joetsu, Eastern Margin of Japan Sea. Earth. Planet. Sci. Lett. 2008, 272, 89–96. [Google Scholar] [CrossRef]
- Hiruta, A.; Klügel, A.; Matsumoto, R. Increase in methane flux and dissociation of iron and manganese oxides recorded in a methane-derived carbonate nodule in the eastern margin of the Sea of Japan. GeoResJ 2016, 9, 104–116. [Google Scholar] [CrossRef]
- Mason, O.U.; Case, D.H.; Naehr, T.H.; Lee, R.W.; Thomas, R.B.; Bailey, J.V.; Orphan, V.J. Comparison of archaeal and bacterial diversity in methane seep carbonate nodules and host sediments, Eel River Basin and Hydrate Ridge, USA. Microb. Ecol. 2015, 70, 766–784. [Google Scholar] [CrossRef] [PubMed]
- Aloisi, G.; Bouloubassi, I.; Heijs, S.K.; Pancost, R.D.; Pierre, C.; Sinninghe Damsté, J.S.; Gottschal, J.C.; Forney, L.J.; Rouchy, J.-M. CH4-consuming microorganisms and the formation of carbonate crusts at cold seeps. Earth Planet. Sci. Lett. 2002, 203, 195–203. [Google Scholar] [CrossRef]
- Valentine, D.L. Biogeochemistry and microbial ecology of methane oxidation in anoxic environments: A review. Anton. Leeuw. Int. J. G. 2002, 81, 271–282. [Google Scholar] [CrossRef]
- Valentine, D.L.; Reeburgh, W.S. New perspectives on anaerobic methane oxidation. Environ. Microbiol. 2000, 2, 477–484. [Google Scholar] [CrossRef]
- Knittel, K.; Boetius, A. Anaerobic oxidation of methane: Progress with an unknown process. Annu. Rev. Microbiol. 2009, 63, 311–334. [Google Scholar] [CrossRef]
- Lösekann, T.; Knittel, K.; Nadalig, T.; Fuchs, B.; Niemann, H.; Boetius, A.; Amann, R. Diversity and abundance of aerobic and anaerobic methane oxidizers at the Haakon Mosby mud volcano, Barents Sea. Appl. Environ. Microbiol. 2007, 73, 3348–3362. [Google Scholar] [CrossRef]
- Scheller, S.; Yu, H.; Chadwick, G.L.; McGlynn, S.E.; Orphan, V.J. Artificial electron acceptors decouple archaeal methane oxidation from sulfate reduction. Science 2016, 351, 703–707. [Google Scholar] [CrossRef]
- Milucka, J.; Ferdelman, T.G.; Polerecky, L.; Franzke, D.; Wegener, G.; Schmid, M.; Lieberwirth, I.; Wagner, M.; Widdel, F.; Kuypers, M.M.M. Zero-valent sulphur is a key intermediate in marine methane oxidation. Nature 2012, 491, 541. [Google Scholar] [CrossRef] [PubMed]
- McGlynn, S.E.; Chadwick, G.L.; Kempes, C.P.; Orphan, V.J. Single cell activity reveals direct electron transfer in methanotrophic consortia. Nature 2015, 526, 531. [Google Scholar] [CrossRef] [PubMed]
- Wegener, G.; Krukenberg, V.; Riedel, D.; Tegetmeyer, H.E.; Boetius, A. Intercellular wiring enables electron transfer between methanotrophic archaea and bacteria. Nature 2015, 526, 587. [Google Scholar] [CrossRef] [PubMed]
- Hallam, S.J.; Putnam, N.; Preston, C.M.; Detter, J.C.; Rokhsar, D.; Richardson, P.M.; DeLong, E.F. Reverse methanogenesis: Testing the hypothesis with environmental genomics. Science 2004, 305, 1457–1462. [Google Scholar] [CrossRef] [PubMed]
- Hinrichs, K.U.; Hayes, J.M.; Sylva, S.P.; Brewer, P.G.; DeLong, E.F. Methane-consuming archaebacteria in marine sediments. Nature 1999, 398, 802–805. [Google Scholar] [CrossRef]
- Meyerdierks, A.; Kube, M.; Lombardot, T.; Knittel, K.; Bauer, M.; Glockner, F.O.; Reinhardt, R.; Amann, R. Insights into the genomes of archaea mediating the anaerobic oxidation of methane. Environ. Microbiol. 2005, 7, 1937–1951. [Google Scholar] [CrossRef] [PubMed]
- Orphan, V.J.; House, C.H.; Hinrichs, K.U.; McKeegan, K.D.; DeLong, E.F. Multiple archaeal groups mediate methane oxidation in anoxic cold seep sediments. Proc. Natl. Acad. Sci. USA 2002, 99, 7663–7668. [Google Scholar] [CrossRef]
- Treude, T.; Krüger, M.; Boetius, A.; Jørgensen, B.B. Environmental control on anaerobic oxidation of methane in the gassy sediments of Eckernförde Bay (German Baltic). Limnol. Oceanogr. 2005, 50, 1771–1786. [Google Scholar] [CrossRef]
- Yanagawa, K.; Tomaru, H.; Sunamura, M.; Matsumoto, R. Thermodynamic control on anaerobic oxidation of methane below the sulphate-methane interface. In Proceedings of the 7th International Conference on Gas Hydrates, Scotland, UK, 17–21 July 2011. [Google Scholar]
- Orphan, V.J.; House, C.H.; Hinrichs, K.U.; McKeegan, K.D.; DeLong, E.F. Methane-consuming archaea revealed by directly coupled isotopic and phylogenetic analysis. Science 2001, 293, 484–487. [Google Scholar] [CrossRef]
- Yanagawa, K.; Morono, Y.; Beer, D.d.; Haeckel, M.; Sunamura, M.; Futagami, T.; Hoshino, T.; Terada, T.; Nakamura, K.-L.; Urabe, T.; et al. Metabolically active microbial communities in marine sediment under high-CO2 and low-pH extremes. ISME J. 2013, 7, 555–567. [Google Scholar] [CrossRef]
- Haroon, M.F.; Hu, S.; Shi, Y.; Imelfort, M.; Keller, J.; Hugenholtz, P.; Yuan, Z.; Tyson, G.W. Anaerobic oxidation of methane coupled to nitrate reduction in a novel archaeal lineage. Nature 2013, 500, 567. [Google Scholar] [CrossRef] [PubMed]
- Ino, K.; Hernsdorf, A.W.; Konno, U.; Kouduka, M.; Yanagawa, K.; Kato, S.; Sunamura, M.; Hirota, A.; Togo, Y.S.; Ito, K.; et al. Ecological and genomic profiling of anaerobic methane-oxidizing archaea in a deep granitic environment. ISME J. 2018, 12, 31–47. [Google Scholar] [CrossRef] [PubMed]
- Boetius, A.; Ravenschlag, K.; Schubert, C.J.; Rickert, D.; Widdel, F.; Gieseke, A.; Amann, R.; Jørgensen, B.B.; Witte, U.; Pfannkuche, O. A marine microbial consortium apparently mediating anaerobic oxidation of methane. Nature 2000, 407, 623–626. [Google Scholar] [CrossRef]
- Kleindienst, S.; Ramette, A.; Amann, R.; Knittel, K. Distribution and in situ abundance of sulfate-reducing bacteria in diverse marine hydrocarbon seep sediments. Environ. Microbiol. 2012, 14, 2689–2710. [Google Scholar] [CrossRef] [PubMed]
- Pernthaler, A.; Dekas, A.E.; Brown, C.T.; Goffredi, S.K.; Embaye, T.; Orphan, V.J. Diverse syntrophic partnerships from deep-sea methane vents revealed by direct cell capture and metagenomics. Proc. Natl. Acad. Sci. USA 2008, 105, 7052–7057. [Google Scholar] [CrossRef] [PubMed]
- Hatzenpichler, R.; Connon, S.A.; Goudeau, D.; Malmstrom, R.R.; Woyke, T.; Orphan, V.J. Visualizing in situ translational activity for identifying and sorting slow-growing archaeal−bacterial consortia. Proc. Natl. Acad. Sci. USA 2016, 113, E4069–E4078. [Google Scholar] [CrossRef]
- Peckmann, J.; Thiel, V. Carbon cycling at ancient methane–seeps. Chem. Geol. 2004, 205, 443–467. [Google Scholar] [CrossRef]
- Blumenberg, M.; Seifert, R.; Reitner, J.; Pape, T.; Michaelis, W. Membrane lipid patterns typify distinct anaerobic methanotrophic consortia. Proc. Natl. Acad. Sci. USA 2004, 101, 11111–11116. [Google Scholar] [CrossRef]
- Stadnitskaia, A.; Nadezhkin, D.; Abbas, B.; Blinova, V.; Ivanov, M.K.; Damste, J.S.S. Carbonate formation by anaerobic oxidation of methane: Evidence from lipid biomarker and fossil 16S rDNA. Geochimi. Cosmochimi. Acta 2008, 72, 1824–1836. [Google Scholar] [CrossRef]
- Thiel, V.; Peckmann, J.; Seifert, R.; Wehrung, P.; Reitner, J.; Michaelis, W. Highly isotopically depleted isoprenoids: Molecular markers for ancient methane venting. Geochimi. Cosmochimi. Acta 1999, 63, 3959–3966. [Google Scholar] [CrossRef]
- Reitner, J.; Peckmann, J.; Blumenberg, M.; Michaelis, W.; Reimer, A.; Thiel, V. Concretionary methane-seep carbonates and associated microbial communities in Black Sea sediments. Palaeogeogr. Palaeoclimatol. Palaeoecol. 2005, 227, 18–30. [Google Scholar] [CrossRef]
- Marlow, J.J.; Steele, J.A.; Ziebis, W.; Thurber, A.R.; Levin, L.A.; Orphan, V.J. Carbonate-hosted methanotrophy represents an unrecognized methane sink in the deep sea. Nat. Commun. 2014, 5, 5094. [Google Scholar] [CrossRef] [PubMed]
- Ruff, S.E.; Biddle, J.F.; Teske, A.P.; Knittel, K.; Boetius, A.; Ramette, A. Global dispersion and local diversification of the methane seep microbiome. Proc. Natl. Acad. Sci. USA 2015, 112, 4015–4020. [Google Scholar] [CrossRef] [PubMed]
- Gieskes, J.; Mahn, C.; Day, S.; Martin, J.B.; Greinert, J.; Rathburn, T.; McAdoo, B. A study of the chemistry of pore fluids and authigenic carbonates in methane seep environments: Kodiak Trench, Hydrate Ridge, Monterey Bay, and Eel River Basin. Chem. Geol. 2005, 220, 329–345. [Google Scholar] [CrossRef]
- Teichert, B.M.A.; Bohrmann, G.; Suess, E. Chemoherms on Hydrate Ridge — Unique microbially-mediated carbonate build-ups growing into the water column. Palaeogeogr. Palaeoclimatol. Palaeoecol. 2005, 227, 67–85. [Google Scholar] [CrossRef]
- Birgel, D.; Himmler, T.; Freiwald, A.; Peckmann, J. A new constraint on the antiquity of anaerobic oxidation of methane: Late Pennsylvanian seep limestones from southern Namibia. Geology 2008, 36, 543–546. [Google Scholar] [CrossRef]
- Marlow, J.J.; Steele, J.A.; Case, D.H.; Connon, S.A.; Levin, L.A.; Orphan, V.J. Microbial abundance and diversity patterns associated with sediments and carbonates from the methane seep environments of Hydrate Ridge, OR. Front. Mar. Sci. 2014, 1, 44. [Google Scholar] [CrossRef]
- Case, D.H.; Pasulka, A.L.; Marlow, J.J.; Grupe, B.M.; Levin, L.A.; Orphan, V.J. Methane seep carbonates host distinct, diverse, and dynamic microbial assemblages. MBio 2015, 6, e01348-15. [Google Scholar] [CrossRef]
- Matsumoto, R.; Okuda, Y.; Aoyama, C.; Hiruta, A.; Ishida, Y.; Sunamura, M.; Numanami, H.; Tomaru, H.; Snyder, G.T.; Komatsubara, J.; et al. Methane plumes over a marine gas hydrate system in the eastern margin of Japan Sea: A possible mechanism for the transportation of subsurface methane to shallow waters. In Proceedings of the Fifth International Conference on Gas Hydrate, Trondheim, Norway, 13–16 June 2005. [Google Scholar]
- Matsumoto, R.; Kakuwa, Y.; Tanahashi, M. Occurrence and origin of shallow gas hydrates of the eastern margin of Japan Sea as revealed by Calypso and CASQ corings of R/V Marion Dufresne. In Proceedings of the 7th International Conference on Gas Hydrates, Scotland, UK, 17–21 July 2011. [Google Scholar]
- Tomaru, H.; Lu, Z.L.; Snyder, G.T.; Fehn, U.; Hiruta, A.; Matsumoto, R. Origin and age of pore waters in an actively venting gas hydrate field near Sado Island, Japan Sea: Interpretation of halogen and I-129 distributions. Chem. Geol. 2007, 236, 350–366. [Google Scholar] [CrossRef]
- Hachikubo, A.; Yanagawa, K.; Tomaru, H.; Lu, H.; Matsumoto, R. Molecular and isotopic composition of volatiles in gas hydrates and in sediment from the Joetsu Basin, eastern margin of the Japan Sea. Energies 2015, 8, 4647–4666. [Google Scholar] [CrossRef]
- Freire, A.F.M.; Matsumoto, R.; Santos, L.A. Structural-stratigraphic control on the Umitaka Spur gas hydrates of Joetsu Basin in the eastern margin of Japan Sea. Mar. Pet. Geol. 2011, 28, 1967–1978. [Google Scholar] [CrossRef]
- Kano, A.; Miyahara, R.; Yanagawa, K.; Mori, T.; Owari, S.; Tomaru, H.; Kakizaki, Y.; Snyder, G.; Shimono, T.; Kakuwa, Y.; et al. Gas hydrate estimates in muddy sediments from the oxygen isotope of water fraction. Chem. Geol. 2017, 470, 107–115. [Google Scholar] [CrossRef]
- Kouduka, M.; Tanabe, A.S.; Yamamoto, S.; Yanagawa, K.; Nakamura, Y.; Akiba, F.; Tomaru, H.; Toju, H.; Suzuki, Y. Eukaryotic diversity in late Pleistocene marine sediments around a shallow methane hydrate deposit in the Japan Sea. Geobiology 2017, 15, 715–727. [Google Scholar] [CrossRef] [PubMed]
- Yanagawa, K.; Kouduka, M.; Nakamura, Y.; Hachikubo, A.; Tomaru, H.; Suzuki, Y. Distinct microbial communities thriving in gas hydrate-associated sediments from the eastern Japan Sea. J. Asian Earth Sci. 2014, 90, 243–249. [Google Scholar] [CrossRef]
- Yanagawa, K.; Sunamura, M.; Lever, M.A.; Morono, Y.; Hiruta, A.; Ishizaki, O.; Matsumoto, R.; Urabe, T.; Inagaki, F. Niche separation of methanotrophic archaea (ANME-1 and -2) in methane-seep sediments of the eastern Japan Sea offshore Joetsu. Geomicrobiol. J. 2011, 28, 118–129. [Google Scholar] [CrossRef]
- Yanagawa, K.; Tani, A.; Yamamoto, N.; Hachikubo, A.; Kano, A.; Matsumoto, R.; Suzuki, Y. Biogeochemical Cycle of Methanol in Anoxic Deep-Sea Sediments. Microbes Environ. 2016, 31, 190–193. [Google Scholar] [CrossRef]
- Manheim, F.T.; Brooks, E.G.; Winters, W.J. Description of A Hydraulic Sediment Squeezer; Citeseer: Princeton, NJ, USA, 1994; pp. 94–584. [Google Scholar]
- Tomaru, H.; Fehn, U.; Lu, Z.; Takeuchi, R.; Inagaki, F.; Imachi, H.; Kotani, R.; Matsumoto, R.; Aoike, K. Dating of Dissolved Iodine in Pore Waters from the Gas Hydrate Occurrence Offshore Shimokita Peninsula, Japan: 129I Results from the D/V Chikyu Shakedown Cruise. Resour. Geol. 2009, 59, 359–373. [Google Scholar] [CrossRef]
- Sarazin, G.; Michard, G.; Prevot, F. A rapid and accurate spectroscopic method for alkalinity measurements in sea water samples. Water Res. 1999, 33, 290–294. [Google Scholar] [CrossRef]
- Yanagawa, K.; Morono, Y.; Yoshida-Takashima, Y.; Eitoku, M.; Sunamura, M.; Inagaki, F.; Imachi, H.; Takai, K.; Nunoura, T. Variability of subseafloor viral abundance at the geographically and geologically distinct continental margins. FEMS Microbiol. Ecol. 2014, 88, 60–68. [Google Scholar] [CrossRef]
- Takai, K.; Horikoshi, K. Rapid detection and quantification of members of the archaeal community by quantitative PCR using fluorogenic probes. Appl. Environ. Microbiol. 2000, 66, 5066–5072. [Google Scholar] [CrossRef]
- Nunoura, T.; Oida, H.; Miyazaki, J.; Miyashita, A.; Imachi, H.; Takai, K. Quantification of mcrA by fluorescent PCR in methanogenic and methanotrophic microbial communities. FEMS Microbiol. Ecol. 2008, 64, 240–247. [Google Scholar] [CrossRef] [PubMed]
- Ludwig, W.; Strunk, O.; Westram, R.; Richter, L.; Meier, H.; Yadhukumar; Buchner, A.; Lai, T.; Steppi, S.; Jobb, G.; et al. ARB: A software environment for sequence data. Nucleic. Acids. Res. 2004, 32, 1363–1371. [Google Scholar] [CrossRef] [PubMed]
- Caporaso, J.G.; Lauber, C.L.; Walters, W.A.; Berg-Lyons, D.; Lozupone, C.A.; Turnbaugh, P.J.; Fierer, N.; Knight, R. Global patterns of 16S rRNA diversity at a depth of millions of sequences per sample. Proc. Natl. Acad. Sci. USA 2011, 108, 4516–4522. [Google Scholar] [CrossRef] [PubMed]
- Bolyen, E.; Rideout, J.R.; Dillon, M.R.; Bokulich, N.A.; Abnet, C.C.; Al-Ghalith, G.A.; Alexander, H.; Alm, E.J.; Arumugam, M.; Asnicar, F.; et al. Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat. Biotechnol. 2019, 37, 852–857. [Google Scholar] [CrossRef] [PubMed]
- Reeburgh, W.S. Oceanic methane biogeochemistry. Chem. Rev. 2007, 107, 486–513. [Google Scholar] [CrossRef] [PubMed]
- Borowski, W.S.; Paull, C.K.; Ussler, W. Marine pore-water sulfate profiles indicate in situ methane flux from underlying gas hydrate. Geology 1996, 24, 655–658. [Google Scholar] [CrossRef]
- Naehr, T.H.; Eichhubl, P.; Orphan, V.J.; Hovland, M.; Paull, C.K.; Ussler, W.; Lorenson, T.D.; Greene, H.G. Authigenic carbonate formation at hydrocarbon seeps in continental margin sediments: A comparative study. Deep Sea Res. Part II Top. Stud. Oceanogr. 2007, 54, 1268–1291. [Google Scholar] [CrossRef]
- Reitner, J.; Peckmann, J.; Reimer, A.; Schumann, G.; Thiel, V. Methane-derived carbonate build-ups and associated microbial communities at cold seeps on the lower Crimean shelf (Black Sea). Facies 2005, 51, 66–79. [Google Scholar] [CrossRef]
- Hallam, S.J.; Girguis, P.R.; Preston, C.M.; Richardson, P.M.; DeLong, E.F. Identification of methyl coenzyme M reductase A (mcrA) genes associated with methane-oxidizing archaea. Appl. Environ. Microbiol. 2003, 69, 5483–5491. [Google Scholar] [CrossRef]
- Knittel, K.; Lösekann, T.; Boetius, A.; Kort, R.; Amann, R. Diversity and distribution of methanotrophic archaea at cold seeps. Appl. Environ. Microbiol. 2005, 71, 467–479. [Google Scholar] [CrossRef]
- Torres, M.E.; Wallmann, K.; Tréhu, A.M.; Bohrmann, G.; Borowski, W.S.; Tomaru, H. Gas hydrate growth, methane transport, and chloride enrichment at the southern summit of Hydrate Ridge, Cascadia margin off Oregon. Earth. Planet. Sci. Lett. 2004, 226, 225–241. [Google Scholar] [CrossRef]
- Blumenberg, M.; Walliser, E.-O.; Taviani, M.; Seifert, R.; Reitner, J. Authigenic carbonate formation and its impact on the biomarker inventory at hydrocarbon seeps—A case study from the Holocene Black Sea and the Plio-Pleistocene Northern Apennines (Italy). Mar. Pet. Geol. 2015, 66, 532–541. [Google Scholar] [CrossRef]
- Boivin-Jahns, V.; Ruimy, R.; Bianchi, A.; Daumas, S.; Christen, R. Bacterial diversity in a deep-subsurface clay environment. Appl. Environ. Microbiol. 1996, 62, 3405–3412. [Google Scholar] [PubMed]
- Zhou, J.; Xia, B.; Huang, H.; Palumbo, A.V.; Tiedje, J.M. Microbial diversity and heterogeneity in sandy subsurface soils. Appl. Environ. Microbiol. 2004, 70, 1723–1734. [Google Scholar] [CrossRef] [PubMed]
- Niemann, H.; Lösekann, T.; de Beer, D.; Elvert, M.; Nadalig, T.; Knittel, K.; Amann, R.; Sauter, E.J.; Schluter, M.; Klages, M.; et al. Novel microbial communities of the Haakon Mosby mud volcano and their role as a methane sink. Nature 2006, 443, 854–858. [Google Scholar] [CrossRef] [PubMed]
- Felden, J.; Ruff, S.E.; Ertefai, T.; Inagaki, F.; Hinrichs, K.-U.; Wenzhöfer, F. Anaerobic methanotrophic community of a 5346-m-deep vesicomyid clam colony in the Japan Trench. Geobiology 2014, 12, 183–199. [Google Scholar] [CrossRef] [PubMed]
- Pop Ristova, P.; Wenzhöfer, F.; Ramette, A.; Felden, J.; Boetius, A. Spatial scales of bacterial community diversity at cold seeps (Eastern Mediterranean Sea). ISME J. 2015, 9, 1306–1318. [Google Scholar] [CrossRef] [PubMed]
- Fry, J.C.; Parkes, R.J.; Cragg, B.A.; Weightman, A.J.; Webster, G. Prokaryotic biodiversity and activity in the deep subseafloor biosphere. FEMS Microbiol. Ecol. 2008, 66, 181–196. [Google Scholar] [CrossRef] [PubMed]
- Girguis, P.R.; Cozen, A.E.; DeLong, E.F. Growth and population dynamics of anaerobic methane-oxidizing archaea and sulfate-reducing bacteria in a continuous-flow bioreactor. Appl. Environ. Microbiol. 2005, 71, 3725–3733. [Google Scholar] [CrossRef]
- Timmers, P.; Widjaja-Greefkes, H.C.; Ramiro-Garcia, J.; Plugge, C.; Stams, A. Growth and activity of ANME clades with different sulfate and sulfide concentrations in the presence of methane. Front. Microbiol. 2015, 6, 998. [Google Scholar] [CrossRef]
- Harrison, B.K.; Zhang, H.; Berelson, W.; Orphan, V.J. Variations in archaeal and bacterial diversity associated with the sulfate-methane transition zone in continental margin sediments (Santa Barbara Basin, California). Appl. Environ. Microbiol. 2009, 75, 1487–1499. [Google Scholar] [CrossRef] [PubMed]
- Krüger, M.; Blumenberg, M.; Kasten, S.; Wieland, A.; Kanel, L.; Klock, J.H.; Michaelis, W.; Seifert, R. A novel, multi-layered methanotrophic microbial mat system growing on the sediment of the Black Sea. Environ. Microbiol. 2008, 10, 1934–1947. [Google Scholar] [CrossRef] [PubMed]
- Niu, M.; Fan, X.; Zhuang, G.; Liang, Q.; Wang, F. Methane-metabolizing microbial communities in sediments of the Haima cold seep area, northwest slope of the South China Sea. FEMS Microbiol. Ecol. 2017, 93, fix101. [Google Scholar] [CrossRef] [PubMed]
- Nunoura, T.; Oida, H.; Toki, T.; Ashi, J.; Takai, K.; Horikoshi, K. Quantification of mcrA by quantitative fluorescent PCR in sediments from methane seep of the Nankai Trough. FEMS Microbiol. Ecol. 2006, 57, 149–157. [Google Scholar] [CrossRef] [PubMed]
- Orcutt, B.; Boetius, A.; Elvert, M.; Samarkin, V.; Joye, S.B. Molecular biogeochemistry of sulfate reduction, methanogenesis and the anaerobic oxidation of methane at Gulf of Mexico cold seeps. Geochimi. Cosmochimi. Acta 2005, 69, 4267–4281. [Google Scholar] [CrossRef]
- Hinrichs, K.U.; Boetius, A. The anaerobic oxidation of methane: new insights in microbial ecology and biogeochemistry. In Ocean Margin Systems; Wefer, G., Billett, D., Hebbeln, D., Jørgensen, B.B., Schlüter, M., Weering, T.V., Eds.; Springer: Berlin, Germany, 2002; pp. 457–477. [Google Scholar]
- Boetius, A.; Wenzhöfer, F. Seafloor oxygen consumption fuelled by methane from cold seeps. Nat. Geosci. 2013, 6, 725. [Google Scholar] [CrossRef]
- Klaucke, I.; Masson, D.G.; Petersen, C.J.; Weinrebe, W.; Ranero, C.R. Multifrequency geoacoustic imaging of fluid escape structures offshore Costa Rica: Implications for the quantification of seep processes. Geochem. Geophys. Geosyst. 2008, 9. [Google Scholar] [CrossRef]
| Core | Depth (mbsf) | Sample | δ13C (‰) | δ18O (‰) | Cell Count (cells/g) |
|---|---|---|---|---|---|
| PC1406 | 2.2 | Nodule outside | −37.32 | +4.56 | 1.75 × 107 (± 0.37 × 107) |
| Nodule inside | −38.05 | +4.50 | 0.99 × 107 (± 0.23 × 107) | ||
| Sediment | — | — | 2.30 × 107 (± 0.11 × 107) | ||
| PC1407 | 0.7 | Nodule outside | −54.91 | +5.12 | 1.45 × 107 (± 0.33 × 107) |
| Nodule inside | −52.97 | +5.31 | 0.85 × 107 (± 0.14 × 107) | ||
| Sediment | — | — | 1.08 × 107 (± 0.23 × 107) |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yanagawa, K.; Shiraishi, F.; Tanigawa, Y.; Maeda, T.; Mustapha, N.A.; Owari, S.; Tomaru, H.; Matsumoto, R.; Kano, A. Endolithic Microbial Habitats Hosted in Carbonate Nodules Currently Forming within Sediment at a High Methane Flux Site in the Sea of Japan. Geosciences 2019, 9, 463. https://doi.org/10.3390/geosciences9110463
Yanagawa K, Shiraishi F, Tanigawa Y, Maeda T, Mustapha NA, Owari S, Tomaru H, Matsumoto R, Kano A. Endolithic Microbial Habitats Hosted in Carbonate Nodules Currently Forming within Sediment at a High Methane Flux Site in the Sea of Japan. Geosciences. 2019; 9(11):463. https://doi.org/10.3390/geosciences9110463
Chicago/Turabian StyleYanagawa, Katsunori, Fumito Shiraishi, Yusuke Tanigawa, Toshinari Maeda, Nurul Asyifah Mustapha, Satoko Owari, Hitoshi Tomaru, Ryo Matsumoto, and Akihiro Kano. 2019. "Endolithic Microbial Habitats Hosted in Carbonate Nodules Currently Forming within Sediment at a High Methane Flux Site in the Sea of Japan" Geosciences 9, no. 11: 463. https://doi.org/10.3390/geosciences9110463
APA StyleYanagawa, K., Shiraishi, F., Tanigawa, Y., Maeda, T., Mustapha, N. A., Owari, S., Tomaru, H., Matsumoto, R., & Kano, A. (2019). Endolithic Microbial Habitats Hosted in Carbonate Nodules Currently Forming within Sediment at a High Methane Flux Site in the Sea of Japan. Geosciences, 9(11), 463. https://doi.org/10.3390/geosciences9110463





