Relationship between SNPs of POU1F1 Gene and Litter Size and Growth Traits in Shaanbei White Cashmere Goats
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Animal, Data Collection, and DNA Extraction
2.2. Primer Design, Amplification, and SNPs Scanning
2.3. Identification of c.682G > T Locus by HinfI Restriction Endonuclease
2.4. Statistical Analysis
3. Results
3.1. Identification of POU1F1 Gene Polymorphisms
3.2. Genetic Parameter of POU1F1 Gene Polymorphisms
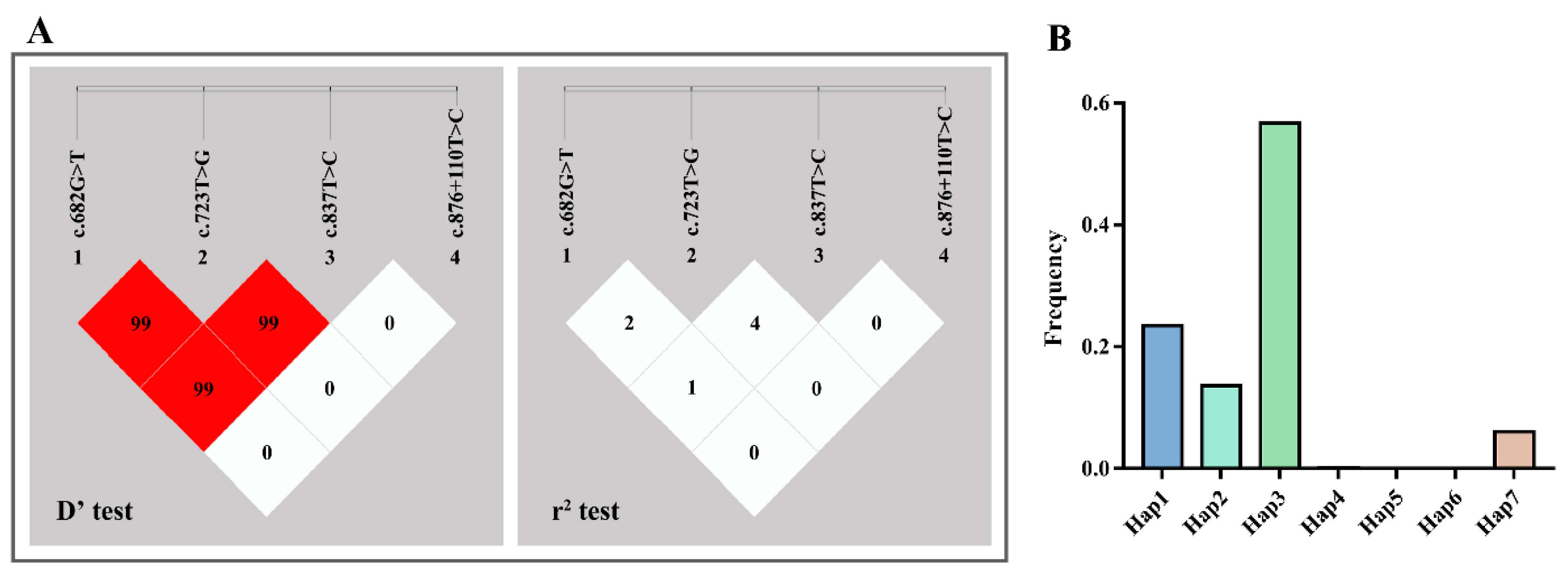
3.3. LD and Haplotype Analysis
3.4. Association between SNPs of POU1F1 and Litter Size
3.5. Association between Diplotypes of POU1F1 and Litter Size
3.6. SNPs and Diplotypes of POU1F1 Associated with Growth Parameters
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Lai, F.N.; Zhai, H.L.; Cheng, M.; Ma, J.Y.; Cheng, S.F.; Ge, W.; Zhang, G.L.; Wang, J.J.; Zhang, R.Q.; Wang, X.; et al. Whole-genome scanning for the litter size trait associated genes and SNPs under selection in dairy goat (Capra hircus). Sci. Rep. 2016, 6, 38096. [Google Scholar] [CrossRef]
- Misztal, I.; Legarra, A. Invited review: Efficient computation strategies in genomic selection. Animal 2017, 11, 731–736. [Google Scholar] [CrossRef]
- Meuwissen, T.H.; Hayes, B.J.; Goddard, M.E. Prediction of total genetic value using genome-wide dense marker maps. Genetics 2001, 157, 1819–1829. [Google Scholar]
- VanRaden, P.M. Efficient methods to compute genomic predictions. J. Dairy Sci. 2008, 91, 4414–4423. [Google Scholar] [CrossRef]
- Azevedo, C.F.; de Resende, M.D.V.; Silva, F.F.E.; Viana, J.M.S.; Valente, M.S.F.; Resende, M.F.R.; Muñoz, P. Ridge, Lasso and Bayesian additive-dominance genomic models. BMC Genet. 2015, 16, 105. [Google Scholar] [CrossRef] [PubMed]
- Andersen, B.; Rosenfeld, M.G. POU domain factors in the neuroendocrine system: Lessons from developmental biology provide insights into human disease. Endocr. Rev. 2001, 22, 2–35. [Google Scholar] [CrossRef] [PubMed]
- Sobrier, M.L.; Tsai, Y.C.; Pérez, C.; Leheup, B.; Bouceba, T.; Duquesnoy, P.; Copin, B.; Sizova, D.; Penzo, A.; Stanger, B.Z.; et al. Functional characterization of a human POU1F1 mutation associated with isolated growth hormone deficiency: A novel etiology for IGHD. Hum. Mol. Genet. 2016, 25, 472–483. [Google Scholar] [CrossRef]
- Li, S.; Crenshaw, E.B., III; Rawson, E.J.; Simmons, D.M.; Swanson, L.W.; Rosenfeld, M.G. Dwarf locus mutants lacking three pituitary cell types result from mutations in the POU-domain gene pit-1. Nature 1990, 347, 528–533. [Google Scholar] [CrossRef] [PubMed]
- Giordano, M. Genetic causes of isolated and combined pituitary hormone deficiency. Best. Pract. Res. Clin. Endocrinol. Metab. 2016, 30, 679–691. [Google Scholar] [CrossRef] [PubMed]
- Stanceková, K.; Vasícek, D.; Peskovicová, D.; Bulla, J.; Kúbek, A. Effect of genetic variability of the porcine pituitary-specific transcription factor (PIT-1) on carcas traits in pigs. Anim. Genet. 1999, 30, 313–315. [Google Scholar] [CrossRef] [PubMed]
- Kim, G.W.; Yoo, J.Y.; Kim, H.Y. Association of genotype of POU1F1 intron 1 with carcass characteristics in crossbred pigs. J. Anim. Sci. Technol. 2014, 56, 25. [Google Scholar] [CrossRef]
- Huang, W.; Maltecca, C.; Khatib, H. A proline-to-histidine mutation in POU1F1 is associated with production traits in dairy cattle. Anim. Genet. 2008, 39, 554–557. [Google Scholar] [CrossRef]
- Viale, E.; Tiezzi, F.; Maretto, F.; De Marchi, M.; Penasa, M.; Cassandro, M. Association of candidate gene polymorphisms with milk technological traits, yield, composition, and somatic cell score in Italian Holstein-Friesian sires. J. Dairy Sci. 2017, 100, 7271–7281. [Google Scholar] [CrossRef]
- Jalil-Sarghale, A.; Shahrbabak, M.M.; Sharbabak, H.M.; Sadeghi, M.; Mura, M.C. Association of pituitary specific transcription factor-1 (POU1F1) gene polymorphism with growth and biometric traits and blood metabolites in Iranian Zel and Lori-Bakhtiari sheep. Mol. Biol. Rep. 2014, 41, 5787–5792. [Google Scholar] [CrossRef] [PubMed]
- Ozmen, O.; Kul, S.; Unal, E.O. Polymorphism of sheep POU1F1 gene exon 6 and 3′UTR region and their association with milk production traits. Iran. J. Vet. Res. 2014, 15, 331–335. [Google Scholar] [PubMed]
- Nazari-Ghadikolaei, A.; Mehrabani-Yeganeh, H.; Miarei-Aashtiani, S.R.; Staiger, E.A.; Rashidi, A.; Huson, H.J. Genome-wide association studies identify candidate genes for coat color and mohair traits in the Iranian Markhoz goat. Front. Genet. 2018, 9, 105. [Google Scholar] [CrossRef] [PubMed]
- Lan, X.Y.; Pan, C.Y.; Chen, H.; Zhang, C.L.; Li, J.Y.; Zhao, M.; Lei, C.Z.; Zhang, A.L.; Zhang, L. An AluI PCR-RFLP detecting a silent allele at the goat POU1F1 locus and its association with production traits. Small Rumin. Res. 2007, 73, 8–12. [Google Scholar] [CrossRef]
- Lan, X.Y.; Shu, J.H.; Chen, H.; Pan, C.Y.; Lei, C.Z.; Wang, X.; Liu, S.Q.; Zhang, Y.B. A PstI polymorphism at 3′UTR of goat POU1F1 gene and its effect on cashmere production. Mol. Biol. Rep. 2009, 36, 1371–1374. [Google Scholar] [CrossRef]
- Lan, X.Y.; Pan, C.Y.; Chen, H.; Lei, C.Z. A Ddel PCR-RFLP detecting genetic variation of goat POU1F1 gene. Can. J. Anim. Sci. 2007, 87, 13–14. [Google Scholar] [CrossRef]
- Lan, X.Y.; Pan, C.Y.; Li, J.Y.; Guo, Y.W.; Hu, S.; Wang, J.; Liu, Y.B.; Hu, S.R.; Lei, C.Z.; Chen, H. Twelve novel SNPs of the goat POU1F1 gene and their associations with cashmere traits. Small Rumin. Res. 2009, 85, 116–121. [Google Scholar] [CrossRef]
- Lan, X.Y.; Pan, C.Y.; Chen, H.; Lei, C.Z.; Hua, L.S.; Yang, X.B.; Qiu, G.Y.; Zhang, R.F.; Lun, Y.Z. Ddel polymorphism in coding region of goat POU1F1 gene and its association with production traits. Asian Aust. J. Anim. 2007, 20, 1342–1348. [Google Scholar] [CrossRef]
- Feng, T.; Chu, M.X.; Cao, G.L.; Tang, Q.Q.; Di, R.; Fang, L.; Li, N. Polymorphisms of caprine POU1F1 gene and their association with litter size in Jining Grey goats. Mol. Biol. Rep. 2012, 39, 4029–4038. [Google Scholar] [CrossRef]
- Daga, C.; Paludo, M.; Luridiana, S.; Mura, M.C.; Bodano, S.; Pazzola, M.; Dettori, M.L.; Vacca, G.M.; Carcangiu, V. Identification of novel SNPs in the Sarda breed goats POU1F1 gene and their association with milk productive performance. Mol. Biol. Rep. 2013, 40, 2829–2835. [Google Scholar] [CrossRef]
- Zhou, F.Y.; Yang, Q.; Lei, C.Z.; Chen, H.; Lan, X.Y. Relationship between genetic variants of POUIF1, PROP1, IGFBP3 genes and milk performance in Guanzhong dairy goats. Small Rumin. Res. 2016, 140, 40–45. [Google Scholar] [CrossRef]
- Lan, X.Y.; Li, M.J.; Chen, H.; Zhang, L.Z.; Jing, Y.J.; Wei, T.B.; Ren, G.; Wang, X.; Fang, X.T.; Zhang, C.L.; et al. Analysis of caprine pituitary specific transcription factor-1 gene polymorphism in indigenous Chinese goats. Mol. Biol. Rep. 2009, 36, 705–709. [Google Scholar] [CrossRef] [PubMed]
- Li, M.J.; Zhang, C.M.; Lan, X.Y.; Fang, X.T.; Lei, C.Z.; Chen, H. Analysis of POU1F1 gene DdeI polymorphism in Chinese goats. Genet. Mol. Res. 2016, 15, 15017747. [Google Scholar] [CrossRef] [PubMed]
- Wang, K.; Yan, H.L.; Xu, H.; Yang, Q.; Zhang, S.H.; Pan, C.Y.; Chen, H.; Zhu, H.J.; Liu, J.W.; Qu, L.; et al. A novel indel within goat casein alpha S1 gene is significantly associated with litter size. Gene 2018, 671, 161–169. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.Y.; Yang, Q.; Wang, K.; Yan, H.L.; Pan, C.Y.; Chen, H.; Liu, J.W.; Zhu, H.J.; Qu, L.; Lan, X.Y. Two strongly linked single nucleotide polymorphisms (Q320P and V397I) in GDF9 gene are associated with litter size in cashmere goats. Theriogenology 2019, 125, 115–121. [Google Scholar] [CrossRef]
- Li, J.; Zhu, X.; Ma, L.; Xu, H.; Cao, X.; Luo, R.; Chen, H.; Sun, X.; Cai, Y.; Lan, X. Detection of a new 20-bp insertion/deletion (indel) within sheep PRND gene using mathematical expectation (ME) method. Prion 2017, 11, 143–150. [Google Scholar] [CrossRef]
- Chen, F.; Shi, J.; Luo, Y.Q.; Sun, S.Y.; Pu, M. Genetic characterization of the gypsy moth from China (Lepidoptera, Lymantriidae) using inter simple sequence repeats markers. PLoS ONE 2013, 8, e73017. [Google Scholar] [CrossRef]
- Shi, Y.Y.; He, L. SHEsis, a powerful software platform for analyses of linkage disequilibrium, haplotype construction, and genetic association at polymorphism loci. Cell Res. 2005, 15, 97–98. [Google Scholar] [CrossRef]
- Li, Z.Q.; Zhang, Z.; He, Z.D.; Tang, W.; Li, T.; Zeng, Z.; He, L.; Shi, Y.Y. A partition-ligation-combination-subdivision EM algorithm for haplotype inference with multiallelic markers: Update of the SHEsis (http://analysis.bio-x.cn). Cell Res. 2009, 19, 519–523. [Google Scholar] [CrossRef] [PubMed]
- Cui, Y.; Yan, H.L.; Wang, K.; Xu, H.; Zhang, X.L.; Zhu, H.J.; Liu, J.W.; Qu, L.; Lan, X.Y.; Pan, C.Y. Insertion/deletion within the KDM6A gene is significantly associated with litter size in goat. Front. Genet. 2018, 9, 91. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.Y.; Yang, Q.; Wang, K.; Zhang, S.H.; Pan, C.Y.; Chen, H.; Qu, L.; Yan, H.L.; Lan, X.Y. A novel 12-bp indel polymorphism within the GDF9 gene is significantly associated with litter size and growth traits in goats. Anim. Genet. 2017, 48, 735–736. [Google Scholar] [CrossRef]
- Zhang, Y.H.; Cui, Y.; Zhang, X.L.; Wang, Y.M.; Gao, J.Y.; Yu, T.; Lv, X.Y.; Pan, C.Y. Pig StAR: mRNA expression and alternative splicing in testis and Leydig cells, and association analyses with testicular morphology traits. Theriogenology 2018, 118, 46–56. [Google Scholar] [CrossRef] [PubMed]
- Bland, J.M.; Altman, D.G. Multiple significance tests: The Bonferroni method. BMJ 1995, 310, 170. [Google Scholar] [CrossRef]
- Misrianti, R.; Sumantri, C.; Farajallah, A. Polymorphism identification of Pit1 gene in Indonesian buffaloes (Bubalus bubalis) and Holstein-Friesian cows. Media Peternak. 2010, 33, 131–136. [Google Scholar] [CrossRef]
- Chang, H.L.; Lai, Y.Y.; Wu, M.C.; Sasaki, O. Genetic correlations between male reproductive traits and growth traits in growth performance tested Duroc, Landrace and Yorkshire breed boars. Anim. Sci. J. 2017, 88, 1258–1268. [Google Scholar] [CrossRef]
- Yang, Q.; Yan, H.L.; Li, J.; Xu, H.; Wang, K.; Zhu, H.J.; Chen, H.; Qu, L.; Lan, X.Y. A novel 14-bp duplicated deletion within goat GHR gene is significantly associated with growth traits and litter size. Anim. Genet. 2017, 48, 499–500. [Google Scholar] [CrossRef]
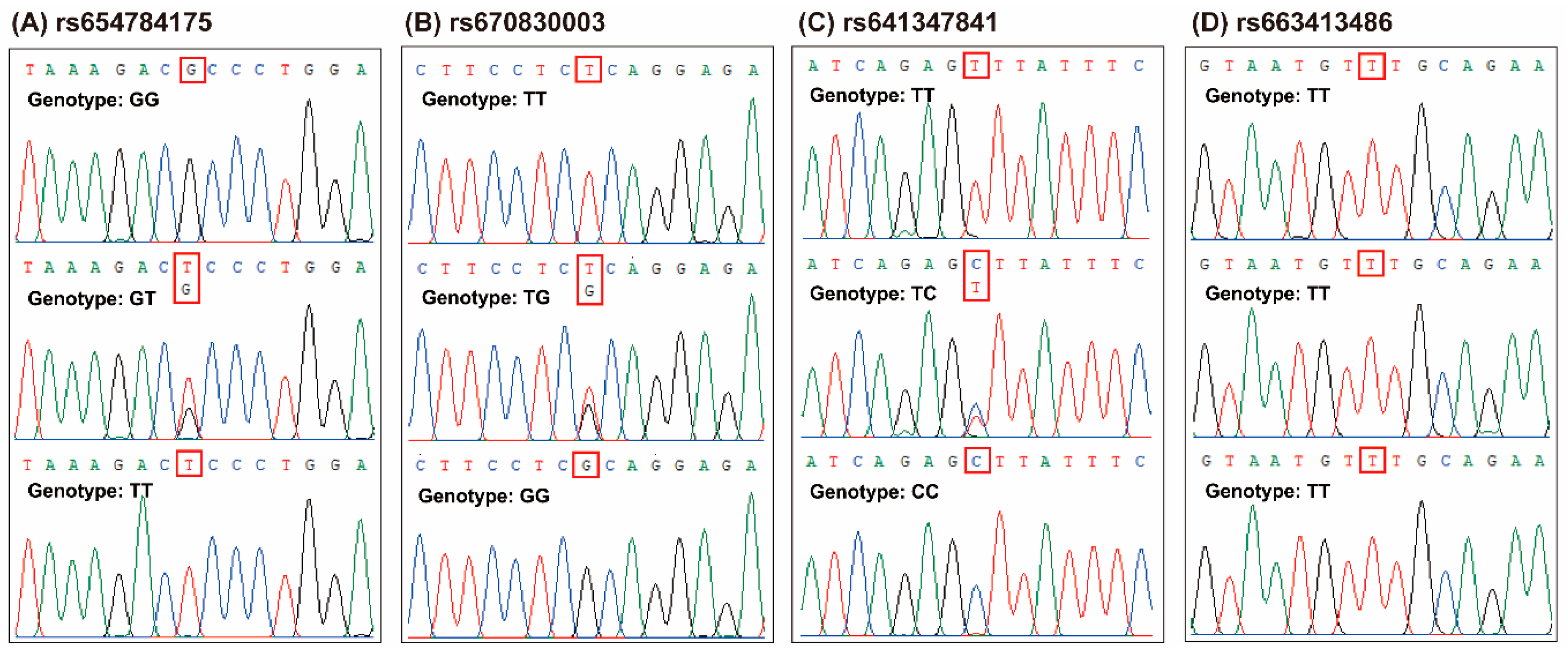

| refSNP No. | HGVS Names | Other Names | Reference | |||
|---|---|---|---|---|---|---|
| rs654784175 | NC_030808.1:g.34236325C > A | c.682G > T | p.A228S | Ex 6 17 G > T | HinfI * | [19] |
| rs670830003 | NC_030808.1:g.34236284A > C | c.723T > G | p.S241S | Ex 6 58 T > G | DdeI | [16] |
| rs641347841 | NC_030808.1:g.34236170A > G | c.837T > C | p.S279S | Ex 6 172 T > C | AluI | [14] |
| rs663413486 | NC_030808.1:g.34236021A > G | c.876 + 110T > C | 3′UTR 110 T > C | PstI | [15] | |
| Loci Names | Observed Genotypes (N) | Genotype Frequencies | Allele Frequencies | Ho | He | Ne | PIC | χ2 |
|---|---|---|---|---|---|---|---|---|
| (p Value) | ||||||||
| c.682G > T | GG (526) | 0.884 | 0.938 (G) | 0.883 | 0.117 | 1.132 | 0.110 | 3.600 (p = 0.058) |
| GT (64) | 0.108 | 0.062 (T) | ||||||
| TT (5) | 0.008 | |||||||
| c.723T > G | TT (354) | 0.581 | 0.767 (T) | 0.642 | 0.358 | 1.557 | 0.294 | 0.868 (p = 0.352) |
| TG (226) | 0.371 | 0.233 (G) | ||||||
| GG (29) | 0.048 | |||||||
| c.837T > C | TT (456) | 0.750 | 0.860 (T) | 0.759 | 0.241 | 1.317 | 0.212 | 4.255 (p = 0.039) |
| TC (134) | 0.220 | 0.140 (C) | ||||||
| CC (18) | 0.030 | |||||||
| c.876 + 110T > C | TT (609) | 1.000 | 1.000 (T) | 1.000 | 1.000 | 1.000 | 1.000 | - |
| TC (0) | 0.000 | 0.000 (C) | ||||||
| CC (0) | 0.000 |
| D’\r2 | c.682G > T | c.723T > G | c.837T > C | c.876 + 110T > C |
|---|---|---|---|---|
| c.682G > T | 0.020 | 0.010 | 0.000 | |
| c.723T > G | 0.991 | 0.049 | 0.000 | |
| c.837T > C | 0.990 | 0.998 | 0.000 | |
| c.876 + 110T > C | 0.000 | 0.000 | 0.000 |
| Haplotype | c.682G > T | c.723T > G | c.837T > C | c.876 + 110T > C | Frequencies |
|---|---|---|---|---|---|
| Hap1 | G | G | T | T | 0.235 |
| Hap2 | G | T | C | T | 0.136 |
| Hap3 | G | T | T | T | 0.567 |
| Hap4 | T | G | C | T | 0.001 |
| Hap5 | T | G | T | T | 3.03 × 10−4 |
| Hap6 | T | T | C | T | 1.60 × 10−4 |
| Hap7 | T | T | T | T | 0.061 |
| Loci Names | Genotypes (N) | p Value | ||
|---|---|---|---|---|
| c.682G > T | GG | GT | TT | |
| 1.27 ± 0.03 (437) | 1.49 ± 0.11 (37) | 1.00 ± 0.00 (3) | ≥0.061 | |
| c.723T > G | TT | TG | GG | |
| 1.32 ± 0.03 (275) | 1.25 ± 0.04 (185) | 1.21 ± 0.09 (28) | ≥0.476 | |
| c.837T > C | TT | TC | CC | |
| 1.34 A ± 0.03 (360) | 1.13 B ± 0.04 (111) | 1.18 AB ± 0.13 (17) | =0.001 | |
| Loci Names | Types | Sample Sizes | Genotypes | Independent χ2, p Value | ||
|---|---|---|---|---|---|---|
| c.682G > T | GG | GT | ||||
| Mothers of single lamb | 354 | 332 | 22 | χ2 = 4.920; p = 0.027 | ||
| Mothers of multi-lamb (≥2) | 120 | 105 | 15 | |||
| c.723T > G | TT | TG | GG | |||
| Mothers of single lamb | 366 | 198 | 145 | 23 | χ2 = 3.208; p = 0.201 | |
| Mothers of multi-lamb (≥2) | 122 | 77 | 40 | 5 | ||
| c.837T > C | TT | TC | CC | |||
| Mothers of single lamb | 366 | 252 | 99 | 15 | χ2 = 18.307; p = 1.04 × 10−4 | |
| Mothers of multi-lamb (≥2) | 122 | 108 | 12 | 2 | ||
| Trait | H3H1 (N) | H3H2 (N) | H3H3 (N) | H3H7 (N) | p Value |
|---|---|---|---|---|---|
| Litter size | 1.28 bc ± 0.05 (137) | 1.12 c ± 0.04 (67) | 1.40 ab ± 0.05 (155) | 1.62 a ± 0.15 (21) | ≤0.039 |
| Types | Sample Sizes | Diplotypes | Independentχ2, p Value | |||
|---|---|---|---|---|---|---|
| H3H1 | H3H2 | H3H3 | H3H7 | |||
| Mothers of single lamb | 273 | 104 | 59 | 100 | 10 | χ2 = 54.647; |
| Mothers of multi-lamb (≥2) | 77 | 33 | 8 | 55 | 11 | p = 8.17 × 10−12 |
| Loci Names | Parameters | Genotypes (N) | p Value | ||
|---|---|---|---|---|---|
| c.682G > T | GG | GT | TT | ||
| BH (cm) | 57.10 b ± 0.21 (466) | 58.67 a ± 0.59 (57) | 57.80 ab ± 1.71 (5) | =0.044 | |
| HHC (cm) | 60.24 b ± 0.23 (465) | 62.27 a ± 0.64 (57) | 59.90 ab ± 2.06 (5) | =0.011 | |
| BL (cm) | 63.27 B ± 0.26 (466) | 65.70 A ± 0.71 (57) | 63.80 AB ± 1.63 (5) | =0.006 | |
| CC (cm) | 7.89 b ± 0.05 (467) | 8.25 a ± 0.14 (56) | 7.40 ab ± 0.53 (5) | =0.028 | |
| ChW (cm) | 18.75 B ± 0.17 (467) | 20.23 A ± 0.51 (57) | 17.90 AB ± 1.89 (5) | =0.015 | |
| ChWI (%) | 68.24 B ± 0.47 (467) | 72.70 A ± 1.75 (57) | 64.26 AB ± 5.46 (5) | =0.008 | |
| c.723T > G | TT | TG | GG | ||
| HW (cm) | 20.98 ab ± 0.23 (99) | 20.56 b ± 0.40 (53) | 23.42 a ± 0.75 (6) | =0.024 | |
| BI (%) | 138.69 B ± 0.87 (303) | 142.94 A ± 1.12 (202) | 141.72 AB ± 3.66 (28) | =0.009 | |
| ChCI (%) | 153.90 B ± 1.03 (303) | 158.98 A ± 1.31 (203) | 157.78 AB ± 4.50 (27) | =0.008 | |
| CCI (%) | 13.71 b ± 0.10 (306) | 14.11 a ± 0.13 (203) | 13.93 ab ± 0.34 (27) | =0.035 | |
| c.837T > C | TT | TC | CC | ||
| HHC (cm) | 60.89 A ± 0.26 (400) | 59.11 B ± 0.40 (121) | 58.29 AB ± 1.10 (17) | =0.002 | |
| ChC (cm) | 89.97 A ± 0.51 (400) | 86.80 B ± 1.01 (121) | 81.33 B ± 2.33 (15) | ≤0.010 | |
| CC (cm) | 8.04 A ± 0.05 (401) | 7.60 B ± 0.08 (121) | 7.21 B ± 0.26 (17) | ≤0.002 | |
| ChD (cm) | 27.64 A ± 0.14 (402) | 26.75 B ± 0.25 (121) | 27.32 AB ± 0.66 (17) | =0.007 | |
| ChW (cm) | 19.15 a ± 0.19 (402) | 18.06 b ± 0.31 (121) | 17.97 ab ± 0.71 (17) | =0.015 | |
| BI (%) | 141.58 a ± 0.77 (398) | 138.41 a ± 1.54 (120) | 127.21 b ± 2.83 (15) | ≤0.027 | |
| CCI (%) | 14.02 a ± 0.09 (399) | 13.50 b ± 0.16 (120) | 13.04 ab ± 0.43 (17) | =0.012 | |
| Parameters | H3H1 (N) | H3H2 (N) | H3H3 (N) | H3H7 (N) | p Value |
|---|---|---|---|---|---|
| BH (cm) | 56.93 ab ± 0.40 (148) | 56.62 b ± 0.49 (73) | 57.96 ab ± 0.36 (166) | 59.21 a ± 0.76 (33) | =0.048 |
| HHC (cm) | 60.51 AB ± 0.42 (147) | 58.81 B ± 0.49 (73) | 61.07 A ± 0.41 (165) | 62.94 A ± 0.88 (33) | ≤0.008 |
| BL (cm) | 63.35 ab ± 0.47 (147) | 62.57 b ± 0.56 (73) | 63.84 ab ± 0.46 (166) | 65.94 a ± 1.02 (33) | =0.029 |
| ChC (cm) | 91.16 A ± 0.80 (147) | 86.34 B ± 1.32 (73) | 89.52 AB ± 0.81 (166) | 90.66 AB ± 1.78 (32) | =0.007 |
| CC (cm) | 8.09 A ± 0.08 (147) | 7.51 B ± 0.10 (73) | 8.01 A ± 0.08 (167) | 8.45 A ± 0.19 (32) | ≤0.002 |
| ChD (cm) | 27.57 ab ± 0.23 (147) | 26.60 b ± 0.32 (73) | 27.69 a ± 0.23 (167) | 28.32 a ± 0.53 (33) | ≤0.042 |
| ChW(cm) | 19.14 a ± 0.32 (147) | 17.50 b ± 0.31 (73) | 19.13 a ± 0.30 (167) | 20.21 a ± 0.69 (33) | ≤0.013 |
| ChCI (%) | 160.78 a ± 1.48 (147) | 153.39 b ± 2.48 (72) | 154.95 b ± 1.30 (165) | 153.13 ab ± 2.78 (32) | =0.026 |
| CCI (%) | 14.27 A ± 0.15 (147) | 13.33 B ± 0.19 (72) | 13.86 AB ± 0.13 (166) | 14.28 AB ± 0.30 (32) | =0.001 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhu, H.; Zhang, Y.; Bai, Y.; Yang, H.; Yan, H.; Liu, J.; Shi, L.; Song, X.; Li, L.; Dong, S.; et al. Relationship between SNPs of POU1F1 Gene and Litter Size and Growth Traits in Shaanbei White Cashmere Goats. Animals 2019, 9, 114. https://doi.org/10.3390/ani9030114
Zhu H, Zhang Y, Bai Y, Yang H, Yan H, Liu J, Shi L, Song X, Li L, Dong S, et al. Relationship between SNPs of POU1F1 Gene and Litter Size and Growth Traits in Shaanbei White Cashmere Goats. Animals. 2019; 9(3):114. https://doi.org/10.3390/ani9030114
Chicago/Turabian StyleZhu, Haijing, Yanghai Zhang, Yangyang Bai, Han Yang, Hailong Yan, Jinwang Liu, Lei Shi, Xiaoyue Song, Longping Li, Shuwei Dong, and et al. 2019. "Relationship between SNPs of POU1F1 Gene and Litter Size and Growth Traits in Shaanbei White Cashmere Goats" Animals 9, no. 3: 114. https://doi.org/10.3390/ani9030114
APA StyleZhu, H., Zhang, Y., Bai, Y., Yang, H., Yan, H., Liu, J., Shi, L., Song, X., Li, L., Dong, S., Pan, C., Lan, X., & Qu, L. (2019). Relationship between SNPs of POU1F1 Gene and Litter Size and Growth Traits in Shaanbei White Cashmere Goats. Animals, 9(3), 114. https://doi.org/10.3390/ani9030114






