Expression Analysis of the Prolific Candidate Genes, BMPR1B, BMP15, and GDF9 in Small Tail Han Ewes with Three Fecundity (FecB Gene) Genotypes
Abstract
Simple Summary
Abstract
1. Introduction
2. Material and Methods
2.1. Selection and Treatment of Experimental Sheep
2.2. Sample Collection
2.3. Total RNA Extraction and cDNA Synthesis
2.4. Primer Design
2.5. Exploration of Gene Amplification Parameters
2.6. RT-PCR Program and Amplification System
2.7. qPCR
2.7.1. System and Program for Real-Time qPCR Analyses
2.7.2. Establishment of the Standard Curve
2.8. Data Statistics
3. Results
3.1. RNA Extraction and cDNA Synthesis
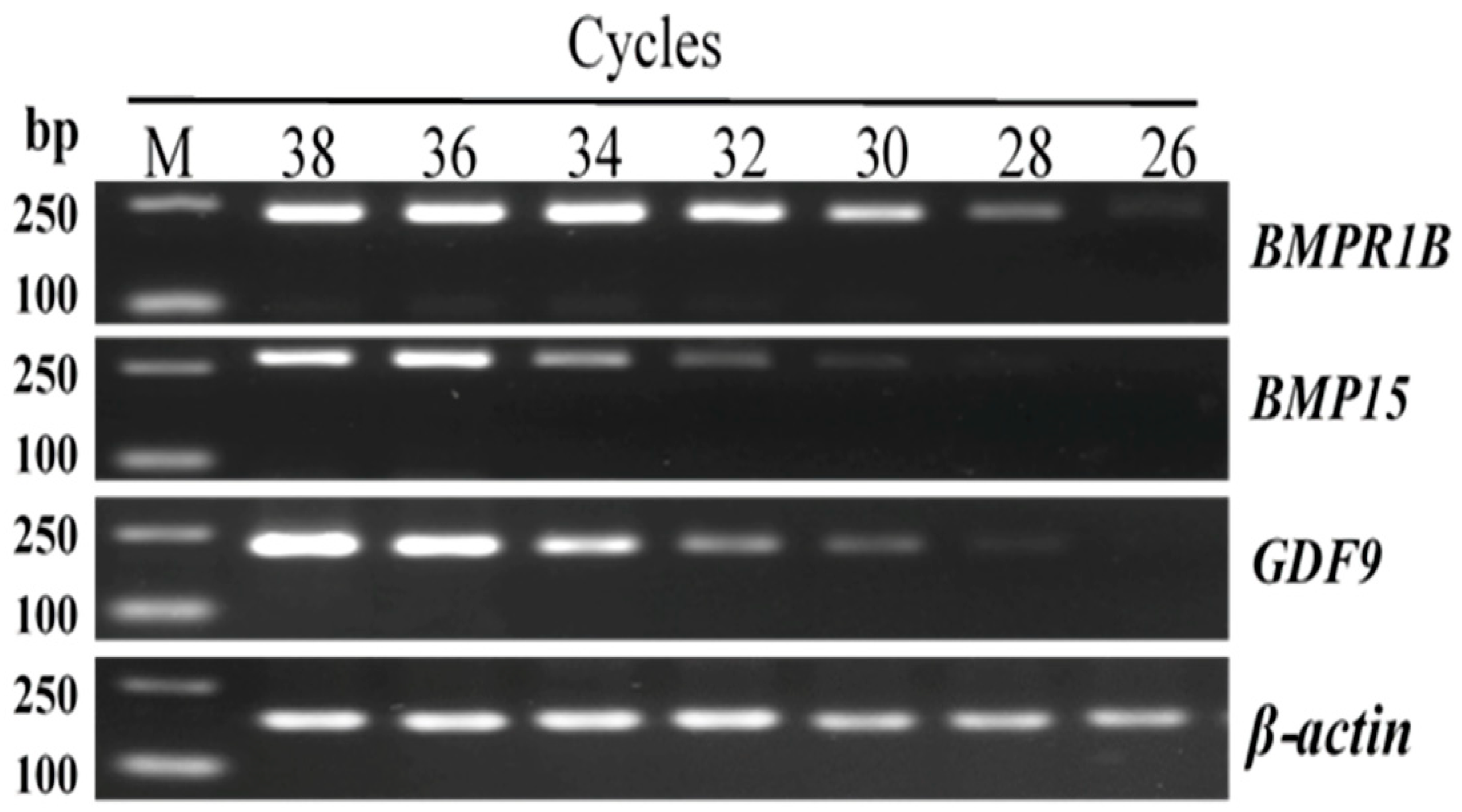
3.2. Exploration of the Amplification Parameters
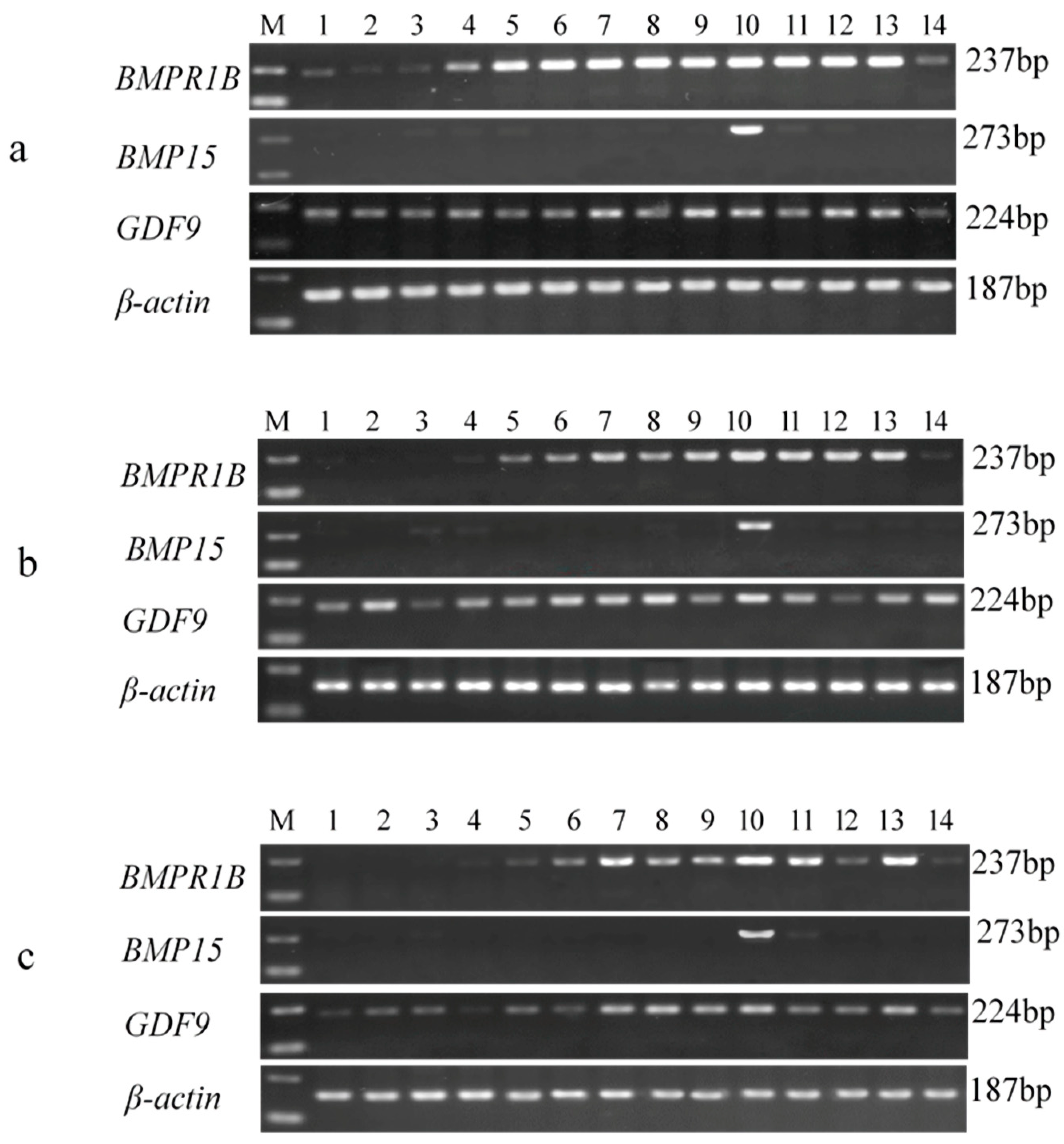
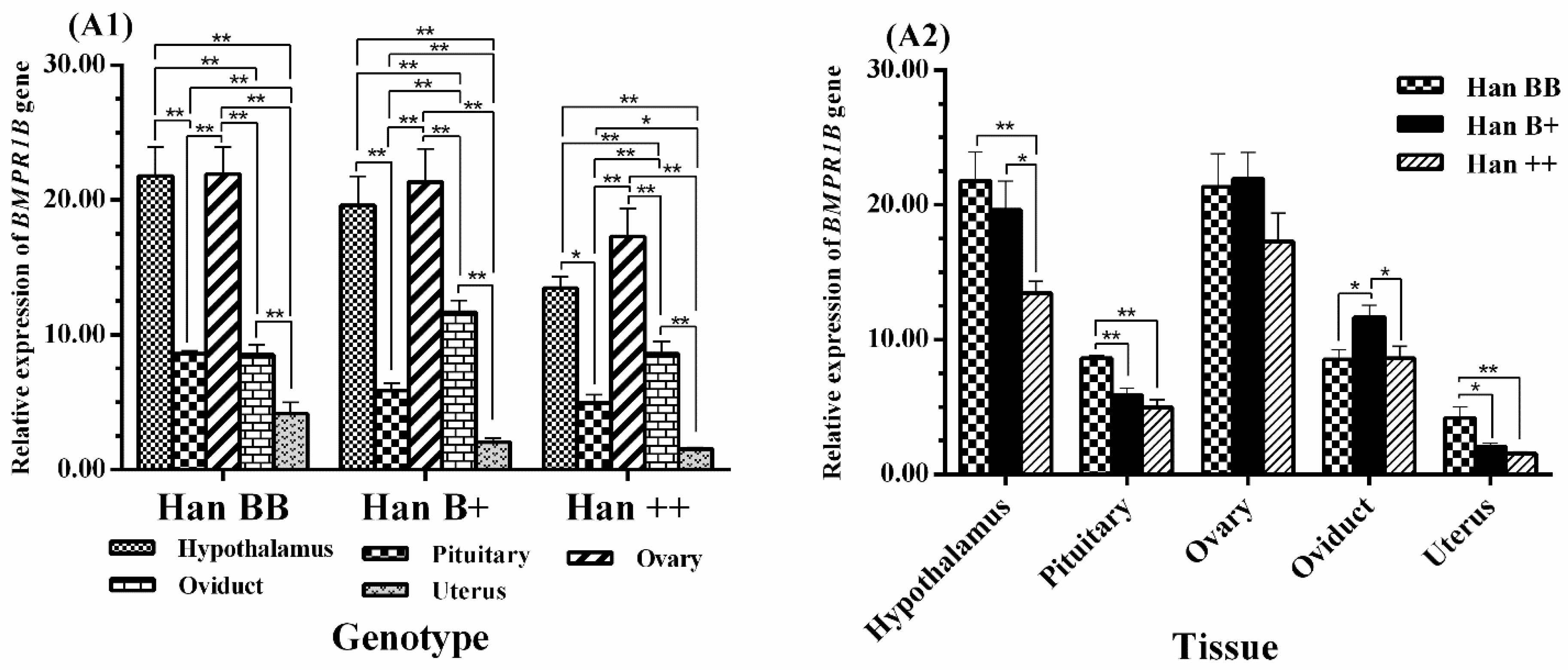
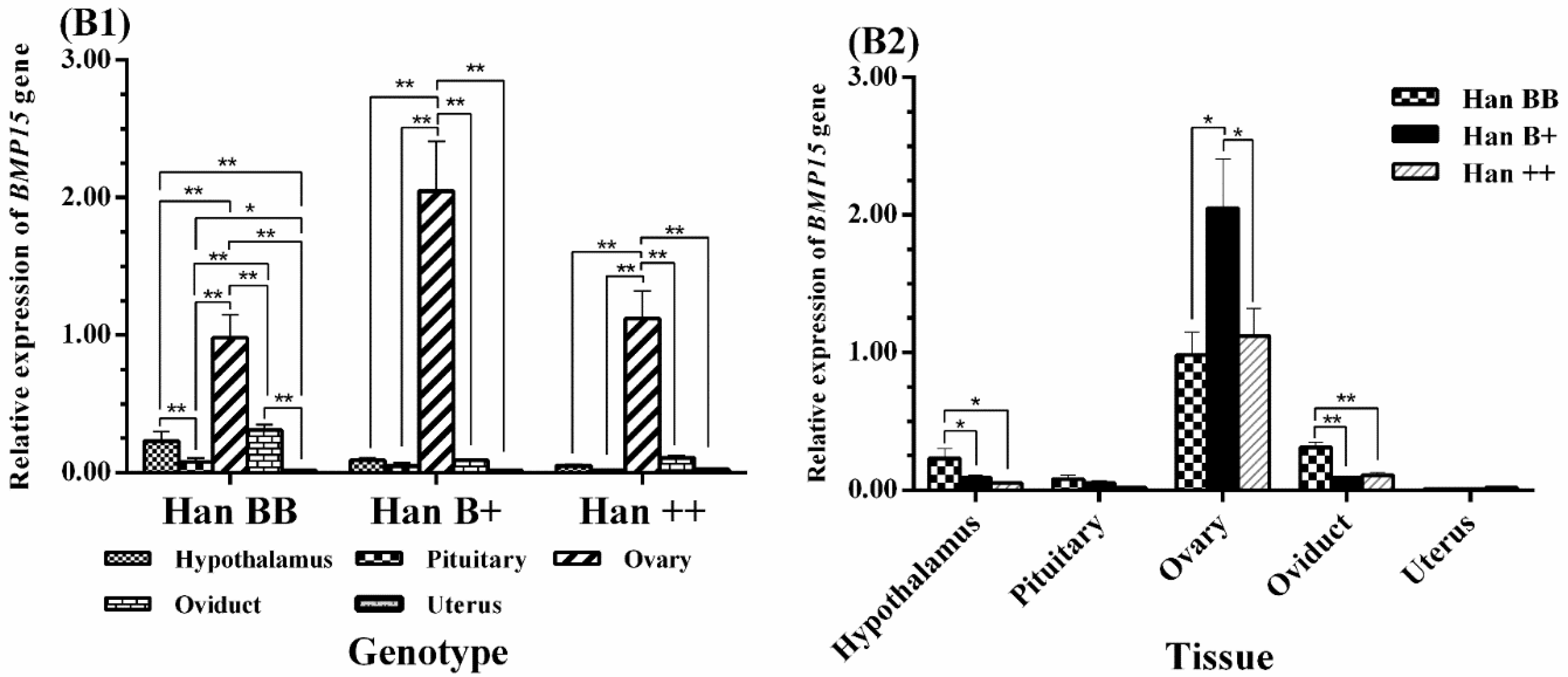
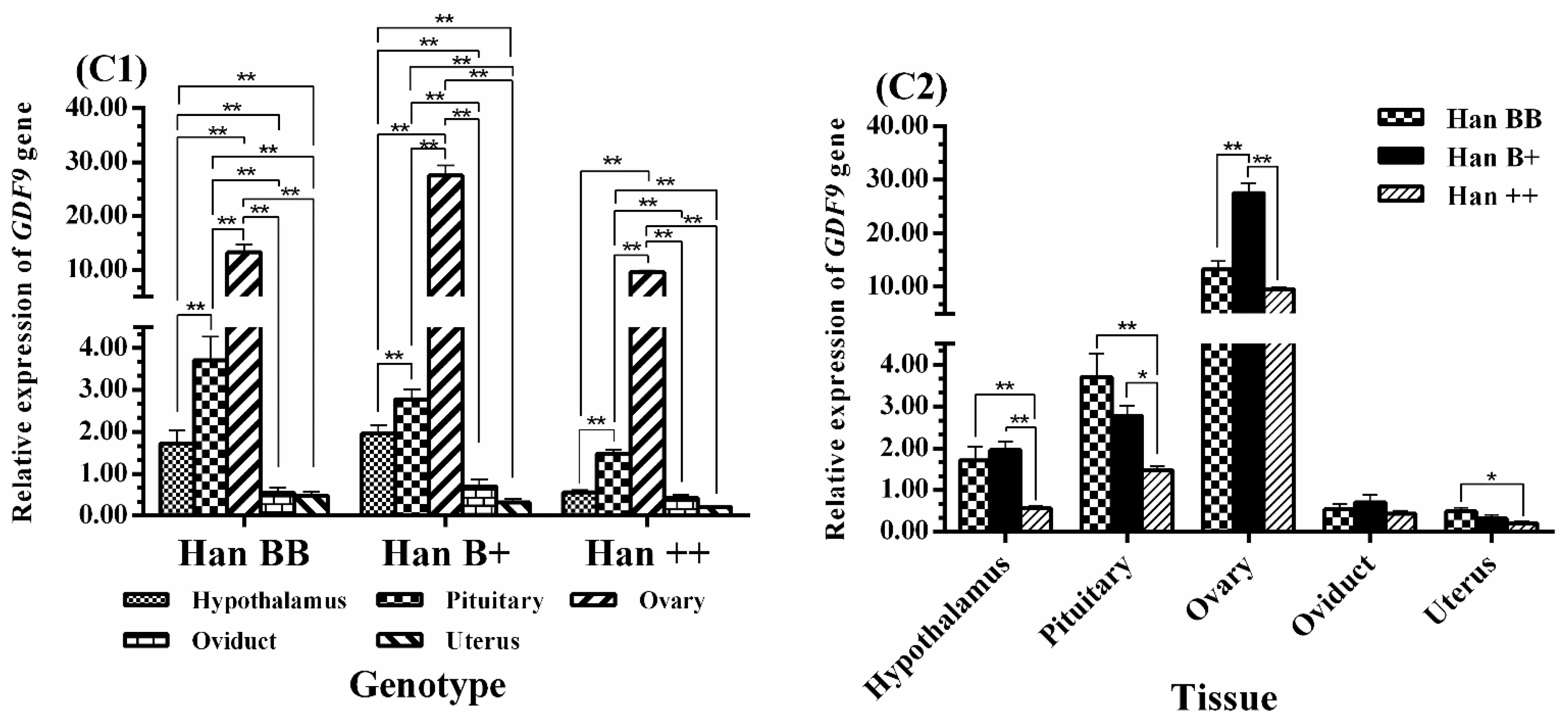
3.3. Tissue-Specific Expression Analysis of BMPR1B, BMP15, and GDF9
3.4. Expression Levels of BMPR1B, BMP15, and GDF9 in the HPG Axis
4. Discussion
4.1. BMPR1B Expression
4.2. BMP15 Expression
4.3. GDF9 Expression
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Gabina, D. Improvement of the reproductive performance of Rasa Aragonesa flocks in frequent lambing systems. II. Repeatability and heritability of sexual precocity, fertility and litter size: Selection strategies. Livest. Prod. Sci. 1989, 22, 87–98. [Google Scholar] [CrossRef]
- Davis, G.H.; Montgomery, G.W.; Allison, A.J.; Kelly, R.W.; Bray, A.R. Segregation of a major gene influencing fecundity in progeny of Booroola sheep. N. Z. J. Agric. Res. 1982, 25, 525–529. [Google Scholar] [CrossRef]
- Wang, J.Q.; Cao, W.G. Progress in exploring genes for high fertility in ewes. Hereditas 2011, 33, 953–961. [Google Scholar] [CrossRef] [PubMed]
- Chu, M.X.; Sang, L.H.; Wang, J.Y.; Fang, L.; Ye, S.C. Study on BMP15 and GDF9 as candidate genes for prolificacy of Small Tail Han sheep. Acta Genet. Sin. 2005, 32, 38–45. [Google Scholar] [PubMed]
- Bodin, L.; Di, P.E.; Fabre, S.; Bontoux, M.; Monget, P.; Persani, L.; Mulsant, P. A novel mutation in the bone morphogenetic protein 15 gene causing defective protein secretion is associated with both increased ovulation rate and sterility in Lacaune sheep. Endocrinology 2007, 148, 393–400. [Google Scholar] [CrossRef] [PubMed]
- Polley, S.; De, S.; Brahma, B.; Mukherjee, A.; Vinesh, P.V.; Batabyal, S.; Arora, J.S.; Pan, S.; Samanta, A.K.; Datta, T.K. Polymorphism of BMPR1B, BMP15 and GDF9 fecundity genes in prolific Garole sheep. Trop. Anim. Health Prod. 2010, 42, 985–993. [Google Scholar] [CrossRef] [PubMed]
- Souza, C.J.; McNeilly, A.S.; Benavides, M.V.; Melo, E.O.; Moraes, J.C. Mutation in the protease cleavage site of GDF9 increases ovulation rate and litter size in heterozygous ewes and causes infertility in homozygous ewes. Anim. Genet. 2014, 45, 732–739. [Google Scholar] [CrossRef] [PubMed]
- EI Fiky, Z.A.; Hassan, G.M.; Nassar, M.I. Genetic polymorphism of growth differentiation factor 9 (GDF9) gene related to fecundity in two Egyptian sheep breeds. J. Assist. Reprod. Genet. 2017, 34, 1683–1690. [Google Scholar] [CrossRef] [PubMed]
- Paz, E.; Quinones, J.; Bravo, S.; Montaldo, H.H.; Sepulveda, N. Genotyping of BMPR1B, BMP15 and GDF9 genes in chilean sheep breeds and association with prolificacy. Anim. Genet. 2015, 46, 98–99. [Google Scholar] [CrossRef] [PubMed]
- Shimasaki, S.; Moore, R.K.; Otsuka, F.; Erickson, G.F. The bone morphogenetic protein system in mammalian reproduction. Endocr. Rev. 2004, 25, 72–101. [Google Scholar] [CrossRef] [PubMed]
- Wilson, T.; Wu, X.Y.; Juengel, J.L.; Ross, I.K.; Lumsden, J.M.; Lord, E.A.; Dodds, K.G.; Walling, G.A.; Mcewan, J.C.; O’Connell, A.R. Highly prolific Booroola sheep have a mutation in the intracellular kinase domain of bone morphogenetic protein ib receptor (ALK-6) that is expressed in both oocytes and granulosa cells. Biol. Reprod. 2001, 64, 1225–1235. [Google Scholar] [CrossRef] [PubMed]
- Souza, C.J.; Macdougall, C.; Macdougall, C.; Campbell, B.K.; McNeilly, A.S.; Baird, D.T. The Booroola (FecB) phenotype is associated with a mutation in the bone morphogenetic receptor type 1b (bmpr1b) gene. J. Endocr. 2001, 169, 1–6. [Google Scholar] [CrossRef]
- Mulsant, P.; Lecerf, F.; Fabre, S.; Schibler, L.; Monget, P.; Lanneluc, I.; Pisselet, C.; Riquet, J.; Monniaux, D.; Callebaut, I.; et al. Mutation in bone morphogenetic protein receptor-ib is associated with increased ovulation rate in booroola merino ewes. Proc. Natl. Acad. Sci. USA 2001, 98, 5104–5109. [Google Scholar] [CrossRef] [PubMed]
- Galloway, S.M.; McNatty, K.P.; Cambridge, L.M.; Laitinen, M.P.E.; Juengel, J.L.; Jokiranta, T.S.; Mclaren, R.J.; Luiro, K.; Dodds, K.G.; Montgomery, G.W. Mutations in an oocyte-derived growth factor gene (BMP15) cause increased ovulation rate and infertility in a dosage-sensitive manner. Nat. Genet. 2000, 25, 279–283. [Google Scholar] [CrossRef] [PubMed]
- Hanrahan, J.P.; Gregan, S.M.; Mulsant, P.; Mullen, M.; Davis, G.H.; Powell, R.; Galloway, S.M. Mutations in the genes for oocyte-derived growth factors GDF9 and BMP15 are associated with both increased ovulation rate and sterility in Cambridge and Belclare sheep (Ovis aries). Biol. Reprod. 2004, 70, 900–909. [Google Scholar] [CrossRef] [PubMed]
- Pramod, R.K.; Sharma, S.K.; Singhi, A.; Pan, S.; Mitra, A. Differential ovarian morphometry and follicular expression of BMP15, GDF9 and BMPR1B influence the prolificacy in goat. Reprod. Domest. Anim. 2013, 48, 803–809. [Google Scholar] [CrossRef] [PubMed]
- Otsuka, F.; Yao, Z.; Lee, T.; Yamamoto, S.; Erickson, G.F.; Shimasaki, S. Bone morphogenetic protein-15. Identification of target cells and biological functions. J. Biol. Chem. 2000, 275, 39523–39528. [Google Scholar] [CrossRef] [PubMed]
- Vitt, U.A.; Hayashi, M.; Klein, C.; Hsueh, A.J.W. Growth differentiation factor-9 stimulates proliferation but suppresses the follicle-stimulating hormone-induced differentiation of cultured granulosa cells from small antral and preovulatory rat follicles. Bio. Reprod. 2000, 62, 370–377. [Google Scholar] [CrossRef] [PubMed]
- Davis, G.H.; Galloway, S.M.; Ross, I.K.; Gregan, S.M.; Ward, J.; Nimbkar, B.V.; Ghalsasi, P.M.; Nimbkar, C.; Gray, G.D.; Subandriyo; et al. DNA tests in prolific sheep from eight countries provide new evidence on origin of the Booroola (FecB) mutation1. Biol. Reprod. 2002, 66, 1869–1874. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Mishra, A.K.; Kolte, A.P.; Dash, S.K.; Karim, S.A. Screening for Booroola (FecB) and Galway (FecXG) mutations in Indian sheep. Small Rumin. Res. 2008, 80, 57–61. [Google Scholar] [CrossRef]
- Guan, F.; Liu, S.R.; Shi, G.Q.; Jun-Tao, A.I.; Mao, D.G.; Yang, L.G. Polymorphism of gene in nine sheep breeds or strains and its effects on litter size, lamb growth and development. Acta Genet. Sin. 2006, 33, 117–124. [Google Scholar] [CrossRef]
- Chu, M.X.; Liu, Z.H.; Jiao, C.L.; He, Y.Q.; Fang, L.; Ye, S.C.; Chen, G.H.; Wang, J.Y. Mutations in BMPR-IB and BMPp-15 genes are associated with litter size in Small Tailed Han sheep (Ovis aries). J. Anim. Sci. 2007, 85, 598–603. [Google Scholar] [CrossRef] [PubMed]
- Miao, X.Y.; Luo, Q.M.; Zhao, H.J.; Qin, X.Y. Co-expression analysis and identification of fecundity-related long non-coding rnas in sheep ovaries. Sci. Rep. 2016, 6, 39398–39407. [Google Scholar] [CrossRef] [PubMed]
- Liu, Q.Y.; Hu, W.P.; He, X.Y.; Pan, Z.Y.; Guo, X.F.; Feng, T.; Cao, G.L.; Huang, D.W.; He, J.N.; Di, R. Establishment of high-throughput molecular detection methods for ovine high fecundity major gene FecB and their application. Acta Veterinaria et Zootechnica Sinica 2017, 48, 39–51. [Google Scholar]
- Li, H.K.; Kuo, T.Y.; Yang, H.S.; Chen, L.R.; Li, S.L.; Huang, H.W. Differential gene expression of bone morphogenetic protein 15 and growth differentiation factor 9 during in vitro maturation of porcine oocytes and early embryos. Anim. Reprod. Sci. 2008, 103, 312–322. [Google Scholar] [CrossRef] [PubMed]
- Kathirvel, M.; Soundian, E.; Kumanan, V. Differential expression dynamics of growth differentiation factor 9 (GDF9) and bone morphogenetic factor15 (BMP15) mRNA transcripts during in vitro maturation of buffalo (Bubalus bubalis) cumulus–oocyte complexes. Springerplus 2013, 2, 206. [Google Scholar] [CrossRef] [PubMed]
- Jain, T.; Haldkar, M.; Sarkhel, B.C. Expression profile of BMPR1B gene in goat reproductive tract. Indian J. Anim. Res. 2014, 48, 329–335. [Google Scholar] [CrossRef]
- Mandon-Pepin, B.; Oustry-Vaiman, A.; Vigier, B.; Piumi, F.; Cribiu, E.; Cotinot, C. Expression profiles and chromosomal localization of genes controlling meiosis and follicular development in the sheep ovary. Biol. Reprod. 2003, 68, 985–995. [Google Scholar] [CrossRef] [PubMed]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-delta delta c(t)) method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
- Schmittgen, T.D.; Livak, K.J. Analyzing real-time PCR data by the comparative c(t) method. Nat. Protoc. 2008, 3, 1101–1108. [Google Scholar] [CrossRef] [PubMed]
- Meier, U. A note on the power of fisher’s least significant difference procedure. Pharm. Stat. 2006, 5, 253–263. [Google Scholar] [CrossRef] [PubMed]
- Foroughinia, G.; Fazileh, A.; Eghbalsaied, S. Expression of genes involved in BMP and estrogen signaling and AMPK production can be important factors affecting total number of antral follicles in ewes. Theriogenology 2017, 91, 36–43. [Google Scholar] [CrossRef] [PubMed]
- Yang, H.; Liu, S.R.; ZHong, F.G.; Yang, Y.L.; Zhang, Y.S. Different expression of BMPR-IB in tissues of sheep. Chin. J. Anim. Sci. 2009, 45, 6–8. [Google Scholar]
- Xu, Y.F.; Li, E.L.; Han, Y.D.; Chen, L.; Xie, Z.A. Differential expression of mRNAs encoding BMP/SMAD pathway molecules in antral follicles of high- and low-fecundity Hu sheep. Anim. Reprod. Sci. 2010, 120, 47–55. [Google Scholar] [CrossRef] [PubMed]
- Pan, Z.Y.; Di, R.; Tang, Q.Q.; Jin, H.H.; Chu, M.X.; Huang, D.W.; He, J.N.; Liu, Q.Y.; Hu, W.P.; Wang, X.Y. Tissue-specific mRNA expression profiles of GDF9, BMP15, and BMPR1B genes in prolific and non-prolific goat breeds. Czech J. Anim. Sci. 2015, 60, 452–458. [Google Scholar] [CrossRef]
- Yang, C.X.; Zi, X.D.; Wang, Y.; Yang, D.Q.; Ma, L.; Lu, J.Y.; Niu, H.R.; Xiao, X.L. Cloning and mRNA expression levels of GDF9, BMP15, and BMPR1B genes in prolific and non-prolific goat breeds. Mol. Reprod. Dev. 2012, 79, 2. [Google Scholar] [CrossRef] [PubMed]
- Valdecantos, P.; Bravo, M.R.; Garcia, E.; Garcia, D.; Roldan-Olarte, M.; Miceli, D. Expression of bone morphogenetic protein receptors in bovine oviductal epithelial cells: Evidence of autocrine BMP signaling. Anim. Reprod. Sci. 2017, 185, 89–96. [Google Scholar] [CrossRef] [PubMed]
- Galloway, S.M.; Gregan, S.M.; Wilson, T.; Mcnatty, K.P.; Juengel, J.L.; Ritvos, O.; Davis, G.H. BMP15 mutations and ovarian function. Mol. Cell. Endocr. 2002, 191, 15–18. [Google Scholar] [CrossRef]
- Hosoe, M.; Kaneyama, K.; Ushizawa, K.; Hayashi, K.G.; Takahashi, T. Quantitative analysis of bone morphogenetic protein 15 (BMP15) and growth differentiation factor 9 (GDF9) gene expression in calf and adult bovine ovaries. Reprod. Biol. Endocr. 2011, 9, 33. [Google Scholar] [CrossRef] [PubMed]
- Gupta, S.; Pandey, S.; Parmar, M.S.; Somal, A.; Paul, A.; Bsk, P.; Bhat, I.A.; Baiju, I.; Bharti, M.K.; Saikumar, G. Impact of oocyte-secreted factors on its developmental competence in buffalo. Zygote 2017, 25, 313–320. [Google Scholar] [CrossRef] [PubMed]
- Sugiura, K.; Su, Y.Q.; Diaz, F.J.; Pangas, S.A.; Sharma, S.; Wigglesworth, K.; O’Brien, M.J.; Matzuk, M.M.; Shimasaki, S.; Eppig, J.J. Oocyte-derived BMP15 and FGFs cooperate to promote glycolysis in cumulus cells. Development 2007, 134, 2593–2603. [Google Scholar] [CrossRef] [PubMed]
- Otsuka, F.; Shimasaki, S. A negative feedback system between oocyte bone morphogenetic protein 15 and granulosa cell kit ligand: Its role in regulating granulosa cell mitosis. Proc. Natl. Acad. Sci. USA 2002, 99, 8060–8065. [Google Scholar] [CrossRef] [PubMed]
- Hussein, T.S.; Froiland, D.A.; Amato, F.; Thompson, J.G.; Gilchrist, R.B. Oocytes prevent cumulus cell apoptosis by maintaining a morphogenic paracrine gradient of bone morphogenetic proteins. J. Cell Sci. 2005, 118, 5257–5268. [Google Scholar] [CrossRef] [PubMed]
- Dube, J.L.; Wang, P.; Elvin, J.; Lyons, K.M.; Celeste, A.J.; Matzuk, M.M. The bone morphogenetic protein 15 gene is X-linked and expressed in oocytes. Mol. Endocr. 1998, 12, 1809–1817. [Google Scholar] [CrossRef] [PubMed]
- Crawford, J.L.; Heath, D.A.; Reader, K.L.; Quirke, L.D.; Hudson, N.L.; Juengel, J.L.; McNatty, K.P. Oocytes in sheep homozygous for a mutation in bone morphogenetic protein receptor 1B express lower mRNA levels of bone morphogenetic protein 15 but not growth differentiation factor 9. Reproduction 2011, 142, 53–61. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.J.; Pan, X.Y.; LI, F.D.; Wang, W.M.; Li, T.F. Characteristics and expression of BMP15 gene and its association with litter size in sheep. J. Gansu Agric. Univ. 2015, 50, 40–48. [Google Scholar]
- Cui, H.X.; Zhao, S.M.; Cheng, M.L.; Guo, L.; Ye, R.Q.; Liu, W.Q.; Gao, S.Z. Cloning and expression levels of genes relating to the ovulation rate of the Yunling black goat. Biol. Reprod. 2009, 80, 219–226. [Google Scholar] [CrossRef] [PubMed]
- Almusawi, S.L.; Walton, K.L.; Heath, D.; Simpson, C.M.; Harrison, C.A. Species differences in the expression and activity of bone morphogenetic protein 15. Endocrinology 2013, 154, 888–899. [Google Scholar] [CrossRef] [PubMed]
- Zamani, P.; Nadri, S.; Saffaripour, R.; Ahmadi, A.; Dashti, F.; Abdoli, R. A new mutation in exon 2 of the bone morphogenetic protein 15 gene is associated with increase in prolificacy of Mehraban and Lori sheep. Trop. Anim. Health Prod. 2015, 47, 855–860. [Google Scholar] [CrossRef] [PubMed]
- Kaivo-Oja, N.; Bondestam, J.; Kamarainen, M.; Koskimies, J.; Vitt, U.; Cranfield, M.; Vuojolainen, K.; Kallio, J.P.; Olkkonen, V.M.; Hayashi, M. Growth differentiation factor-9 induces Smad2 activation and inhibin B production in cultured human granulosa-luteal cells. J. Clin. Endocr. Metab. 2003, 88, 755–762. [Google Scholar] [CrossRef] [PubMed]
- Elvin, J.A.; Clark, A.T.; Wang, P.; Wolfman, N.M.; Matzuk, M.M. Paracrine actions of growth differentiation factor-9 in the mammalian ovary. Mol. Endocr. 1999, 13, 1035–1048. [Google Scholar] [CrossRef] [PubMed]
- Hreinsson, J.G.; Scott, J.E.; Rasmussen, C.; Swahn, M.L.; Hsueh, A.J.W.; Hovatta, O. Growth differentiation factor-9 promotes the growth, development, and survival of human ovarian follicles in organ culture. J. Clin. Endocr. Metab. 2002, 87, 316–321. [Google Scholar] [CrossRef] [PubMed]
- Hsueh, A.J.; Mcgee, E.A.; Hayashi, M.; Hsu, S.Y. Hormonal regulation of early follicle development in the rat ovary. Mol. Cell. Endocr. 2000, 163, 95–100. [Google Scholar] [CrossRef]
- Paradis, F.; Novak, S.; Murdoch, G.K.; Dyck, M.K.; Dixon, W.T.; Foxcroft, G.R. Temporal regulation of BMP2, BMP6, BMP15, GDF9, BMPR1A, BMPR1B, BMPR2 and TGFBR1 mRNA expression in the oocyte, granulosa and theca cells of developing preovulatory follicles in the pig. Reproduction 2009, 138, 115–129. [Google Scholar] [CrossRef] [PubMed]
- Hu, D.L.; Li, Q.F.; Xu, Y.F.; Li, E.L.; Han, Y.D.; Tu, F.; Xie, Z. The tissue expression profile, mRNA expression level and SNPs analysis on GDF9 gene in Hu sheep. J. Agric. Biotechnol. 2010, 18, 533–538. [Google Scholar]
- Goyal, S.; Aggarwal, J.; Dubey, P.K.; Mishra, B.P.; Ghalsasi, P.; Nimbkar, C.; Joshi, B.K.; Kataria, R.S. Expression analysis of genes associated with prolificacy in fecB carrier and noncarrier Indian sheep. Anim. Biotechnol. 2017, 28, 220–227. [Google Scholar] [CrossRef] [PubMed]
| Method | Gene Name | Primer Sequence (5′→3′) | Length (bp) | Tm (°C) | Accession No. |
|---|---|---|---|---|---|
| qPCR primer | BMPR1B | F: 5′-TGACGGACCTATACACCACA-3′ R: 5′-GTACCGAGGTCTGGCTTCTT-3′ | 121 | 60 | NM_001142888.2 |
| BMP15 | F: 5′-TGTTGGGCAAAAGCTCTGGA-3′ R: 5′-GCCATGCCACCAGAACTCAA-3′ | 106 | 60 | NM_001114767.1 | |
| GDF9 | F: 5′-AACAGACGCCACCTCTACAA-3′ R: 5′-CACGATCCAGGTTAAACAGCA-3′ | 124 | 60 | NM_001009431.1 | |
| β-actin | F: 5′-ACCCAGCACGATGAAGATCA-3′ R: 5′-GTAACGCAGCTAACAGTCCG-3 | 97 | 60 | NM_001009784.1 | |
| RT-PCR primer | BMPR1B | F: 5′-GGGTTCTACGACTCCGCTTC-3′ R: 5′-GGTTACTTTCAGGCCCATCAT-3 | 237 | 60 | NM_001142888.2 |
| BMP15 | F: 5′-GGGTTCTACGACTCCGCTTC-3′ R: 5′-GGTTACTTTCAGGCCCATCAT-3′ | 273 | 62 | NM_001114767.1 | |
| GDF9 | F: 5′-TAGTCAGCTGAAGTGGGACA-3′ R: 5′-AGCCATCAGGCTCGATGGCC-3′ | 224 | 61 | NM_001009431.1 | |
| β-actin | F: 5′-ACCCAGCACGATGAAGATCA-3′ R: 5′-GTAACGCAGCTAACAGTCCG-3′ | 187 | 61 | NM_001009784.1 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tang, J.; Hu, W.; Di, R.; Liu, Q.; Wang, X.; Zhang, X.; Zhang, J.; Chu, M. Expression Analysis of the Prolific Candidate Genes, BMPR1B, BMP15, and GDF9 in Small Tail Han Ewes with Three Fecundity (FecB Gene) Genotypes. Animals 2018, 8, 166. https://doi.org/10.3390/ani8100166
Tang J, Hu W, Di R, Liu Q, Wang X, Zhang X, Zhang J, Chu M. Expression Analysis of the Prolific Candidate Genes, BMPR1B, BMP15, and GDF9 in Small Tail Han Ewes with Three Fecundity (FecB Gene) Genotypes. Animals. 2018; 8(10):166. https://doi.org/10.3390/ani8100166
Chicago/Turabian StyleTang, Jishun, Wenping Hu, Ran Di, Qiuyue Liu, Xiangyu Wang, Xiaosheng Zhang, Jinlong Zhang, and Mingxing Chu. 2018. "Expression Analysis of the Prolific Candidate Genes, BMPR1B, BMP15, and GDF9 in Small Tail Han Ewes with Three Fecundity (FecB Gene) Genotypes" Animals 8, no. 10: 166. https://doi.org/10.3390/ani8100166
APA StyleTang, J., Hu, W., Di, R., Liu, Q., Wang, X., Zhang, X., Zhang, J., & Chu, M. (2018). Expression Analysis of the Prolific Candidate Genes, BMPR1B, BMP15, and GDF9 in Small Tail Han Ewes with Three Fecundity (FecB Gene) Genotypes. Animals, 8(10), 166. https://doi.org/10.3390/ani8100166





