Inclusion of Oat in Feeding Can Increase the Potential Probiotic Bifidobacteria in Sow Milk
Simple Summary
Abstract
1. Introduction
2. Experimental Section
2.1. Animals, Experimental Diet, and Milk Collection
| Feed Ingredients | Gestation (% of Feed) | |
| Control | Oat | |
| 11102 NCDA Corn (Rolled) | 80.3 | 65.38 |
| Soybean Meal | 13.8 | 13.8 |
| Ground Whole Oat 15% | 0 | 15 |
| Corn (1/8) Micro-Flush | 1 | 1 |
| Limestone Fine | 1.11 | 1.11 |
| MON-CAL 21% P | 2.05 | 2.05 |
| Salt | 0.5 | 0.5 |
| Swine TM PX (KSU) | 0.15 | 0.15 |
| Swine Sow-Pig VIT | 0.04 | 0.04 |
| Poultry Fat | 1 | 1 |
| Feed Ingredients | Lactation (% of Feed) | |
| Control | Oat | |
| Corn NCDA | 73 | 64.8 |
| Soybean Meal 48% | 17.6 | 10.8 |
| Ground Whole Oat 15% | 0 | 15 |
| Corn (1/8) Micro-Flush | 1 | 1 |
| Limestone Fine | 1.08 | 1.08 |
| MON-CAL 21% P | 2.38 | 2.38 |
| L-Lysine 50% | 0.25 | 0.25 |
| Salt | 0.5 | 0.5 |
| Threonine | 0.01 | 0.01 |
| Swine TM Prmx (KSU) | 0.15 | 0.15 |
| Swine VTM Prmx | 0.04 | 0.04 |
| Poultry Fat | 3.99 | 3.99 |
2.2. Enumeration of Bacterial Population
2.3. Identification of Bifidobacterium from Sow Milk
2.3.1. Phenotypical Characterization
2.3.1.1. Aggregation Assay
2.3.1.2. Antimicrobial Activity Assay
2.3.1.3. Determination of Bile Tolerance
2.3.1.4. Tolerance of Isolates at Low pH
2.3.1.5. Antibiotic Susceptibility Test
2.4. Data Preparation and Statistical Analysis
3. Results
3.1. Population of Lactobacillus and Bifidobacterium Spp.
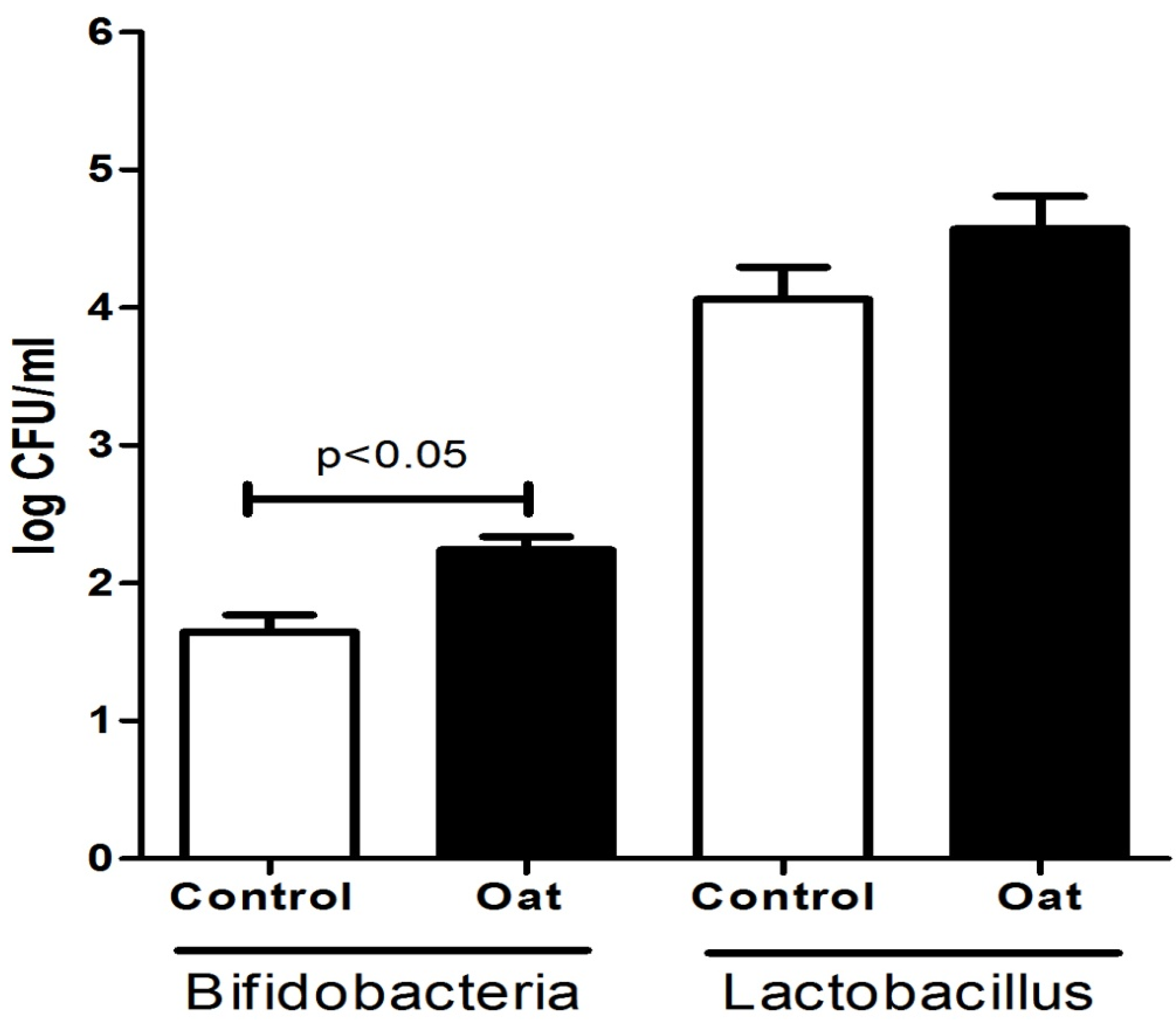
3.2. Phenotypic Characterization
3.2.1. Autoaggregation and Antimicrobial Activity

3.2.2. Bile and Acid Tolerance
| Isolate No. | Autoaggregation (%) | Zone of Inhibition (mm) |
|---|---|---|
| 1 | Nonaggregating | 10.0 ± 0.06 |
| 2 | Nonaggregating | 8.0 ± 0.07 |
| 3 | 17.5 ± 2.12 | 9.0 ± 0.04 |
| 4 | 25.5 ± 2.12 | 9.0 ± 0.06 |
| 5 | 68.0 ± 1.41 | 11.0 ± 0.04 |
| 6 | 45.5 ± 3.53 | 9.0 ± 0.08 |
| 7 | 62.0 ± 2.82 | 8.0 ± 0.06 |
| 8 | 73.0 ± 2.80 | 8.0 ± 0.04 |
| 9 | 66.0 ± 1.41 | 10.0 ± 0.05 |
| 10 | 60.5 ± 2.12 | 10.0 ± 0.03 |
| 11 | 51.0 ± 2.82 | 10.0 ± 0.04 |
| 12 | 63.5 ± 2.12 | 9.0 ± 0.05 |
| 13 | 64.0 ± 1.41 | 6.0 ± 0.06 |
| 14 | 25.0 ± 3.50 | 10.0 ± 0.03 |
| Isolate No. | Bile (%) | ||||
|---|---|---|---|---|---|
| 0% | 0.25% | 0.50% | 1.00% | 2.00% | |
| 1 | 0.98 ± 0.04 | 0.73 ± 0.04 | 0.54 ± 0.08 | 0.48±0.07 | 0.41 ± 0.11 |
| 2 | 0.95 ± 0.04 | 0.70 ± 0.08 | 0.47 ± 0.04 | 0.41±0.08 | 0.35 ± 0.12 |
| 3 | 0.93 ± 0.01 | 0.71 ± 0.08 | 0.51 ± 0.01 | 0.36±0.07 | 0.27 ± 0.08 |
| 4 | 0.82 ± 0.11 | 0.65 ± 0.01 | 0.46 ± 0.01 | 0.37±0.04 | 0.33 ± 0.10 |
| 5 | 0.93 ± 0.06 | 0.81 ± 0.03 | 0.68 ± 0.03 | 0.62±0.04 | 0.45 ± 0.11 |
| 6 | 1.02 ± 0.04 | 0.73 ± 0.04 | 0.72 ± 0.02 | 0.60±0.04 | 0.51 ± 0.05 |
| 7 | 1.01 ± 0.08 | 0.82 ± 0.04 | 0.74 ± 0.04 | 0.61±0.03 | 0.48 ± 0.04 |
| 8 | 0.96 ± 0.02 | 0.82 ± 0.03 | 0.73 ± 0.04 | 0.68±0.05 | 0.51±0.05 |
| 9 | 0.94 ± 0.06 | 0.74 ± 0.04 | 0.64 ± 0.04 | 0.65 ± 0.07 | 0.57 ± 0.04 |
| 10 | 0.89 ± 0.07 | 0.69 ± 0.04 | 0.53 ± 0.04 | 0.48 ± 0.01 | 0.43 ± 0.02 |
| 11 | 0.90 ± 0.05 | 0.72 ± 0.01 | 0.52 ± 0.05 | 0.43 ± 0.04 | 0.38 ± 0.04 |
| 12 | 0.85 ± 0.05 | 0.69 ± 0.01 | 0.54 ± 0.03 | 0.42 ± 0.04 | 0.36 ± 0.04 |
| 13 | 1.03 ± 0.18 | 0.95 ± 0.04 | 0.58 ± 0.02 | 0.53 ± 0.03 | 0.34 ± 0.04 |
| 14 | 1.01± 0.11 | 0.71 ± 0.04 | 0.56 ± 0.02 | 0.47 ± 0.02 | 0.41 ± 0.04 |
| Average | 0.94 ± 0.07 a | 0.75 ± 0.03 b | 0.59 ± 0.04 c | 0.51 ± 0.04 d | 0.41 ± 0.03 e |
| Isolate No. | pH 6.5 | pH 2.5 | ||
|---|---|---|---|---|
| 0 min | 90 min | 0 min | 90 min | |
| 1 | 7.32 ± 0.13 | 7.72 ± 0.12 | 7.31 ± 0.02 | 2.83 ± 0.03 |
| 2 | 7.07 ± 0.06 | 7.74 ± 0.04 | 6.84 ± 0.06 | 2.63 ± 0.14 |
| 3 | 7.14 ± 0.03 | 7.41 ± 0.02 | 6.48 ± 0.07 | 2.35 ± 0.15 |
| 4 | 6.63 ± 0.04 | 7.21 ± 0.01 | 6.48 ± 0.02 | 2.65 ± 0.14 |
| 5 | 9.37 ± 0.24 | 9.41 ± 0.10 | 9.24 ± 0.05 | 3.87 ± 0.04 |
| 6 | 9.56 ± 0.08 | 9.77 ± 0.02 | 9.42 ± 0.04 | 3.31 ± 0.19 |
| 7 | 9.47 ± 0.11 | 9.56 ± 0.17 | 9.41 ± 0.09 | 2.70 ± 0.15 |
| 8 | 9.49 ± 0.04 | 9.51 ± 0.05 | 9.27 ± 0.10 | 3.30 ± 0.04 |
| 9 | 8.46 ± 0.02 | 8.73 ± 0.07 | 8.39 ± 0.10 | 2.64 ± 0.09 |
| 10 | 8.36 ± 0.04 | 8.62 ± 0.19 | 8.17 ± 0.05 | 2.67 ± 0.16 |
| 11 | 8.74 ± 0.05 | 8.80 ± 0.03 | 8.26 ± 0.06 | 3.34 ± 0.06 |
| 12 | 8.33 ± 0.10 | 8.88 ± 0.19 | 8.08 ± 0.13 | 4.06 ± 0.02 |
| 13 | 9.01 ± 0.01 | 9.18 ± 0.07 | 8.11 ± 0.04 | 4.93 ± 0.03 |
| 14 | 8.18 ± 0.02 | 9.00 ± 0.08 | 7.93 ± 0.10 | 4.01 ± 0.07 |
| Average | 8.36 ± 0.07 | 8.68 ± 0.08 | 8.10 ± 0.07 | 3.23 ± 0.09 |
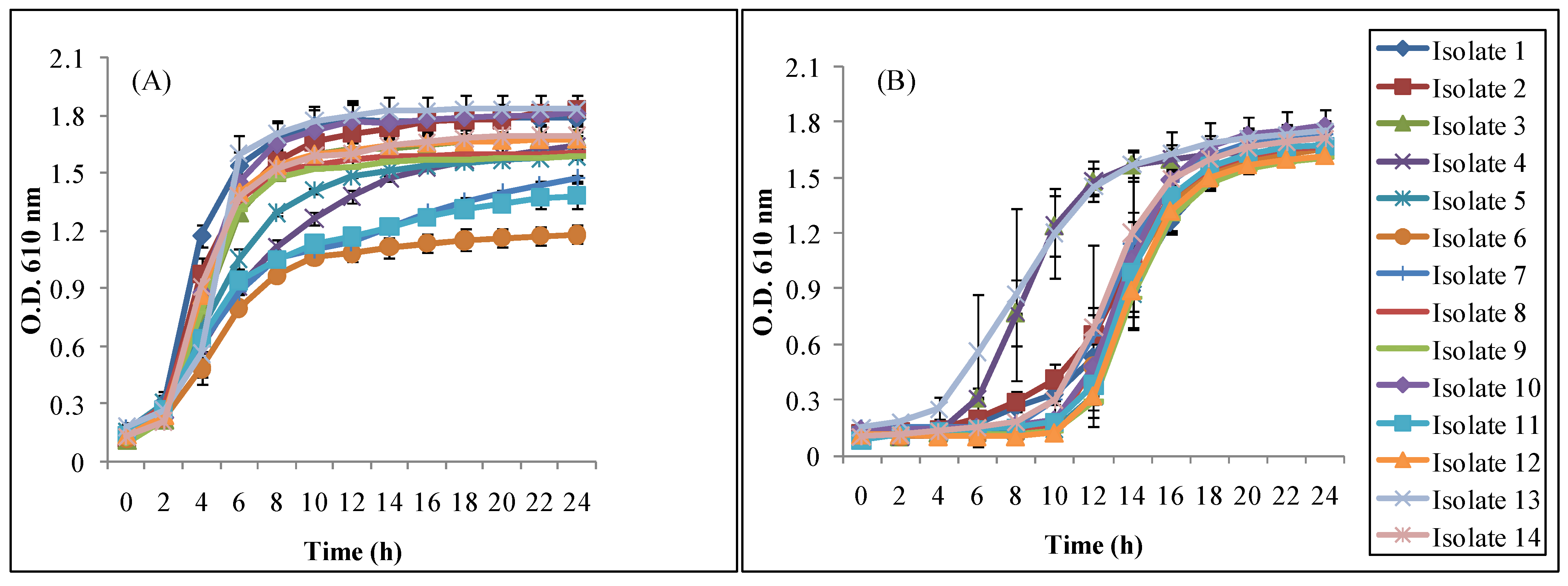
3.2.3. Antibiotic Susceptibility
| Antibiotics (mg/L) | ||||||||
|---|---|---|---|---|---|---|---|---|
| Isolate No. | Ampicillin | Chloramphenicol | Erythromycin | Gentamicin | ||||
| 4 | 5 | 5 | 6 | 1 | 2 | 65 | 66 | |
| 1 | S | S | S | S | S | S | R | R |
| 2 | S | S | S | S | R | R | R | R |
| 3 | S | S | S | S | R | R | R | R |
| 4 | S | S | S | S | R | R | R | R |
| 5 | S | S | S | S | R | R | R | R |
| 6 | S | S | S | S | R | R | R | R |
| 7 | S | S | S | S | R | R | R | R |
| 8 | S | S | S | S | S | S | R | R |
| 9 | S | S | S | S | R | R | R | R |
| 10 | S | S | S | S | R | R | R | R |
| 11 | S | S | S | S | R | R | R | R |
| 12 | S | S | S | S | S | S | R | R |
| 13 | S | S | S | S | S | S | R | R |
| 14 | S | S | S | S | S | S | R | R |
4. Discussion
5. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Guthrie, H.D.; Pursel, V.G.; Frobish, L.T. Attempts to induce conception in lactating sows. J. Anim. Sci. 1978, 47, 1145–1151. [Google Scholar] [PubMed]
- Blecha, F.; Pollman, D.S.; Nichols, D.A. Weaning pigs at an early age decreases cellular immunity. J. Anim. Sci. 1983, 56, 396–400. [Google Scholar] [PubMed]
- Van der Fels-Klerx, H.J.; Puister-Jansen, L.F.; van Asselt, E.D.; Burgers, S.L. Farm factors associated with the use of antibiotics in pig production. J. Anim. Sci. 2011, 89, 1922–1929. [Google Scholar]
- Vondruskova, H.; Slamova, R.; Trckova, M.; Zraly, Z.; Pavlik, I. Alternatives to antibiotic growth promoters in prevention of diarrhoea in weaned piglets: A review. Vet. Med. 2010, 55, 199–224. [Google Scholar]
- Alexopoulos, C.; Georgoulakis, I.E.; Tzivara, A.; Kritas, S.K.; Siochu, A.; Kyriakis, S.C. Field evaluation of the efficacy of a probiotic containing bacillus licheniformis and Bacillus subtilis spores, on the health status and performance of sows and their litters. J. Anim. Physiol. Anim. Nutr. 2004, 88, 381–392. [Google Scholar] [CrossRef] [PubMed]
- Cho, J.H.; Zhao, P.Y.; Kim, I.H. Probiotics as a dietary additive for pigs: A review. J. Anim. Vet. Adv. 2011, 10, 2127–2134. [Google Scholar] [CrossRef]
- Ibrahim, S.A.; Dharmavaram, S.R.; Seo, C.W.; Shahbazi, G. Antimicrobial activity of Bifidobacterium longum (NCFB 2259) as influenced by spices. Internet J. Food Saf. 2003, 2, 6–8. [Google Scholar]
- Sanz, Y. Gut microbiota and probiotics in maternal and infant health. Am. J. Clin. Nutr. 2011, 94, 2000S–2005S. [Google Scholar] [CrossRef] [PubMed]
- Munoa, F.J.; Pares, R. Selective medium for isolation and enumeration of Bifidobacterium spp. Appl. Environ. Microbiol. 1988, 54, 1715–1718. [Google Scholar] [PubMed]
- Ibrahim, S.A.; Hassan, O.A.; Salameh, M.M.; Shahbazi, A. Effect of media composition and incubation temperatures on autoaggregation behavior of bifidobacteria. Milchwissenschaft 2005, 60, 127–129. [Google Scholar]
- Awaisheh, S.S.; Ibrahim, S.A. Screening of antibacterial activity of lactic acid bacteria against different pathogens found in vacuum-packaged meat products. Foodborne Pathog. Dis. 2009, 6, 1125–1132. [Google Scholar] [CrossRef] [PubMed]
- European Food Safety Authority (EFSA). Technical guidance prepared by the panel on additives and products or substances used in animal feed (FEEDAP) on the update of the criteria used in the assessment of bacterial resistance to antibiotics of human or veterinary importance. EFSA J. 2008, 732, 1–15. [Google Scholar]
- Fairbrother, J.M.; Nadeau, É.; Gyles, C.L. Escherichia coli in postweaning diarrhea in pigs: An update on bacterial types, pathogenesis, and prevention strategies. Anim. Health Res. Rev. 2005, 6, 17–39. [Google Scholar] [PubMed]
- Jacobson, M.; af Segerstad, C.H.; Gunnarsson, A.; Fellström, C.; de Verdier Klingenberg, K.; Wallgren, P.; Jensen-Waern, M. Diarrhoea in the growing pig—A comparison of clinical, morphological and microbial findings between animals from good and poor performance herds. Res. Vet. Sci. 2003, 74, 163–169. [Google Scholar] [CrossRef]
- Frydendahl, K. Prevalence of serogroups and virulence genes in Escherichia coli associated with postweaning diarrhoea and edema disease in pigs and a comparison of diagnostic approaches. Vet. Microbiol. 2002, 85, 169–182. [Google Scholar] [CrossRef]
- Casewell, M.; Friis, C.; Marco, E.; McMullin, P.; Phillips, I. The European ban on growth-promoting antibiotics and emerging consequences for human and animal health. J. Antimicrob. Chemother. 2003, 52, 159–161. [Google Scholar] [CrossRef] [PubMed]
- Gavini, F.; Delcenserie, V.; Kopeinig, K.; Pollinger, S.; Beerens, H.; Bonaparte, C.; Upmann, M. Bifidobacterium species isolated from animal feces and from beef and pork meat. J. Food Prot. 2006, 69, 871–877. [Google Scholar] [PubMed]
- Kontula, P.; von Wright, A.; Mattila-Sandholm, T. Oat bran β-gluco-and xylo-oligosaccharides as fermentative substrates for lactic acid bacteria. Int. J. Food Microbiol. 1999, 45, 163–169. [Google Scholar] [CrossRef]
- Tuohy, K.M.; Brown, D.T.; Klinder, A.; Costabile, A. Shaping the human microbiome with prebiotic foods: Current perspectives for continued development. Food Sci. Technol. Bull. 2010, 7, 49–64. [Google Scholar] [CrossRef]
- Hinz, S.W.; Pastink, M.I.; van den Broek, L.A.; Vincken, J.-P.; Voragen, A.G. Bifidobacterium longum endogalactanase liberates galactotriose from type I galactans. Appl. Environ. Microbiol. 2005, 71, 5501–5510. [Google Scholar] [CrossRef] [PubMed]
- Connolly, M.L.; Lovegrove, J.A.; Tuohy, K.M. In vitro evaluation of the microbiota modulation abilities of different sized whole oat grain flakes. Anaerobe 2010, 16, 483–488. [Google Scholar] [CrossRef] [PubMed]
- Jost, T.; Lacroix, C.; Braegger, C.P.; Rochat, F.; Chassard, C. Vertical mother–neonate transfer of maternal gut bacteria via breastfeeding. Environ. Microbiol. 2014, 16, 2891–2904. [Google Scholar] [CrossRef] [PubMed]
- Argyri, A.A.; Zoumpopoulou, G.; Karatzas, K.-A.G.; Tsakalidou, E.; Nychas, G.-J.E.; Panagou, E.Z.; Tassou, C.C. Selection of potential probiotic lactic acid bacteria from fermented olives by in vitro tests. Food Microbiol. 2013, 33, 282–291. [Google Scholar] [CrossRef] [PubMed]
- Delgado, S.; O’sullivan, E.; Fitzgerald, G.; Mayo, B. Subtractive screening for probiotic properties of lactobacillus species from the human gastrointestinal tract in the search for new probiotics. J. Food Sci. 2007, 72, M310–M315. [Google Scholar] [CrossRef] [PubMed]
- Salyers, A.A.; Gupta, A.; Wang, Y. Human intestinal bacteria as reservoirs for antibiotic resistance genes. Trends. Microbiol. 2004, 12, 412–416. [Google Scholar] [CrossRef] [PubMed]
- Yun, J.H.; Lee, K.B.; Sung, Y.K.; Kim, E.B.; Lee, H.G.; Choi, Y.J. Isolation and characterization of potential probiotic lactobacilli from pig feces. J. Basic Microbiol. 2009, 49, 220–226. [Google Scholar] [CrossRef] [PubMed]
- Ibrahim, S.A.; Bezkorovainy, A. Survival of bifidobacteria in the presence of bile salt. J. Sci. Food Agric. 1993, 62, 351–354. [Google Scholar] [CrossRef]
- Lee, J.-H.; O’Sullivan, D.J. Genomic insights into bifidobacteria. Microbiol. Mol. Biol. Rev. 2010, 74, 378–416. [Google Scholar] [CrossRef] [PubMed]
- Ruiz, L.; Ruas-Madiedo, P.; Gueimonde, M.; de los Reyes-Gavilán, C.G.; Margolles, A.; Sánchez, B. How do bifidobacteria counteract environmental challenges? Mechanisms involved and physiological consequences. Genes Nutr. 2011, 6, 307–318. [Google Scholar] [CrossRef] [PubMed]
- Zárate, G.; Morata de Ambrosini, V.I.; Perez Chaia, A.; González, S.N. Adhesion of dairy propionibacteria to intestinal epithelial tissue in vitro and in vivo. J. Food Prot. 2002, 65, 534–539. [Google Scholar] [PubMed]
- Agaliya, P.; Jeevaratnam, K. Screening of lactobacillus plantarum isolated from fermented idli batter for probiotic properties. Afr. J. Biotechnol. 2012, 11, 12856–12864. [Google Scholar]
- Moubareck, C.; Gavini, F.; Vaugien, L.; Butel, M.J.; Doucet-Populaire, F. Antimicrobial susceptibility of bifidobacteria. J. Antimicrob. Chemother. 2005, 55, 38–44. [Google Scholar] [CrossRef] [PubMed]
- D’Aimmo, M.R.; Modesto, M.; Biavati, B. Antibiotic resistance of lactic acid bacteria and Bifidobacterium spp. Isolated from dairy and pharmaceutical products. Int. J. Food Microbiol. 2007, 115, 35–42. [Google Scholar] [CrossRef] [PubMed]
- Charteris, W.P.; Kelly, P.M.; Morelli, L.; Collins, J.K. Antibiotic susceptibility of potentially probiotic lactobacillus species. J. Food Prot. 1998, 61, 1636–1643. [Google Scholar] [PubMed]
- Curragh, H.J.; Collins, M. High levels of spontaneous drug resistance in lactobacillus. J. Appl. Bacterial. 1992, 73, 31–36. [Google Scholar] [CrossRef]
- Dicks, L.; Todorov, S.; Franco, B. Current Status of Antibiotic Resistance in Lactic Acid Bacteria. Antibiotic Resistance: Causes and Risk Factors, Mechanisms and Alternatives; Bonilla, A.R., Muniz, K.P., Eds.; Nova Publications: Delhi, Indian, 2009; pp. 379–425. [Google Scholar]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gyawali, R.; Minor, R.C.; Donovan, B.; Ibrahim, S.A. Inclusion of Oat in Feeding Can Increase the Potential Probiotic Bifidobacteria in Sow Milk. Animals 2015, 5, 610-623. https://doi.org/10.3390/ani5030375
Gyawali R, Minor RC, Donovan B, Ibrahim SA. Inclusion of Oat in Feeding Can Increase the Potential Probiotic Bifidobacteria in Sow Milk. Animals. 2015; 5(3):610-623. https://doi.org/10.3390/ani5030375
Chicago/Turabian StyleGyawali, Rabin, Radiah C. Minor, Barry Donovan, and Salam A. Ibrahim. 2015. "Inclusion of Oat in Feeding Can Increase the Potential Probiotic Bifidobacteria in Sow Milk" Animals 5, no. 3: 610-623. https://doi.org/10.3390/ani5030375
APA StyleGyawali, R., Minor, R. C., Donovan, B., & Ibrahim, S. A. (2015). Inclusion of Oat in Feeding Can Increase the Potential Probiotic Bifidobacteria in Sow Milk. Animals, 5(3), 610-623. https://doi.org/10.3390/ani5030375






