Integrating Genomic Selection and Genome-Wide Association Study to Enhance Reproductive Traits in Thai Swamp Buffalo
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Data Collection
2.2. Genetic Parameter Estimation
2.3. Estimation of GEBVs
2.4. GWAS
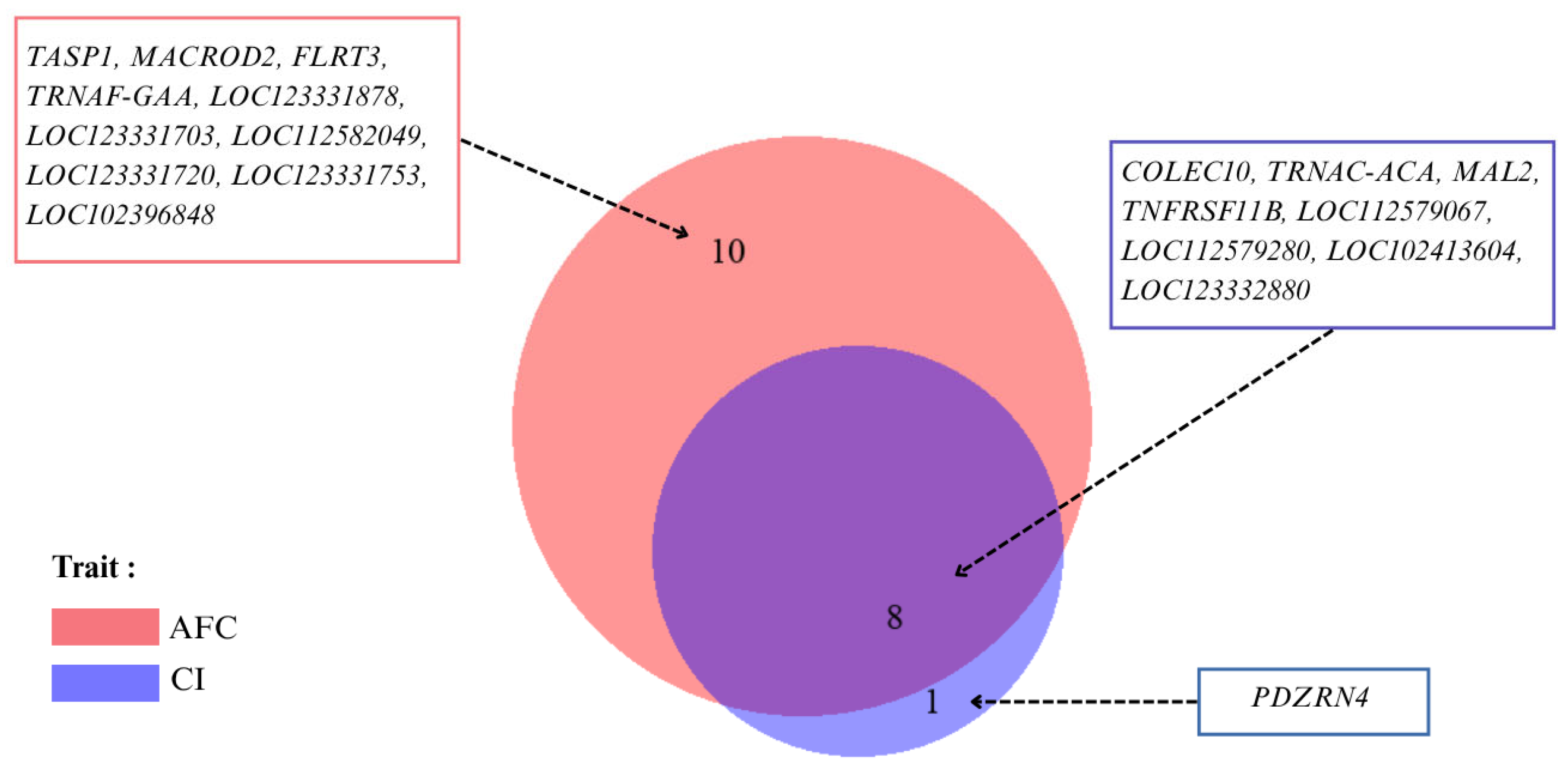
2.5. Identification of Candidate and Pleiotropic Genes
3. Results
3.1. Descriptive Statistics of Reproductive Traits
3.2. Variance Components and Genetic Parameter Estimates
3.3. Comparison of EBVs and GEBVs for Reproductive Traits
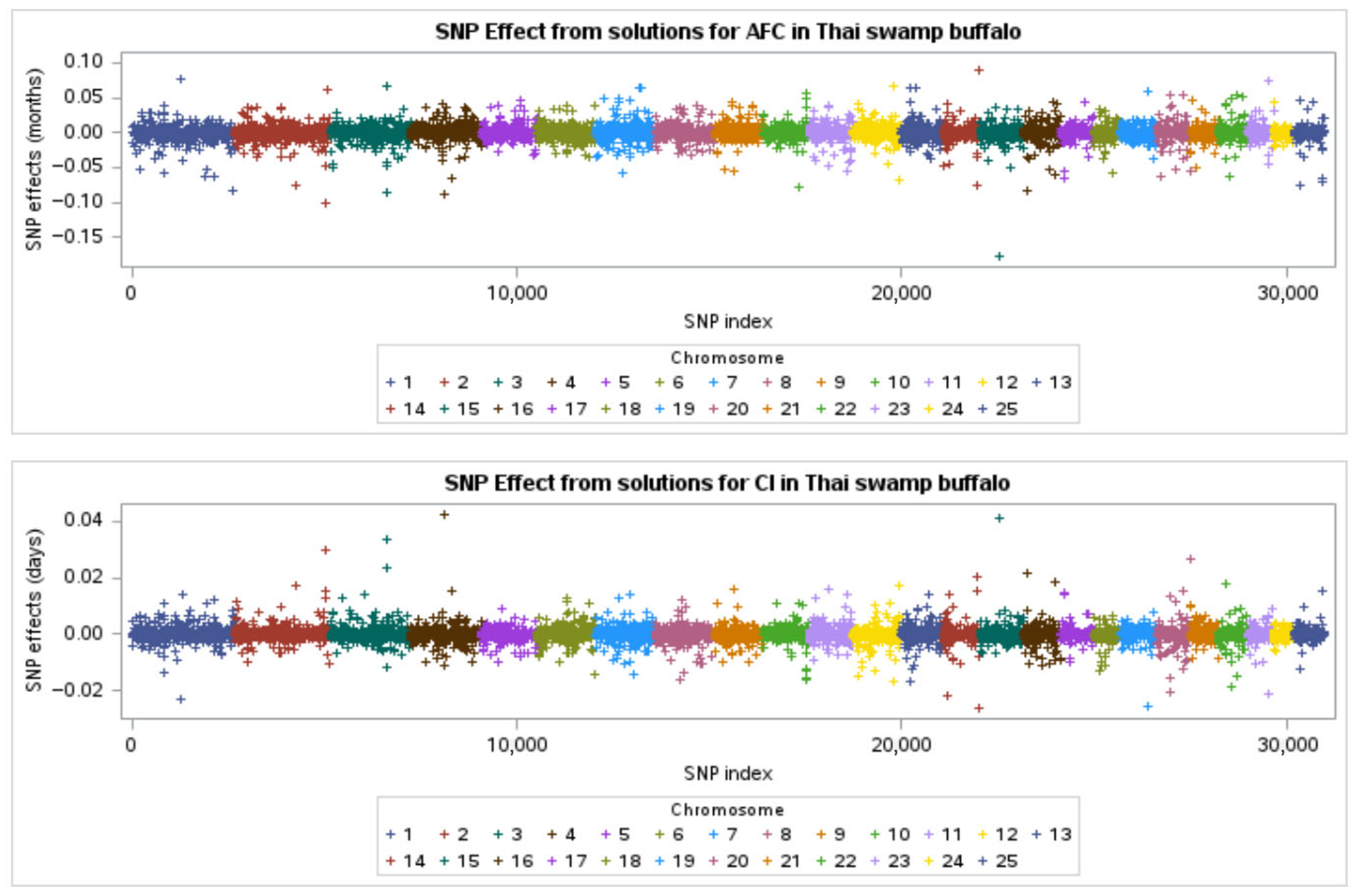
3.4. SNP Effects and Genomic Regions Associated with Reproductive Traits
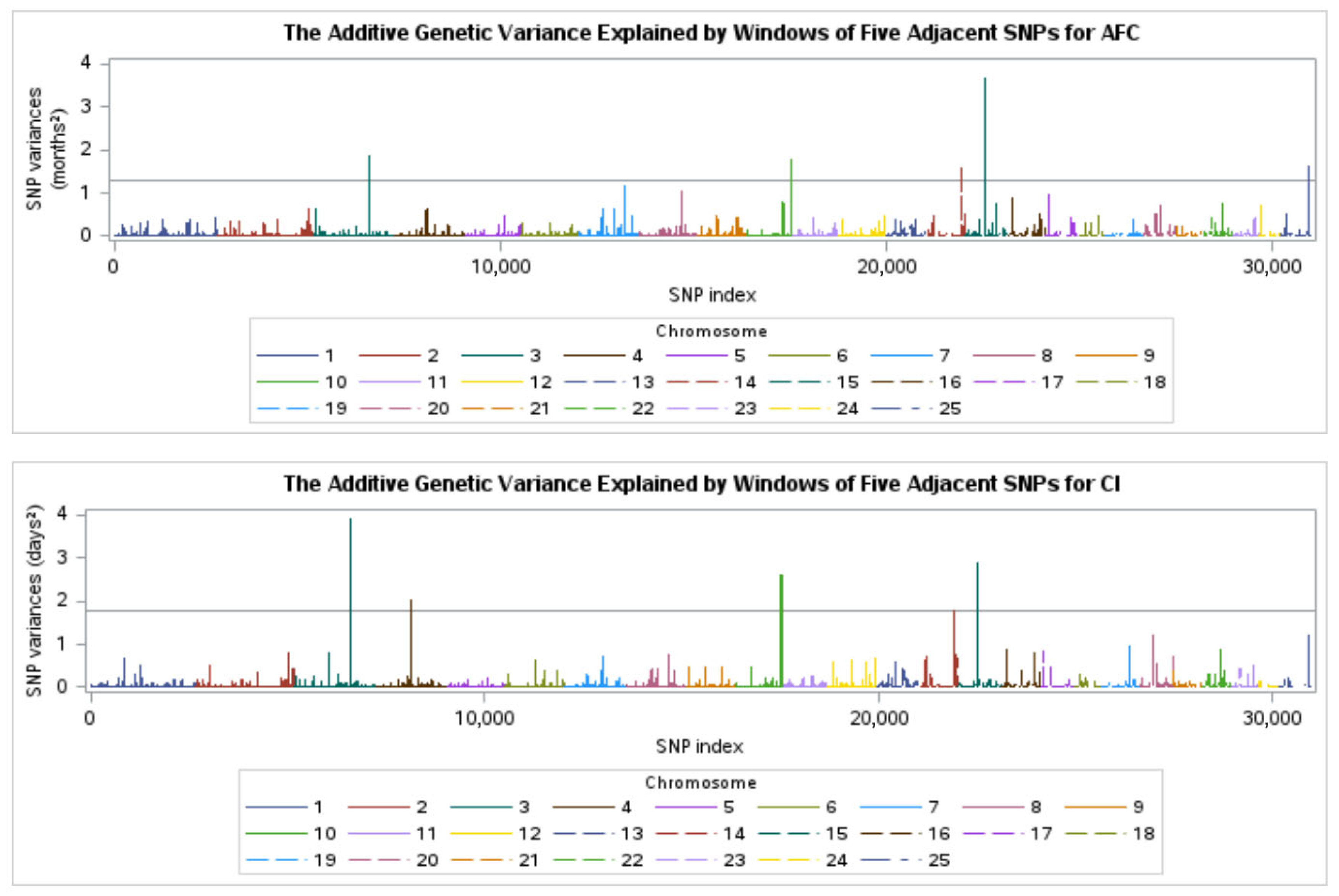
3.5. WssGWAS
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Wanapat, M.; Chanthakhoun, V. Buffalo production for emerging market as a potential animal protein source for global population. Buffalo Bull. 2015, 34, 169–180. [Google Scholar]
- Siddiky, M.; Faruque, M. Buffaloes for dairying in South Asia: Potential, challenges and way forward. SAARC J. Agric. 2018, 15, 227–239. [Google Scholar] [CrossRef]
- Kenchaiwong, W.; Pongthaisong, P.; Kananit, S.; Duangjinda, M.; Boonkum, W. An analysis of DNA sequence polymorphism in the swamp buffalo toll-like receptor (TLR2) gene. Animals 2023, 13, 2012. [Google Scholar] [CrossRef]
- Chantalakhana, C. Buffalo breeding program in Thailand. In Proceedings of the Workshop on Developing Breeding Strategies for Lower Input Animal Production Environments, Bella, Italy, 22–25 September 1999. [Google Scholar]
- Suphachavalit, S.; Sricharoen, P.; Luesopha, T.; Srisakdi, T.; Na-Chiangmai, A.; Boonprong, S. Swamp buffalo production system and needs for extension on local scale farmers in the lower northeast of Thailand. Buffalo Bull. 2013, 32, 1204–1207. [Google Scholar]
- IBIC (International Buffalo Information Center). 2022. Available online: https://ibic.lib.ku.ac.th/buf-th/61-statistic-buffalo (accessed on 25 June 2025).
- DLD (Department of Livestock Development). 2024. Available online: https://ict.dld.go.th/webnew/images/stories/report/regislives/2567/eRegist2567.pdf (accessed on 25 June 2025).
- Kanloung, T.; Taemchuay, D.; Hengtrakunsin, R.; Tavitchasri, P. The reproductive performances of Murrah and swamp buffaloes in Thailand. Buffalo Bull. 2021, 40, 495–498. [Google Scholar]
- Jongsuwanwattana, R.; Swangchan-uthai, T.; Sirivaidyapong, S. Reproduction and in vitro technologies of swamp buffalo; past, present, and future. Thai J. Vet. Med. 2024, 54, 115–124. [Google Scholar] [CrossRef]
- Srirattana, K.; Hufana-Duran, D.; Atabay, E.P.; Duran, P.G.; Atabay, E.C.; Lu, K.; Liang, Y.; Chaikhun-Marcou, T.; Theerakittayakorn, K.; Parnpai, R. Current status of assisted reproductive technologies in buffaloes. Anim. Sci. J. 2022, 93, e13767. [Google Scholar] [CrossRef]
- Warriach, H.M.; McGill, D.; Bush, R.D.; Wynn, P.C. Production and reproduction performance of Nili-Ravi buffaloes under field conditions of Pakistan. J. Anim. Plant Sci. 2012, 22, 121–124. [Google Scholar]
- Devkota, B.; Shah, S.; Gautam, G. Reproduction and fertility of buffaloes in Nepal. Animals 2023, 13, 70. [Google Scholar] [CrossRef]
- Rautela, R.; Kumar, S.; Sharma, R.K.; Phulia, S.K.; Kumar, R.; Singh, M.; Katiyar, R.; Bharadwaj, A.; Datta, T.K. Impact of age at first calving on fertility and production performance in Murrah buffalo. Reprod. Domest. Anim. 2024, 59, e14691. [Google Scholar] [CrossRef]
- Bhat, G.R.; Dhaliwal, G.S. Estrus and ovulation synchrony of buffaloes (bubalus bubalis): A review. Buffalo Bull. 2023, 42, 239–261. [Google Scholar] [CrossRef]
- Verma, R.; Vijayalakshmy, K.; Virmani, M.; Kumar, S.; Verma, A. Seasonal influence of age at first calving on genetic variation and subsequent reproductive performances in Murrah buffaloes. Biol. Rhythm. Res. 2019, 52, 622–635. [Google Scholar] [CrossRef]
- El-Awady1, H.G.; Ibrahim, A.F.; El-Naser, I.A.M.A. The effect of age at first calving on productive life and lifetime profit in lactating Egyptian buffaloes. Buffalo Bull. 2021, 40, 71–85. [Google Scholar]
- Devendra, C. Rainfed areas and animal agriculture in Asia: The wanting agenda for transforming productivity growth and rural poverty. Asian-Australas. J. Anim. Sci. 2012, 25, 122–142. [Google Scholar] [CrossRef] [PubMed]
- Van Eenennaam, A.L.; Silva, F.D.F.; Trott, J.F.; Zilberman, D. Genetic engineering of livestock: The opportunity cost of regulatory delay. Annu. Rev. Anim. Biosci. 2021, 9, 453–478. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Zhang, Y.; Zhou, F.; Yao, Z.; Zhan, Y.; Fan, Z.; Meng, X.; Zhang, Z.; Liu, L.; Yang, J.; et al. Increased accuracy of genomic prediction using preselected SNPs from GWAS with imputed whole-genome sequence data in pigs. Animals 2023, 13, 3871. [Google Scholar] [CrossRef] [PubMed]
- Teng, J.-Y.; Ye, S.-P.; Gao, N.; Chen, Z.-T.; Diao, S.-Q.; Li, X.-J.; Yuan, X.-L.; Zhang, H.; Li, J.-Q.; Zhang, X.-Q.; et al. Incorporating genomic annotation into single-step genomic prediction with imputed whole-genome sequence data. J. Integr. Agric. 2022, 21, 1126–1136. [Google Scholar] [CrossRef]
- Gutierrez-Reinoso, M.A.; Aponte, P.M.; Garcia-Herreros, M. Genomic analysis, progress and future perspectives in dairy cattle selection: A review. Animals 2021, 11, 599. [Google Scholar] [CrossRef]
- Liang, H.; Xuan, J. Feature review on the use of genomic selection in chicken breeding: Current practices and future prospects. Int. J. Mol. Zool. 2024, 14, 334–343. [Google Scholar] [CrossRef]
- Chen, S.Y.; Schenkel, F.S.; Melo, A.L.P.; Oliveira, H.R.; Pedrosa, V.B.; Araujo, A.C.; Melka, M.G.; Brito, L.F. Identifying pleiotropic variants and candidate genes for fertility and reproduction traits in Holstein cattle via association studies based on imputed whole-genome sequence genotypes. BMC Genom. 2022, 23, 331. [Google Scholar] [CrossRef]
- Wang, H.; Wang, X.; Yang, Y.; Zhu, Y.; Wang, S.; Chen, Q.; Yan, D.; Dong, X.; Li, M.; Lu, S. Genome-wide identification of quantitative trait loci and candidate genes for seven carcass traits in a four-way intercross porcine population. BMC Genom. 2024, 25, 582. [Google Scholar] [CrossRef]
- Gowane, G.R.; Vohra, V. Advances in buffalo breeding: A journey from classical breeding to genomic selection. In Biotechnological Applications in Buffalo Research; Chauhan, M.S., Selokar, N., Eds.; Springer: Singapore, 2022. [Google Scholar]
- Saravanan, K.A.; Rajawat, D.; Kumar, H.; Nayak, S.S.; Bhushan, B.; Dutt, T.; Panigrahi, M. Signatures of selection in riverine buffalo populations revealed by genome-wide SNP data. Anim. Biotechnol. 2022, 34, 3343–3354. [Google Scholar] [CrossRef]
- Rehman, S.U.; Hassan, F.-U.; Luo, X.; Li, Z.; Liu, Q. Whole-genome sequencing and characterization of buffalo genetic resources: Recent advances and future challenges. Animals 2021, 11, 904. [Google Scholar] [CrossRef]
- George, L.; Alex, R.; Sukhija, N.; Jaglan, K.; Vohra, V.; Kumar, R.; Verma, A. Genetic improvement of economic traits in Murrah buffalo using significant SNPs from genome-wide association study. Trop. Anim. Health Prod. 2023, 55, 199. [Google Scholar] [CrossRef]
- Mohan Jeena, L.; Kumar, D.; Rahangdale, S.; Singh, A.P.; Sarkhel, B.C. Transcriptional profile of cumulus associated GJA1, PTX3, PRSS35, and SERPINE2 genes with oocytes and embryonic development in water buffalo. Mol. Biol. Rep. 2022, 49, 6285–6293. [Google Scholar] [CrossRef]
- Misztal, I.; Tsuruta, S.; Lourenco, D.; Masuda, Y.; Aguilar, I.; Legarra, A.; Vitezica, Z. Manual for BLUPF90 Family of Programs. 2024. Available online: https://nce.ads.uga.edu/html/projects/programs/docs/blupf90_all8.pdf (accessed on 15 February 2025).
- Wang, H.; Misztal, I.; Aguilar, I.; Legarra, A.; Muir, W.M. Genome-wide association mapping including phenotypes from relatives without genotypes. Genet. Res. 2012, 94, 73–83. [Google Scholar] [CrossRef]
- Aguilar, I.; Misztal, I.; Johnson, D.L.; Legarra, A.; Tsuruta, S.; Lawlor, T.J. Hot topic: A unified approach to utilize phenotypic, full pedigree, and genomic information for genetic evaluation of Holstein final score. J. Dairy Sci. 2010, 93, 743–752. [Google Scholar] [CrossRef]
- VanRaden, P.M. Efficient methods to compute genomic predictions. J. Dairy Sci. 2008, 91, 4414–4423. [Google Scholar] [CrossRef]
- Chen, Y.; Atashi, H.; Grelet, C.; Gengler, N. Weighted single-step genomic best linear unbiased predictor enhances the genomic prediction accuracy for milk citrate predicted by milk mid-infrared spectra of Holstein cows in early lactation. JDS Commun. 2024, 6, 90–94. [Google Scholar] [CrossRef]
- Mrode, R.A. Linear Models for the Prediction of Animal Breeding Values; Cabi: Cambridge, UK, 2014. [Google Scholar]
- Gondro, C.; van der Werf, J.; Hayes, B. Genome-Wide Association Studies and Genomic Prediction; Springer: New York, NY, USA; Heidelberg, Germany; Dordrecht, The Netherlands; London, UK, 2013. [Google Scholar]
- Wang, H.; Misztal, I.; Aguilar, I.; Legarra, A.; Fernando, R.L.; Vitezica, Z.; Okimoto, R.; Wing, T.; Hawken, R.; Muir, W.M. Genome-wide association mapping including phenotypes from relatives without genotypes in a single-step (ssGWAS) for 6-week body weight in broiler chickens. Front. Genet. 2014, 5, 134. [Google Scholar] [CrossRef]
- Aguilar, I.; Misztal, I.; Tsuruta, S.; Legarra, A.; Wang, H. PREGSF90—POSTGSF90: Computational tools for the implementation of Single-step Genomic Selection and Genome-wide as sociation with ungenotyped individuals in BLUPF90 programs. In Proceedings of the 10th World Congress of Genetics Applied to Livestock Production, Vancouver, BC, Canada, 17–22 August 2014; pp. 1–3. [Google Scholar]
- Zhang, Z.; Liu, J.; Ding, X.; Bijma, P.; De Koning, D.-J.; Zhang, Q. Best linear unbiased prediction of genomic breeding values using a trait-specific marker-derived relationship matrix. PLoS ONE 2010, 5, e12648. [Google Scholar] [CrossRef]
- Huang, M.; Liu, X.; Zhou, Y.; Summers, R.M.; Zhang, Z. BLINK: A package for the next level of genome-wide association studies with both individuals and markers in the millions. GigaScience 2019, 8, giy154. [Google Scholar] [CrossRef]
- Cai, Z.; Iso-Touru, T.; Sanchez, M.-P.; Kadri, N.; Bouwman, A.C.; Chitneedi, P.K.; MacLeod, I.M.; Vander Jagt, C.J.; Chamberlain, A.J.; Gredler-Grandl, B.; et al. Meta-analysis of six dairy cattle breeds reveals biologically relevant candidate genes for mastitis resistance. Genet. Sel. Evol. 2024, 56, 54. [Google Scholar] [CrossRef]
- Singh, P. Genetic parameters for first lactation and lifetime traits of Nili Ravi buffaloes. Front. Anim. Sci. 2022, 2, 8242160. [Google Scholar]
- Easa, A.A.; Abd El Aziz, A.H.; El Barbary, A.S.A.; Kostomakhin, N.M.; Nasr, M.A.F.; Imbabi, T.A. Genetic parameters of production and reproduction traits of Egyptian buffaloes under subtropical conditions. Trop. Anim. Health Prod. 2022, 54, 270. [Google Scholar] [CrossRef]
- Amin, M.S.; Salem, M.M.I.; El-Zarkouny, S.Z. Genetic parameters and genetic trends for reproductive traits in Egyptian buffaloes. Anim. Reprod. Sci. 2021, 231, 106800. [Google Scholar] [CrossRef]
- Shao, B.; Sun, H.; Ahmad, M.J.; Ghanem, N.; Abdel Shafy, H.; Du, C.; Deng, T.; Mansoor, S.; Zhou, Y.; Yang, Y.; et al. Genetic features of reproductive traits in bovine and buffalo: Lessons from bovine to buffalo. Front. Genet. 2021, 12, 617128. [Google Scholar] [CrossRef]
- Sehar Zaidi, N.; Anwar, M. Effect of calving season and climatic factors on age at puberty, service period and successful mating in Nili-Ravi buffalo. Buffalo Bull. 2021, 3, 475–484. [Google Scholar]
- Dhakad, R.; Chaudhary, A.P.; Gupta, J.P.; Tiwari, S. Effect of non-genetic factors on reproductive efficiency of Mehsana buffaloes maintained at organized farm. Buffalo Bull. 2024, 2, 173–185. [Google Scholar] [CrossRef]
- Gutierrez, J.P.; Alvarez, I.; Fernandez, I.; Royo, L.J.; Dıez, J.; Goyache, F. Genetic relationships between calving date, calving interval, age at first calving and type traits in beef cattle. Livest. Prod. Sci. 2002, 78, 215–222. [Google Scholar] [CrossRef]
- Vergara, O.D.; Elzo, M.A.; Cerón-Muñoz, M.F. Genetic parameters and genetic trends for age at first calving and calving interval in an Angus-Blanco Orejinegro-Zebu multibreed cattle population in Colombia. Livest. Sci. 2009, 126, 318–322. [Google Scholar] [CrossRef]
- Tamboli, P.; Bharadwaj, A.; Chaurasiya, A.; Bangar, Y.C.; Jerome, A. Association between age at first calving, first lactation traits and lifetime productivity in Murrah buffaloes. Anim. Biosci. 2022, 35, 1151–1161. [Google Scholar] [CrossRef] [PubMed]
- Crowe, M.A.; Diskin, M.G.; Williams, E.J. Parturition to resumption of ovarian cyclicity: Comparative aspects of beef and dairy cows. Animal 2014, 8, 40–53. [Google Scholar] [CrossRef] [PubMed]
- Shah, S.N.H. Prolonged calving intervals in the Nili Ravi buffalo. Ital. J. Anim. Sci. 2007, 6, 694–696. [Google Scholar] [CrossRef]
- Yamboue, A.T.; Zongo, M.; Traore, B.; Karimou, B.H. Dynamics of uterine involution and follicular growth during anoestrus postpartum in African zebu Peulh. Int. J. Biol. Chem. Sci. 2020, 14, 1800–1811. [Google Scholar] [CrossRef]
- Gonçalves dos Santos, J.C.; de Araújo Neto, F.R.; Fernandez, G.S.; de Abreu Santos, D.J.; Pereira Cunha, F.; Aspilcueta-Borquis, R.R.; Tonhati, H. Comparison of single-step methods for genomic prediction of age at first calving in dairy buffaloes. J. Agric. Sci. 2024, 162, 377–383. [Google Scholar] [CrossRef]
- George, L.; Alex, R.; Gowane, G.; Vohra, V.; Joshi, P.; Kumar, R.; Verma, A. Weighted single-step GWAS reveals genomic regions associated with economic traits in Murrah buffaloes. Anim. Biotechnol. 2024, 35, 2319622. [Google Scholar] [CrossRef]
- de Araujo Neto, F.R.; Takada, L.; de Abreu Santos, D.J.; Aspilcueta Borquis, R.R.; Cardoso, D.F.; do Nascimento, A.V.; Silva, R.M.O.; Tonhati, H. Identification of genomic regions related to age at first calving and first calving interval in water buffalo using single step GBLUP. Reprod. Domest. Anim. 2020, 55, 1565–1572. [Google Scholar] [CrossRef]
- Świerzko, A.S.; Cedzyński, M. The influence of the lectin pathway of complement activation on infections of the respiratory system. Front. Immunol. 2020, 11, 585243. [Google Scholar] [CrossRef]
- Ding, M.; Ding, Q.; Liu, Z.; Wang, L.; Pei, K.; Hu, J.; Liao, Y.; Zhang, J.V. TNFRSF11B modified umbilical cord mesenchymal stem cells as a novel strategy for bone related diseases by suppressing osteoclast activity. J. Orthop. Surg. Res. 2025, 20, 478. [Google Scholar] [CrossRef]
- MedlinePlus Genetics. TNFRSF11B Gene: Tumor Necrosis Factor Receptor Superfamily Member 11B; U.S. National Library of Medicine: Bethesda, MD, USA, 2021. Available online: https://www.medlineplus.gov/genetics/gene/tnfrsf11b/ (accessed on 5 July 2025).
- Lee, J.; Mun, H.; Koo, Y.; Park, S.; Kim, J.; Yu, S.; Shin, J.; Lee, J.; Son, J.; Park, C.; et al. Enhancing genomic prediction accuracy for body conformation traits in Korean Holstein cattle. Animals 2024, 14, 1052. [Google Scholar] [CrossRef]
- GeneCards. PDZRN4 Gene: PDZ Domain Containing Ring Finger 4; Weizmann Institute of Science: Rehovot, Israel, 2025; Available online: https://www.genecards.org/cgi-bin/carddisp.pl?gene=PDZRN4 (accessed on 5 July 2025).
- Shi, L.; Zhang, P.; Liu, Q.; Liu, C.; Cheng, L.; Yu, B.; Chen, H. Genome-wide analysis of genetic diversity and selection signatures in Zaobei beef cattle. Animals 2024, 14, 2447. [Google Scholar] [CrossRef]
- Li, J.; Liu, J.; Campanile, G.; Plastow, G.; Zhang, C.; Wang, Z.; Cassandro, M.; Gasparrini, B.; Salzano, A.; Hua, G.; et al. Novel insights into the genetic basis of buffalo reproductive performance using integrated GWAS and RNA seq. BMC Genom. 2018, 19, 814. [Google Scholar] [CrossRef]
- Xu, X.; Jiang, H.; Wang, D.; Rehman, S.U.; Li, Z.; Song, X.; Cui, K.; Luo, X.; Yang, C.; Liu, Q. Exploration of the transcriptional regulation network between buffalo granulosa cells and follicle development using integrated RNA seq and GWAS. BMC Genom. 2024, 25, 10912. [Google Scholar] [CrossRef]
- Macciotta, N.P.P.; Colli, L.; Cesarani, A.; Ajmone Marsan, P.; Low, W.Y.; Tearle, R.; Williams, J.L. The distribution of runs of homozygosity in the genome of river and swamp buffaloes reveals a history of adaptation, migration and crossbred events. Genet. Sel. Evol. 2021, 53, 20. [Google Scholar] [CrossRef]
| Traits | Mean | SD | Min. | Max. | %CV |
|---|---|---|---|---|---|
| AFC (months) | 46.31 | 10.01 | 32.00 | 66.33 | 21.62 |
| CI (days) | 520.87 | 103.68 | 447.64 | 624.55 | 19.91 |
| Methods | ABLUP | WssGBLUP | ||
|---|---|---|---|---|
| Parameters | AFC | CI | AFC | CI |
| 72.27 | 6211.20 | 88.91 | 5223.90 | |
| 130.37 | 114,510.00 | 109.03 | 114,920.00 | |
| 0.36 (0.027) | 0.051 (0.004) | 0.45 (0.026) | 0.043 (0.003) | |
| 0.495 | 0.517 | |||
| 0.467 | 0.473 | |||
| Dataset | CORR | Bias Reduction (%) | Accuracy Increase (%) |
|---|---|---|---|
| AFC | |||
| All animal datasets | 0.984 | −12.939 | 28.342 |
| Bull dataset | 0.991 | −21.538 | 25.858 |
| Dam dataset | 0.999 | −30.140 | 26.606 |
| Top 20% of all animals | 0.968 | −35.334 | 26.076 |
| Top 20% of bulls | 0.989 | −12.950 | 12.481 |
| Top 20% of dams | 0.994 | −2.196 | 25.949 |
| CI | |||
| All animal datasets | 0.984 | −1.339 | 41.070 |
| Bull dataset | 0.991 | −5.945 | 38.224 |
| Dam dataset | 0.999 | −14.627 | 39.196 |
| Top 20% of all animals | 0.977 | −12.105 | 39.571 |
| Top 20% of bulls | 0.996 | −2.533 | 23.576 |
| Top 20% of dams | 0.988 | −7.684 | 39.057 |
| No. | SNP Name | SNP Variance | Chromosome | Location (bp) | Gene | Size (bp) | Distance to SNP (bp) | Putative Function | Associated Traits |
|---|---|---|---|---|---|---|---|---|---|
| 1 | AX-85068166 | 3.653 2.884 | 15 | 36,985,581 | COLEC10 | 46,505 | −5651 | Innate immune response via lectin complement pathway; involved in embryonic development (neural crest cell migration). | AFC, CI |
| LOC112579067 | 62,540 | 20,530 | Likely acts as a regulatory ncRNA involved in gene expression modulation via chromatin interactions, transcriptional regulation, or as a molecular scaffold; may influence developmental and physiological pathways relevant to reproductive traits. | ||||||
| 2 | AX-85072819 | 3.653 2.885 | 15 | 37,187,614 | TRNAC-ACA | 73 | 4031 | Involved in the delivery of amino acid cysteine to the ribosome during protein translation; plays a crucial role in maintaining translational fidelity and efficiency. | AFC, CI |
| MAL2 | 33,379 | 24,124 | Encodes a multispan transmembrane proteolipid involved in lipid raft-mediated transcytosis in polarized epithelial cells; may regulate membrane protein sorting and vesicular transport. | ||||||
| 3 | AX-85075636 | 3.652 2.886 | 15 | 37,227,004 | TRNAC-ACA | 73 | 43,421 | Involved in the delivery of amino acid cysteine to the ribosome during protein translation; plays a crucial role in maintaining translational fidelity and efficiency. | AFC, CI |
| 4 | AX-85129152 | 3.652 2.880 | 15 | 36,861,303 | TNFRSF11B | 28,196 | 9794 | Acts as a secreted decoy receptor for RANKL and TRAIL; inhibits osteoclastogenesis and bone resorption and regulates bone remodeling; modulates NF-κB-mediated immune signaling and may prevent vascular calcification. | AFC, CI |
| LOC112579280 | 135 | 17,407 | Acts as an H/ACA box snoRNA guiding pseudouridylation of rRNA and potentially other RNAs; may regulate RNA processing, ribosome function, and cellular proliferation. | ||||||
| LOC102413604 | 347,952 | 14,148 | Likely functions as a regulatory ncRNA involved in modulation of gene expression via chromatin interactions or transcription regulation; may act as a molecular scaffold and influence developmental and reproductive pathways. | ||||||
| LOC112579067 | 62,540 | −41,209 | Likely acts as a regulatory ncRNA involved in gene expression modulation via chromatin interactions, transcriptional regulation, or as a molecular scaffold; may influence developmental and physiological pathways relevant to reproductive traits. | ||||||
| 5 | AX-85102487 AX-85149742 | 3.651 2.880 | 15 | 36,559,391 | LOC102413604 | 347,952 | On target | Likely functions as a regulatory ncRNA involved in modulation of gene expression through chromatin interactions or transcription regulation; may act as molecular scaffold and influence developmental and reproductive pathways. | AFC, CI |
| 6 | AX-85087929 AX-85133105 | 1.862 3.896 | 3 | 130,355,532 | LOC123332880 | 76,994 | On target | Likely functions as a regulatory lncRNA involved in transcriptional and epigenetic regulation through chromatin remodeling, enhancer–promoter interactions, splicing, or acting as a molecular scaffold; may influence reproductive developmental pathways. | AFC, CI |
| 7 | AX-85170264 | 1.625 | 25 | 135,219,084 | LOC123331878 | 95,019 | On target | Likely functions as a regulatory lncRNA implicated in transcriptional or post-transcriptional regulation via chromatin remodeling, RNA splicing, RNA stabilization, or scaffold formation; potentially influences genes involved in reproductive development and physiological pathways. | AFC |
| 8 | AX-85125727 AX-85182553 | 1.621 | 25 | 135,311,866 | LOC123331703 | 1118 | −6735 | May function as a processed pseudogene derived from MORF4L1; could act as a regulatory RNA influencing expression of the MORF/MRG gene family involved in chromatin remodeling, cell proliferation, senescence, and transcriptional regulation. | AFC |
| LOC112582049 | 2037 | −8970 | Likely acts as a regulatory pseudogene derived from MORF4L1; may produce ncRNAs that serve as sponges for microRNAs or siRNA precursors; modulates expression of MORF/MRG chromatin remodeling complex, influencing cell proliferation, senescence, and transcriptional control. | ||||||
| LOC123331720 | 114,704 | −17,294 | Likely encodes a novel protein with potential enzyme or binding functions; may participate in intracellular signaling or protein complex formation; predicted involvement in developmental or reproductive pathways. | ||||||
| LOC123331753 | 7170 | 17,530 | Likely functions as a regulatory lncRNA involved in transcriptional and post-transcriptional regulation (via chromatin remodeling, splicing, RNA stabilization, or molecular scaffolding); may influence reproductive pathway genes. | ||||||
| LOC123331878 | 95,019 | 38,369 | Likely functions as a regulatory lncRNA implicated in transcriptional or post-transcriptional regulation; may act via chromatin remodeling, RNA splicing, RNA stabilization, or scaffold formation, potentially influencing genes involved in reproductive development and physiological pathways. | ||||||
| 9 | AX-85048696 AX-85138148 | 1.607 | 14 | 76,488,914 | TASP1 | 278,804 | −19,341 | Encodes a conserved threonine endopeptidase that proteolytically cleaves transcriptional regulators such as MLL1/2 and TFIIA, essential for HOX gene expression, embryonic development, hematopoietic stem cell maintenance, craniofacial morphogenesis, and cell cycle progression. | AFC |
| 10 | AX-85123888 | 1.606 | 14 | 76,079,203 | MACROD2 | 2,318,987 | On target | Catalyzes removal of mono-ADP-ribose from proteins, regulating DNA damage repair, chromatin remodeling, and Wnt/β-catenin signaling; implicated in cell proliferation, genome stabilization, and developmental processes. | AFC |
| LOC102396848 | 835 | −49,655 | Likely acts as a regulatory pseudogene derived from SMIM20; may produce RNA transcripts functioning as ceRNA/miRNA sponges or antisense regulators, modulating SMIM20 expression involved in mitochondrial cytochrome c oxidase assembly and reproductive peptide (phoenixin) production influencing ovarian follicle growth and GnRH signaling. | ||||||
| 11 | AX-85116617 | 1.565 | 14 | 75,859,588 | MACROD2 | 2,318,987 | On target | Catalyzes removal of mono-ADP-ribose from proteins, regulating DNA damage repair, chromatin remodeling, and Wnt/β-catenin signaling; implicated in cell proliferation, genome stabilization, and developmental processes. | AFC |
| FLRT3 | 12,568 | On target | Mediates cell–cell adhesion and receptor signaling; modulates FGF-ERK and VEGF-dependent pathways; involved in embryonic morphogenesis, neuronal guidance, angiogenesis, and tissue polarity. | ||||||
| 12 | AX-85072646 | 1.562 | 25 | 135,219,084 | LOC123331878 | 95,019 | On target | Likely functions as a regulatory lncRNA implicated in transcriptional or post-transcriptional regulation; may act through chromatin remodeling, RNA splicing, RNA stabilization, or scaffold formation, potentially influencing genes involved in reproductive development and physiological pathways. | AFC |
| 13 | AX-85089841 | 1.531 | 14 | 75,807,197 | MACROD2 | 2,318,987 | On target | Catalyzes removal of mono-ADP-ribose from proteins, regulating DNA damage repair, chromatin remodeling, and Wnt/β-catenin signaling; implicated in cell proliferation, genome stabilization, and developmental processes. | AFC |
| 14 | AX-85071095 AX-85167220 | 1.518 | 25 | 135,188,395 | LOC123331878 | 95,019 | On target | Likely functions as a regulatory lncRNA implicated in transcriptional or post-transcriptional regulation; may act via chromatin remodeling, RNA splicing, RNA stabilization, or scaffold formation, potentially influencing genes involved in reproductive development and physiological pathways. | AFC |
| 15 | AX-85048217 | 1.308 | 3 | 130,427,805 | LOC123332880 | 76,994 | On target | Likely functions as a regulatory lncRNA involved in transcriptional and epigenetic regulation via chromatin remodeling, enhancer–promoter interactions, splicing, or acting as a molecular scaffold; may influence reproductive developmental pathways. | AFC |
| 16 | AX-85075413 | 1.308 | 3 | 130,503,159 | TRNAF-GAA | 73 | 16,083 | Delivers phenylalanine to the ribosome during protein translation; essential for translational fidelity and efficiency. | AFC |
| 17 | AX-85129182 AX-85187994 | 2.030 2.029 | 4 | 80,876,599 | PDZRN4 | 437,151 | On target | Functions as a tumor suppressor and regulates cell proliferation and protein degradation pathways; involved in neuronal/neural crest development, spermatogenesis, and immune signaling via PDZ domain-mediated scaffold interactions. | CI |
| 18 | AX-85099246 | 2.028 | 4 | 81,019,718 | PDZRN4 | 437,151 | On target | Functions as a tumor suppressor, regulates cell proliferation and protein degradation pathways; involved in neuronal/neural crest development, spermatogenesis, and immune signaling via PDZ domain-mediated scaffold interactions. | CI |
| 19 | AX-85081751 | 2.017 | 4 | 80,809,832 | PDZRN4 | 437,151 | On target | Functions as a tumor suppressor and regulates cell proliferation and protein degradation pathways; involved in neuronal/neural crest development, spermatogenesis, and immune signaling via PDZ domain-mediated scaffold interactions. | CI |
| 20 | AX-85052779 | 1.950 | 4 | 80,872,538 | PDZRN4 | 437,151 | On target | Functions as a tumor suppressor and regulates cell proliferation and protein degradation pathways; involved in neuronal/neural crest development, spermatogenesis, and immune signaling via PDZ domain–mediated scaffold interactions. | CI |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lomngam, R.; Chankitisakul, V.; Duangjinda, M.; Kenchaiwong, W.; Kuha, K.; Sintala, K.; Pothikanit, K.; Boonkum, W. Integrating Genomic Selection and Genome-Wide Association Study to Enhance Reproductive Traits in Thai Swamp Buffalo. Animals 2025, 15, 2333. https://doi.org/10.3390/ani15162333
Lomngam R, Chankitisakul V, Duangjinda M, Kenchaiwong W, Kuha K, Sintala K, Pothikanit K, Boonkum W. Integrating Genomic Selection and Genome-Wide Association Study to Enhance Reproductive Traits in Thai Swamp Buffalo. Animals. 2025; 15(16):2333. https://doi.org/10.3390/ani15162333
Chicago/Turabian StyleLomngam, Rawinan, Vibuntita Chankitisakul, Monchai Duangjinda, Wootichai Kenchaiwong, Kecha Kuha, Kritsanathon Sintala, Kulphat Pothikanit, and Wuttigrai Boonkum. 2025. "Integrating Genomic Selection and Genome-Wide Association Study to Enhance Reproductive Traits in Thai Swamp Buffalo" Animals 15, no. 16: 2333. https://doi.org/10.3390/ani15162333
APA StyleLomngam, R., Chankitisakul, V., Duangjinda, M., Kenchaiwong, W., Kuha, K., Sintala, K., Pothikanit, K., & Boonkum, W. (2025). Integrating Genomic Selection and Genome-Wide Association Study to Enhance Reproductive Traits in Thai Swamp Buffalo. Animals, 15(16), 2333. https://doi.org/10.3390/ani15162333







