mtDNA CR Evidence Indicates High Genetic Diversity of Captive Forest Musk Deer in Shaanxi Province, China
Abstract
:Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Sample Collection
2.2. DNA Extraction and Amplification
2.3. Data Analysis
2.4. Data Acquisition of mtDNA CR
3. Results
3.1. Genetic Diversity
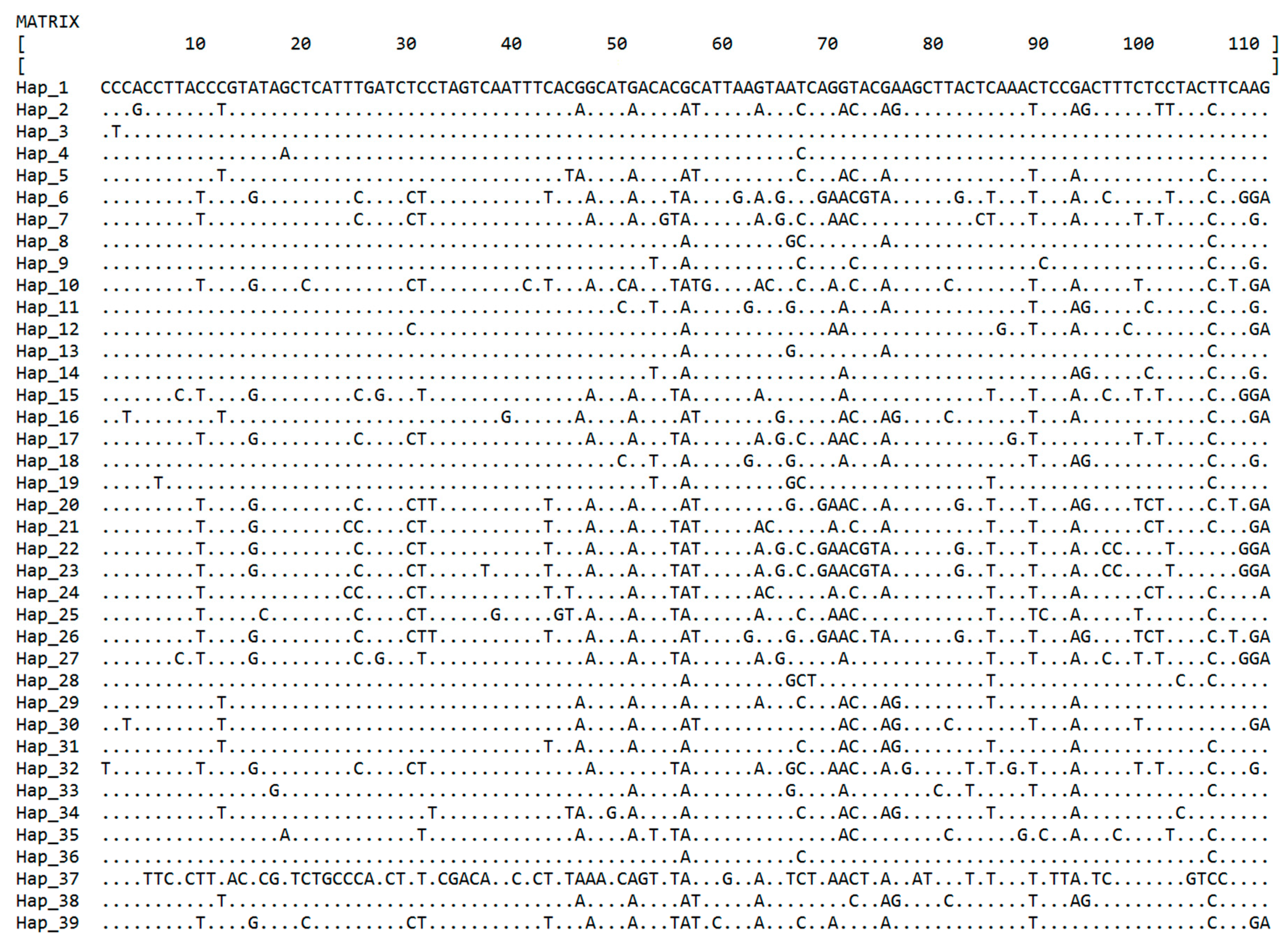
3.2. Haplotype Analyses
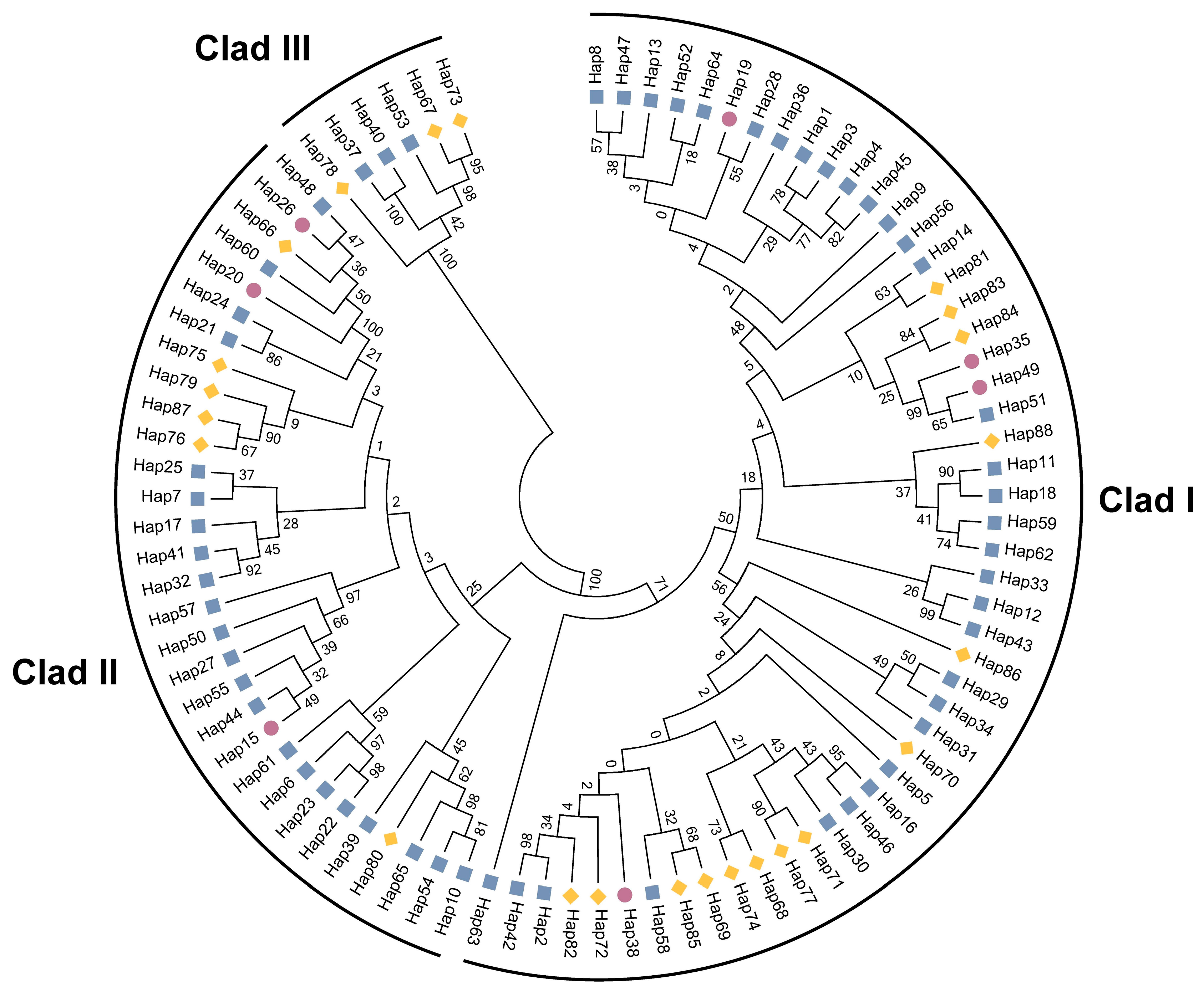
3.3. Genetic Structure
3.4. Demographic History
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Romiguier, J.; Gayral, P.; Ballenghien, M.; Bernard, A.; Cahais, V.; Chenuil, A.; Chiari, Y.; Dernat, R.; Duret, L.; Faivre, N.; et al. Comparative population genomics in animals uncovers the determinants of genetic diversity. Nature 2014, 515, 261–263. [Google Scholar] [CrossRef] [PubMed]
- Pauls, S.U.; Nowak, C.; Balint, M.; Pfenninger, M. The impact of global climate change on genetic diversity within populations and species. Mol. Ecol. 2013, 22, 925–946. [Google Scholar] [CrossRef] [PubMed]
- Leigh, D.M.; Hendry, A.P.; Vazquez-Dominguez, E.; Friesen, V.L. Estimated six per cent loss of genetic variation in wild populations since the industrial revolution. Evol. Appl. 2019, 12, 1505–1512. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Reed, D.H.; Frankham, R. Correlation between fitness and genetic diversity. Conserv. Biol. 2003, 17, 230–237. [Google Scholar] [CrossRef]
- Ellegren, H.; Galtier, N. Determinants of genetic diversity. Nat. Rev. Genet. 2016, 17, 422–433. [Google Scholar] [CrossRef] [Green Version]
- Wu, J.; Wang, W. The Musk Deer of China; The China Forestry Publishing House: Beijing, China, 2006. [Google Scholar]
- Sheng, H.; Liu, Z. The Musk Deer in China; The Shanghai Scientific & Technical Publishers: Shanghai, China, 2007. [Google Scholar]
- Wang, Y.; Harris, R. Moschus berezovskii (Errata Version Published in 2016). The IUCN Red List of Threatened Species 2015: E.T13894A103431781. 2015. Available online: https://dx.doi.org/10.2305/IUCN.UK.2015-4.RLTS.T13894A61976926.en (accessed on 26 March 2023).
- Zheng, C.L.; Zhao, R.H.; Meng, Z.B.; Hu, D.F.; Chen, S.K.; Liu, B.Q.; Wang, J.M. Current situation and future perspectives of musk deer farming in China. Mod. Chin. Med. 2022, 24, 1684–1692. [Google Scholar]
- Frankham, R. Challenges and opportunities of genetic approaches to biological conservation. Biol. Conserv. 2010, 143, 1919–1927. [Google Scholar] [CrossRef]
- Nadeem, M.A.; Nawaz, M.A.; Shahid, M.Q.; Dogan, Y.; Comertpay, G.; Yildiz, M.; Hatipoglu, R.; Ahmad, F.; Alsaleh, A.; Labhane, N.; et al. DNA molecular markers in plant breeding: Current status and recent advancements in genomic selection and genome editing. Biotechnol. Biotechnol. Equip. 2018, 32, 261–285. [Google Scholar] [CrossRef] [Green Version]
- Bruford, M.W.; Bradley, D.G.; Luikart, G. DNA markers reveal the complexity of livestock domestication. Nat. Rev. Genet. 2003, 4, 900–910. [Google Scholar] [CrossRef]
- Galtier, N.; Nabholz, B.; Glemin, S.; Hurst, G.D.D. Mitochondrial DNA as a marker of molecular diversity: A reappraisal. Mol. Ecol. 2009, 18, 4541–4550. [Google Scholar] [CrossRef]
- Allio, R.; Donega, S.; Galtier, N.; Nabholz, B. Large variation in the ratio of mitochondrial to nuclear mutation rate across animals: Implications for genetic diversity and the use of mitochondrial DNA as a molecular marker. Mol. Biol. Evol. 2017, 34, 2762–2772. [Google Scholar] [CrossRef] [Green Version]
- Saccone, C.; Attimonelli, M.; Sbisa, E. Structural elements highly preserved during the evolution of the D-loop-containing region in vertebrate mitochondrial DNA. J. Mol. Evol. 1987, 26, 205–211. [Google Scholar] [CrossRef] [PubMed]
- Sbisa, E.; Tanzariello, F.; Reyes, A.; Pesole, G.; Saccone, C. Mammalian mitochondrial D-loop region structural analysis: Identification of new conserved sequences and their functional and evolutionary implications. Gene 1997, 205, 125–140. [Google Scholar] [CrossRef] [PubMed]
- Alvarez, I.; Fernandez, I.; Lorenzo, L.; Payeras, L.; Cuervo, M.; Goyache, F. Founder and present maternal diversity in two endangered Spanish horse breeds assessed via pedigree and mitochondrial DNA information. J. Anim. Breed. Genet. 2012, 129, 271–279. [Google Scholar] [CrossRef]
- Giontella, A.; Cardinali, I.; Lancioni, H.; Giovannini, S.; Pieramati, C.; Silvestrelli, M.; Sarti, F.M. Mitochondrial DNA survey reveals the lack of accuracy in Maremmano horse Studbook records. Animals 2020, 10, 839. [Google Scholar] [CrossRef] [PubMed]
- Bower, M.A.; Whitten, M.; Nisbet, R.E.R.; Spencer, M.; Dominy, K.M.; Murphy, A.M.; Cassidy, R.; Barrett, E.; Hill, E.W.; Binns, M. Thoroughbred racehorse mitochondrial DNA demonstrates closer than expected links between maternal genetic history and pedigree records. J. Anim. Breed. Genet. 2013, 130, 227–235. [Google Scholar] [CrossRef] [PubMed]
- Tang, J.; Li, J.J.; Wang, B.; Suo, L.J.; Li, F.R.; Liu, W.H.; Wang, Y.; Yan, R.X. Analysis two methods for extracting DNA from musk deer feces. Acta Agric. Boreali-Occident. Sin. 2018, 27, 326–330. [Google Scholar]
- Burland, T.G. DNASTAR’s Lasergene sequence analysis software. Methods Mol. Biol. 2000, 132, 71–91. [Google Scholar]
- Tamura, K.; Peterson, D.; Peterson, N.; Stecher, G.; Nei, M.; Kumar, S. MEGA5: Molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol. Biol. Evol. 2011, 28, 2731–2739. [Google Scholar] [CrossRef] [Green Version]
- Rozas, J.; Ferrer-Mata, A.; Sanchez-DelBarrio, J.C.; Guirao-Rico, S.; Librado, P.; Ramos-Onsins, S.E.; Sanchez-Gracia, A. DnaSP 6: DNA sequence polymorphism analysis of large data sets. Mol. Biol. Evol. 2017, 34, 3299–3302. [Google Scholar] [CrossRef]
- Colwell, R.K. Estimates: Statistical Estimation of Species Richness and Shared Species from Samples. Version 9.1.0. 2019. Available online: https://www.robertkcolwell.org/pages/1407-estimates (accessed on 15 May 2023).
- Leigh, J.W.; Bryant, D. POPART: Full-feature software for haplotype network construction. Methods Ecol. Evol. 2015, 6, 1110–1116. [Google Scholar] [CrossRef]
- Excoffier, L.; Lischer, H.E.L. Arlequin suite ver 3.5: A new series of programs to perform population genetics analyses under Linux and Windows. Mol. Ecol. Resour. 2010, 10, 564–567. [Google Scholar] [CrossRef]
- Liu, G.; Xu, C.Q.; He, L.; Zhao, S.S.; Hu, D.F. Fecal DNA analysis of mitochondrial D-Loop region sequence polymorphism in captive forest musk deer. Proc. Natl. Biogenet. Divers. Summit Forum 2012, 26–27. [Google Scholar]
- Yang, C.; Tang, J.; Bian, K.; Suo, L.J.; Yuan, H.; Wang, Y.; Huang, Y. Next generation sequencing yields the complete mitogenome of captive forest musk deer, Moschus berezovskii (Ruminantia: Moschidae). Mitochondrial DNA Part B-Resour. 2018, 3, 472–473. [Google Scholar] [CrossRef] [Green Version]
- Guo, X.B.; Huang, Z.X.; Zhang, T.X.; Ding, J.H.; Gao, Y.Y.; Fu, Y.J.; Hu, D.F.; Liu, G. Analysis of mtDNA genetic diversity and genetic structure of captive forest musk deer population. J. Beijing For. Univ. 2020, 44, 98–105. [Google Scholar]
- Cai, R.B. Genetic Diversity of MHC Genes and Mitochondrial Sequences in Captive Forest Musk Deer (Moschus berezovskii). Master’s Thesis, Beijing Forestry University, Beijing, China, 2016. [Google Scholar]
- Guo, X.B. Study on Genetic Diversity of Mitochondrial DNA Control Region in Captive Forest Musk Deer Populations in Shaanxi Province. Master’s Thesis, Beijing Forestry University, Beijing, China, 2021. [Google Scholar]
- Peng, H.; Liu, S.; Zou, F.; Zeng, B.; Yue, B. Genetic diversity of captive forest musk deer (Moschus berezovskii) inferred from the mitochondrial DNA control region. Anim. Genet. 2009, 40, 65–72. [Google Scholar] [CrossRef]
- Shan, L.; Hu, Y.B.; Zhu, L.F.; Yan, L.; Wang, C.D.; Li, D.S.; Jin, X.L.; Zhang, C.L.; Wei, F.W. Large-scale genetic survey provides insights into the captive management and reintroduction of giant pandas. Mol. Biol. Evol. 2014, 31, 2663–2671. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Frankham, R. Genetic adaptation to captivity in species conservation programs. Mol. Ecol. 2008, 17, 325–333. [Google Scholar] [CrossRef]
- Liu, G.; Shafer, A.B.A.; Zimmermann, W.; Hu, D.F.; Wang, W.T.; Chu, H.J.; Cao, J.; Zhao, C.X. Evaluating the reintroduction project of Przewalski’s horse in China using genetic and pedigree data. Biol. Conserv. 2014, 171, 288–298. [Google Scholar] [CrossRef]
- Bar-David, S.; Saltz, D.; Dayan, T. Predicting the spatial dynamics of a reintroduced population: The Persian fallow deer. Ecol. Appl. 2005, 15, 1833–1846. [Google Scholar] [CrossRef]
- Zhang, Y.Y.; Bai, J.D.; Zhu, A.N.; Chen, R.S.; Xue, D.Y.; Zhong, Z.Y.; Cheng, Z.B. Reversing extinction in China’s Pere David’s deer. Science 2021, 371, 685. [Google Scholar] [CrossRef]
- Groeneveld, L.F.; Lenstra, J.A.; Eding, H.; Toro, M.A.; Scherf, B.; Pilling, D.; Negrini, R.; Finlay, E.K.; Jianlin, H.; Groeneveld, E.; et al. Genetic diversity in farm animals—A review. Anim. Genet. 2010, 41, 6–31. [Google Scholar] [CrossRef] [Green Version]
- DeWoody, J.A.; Harder, A.M.; Mathur, S.; Willoughby, J.R. The long-standing significance of genetic diversity in conservation. Mol. Ecol. 2021, 30, 4147–4154. [Google Scholar] [CrossRef]
- Hu, Y.B.; Fan, H.Z.; Chen, Y.H.; Chang, J.; Zhan, X.J.; Wu, H.; Zhang, B.W.; Wang, M.; Zhang, W.Y.; Yang, L.; et al. Spatial patterns and conservation of genetic and phylogenetic diversity of wildlife in China. Sci. Adv. 2021, 7, 9. [Google Scholar] [CrossRef] [PubMed]
- Feng, H.; Feng, C.L.; Huang, Y.; Tang, J. Structure of mitochondrial DNA control region and genetic diversity of Moschus berezovskii populations in Shaanxi Province. Genet. Mol. Res. 2016, 15, 11. [Google Scholar] [CrossRef] [PubMed]
- Hu, D.M.; Hou, Z.Z.; Zheng, C.M.; Chen, X.; Wu, J.; Chen, L.; Chen, Q.; Yue, B.S.; Zhang, X.Y. A Genetic diversity study based on microsatellite and mitochondria of forest musk deer in the Baishuihe National Nature Reserve, Sichuan. Sichuan J. Zool. 2021, 40, 641–648. [Google Scholar]
- Was, G.; Bowen, B.W. Shallow population histories in deep evolutionary lineages of marine fishes: Insights from sardines and anchovies and lessons for conservation. J. Hered. 1998, 89, 415–426. [Google Scholar]
- Cacic, M.; Cubric-Curik, V.; Ristov, S.; Curik, I. Computational approach to utilisation of mitochondrial DNA in the verification of complex pedigree errors. Livest. Sci. 2014, 169, 42–47. [Google Scholar] [CrossRef]
- Zhao, L.; Zhang, J.; Liu, Z.J.; Funk, S.M.; Wei, F.W.; Xu, M.Q.; Li, M. Complex population genetic and demographic history of the Salangid, Neosalanx taihuensis, based on cytochrome b sequences. BMC Evol. Biol. 2008, 8, 18. [Google Scholar] [CrossRef] [Green Version]
- Liu, G.; Zhang, B.F.; Chang, J.; Hu, X.L.; Li, C.; Xu, T.T.; Liu, S.Q.; Hu, D.F. Population genomics reveals moderate genetic differentiation between populations of endangered forest musk deer located in Shaanxi and Sichuan. BMC Genom. 2022, 23, 11. [Google Scholar] [CrossRef] [PubMed]
- Tajima, F. Statistical method for testing the neutral mutation hypothesis by DNA polymorphism. Genetics 1989, 123, 585–595. [Google Scholar] [CrossRef] [PubMed]
- Wang, W.X.; He, L.; Liu, S.Q.; Wronski, T.; Hu, D.F. Behavioral and physiological responses of forest musk deer (Moschus berezovskii) to experimental fawn manipulation. Acta Ethol. 2016, 19, 133–141. [Google Scholar] [CrossRef]

| Population | Sample Size (n) | Number of Variable Sites | Number of Haplotypes (H) | Nucleotide Diversity (Pi) | Haplotype Diversity (Hd) | Average Number of Nucleotide Differences (k) |
|---|---|---|---|---|---|---|
| R | 35 | 54 | 10 | 0.02583 | 0.892 | 15.499 |
| FM | 46 | 47 | 9 | 0.01925 | 0.794 | 11.553 |
| PZH | 44 | 70 | 24 | 0.02686 | 0.959 | 18.462 |
| HX | 118 | 61 | 14 | 0.02794 | 0.764 | 16.763 |
| LD | 39 | 56 | 14 | 0.01816 | 0.854 | 10.880 |
| HD | 30 | 84 | 7 | 0.04385 | 0.811 | 26.223 |
| GL | 26 | 51 | 12 | 0.01901 | 0.886 | 11.403 |
| Total | 338 | 111 | 39 | 0.02887 | 0.908 | 17.236 |
| Population | R | FM | PZH | HX | LD | HD | GL | Published | Total | ||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Frequency | |||||||||||
| Haplotype | |||||||||||
| Hap1 | 5 | 5 | 3 | 1 | 11 | 25 | |||||
| Hap2 | 6 | 4 | 6 | 2 | 6 | 5 | 1 | 8 | 38 | ||
| Hap3 | 5 | 15 | 2 | 5 | 9 | 2 | 2 | 29 | 69 | ||
| Hap4 | 6 | 1 | 1 | 11 | 3 | 13 | 35 | ||||
| Hap5 | 1 | 1 | |||||||||
| Hap6 | 2 | 5 | 12 | 19 | |||||||
| Hap7 | 1 | 2 | 3 | ||||||||
| Hap8 | 3 | 2 | 49 | 2 | 11 | 8 | 17 | 92 | |||
| Hap9 | 1 | 3 | 2 | 1 | 2 | 2 | 1 | 12 | |||
| Hap10 | 5 | 3 | 3 | 3 | 7 | 21 | |||||
| Hap11 | 1 | 1 | |||||||||
| Hap12 | 14 | 3 | 1 | 1 | 3 | 1 | 2 | 25 | |||
| Hap13 | 2 | 2 | |||||||||
| Hap14 | 2 | 1 | 2 | 5 | |||||||
| Hap15 | 4 | 13 | 17 | ||||||||
| Hap16 | 1 | 1 | 1 | 9 | 12 | ||||||
| Hap17 | 1 | 1 | |||||||||
| Hap18 | 1 | 1 | 2 | ||||||||
| Hap19 | 2 | 2 | 5 | 9 | |||||||
| Hap20 | 1 | 1 | 1 | 3 | |||||||
| Hap21 | 3 | 2 | 5 | ||||||||
| Hap22 | 1 | 1 | 2 | ||||||||
| Hap23 | 1 | 28 | 5 | 34 | |||||||
| Hap24 | 1 | 1 | 1 | 3 | 6 | ||||||
| Hap25 | 1 | 1 | 2 | ||||||||
| Hap26 | 2 | 2 | 4 | ||||||||
| Hap27 | 1 | 1 | |||||||||
| Hap28 | 1 | 1 | |||||||||
| Hap29 | 1 | 4 | 3 | 14 | 22 | ||||||
| Hap30 | 1 | 1 | 2 | ||||||||
| Hap31 | 4 | 4 | 8 | ||||||||
| Hap32 | 8 | 1 | 9 | ||||||||
| Hap33 | 4 | 1 | 2 | 7 | |||||||
| Hap34 | 1 | 1 | |||||||||
| Hap35 | 1 | 1 | 2 | ||||||||
| Hap36 | 1 | 1 | |||||||||
| Hap37 | 5 | 2 | 7 | ||||||||
| Hap38 | 1 | 1 | 2 | ||||||||
| Hap39 | 1 | 1 | 2 | ||||||||
| Hap40 | 5 | 5 | |||||||||
| Hap41 | 1 | 1 | |||||||||
| Hap42 | 1 | 1 | |||||||||
| Hap43 | 1 | 1 | |||||||||
| Hap44 | 1 | 1 | |||||||||
| Hap45 | 1 | 1 | |||||||||
| Hap46 | 2 | 2 | |||||||||
| Hap47 | 1 | 1 | |||||||||
| Hap48 | 2 | 2 | |||||||||
| Hap49 | 8 | 8 | |||||||||
| Hap50 | 1 | 1 | |||||||||
| Hap51 | 1 | 1 | |||||||||
| Hap52 | 1 | 1 | |||||||||
| Hap53 | 1 | 1 | |||||||||
| Hap54 | 1 | 1 | |||||||||
| Hap55 | 1 | 1 | |||||||||
| Hap56 | 2 | 2 | |||||||||
| Hap57 | 1 | 1 | |||||||||
| Hap58 | 1 | 1 | |||||||||
| Hap59 | 2 | 2 | |||||||||
| Hap60 | 1 | 1 | |||||||||
| Hap61 | 1 | 1 | |||||||||
| Hap62 | 1 | 1 | |||||||||
| Hap63 | 1 | 1 | |||||||||
| Hap64 | 1 | 1 | |||||||||
| Hap65 | 3 | 3 | |||||||||
| Population | R | FM | PZH | HX | LD | HD | GL | Shaanxi | Sichuan |
|---|---|---|---|---|---|---|---|---|---|
| R | 0.027 | ||||||||
| FM | 0.025 | 0.020 | |||||||
| PZH | 0.030 | 0.028 | 0.032 | ||||||
| HX | 0.031 | 0.030 | 0.032 | 0.029 | |||||
| LD | 0.023 | 0.021 | 0.029 | 0.029 | 0.019 | ||||
| HD | 0.040 | 0.039 | 0.042 | 0.042 | 0.038 | 0.047 | |||
| GL | 0.024 | 0.022 | 0.028 | 0.027 | 0.020 | 0.036 | 0.020 | ||
| Shaanxi | 0.032 | ||||||||
| Sichuan | 0.045 | 0.050 |
| Population | R | FM | PZH | HX | LD | HD | GL | Shaanxi | Sichuan |
|---|---|---|---|---|---|---|---|---|---|
| R | 0.00000 | 0.12613 ±0.0309 | 0.00000 | 0.08108 ±0.0286 | 0.01802 ±0.0182 | 0.12613 ±0.0364 | |||
| FM | 0.08076 | 0.00000 | 0.00000 | 0.00000 | 0.00000 | 0.00000 | |||
| PZH | 0.02023 | 0.08415 | 0.00000 | 0.00000 | 0.00000 | 0.00901 ±0.0091 | |||
| HX | 0.07689 | 0.16246 | 0.05156 | 0.00000 | 0.00000 | 0.00000 | |||
| LD | 0.02444 | 0.08726 | 0.10643 | 0.15678 | 0.00000 | 0.02703 ±0.0194 | |||
| HD | 0.06420 | 0.14706 | 0.05357 | 0.09365 | 0.13224 | 0.02703 ±0.0194 | |||
| GL | 0.02497 | 0.10263 | 0.06311 | 0.08097 | 0.03319 | 0.06902 | |||
| Shaanxi | 0.00000 | ||||||||
| Sichuan | 0.10404 |
| Source of Variation | Degrees of Freedom | Sum of Squares | Variance Components | Percentage of Variation |
|---|---|---|---|---|
| Among populations | 6 | 273.781 | 0.83116 Va | 9.47 |
| Within populations | 331 | 2630.452 | 7.94699 Vb | 90.53 |
| Total | 337 | 2904.234 | 8.77814 | |
| Fixation Index | 0.09468 (++) |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wang, Z.; Lu, G.; Gao, Y.; Yan, L.; Li, M.; Hu, D.; Zhang, D. mtDNA CR Evidence Indicates High Genetic Diversity of Captive Forest Musk Deer in Shaanxi Province, China. Animals 2023, 13, 2191. https://doi.org/10.3390/ani13132191
Wang Z, Lu G, Gao Y, Yan L, Li M, Hu D, Zhang D. mtDNA CR Evidence Indicates High Genetic Diversity of Captive Forest Musk Deer in Shaanxi Province, China. Animals. 2023; 13(13):2191. https://doi.org/10.3390/ani13132191
Chicago/Turabian StyleWang, Zhe, Guanjie Lu, Yunyun Gao, Liping Yan, Mingzhe Li, Defu Hu, and Dong Zhang. 2023. "mtDNA CR Evidence Indicates High Genetic Diversity of Captive Forest Musk Deer in Shaanxi Province, China" Animals 13, no. 13: 2191. https://doi.org/10.3390/ani13132191






