Calcium/Calmodulin-Dependent Serine Protein Kinase (CASK) Gene Polymorphisms in Pigeons
Abstract
:Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Animals
2.2. DNA Analyses
2.3. Statistical Analysis
3. Results
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Borg, J.P.; Straight, S.W.; Kaech, S.M.; de Taddéo-Borg, M.; Kroon, D.E.; Karnak, D.; Turner, R.S.; Kim, S.K.; Margolis, B. Identification of an evolutionarily conserved heterotrimeric protein complex involved in protein targeting. J. Biol. Chem. 1998, 273, 31633–31636. [Google Scholar] [CrossRef] [Green Version]
- Maximov, A.; Südhof, T.C.; Bezprozvanny, I. Association of neuronal calcium channels with modular adaptor proteins. J. Biol. Chem. 1999, 274, 24453–24456. [Google Scholar] [CrossRef] [Green Version]
- Hsueh, Y.P.; Wang, T.F.; Yang, F.C.; Sheng, M. Nuclear translocation and transcription regulation by the membrane-associated guanylate kinase CASK/LIN-2. Nature 2000, 404, 298–302. [Google Scholar] [CrossRef] [PubMed]
- Hata, Y.; Butz, S.; Südhof, T.C. CASK: A novel dlg/PSD95 homolog with an N-terminal calmodulin-dependent protein kinase domain identified by interaction with neurexins. J. Neurosci. 1996, 16, 2488–2494. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Butz, S.; Okamoto, M.; Südhof, T.C. A tripartite protein complex with the potential to couple synaptic vesicle exocytosis to cell adhesion in brain. Cell 1998, 94, 773–782. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Laverty, H.G.; Wilson, J.B. Murine CASK is disrupted in a sex-linked cleft palate mouse mutant. Genomics 1998, 53, 29–41. [Google Scholar] [CrossRef]
- Atasoy, D.; Schoch, S.; Ho, A.; Nadasy, K.A.; Liu, X.; Zhang, W.; Mukherjee, K.; Nosyreva, E.D.; Fernandez-Chacon, R.; Missler, M.; et al. Deletion of CASK in mice is lethal and impairs synaptic function. Proc. Natl. Acad. Sci. USA 2007, 104, 2525–2530. [Google Scholar] [CrossRef] [Green Version]
- Sanford, J.L.; Mays, T.A.; Rafael-Fortney, J.A. CASK and Dlg form a PDZ protein complex at the mammalian neuromuscular junction. Muscle Nerve 2004, 30, 164–171. [Google Scholar] [CrossRef]
- Gardner, K.L.; Sanford, J.L.; Mays, T.A.; Rafael-Fortney, J.A. CASK localizes to nuclei in developing skeletal muscle and motor neuron culture models and is agrin-independent. J. Cell. Physiol. 2006, 206, 196–202. [Google Scholar] [CrossRef]
- Oliva, C.; Escobedo, P.; Astorga, C.; Molina, C.; Sierralta, J. Role of the MAGUK protein family in synapse formation and function. Dev. Neurobiol. 2012, 72, 57–72. [Google Scholar] [CrossRef] [PubMed]
- Deschenes, M.R.; Covault, J.; Kraemer, W.J.; Maresh, C.M. The neuromuscular junction. Muscle fibre type differences, plasticity and adaptability to increased and decreased activity. Sports Med. 1994, 17, 358–372. [Google Scholar] [CrossRef] [PubMed]
- Jerolmack, C. Animal archeology: Domestic pigeons and the nature-culture dialectic. Qual. Sociol. Rev. 2007, 3, 74–95. [Google Scholar] [CrossRef]
- New Kim: Racing pigeon from Belgium sold for record €1.6m. Available online: https://www.bbc.com/news/world-europe-54953594 (accessed on 15 May 2023).
- Scullion, F.T.; Scullion, M.G. Profiling Flight Performance of Young Racing Pigeons (Columba livia) in Training. J. Vet. Healthc. 2018, 1, 1–19. [Google Scholar] [CrossRef] [Green Version]
- Wiltschko, R.; Wiltschko, W. Considerations on the role of olfactory input in avian navigation. J. Exp. Biol. 2017, 220, 4347–4350. [Google Scholar] [CrossRef] [Green Version]
- Fleissner, G.; Holtkamp-Rötzler, E.; Hanzlik, M.; Winklhofer, M.; Fleissner, G.; Petersen, N.; Wiltschko, W. Ultrastructural analysis of a putative magnetoreceptor in the beak of homing pigeons. J. Comp. Neurol. 2003, 458, 350–360. [Google Scholar] [CrossRef]
- Mora, C.V.; Davison, M.; Wild, J.M.; Walker, M.M. Magnetoreception and its trigeminal mediation in the homing pigeon. Nature 2004, 432, 508–511. [Google Scholar] [CrossRef]
- Lauwers, M.; Pichler, P.; Edelman, N.B.; Resch, G.P.; Ushakova, L.; Salzer, M.C.; Heyers, D.; Saunders, M.; Shaw, J.; Keays, D.A. An iron-rich organelle in the cuticular plate of avian hair cells. Curr. Biol. 2013, 23, 924–929. [Google Scholar] [CrossRef] [Green Version]
- Treiber, C.D.; Salzer, M.; Breuss, M.; Ushakova, L.; Lauwers, M.; Edelman, N.; Keays, D.A. High resolution anatomical mapping confirms the absence of a magnetic sense system in the rostral upper beak of pigeons. Commun. Integr. Biol. 2013, 6, e24859. [Google Scholar] [CrossRef] [Green Version]
- Malkemper, E.P.; Kagerbauer, D.; Ushakova, L.; Nimpf, S.; Pichler, P.; Treiber, C.D.; de Jonge, M.; Shaw, J.; Keays, D.A. No evidence for a magnetite-based magnetoreceptor in the lagena of pigeons. Curr. Biol. 2019, 29, R14–R15. [Google Scholar] [CrossRef] [Green Version]
- Nimpf, S.; Nordmann, G.C.; Kagerbauer, D.; Malkemper, E.P.; Landler, L.; Papadaki-Anastasopoulou, A.; Ushakova, L.; Wenninger-Weinzierl, A.; Novatchkova, M.; Vincent, P.; et al. A Putative Mechanism for Magnetoreception by Electromagnetic Induction in the Pigeon Inner Ear. Curr. Biol. 2019, 29, 4052–4059.e4. [Google Scholar] [CrossRef] [Green Version]
- Papi, F.; Fiore, L.; Fiaschi, V.; Benvenuti, S. Olfaction and homing in pigeons. Monit. Zool. Ital. 1972, 6, 85–95. [Google Scholar] [CrossRef]
- Gagliardo, A. Forty years of olfactory navigation in birds. J. Exp. Biol. 2013, 216, 2165–2171. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jorge, P.E.; Marques, P.A.; Phillips, J.B. Activational effects of odours on avian navigation. Proc. Biol. Sci. 2010, 277, 45–49. [Google Scholar] [CrossRef] [Green Version]
- Anjum, R.; Ayoubian, H.; Schmitz, F. Differential synaptic distribution of the scaffold proteins Cask and Caskin1 in the bovine retina. Mol. Cell. Neurosci. 2014, 62, 19–29. [Google Scholar] [CrossRef] [PubMed]
- Mukherjee, K.; Patel, P.A.; Rajan, D.S.; LaConte, L.E.W.; Srivastava, S. Survival of a male patient harboring CASK Arg27Ter mutation to adolescence. Mol. Genet. Genom. Med. 2020, 8, e1426. [Google Scholar] [CrossRef]
- Patel, P.A.; Hegert, J.V.; Cristian, I.; Kerr, A.; LaConte, L.E.W.; Fox, M.A.; Srivastava, S.; Mukherjee, K. Complete loss of the X-linked gene CASK causes severe cerebellar degeneration. J. Med. Genet. 2022, 59, 1044–1057. [Google Scholar] [CrossRef] [PubMed]
- LaConte, L.E.W.; Chavan, V.; DeLuca, S.; Rubin, K.; Malc, J.; Berry, S.; Gail Summers, C.; Mukherjee, K. An N-terminal heterozygous missense CASK mutation is associated with microcephaly and bilateral retinal dystrophy plus optic nerve atrophy. Am. J. Med. Genet. A 2019, 179, 94–103. [Google Scholar] [CrossRef]
- Saavedra, M.V.; Smalla, K.H.; Thomas, U.; Sandoval, S.; Olavarria, K.; Castillo, K.; Delgado, M.G.; Delgado, R.; Gundelfinger, E.D.; Bacigalupo, J.; et al. Scaffolding proteins in highly purified rat olfactory cilia membranes. Neuroreport 2008, 19, 1123–1126. [Google Scholar] [CrossRef]
- Dybus, A.; Pijanka, J.; Cheng, Y.H.; Sheen, F.; Grzesiak, W.; Muszyńska, M. Polymorphism within the LDHA gene in the homing and non-homing pigeons. J. Appl. Genet. 2006, 47, 63–66. [Google Scholar] [CrossRef]
- Proskura, W.S.; Cichoń, D.; Grzesiak, W.; Zaborski, D.; Sell-Kubiak, E.; Cheng, Y.-H.; Dybus, A. Single nucleotide polymorphism in the LDHA gene as a potential marker for the racing performance of pigeons. J. Poult. Sci. 2014, 51, 364–368. [Google Scholar] [CrossRef] [Green Version]
- Ramadan, S.; Miyake, T.; Yamaura, J.; Inoue-Murayama, M. LDHA gene is associated with pigeon survivability during racing competitions. PLoS ONE 2018, 13, e0195121. [Google Scholar] [CrossRef] [Green Version]
- Ramadan, S.; Yamaura, J.; Miyake, T.; Inoue-Murayama, M. DNA polymorphism within LDH-A gene in pigeon (Columba livia). J. Poult. Sci. 2013, 50, 194–197. [Google Scholar] [CrossRef] [Green Version]
- Proskura, W.S.; Kustosz, J.; Dybus, A.; Lanckriet, R. Polymorphism in dopamine receptor D4 gene is associated with pigeon racing performance. Anim. Genet. 2015, 46, 586–587. [Google Scholar] [CrossRef] [PubMed]
- Kolvenbag, G.; Scott, M.; de Kloet, A.; de Kloet, E. Prospective study relating genotype profiles with race performance in racing pigeons. J. Appl. Genet. 2022, 63, 563–570. [Google Scholar] [CrossRef] [PubMed]
- Proskura, W.; Łukaszewicz, A.; Dzierzba, E.; Cichoń, D.; Zaborski, D.; Grzesiak, W.; Dybus, A. The Cys83Gly amino acid substitution in feather keratin is associated with pigeon performance in long-distance races. Vet. Med. (Praha) 2017, 62, 221–225. [Google Scholar] [CrossRef] [Green Version]
- Haase, E.; Dybus, A.; Konieczna, A.; Kovalev, A.; Gorb, S. Effects of a FCBP gene polymorphism, location, and sex on Young’s modulus of the tenth primary feather in racing pigeons. Sci. Rep. 2022, 12, 1785. [Google Scholar] [CrossRef] [PubMed]
- Dybus, A.; Kulig, H.; Yu, Y.H.; Lanckriet, R.; Proskura, W.; Cheng, Y.H. CRY1 Gene Polymorphism and Racing Performance of Homing Pigeons. Animals 2021, 11, 2632. [Google Scholar] [CrossRef] [PubMed]
- Dybus, A.; Proskura, W.S.; Sadkowski, S.; Pawlina, E. A single nucleotide polymorphism in exon 3 of the myostatin gene in different breeds of domestic pigeon (Columba livia var. domestica). Vet. Med. (Praha) 2018, 58, 32–38. [Google Scholar] [CrossRef] [Green Version]
- Shao, Y.; Tian, H.Y.; Zhang, J.J.; Kharrati-Koopaee, H.; Guo, X.; Zhuang, X.L.; Li, M.L.; Nanaie, H.A.; Dehghani Tafti, E.; Shojaei, B.; et al. Genomic and Phenotypic Analyses Reveal Mechanisms Underlying Homing Ability in Pigeon. Mol. Biol. Evol. 2020, 37, 134–148. [Google Scholar] [CrossRef]
- Stefaniuk-Szmukier, M.; Piórkowska, K.; Ropka-Molik, K. Current research directions in the genetic basis of flight performance in racing pigeons. Rocz. Nauk. Zoot. 2022, 49, 117–126. [Google Scholar]
- Gazda, M.A.; Andrade, P.; Afonso, S.; Dilyte, J.; Archer, J.P.; Lopes, R.J.; Faria, R.; Carneiro, M. Signatures of Selection on Standing Genetic Variation Underlie Athletic and Navigational Performance in Racing Pigeons. Mol. Biol. Evol. 2018, 35, 1176–1189. [Google Scholar] [CrossRef] [Green Version]
- Untergasser, A.; Cutcutache, I.; Koressaar, T.; Ye, J.; Faircloth, B.C.; Remm, M.; Rozen, S.G. Primer3—New capabilities and interfaces. Nucleic Acids Res. 2012, 40, e115. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Technelysium DNA Sequencing Software. Available online: https://technelysium.com.au/wp/chromas/ (accessed on 12 May 2023).
- Bińkowski, J.; Miks, S. Gene-Calc [Computer Software]. Available online: https://www.gene-calc.pl/ (accessed on 20 September 2018).
- Dario, C.; Dario, M.; Ciotola, F.; Peretti, V.; Carnicella, D.; Selvaggi, M. Analysis of STAT5A/AvaI Gene Polymorphism in Four Italian Cattle Breeds. Biochem. Genet. 2009, 47, 671–679. [Google Scholar] [CrossRef] [PubMed]
- Abdi, H. Bonferroni and Šidák corrections for multiple comparisons. In Encyclopedia of Measurement and Statistics; Sage: Thousand Oaks, CA, USA, 2007; pp. 103–107. [Google Scholar]
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2023; Available online: https://www.R-project.org/ (accessed on 12 May 2023).
- Stefaniuk-Szmukier, M.; Piórkowska, K.; Ropka-Molik, K.; Dybus, A. Molecular characterization and occurrence of variation within the promoter region of CASK gene in racing pigeons. In Proceedings of the 38th International Society for Animal Genetics Conference, Virtual Conference 2021, Abstracts, International Society for Animal Genetics, Virtually, 26–30 July 2021; pp. 36–37. [Google Scholar]
- Rosenberg, M.S.; Subramanian, S.; Kumar, S. Patterns of transitional mutation biases within and among mammalian genomes. Mol. Biol. Evol. 2003, 20, 988–993. [Google Scholar] [CrossRef] [PubMed]
- Mukherjee, K.; Sharma, M.; Urlaub, H.; Bourenkov, G.P.; Jahn, R.; Südhof, T.C.; Wahl, M.C. CASK Functions as a Mg2+-independent neurexin kinase. Cell 2008, 133, 328–339. [Google Scholar] [CrossRef] [Green Version]
- Tibbe, D.; Ferle, P.; Krisp, C.; Nampoothiri, S.; Mirzaa, G.; Assaf, M.; Parikh, S.; Kutsche, K.; Kreienkamp, H.J. Regulation of Liprin-α phase separation by CASK is disrupted by a mutation in its CaM kinase domain. Life Sci. Alliance 2022, 5, e202201512. [Google Scholar] [CrossRef]
- De Klerk, E.; AC’t Hoen, P. Alternative mRNA transcription, processing, and translation: Insights from RNA sequencing. Trends Genet. 2015, 31, 128–139. [Google Scholar] [CrossRef]
- Kozak, M. Point mutations define a sequence flanking the AUG initiator codon that modulates translation by eukaryotic ribosomes. Cell 1986, 44, 283–292. [Google Scholar] [CrossRef]
- Xu, H.; Wang, P.; You, J.; Zheng, Y.; Fu, Y.; Tang, Q.; Zhou, L.; Wei, Z.; Lin, B.; Shu, Y.; et al. Screening of Kozak-motif-located SNPs and analysis of their association with human diseases. Biochem. Biophys. Res. Commun. 2010, 392, 89–94. [Google Scholar] [CrossRef] [PubMed]
- He, Y.; Maier, K.; Leppert, J.; Hausser, I.; Schwieger-Briel, A.; Weibel, L.; Theiler, M.; Kiritsi, D.; Busch, H.; Boerries, M.; et al. Monoallelic Mutations in the Translation Initiation Codon of KLHL24 Cause Skin Fragility. Am. J. Hum. Genet. 2016, 99, 1395–1404. [Google Scholar] [CrossRef] [Green Version]
- Hawley, J.A.; Hargreaves, M.; Zierath, J.R. Signalling mechanisms in skeletal muscle: Role in substrate selection and muscle adaptation. Essays Biochem. 2006, 42, 1–12. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kuo, I.Y.; Ehrlich, B.E. Signaling in muscle contraction. Cold Spring Harb. Perspect. Biol. 2015, 7, a006023. [Google Scholar] [CrossRef] [PubMed]
- Chin, E.R. Role of Ca2+/calmodulin-dependent kinases in skeletal muscle plasticity. J. Appl. Physiol. 2005, 99, 414–423. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yang, C.F.; Tsai, W.C. Calmodulin: The switch button of calcium signaling. Tzu Chi Med. J. 2021, 34, 15–22. [Google Scholar] [CrossRef]
- Jensen, H.H.; Brohus, M.; Nyegaard, M.; Overgaard, M.T. Human Calmodulin Mutations. Front. Mol. Neurosci. 2018, 11, 396. [Google Scholar] [CrossRef] [Green Version]
- Xia, Z.; Storm, D.R. The role of calmodulin as a signal integrator for synaptic plasticity. Nat. Rev. Neurosci. 2005, 6, 267–276. [Google Scholar] [CrossRef]
- Seeger, C.; Talibov, V.O.; Danielson, U.H. Biophysical analysis of the dynamics of calmodulin interactions with neurogranin and Ca2+ /calmodulin-dependent kinase II. J. Mol. Recognit. 2017, 30, e2621. [Google Scholar] [CrossRef] [Green Version]
- Swulius, M.T.; Waxham, M.N. Ca2+/calmodulin-dependent protein kinases. Cell. Mol. Life Sci. 2008, 65, 2637–2657. [Google Scholar] [CrossRef] [Green Version]
- Ye, F.; Zeng, M.; Zhang, M. Mechanisms of MAGUK-mediated cellular junctional complex organization. Curr. Opin. Struct. Biol. 2018, 48, 6–15. [Google Scholar] [CrossRef]
- Won, S.; Levy, J.M.; Nicoll, R.A.; Roche, K.W. MAGUKs: Multifaceted synaptic organizers. Curr. Opin. Neurobiol. 2017, 43, 94–101. [Google Scholar] [CrossRef] [Green Version]
- Zhu, J.; Shang, Y.; Xia, C.; Wang, W.; Wen, W.; Zhang, M. Guanylate kinase domains of the MAGUK family scaffold proteins as specific phospho-protein-binding modules. EMBO J. 2011, 30, 4986–4997. [Google Scholar] [CrossRef] [PubMed]
- Wu, X.; Cai, Q.; Chen, Y.; Zhu, S.; Mi, J.; Wang, J.; Zhang, M. Structural Basis for the High-Affinity Interaction between CASK and Mint1. Structure 2020, 28, 664–673.e3. [Google Scholar] [CrossRef] [PubMed]
- Stafford, R.L.; Ear, J.; Knight, M.J.; Bowie, J.U. The molecular basis of the Caskin1 and Mint1 interaction with CASK. J. Mol. Biol. 2011, 412, 3–13. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Stevenson, D.; Laverty, H.G.; Wenwieser, S.; Douglas, M.; Wilson, J.B. Mapping and expression analysis of the human CASK gene. Mamm. Genome 2000, 11, 934–937. [Google Scholar] [CrossRef]
- Zordan, M.A.; Massironi, M.; Ducato, M.G.; Te Kronnie, G.; Costa, R.; Reggiani, C.; Chagneau, C.; Martin, J.R.; Megighian, A. Drosophila CAKI/CMG protein, a homolog of human CASK, is essential for regulation of neurotransmitter vesicle release. J. Neurophysiol. 2005, 94, 1074–1083. [Google Scholar] [CrossRef] [Green Version]
- Gillespie, J.M.; Hodge, J.J. CASK regulates CaMKII autophosphorylation in neuronal growth, calcium signaling, and learning. Front. Mol. Neurosci. 2013, 6, 27. [Google Scholar] [CrossRef] [Green Version]
- Hackett, A.; Tarpey, P.S.; Licata, A.; Cox, J.; Whibley, A.; Boyle, J.; Rogers, C.; Grigg, J.; Partington, M.; Stevenson, R.E.; et al. CASK mutations are frequent in males and cause X-linked nystagmus and variable XLMR phenotypes. Eur. J. Hum. Genet. 2010, 18, 544–552. [Google Scholar] [CrossRef] [Green Version]
- Burglen, L.; Chantot-Bastaraud, S.; Garel, C.; Milh, M.; Touraine, R.; Zanni, G.; Petit, F.; Afenjar, A.; Goizet, C.; Barresi, S.; et al. Spectrum of pontocerebellar hypoplasia in 13 girls and boys with CASK mutations: Confirmation of a recognizable phenotype and first description of a male mosaic patient. Orphanet J. Rare Dis. 2012, 7, 18. [Google Scholar] [CrossRef] [Green Version]
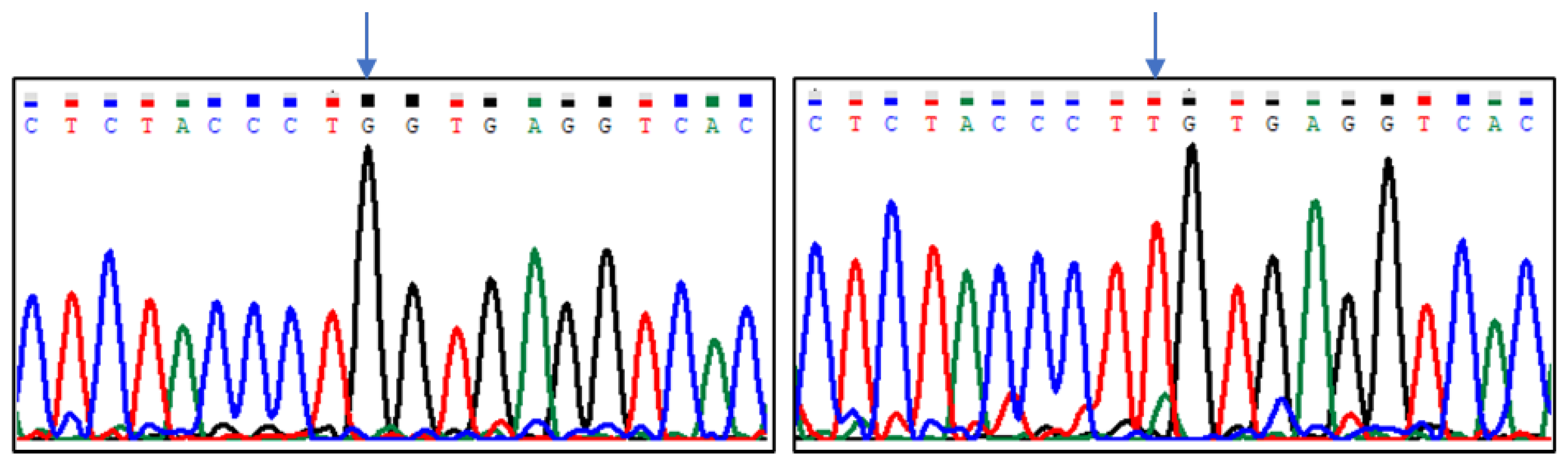
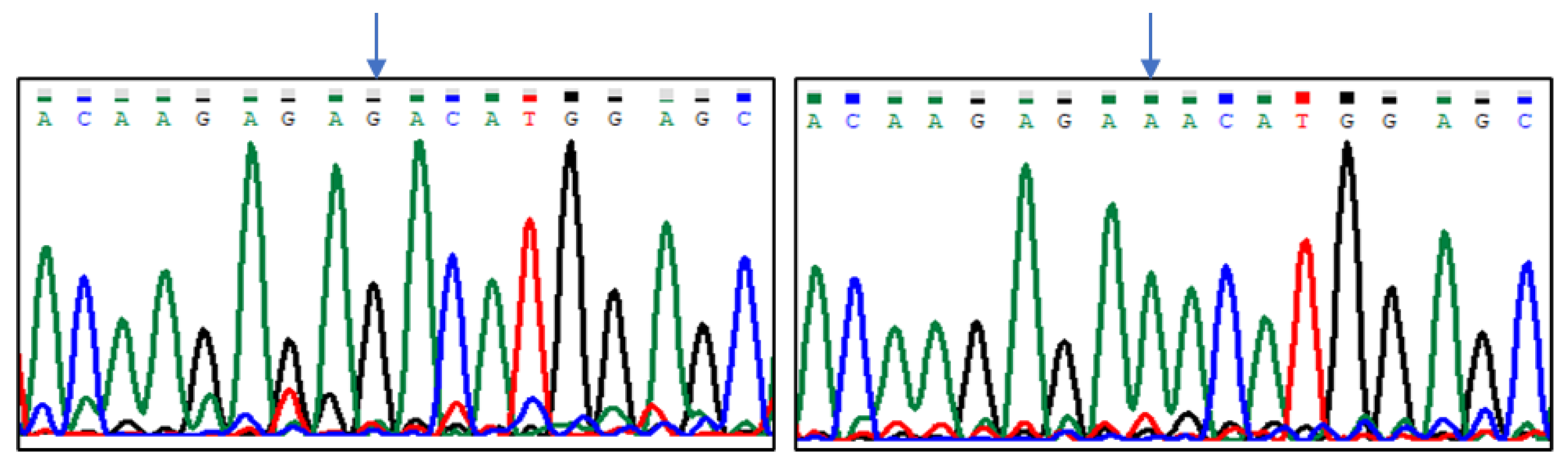

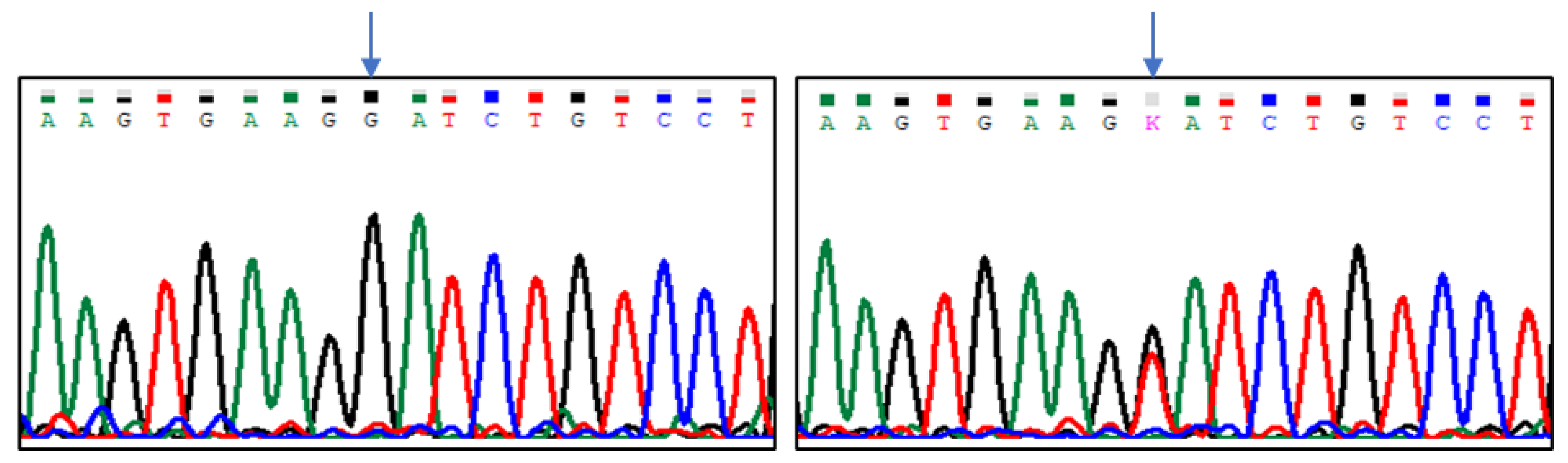

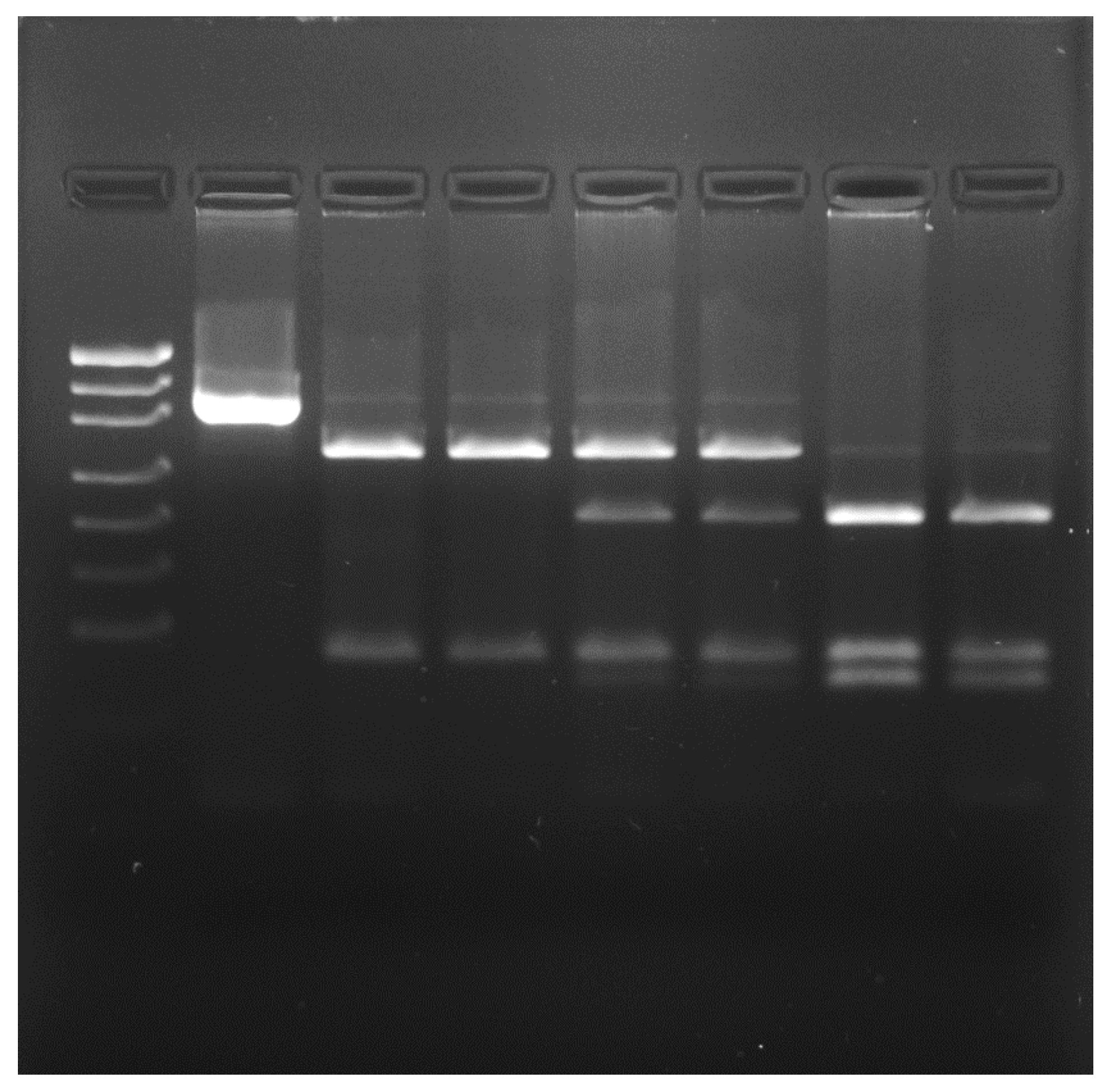
| Group of Pigeons | |||
|---|---|---|---|
| Homing | Non-Homing | ||
| Natural Antwerp (n = 82) | Aces (PiGen vof) (n = 29) | Flying and Fancy (n = 58) | Utility (n = 35) |
| Janssen (40) Bricoux (2) De Smet-Matthys (9) Grondelaers (5) Meulemans (6) Stichelbaut (10) Wanroy (10) | G. Show Homer (6) Bagdad of Nuremberg (4) G. Magpie (5) G. Long Faced Tumbler (10) Polish Barb (4) Vienna Kiebitz (1) Danzig Highflier (9) Carrier (6) Polish Krymka Tumbler (3) Polish Owl (1) Fantail (4) Polish Short-Beaked (5) | King (11) Mondain (4) Strasser (10) Maltese (3) Polish Lynx (3) Cauchois (4) | |
| Group | n | Genotypes | Alleles | |||
|---|---|---|---|---|---|---|
| AA | AG | GG | A | G | ||
| Homing pigeons (Natural Antwerp) | 82 | 0.488 a (n = 40) | 0.463 (n = 38) | 0.049 (n = 4) | 0.720 | 0.280 |
| Homing pigeons (Aces, PiGen vof) | 29 | 0.655 a (n = 19) | 0.345 (n = 10) | - (n = 0) | 0.792 | 0.208 |
| Non-homing pigeons | 93 | 0.194 b (n = 18) | 0.344 (n = 32) | 0.462 (n = 43) | 0.366 | 0.634 |
| Total | 204 | 0.384 (n = 77) | 0.388 (n = 80) | 0.228 (n = 47) | 0.578 | 0.422 |
| Group | n | Genotypes | Alleles | HWE (p-Value) | |||
|---|---|---|---|---|---|---|---|
| AA | AG | GG | A | G | |||
| Homing pigeons (A.S.) | 311 | 0.502 (n = 156) | 0.434 (n = 135) | 0.064 (n = 20) | 0.719 | 0.281 | p > 0.05 |
| Genotype/Sex | RR | AP | SE | ANOVA Test |
|---|---|---|---|---|
| AA | 552 | 29.16 | 1.62 | p > 0.05 |
| AG | 482 | 27.42 | 1.68 | |
| GG | 77 | 24.02 | 4.20 | |
| Females | 511 | 28.59 | 1.67 | p > 0.05 |
| Males | 600 | 27.58 | 1.52 |
| Race Number | RR | AP | SE | ANOVA Test |
|---|---|---|---|---|
| 1 | 227 | 27.14 a | 2.32 | p < 0.05 |
| 2 | 298 | 21.82 a | 2.03 | |
| 3 | 289 | 49.44 b | 2.05 | |
| 4 | 297 | 14.16 c | 2.03 |
| Number of Race | Genotype | RR | AP | SE |
|---|---|---|---|---|
| 1 | AA | 110 | 28.76 | 3.33 |
| AG | 99 | 27.89 | 3.55 | |
| GG | 18 | 16.81 | 8.28 | |
| 2 | AA | 150 | 23.73 | 2.81 |
| AG | 128 | 20.13 | 3.05 | |
| GG | 20 | 19.27 | 7.73 | |
| 3 | AA | 145 | 49.46 | 3.51 |
| AG | 125 | 50.73 | 3.77 | |
| GG | 19 | 41.82 | 9.67 | |
| 4 | AA | 147 | 14.87 | 2.27 |
| AG | 130 | 12.86 | 2.41 | |
| GG | 20 | 15.40 | 6.17 |
| Number of Race | Sex | RR | AP | SE |
|---|---|---|---|---|
| 1 | Female | 102 | 23.82 | 4.77 |
| Male | 125 | 25.15 | 4.27 | |
| 2 | Female | 140 | 19.73 | 4.32 |
| Male | 158 | 22.36 | 3.94 | |
| 3 | Female | 133 | 48.31 | 5.31 |
| Male | 156 | 46.37 | 5.01 | |
| 4 | Female | 136 | 8.88 * | 3.46 |
| Male | 161 | 19.87 * | 3.14 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Dybus, A.; Kulig, H.; Grzesiak, W.; Domke, J.; Yu, Y.-H.; Cheng, Y.-H. Calcium/Calmodulin-Dependent Serine Protein Kinase (CASK) Gene Polymorphisms in Pigeons. Animals 2023, 13, 2070. https://doi.org/10.3390/ani13132070
Dybus A, Kulig H, Grzesiak W, Domke J, Yu Y-H, Cheng Y-H. Calcium/Calmodulin-Dependent Serine Protein Kinase (CASK) Gene Polymorphisms in Pigeons. Animals. 2023; 13(13):2070. https://doi.org/10.3390/ani13132070
Chicago/Turabian StyleDybus, Andrzej, Hanna Kulig, Wilhelm Grzesiak, Justyna Domke, Yu-Hsiang Yu, and Yeong-Hsiang Cheng. 2023. "Calcium/Calmodulin-Dependent Serine Protein Kinase (CASK) Gene Polymorphisms in Pigeons" Animals 13, no. 13: 2070. https://doi.org/10.3390/ani13132070
APA StyleDybus, A., Kulig, H., Grzesiak, W., Domke, J., Yu, Y.-H., & Cheng, Y.-H. (2023). Calcium/Calmodulin-Dependent Serine Protein Kinase (CASK) Gene Polymorphisms in Pigeons. Animals, 13(13), 2070. https://doi.org/10.3390/ani13132070






