RNA Sequencing Analysis Reveals Divergent Adaptive Response to Hypo- and Hyper-Salinity in Greater Amberjack (Seriola dumerili) Juveniles
Abstract
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Ethics Statement
2.2. Experimental Fish, Salinity Development, and Tissue Collection
2.3. Total RNA Extraction, Library Construction, and Illumina Sequencing
2.4. Transcriptome Assembly and Functional Gene Annotation
2.5. Analysis of Differential Expressed Genes (DEGs) Analysis
2.6. Quantitative Real-Time PCR (QPCR) Validation
2.7. Statistical Analysis
3. Results
3.1. Illumina Sequencing
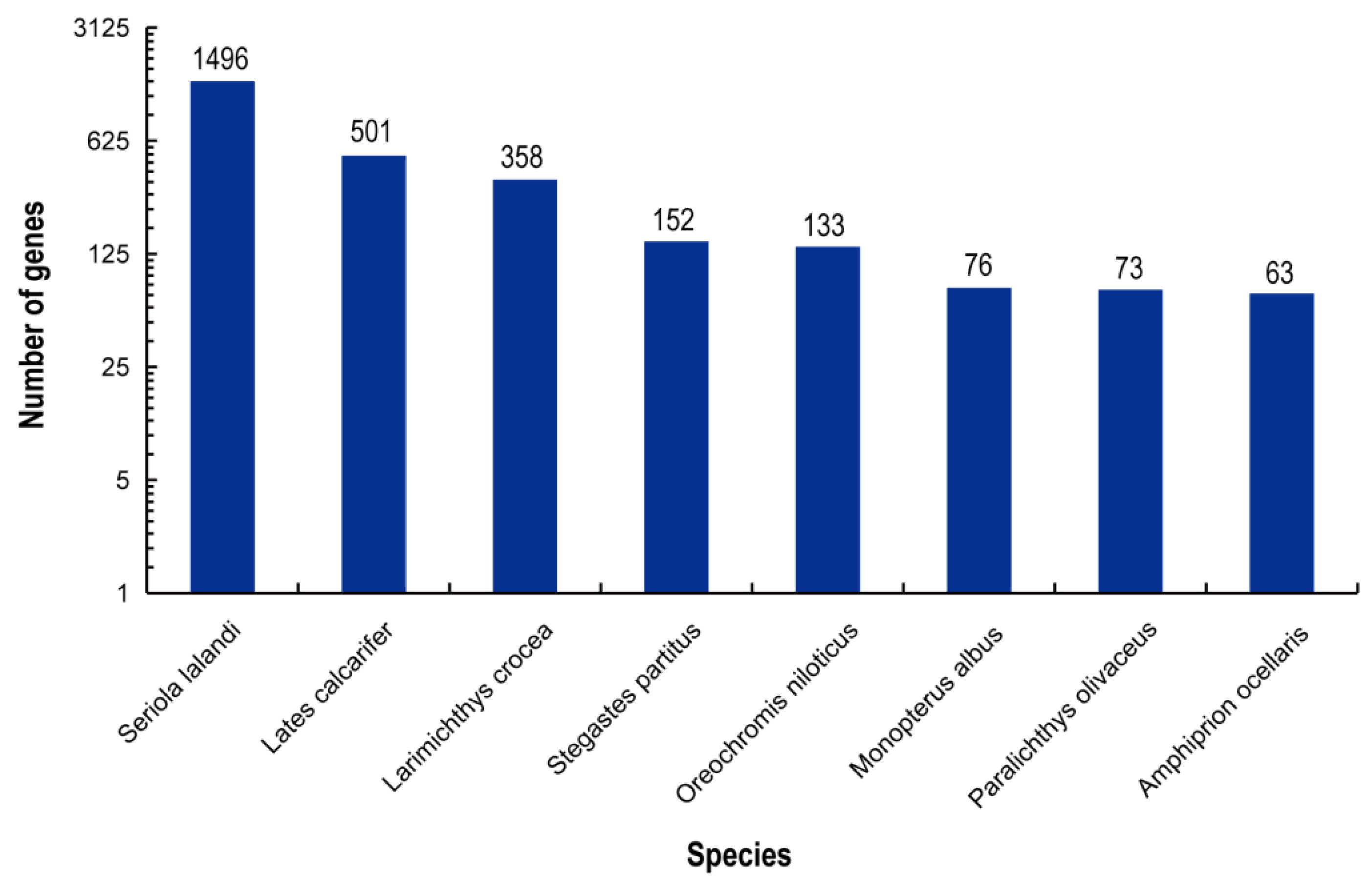
3.2. Annotation and Functional Analysis of the Gill Transcriptome
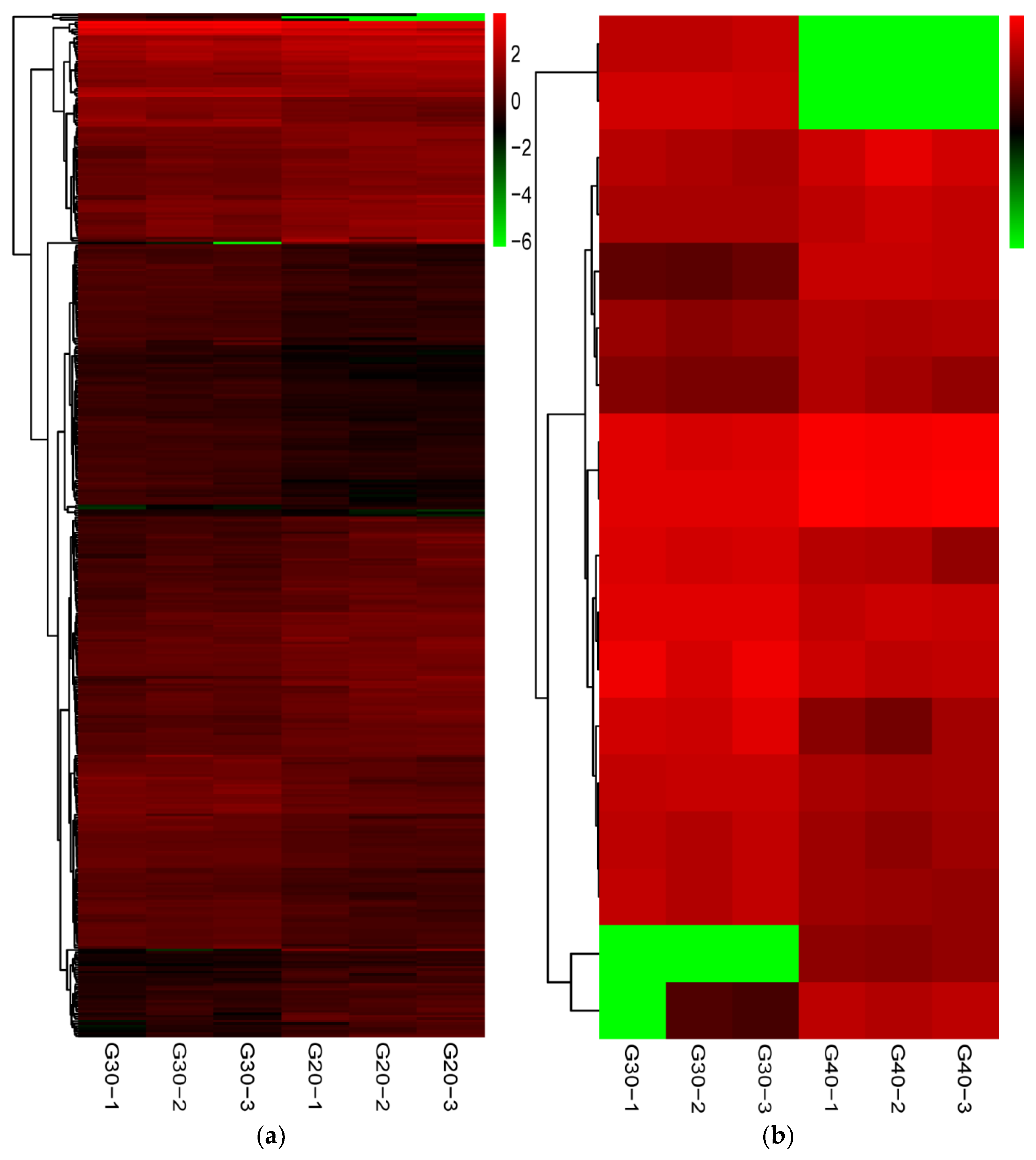
3.3. Identification and Analysis of Differentially Expressed Genes (DEGs)
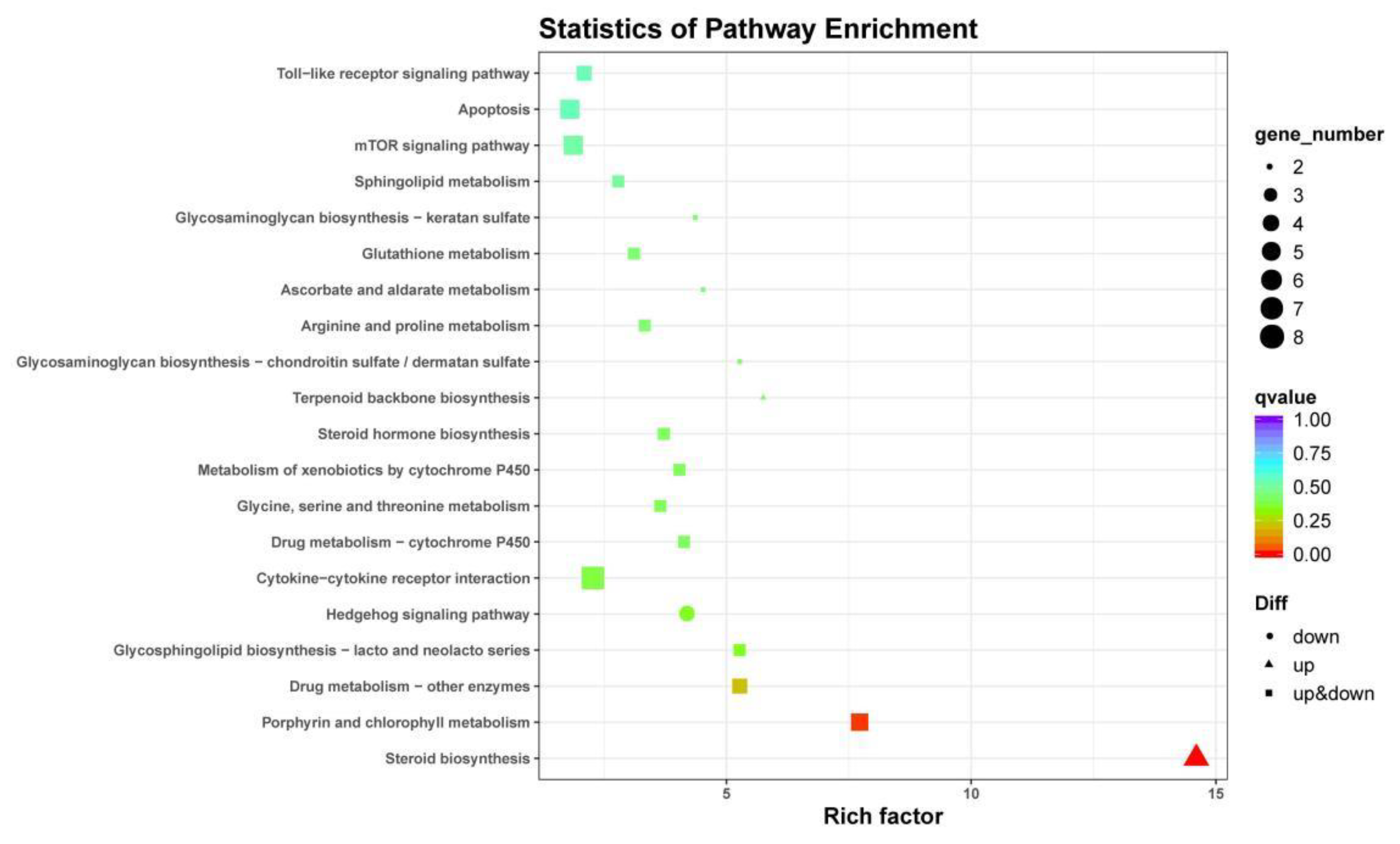
3.4. DEG Annotation and Pathway Analysis
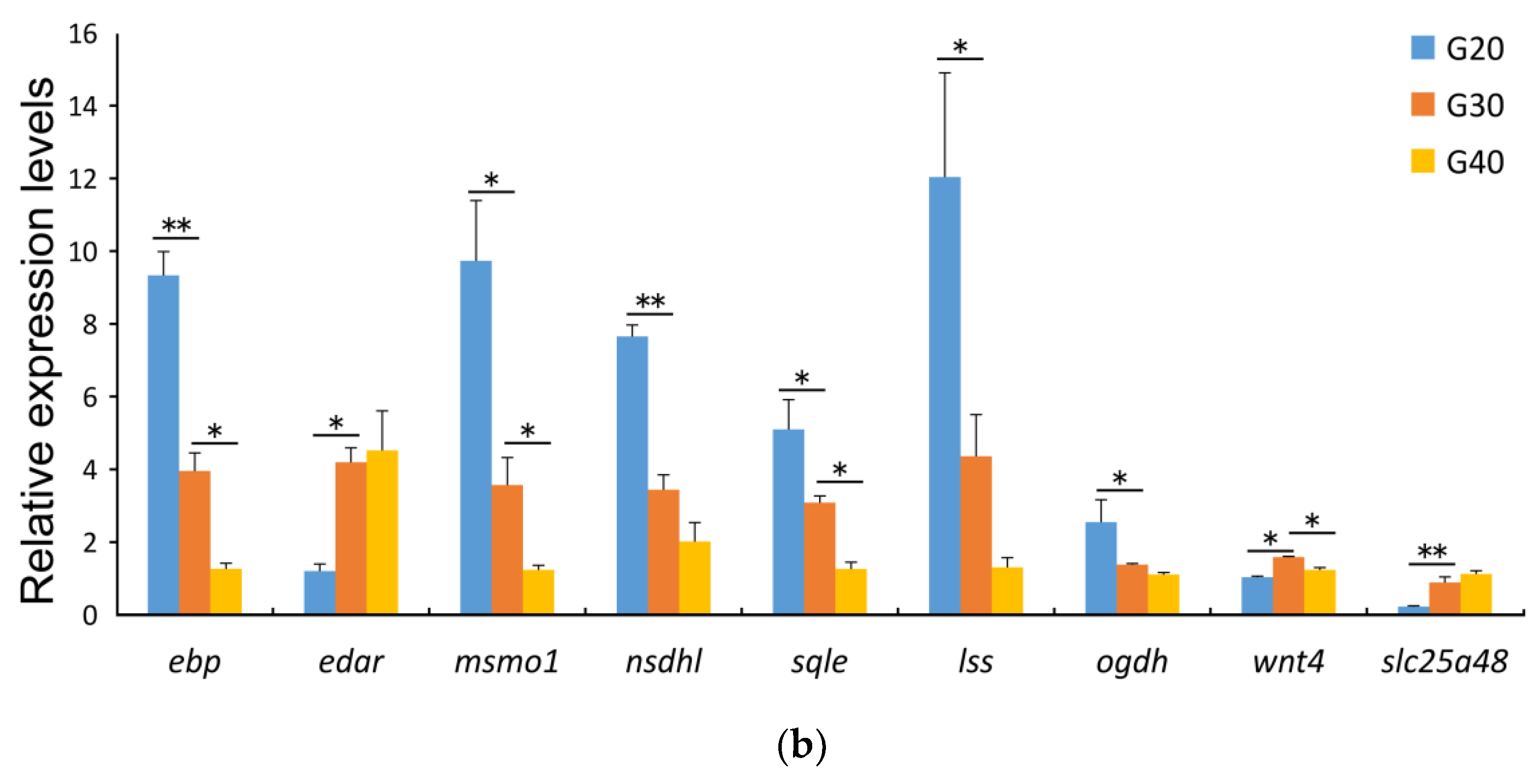
3.5. Validation of RNA Sequencing Data by QPCR
4. Discussion
4.1. Transcriptomic Analysis of Differentially Expressed Genes
4.2. DEGs Involved in Steroid Biosynthesis and Lipid Metabolism
4.3. DEGs Involved in Ion Transport
4.4. DEGs Involved in Immune Response
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Holmer, M. Environmental issues of fish farming in offshore waters: Perspectives, concerns and research needs. Aquac. Environ. Interact. 2010, 1, 57–70. [Google Scholar] [CrossRef]
- Seth, H.; Axelsson, M. Sympathetic, parasympathetic and enteric regulation of the gastrointestinal vasculature in rainbow trout (Oncorhynchus mykiss) under normal and postprandial conditions. J. Exp. Biol. 2010, 213, 3118–3126. [Google Scholar] [CrossRef] [PubMed]
- Perry, S.F.; Capaldo, A. The autonomic nervous system and chromaffin tissue: Neuroendocrine regulation of catecholamine secretion in non-mammalian vertebrates. Auton. Neurosci. 2011, 165, 54–66. [Google Scholar] [CrossRef] [PubMed]
- Jeffrey, J.D.; Esbaugh, A.J.; Vijayan, M.M.; Gilmour, K.M. Modulation of hypothalamic-pituitary-interrenal axis function by social status in rainbow trout. Gen. Comp. Endocrinol. 2012, 176, 201–210. [Google Scholar] [CrossRef] [PubMed]
- Schulte, P.M. What is environmental stress? Insights from fish living in a variable environment. J. Exp. Biol. 2014, 217, 23–34. [Google Scholar] [CrossRef] [PubMed]
- Kammerer, B.D.; Sardella, B.A.; Kultz, D. Salinity stress results in rapid cell cycle changes of tilapia (Oreochromis mossambicus) gill epithelial cells. J. Exp. Zool. Part A Ecol. Genet. Physiol. 2009, 311, 80–90. [Google Scholar] [CrossRef]
- Wen, X.; Chu, P.; Xu, J.; Wei, X.; Fu, D.; Wang, T.; Yin, S. Combined effects of low temperature and salinity on the immune response, antioxidant capacity and lipid metabolism in the pufferfish (Takifugu fasciatus). Aquaculture 2021, 531, 735886. [Google Scholar] [CrossRef]
- Guo, T.; Yang, Y.; Meng, F.; Wang, S.; Xia, S.; Qian, Y.; Li, M.; Wang, R. Effects of low salinity on gill and liver glycogen metabolism of great blue-spotted mudskippers (Boleophthalmus pectinirostris). Comp. Biochem. Physiol. Part C 2020, 230, 108709. [Google Scholar] [CrossRef] [PubMed]
- Cui, Q.; Qiu, L.; Yang, X.; Shang, S.; Yang, B.; Chen, M.; Liu, X.; Chen, B.; Fu, X.; Wang, W.; et al. Transcriptome profiling of the low-salinity stress responses in the gills of the juvenile Pseudopleuronectes yokohamae. Comp. Biochem. Physiol. Part D 2019, 32, 100612. [Google Scholar] [CrossRef]
- Xu, Z.; Gan, L.; Li, T.; Xu, C.; Chen, K.; Wang, X.; Qin, J.G.; Chen, L.; Li, E. Transcriptome profiling and molecular pathway analysis of genes in association with salinity adaptation in nile tilapia Oreochromis niloticus. PLoS ONE 2015, 10, e0136506. [Google Scholar] [CrossRef]
- Zhou, Z.; Hu, F.; Li, W.; Yang, X.; Hallerman, E.; Huang, Z. Effects of salinity on growth, hematological parameters, gill microstructure and transcriptome of fat greenling Hexagrammos otakii. Aquaculture 2021, 531, 735945. [Google Scholar] [CrossRef]
- Xiong, Y.; Dong, S.; Huang, M.; Li, Y.; Wang, X.; Wang, F.; Ma, S.; Zhou, Y. Growth, osmoregulatory response, adenine nucleotide contents, and liver transcriptome analysis of steelhead trout (Oncorhynchus mykiss) under different salinity acclimation methods. Aquaculture 2020, 520, 734937. [Google Scholar] [CrossRef]
- Handayani, K.S.; Soegianto, A.; Chang, C.-F. Effect of salinity on osmoregulation and histopathology in gills of tilapia (Oreochromis niloticus). Asian J. Water Environ. Pollut. 2020, 17, 7–11. [Google Scholar] [CrossRef]
- Lin, Y.T.; Hu, Y.C.; Wang, Y.C.; Hsiao, M.Y.; Lorin-Nebel, C.; Lee, T.H. Differential expression of two ATPases revealed by lipid raft isolation from gills of euryhaline teleosts with different salinity preferences. Comp. Biochem. Physiol. Part B 2021, 253, 110562. [Google Scholar] [CrossRef] [PubMed]
- Mohamed, N.A.; Saad, M.F.; Shukry, M.; El-Keredy, A.M.S.; Nasif, O.; Van Doan, H.; Dawood, M.A.O. Physiological and ion changes of Nile tilapia (Oreochromis niloticus) under the effect of salinity stress. Aquac. Rep. 2021, 19, 100567. [Google Scholar] [CrossRef]
- Fiol, D.F.; Chan, S.Y.; Kultz, D. Identification and pathway analysis of immediate hyperosmotic stress responsive molecular mechanisms in tilapia (Oreochromis mossambicus) gill. Comp. Biochem. Physiol. Part D 2006, 1, 344–356. [Google Scholar] [CrossRef]
- Gjessing, M.C.; Krasnov, A.; Timmerhaus, G.; Brun, S.; Afanasyev, S.; Dale, O.B.; Dahle, M.K. The Atlantic salmon gill transcriptome response in a natural outbreak of salmon gill pox virus infection reveals new biomarkers of gill pathology and suppression of mucosal defense. Front. Immunol. 2020, 11, 2154. [Google Scholar] [CrossRef]
- Cheng, J.X.; Xia, Y.Q.; Liu, Y.F.; Liu, P.F.; Liu, Y. Transcriptome analysis in Takifugu rubripes and Dicentrarchus labrax gills during Cryptocaryon irritans infection. J. Fish Dis. 2021, 44, 249–262. [Google Scholar] [CrossRef]
- Marshall, W.S.; Bellamy, D. The 50 year evolution of in vitro systems to reveal salt transport functions of teleost fish gills. Comp. Biochem. Physiol. Part A 2010, 155, 275–280. [Google Scholar] [CrossRef]
- Evans, T.G.; Kultz, D. The cellular stress response in fish exposed to salinity fluctuations. J. Exp. Zool. Part A 2020, 333, 421–435. [Google Scholar] [CrossRef]
- Jiang, W.; Tian, X.; Fang, Z.; Li, L.; Dong, S.; Li, H.; Zhao, K. Metabolic responses in the gills of tongue sole (Cynoglossus semilaevis) exposed to salinity stress using NMR-based metabolomics. Sci. Total Environ. 2019, 653, 465–474. [Google Scholar] [CrossRef]
- Sun, Z.; Lou, F.; Zhang, Y.; Song, N. Gill transcriptome sequencing and de novo annotation of Acanthogobius ommaturus in response to salinity stress. Genes 2020, 11, 631. [Google Scholar] [CrossRef] [PubMed]
- Farzadfar, F.; Doustshenas, B.; Rezaie, A.; Mousavi, S.M. Salinity induced alterations in ionic concentration of haemolymph and its effects on histopathology of gills and digestive gland in razor clam (Solen dactylus von Cosel, 1989; Bivalvia, Solenidae). Molluscan Res. 2021, 41, 92–102. [Google Scholar] [CrossRef]
- Takata, R.; Mattioli, C.C.; Bazzoli, N.; Júnior, J.D.C.; Luz, R.K. The effects of salinity on growth, gill tissue and muscle cellularity in Lophiosilurus alexandri juvenile, a neotropical freshwater catfish. Aquac. Res. 2021, 52, 4064–4075. [Google Scholar] [CrossRef]
- Cao, Q.; Wang, H.; Fan, C.; Sun, Y.; Li, J.; Cheng, J.; Chu, P.; Yin, S. Environmental salinity influences the branchial expression of TCR pathway related genes based on transcriptome of a catadromous fish. Comp. Biochem. Physiol. Part D 2021, 38, 100815. [Google Scholar] [CrossRef] [PubMed]
- Cao, Q.; Li, J.; Sun, Y.; Geng, D.; Chu, P.; Wang, H.; Yin, S. Time-course studies of osmoregulatory responses in different salinities for the marbled eel (Anguilla marmorata). Aquaculture 2021, 540, 736699. [Google Scholar] [CrossRef]
- Zhang, X.; Wen, H.; Wang, H.; Ren, Y.; Zhao, J.; Li, Y. RNA-Seq analysis of salinity stress-responsive transcriptome in the liver of spotted sea bass (Lateolabrax maculatus). PLoS ONE 2017, 12, e0173238. [Google Scholar] [CrossRef] [PubMed]
- Si, Y.; Wen, H.; Li, Y.; He, F.; Li, J.; Li, S.; He, H. Liver transcriptome analysis reveals extensive transcriptional plasticity during acclimation to low salinity in Cynoglossus semilaevis. BMC Genom. 2018, 19, 464. [Google Scholar] [CrossRef]
- El-Leithy, A.A.A.; Hemeda, S.A.; El Naby, W.; El Nahas, A.F.; Hassan, S.A.H.; Awad, S.T.; El-Deeb, S.I.; Helmy, Z.A. Optimum salinity for nile tilapia (Oreochromis niloticus) growth and mRNA transcripts of ion-regulation, inflammatory, stress- and immune-related genes. Fish Physiol. Biochem. 2019, 45, 1217–1232. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Xue, L.; Cao, M.; Zhang, Y.; Wang, Y.; Xu, S.; Zheng, B.; Lou, Z. Gill transcriptomes reveal expression changes of genes related with immune and ion transport under salinity stress in silvery pomfret (Pampus argenteus). Fish Physiol. Biochem. 2020, 46, 1255–1277. [Google Scholar] [CrossRef]
- Cao, D.; Li, J.; Huang, B.; Zhang, J.; Pan, C.; Huang, J.; Zhou, H.; Ma, Q.; Chen, G.; Wang, Z. RNA-seq analysis reveals divergent adaptive response to hyper- and hypo-salinity in cobia, Rachycentron canadum. Fish Physiol. Biochem. 2020, 46, 1713–1727. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.H.; Thu, B.J.; Skall, H.F.; Vendramin, N.; Evensen, O. A single amino acid mutation (I1012F) of the RNA polymerase of marine viral hemorrhagic septicemia virus changes in vitro virulence to rainbow trout gill epithelial cells. J. Virol. 2014, 88, 7189–7198. [Google Scholar] [CrossRef] [PubMed]
- Deane, E.E.; Woo, N.Y. Tissue distribution, effects of salinity acclimation, and ontogeny of aquaporin 3 in the marine teleost, silver sea bream (Sparus sarba). Mar. Biotechnol. 2006, 8, 663–671. [Google Scholar] [CrossRef] [PubMed]
- Watanabe, S.; Kaneko, T.; Aida, K. Aquaporin-3 expressed in the basolateral membrane of gill chloride cells in Mozambique tilapia Oreochromis mossambicus adapted to freshwater and seawater. J. Exp. Biol. 2005, 208, 2673–2682. [Google Scholar] [CrossRef] [PubMed]
- Fiol, D.F.; Kultz, D. Osmotic stress sensing and signaling in fishes. FEBS J. 2007, 274, 5790–5798. [Google Scholar] [CrossRef] [PubMed]
- Wichmann, L.; Althaus, M. Evolution of epithelial sodium channels: Current concepts and hypotheses. Am. J. Physiol. Regul. Integr. Comp. Physiol. 2020, 319, R387–R400. [Google Scholar] [CrossRef] [PubMed]
- Tang, C.H.; Hwang, L.Y.; Shen, I.D.; Chiu, Y.H.; Lee, T.H. Immunolocalization of chloride transporters to gill epithelia of euryhaline teleosts with opposite salinity-induced Na+/K+-ATPase responses. Fish Physiol. Biochem. 2011, 37, 709–724. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.Y.; Lee, H.J.; Kim, Y.K. Comparative transcriptome profiling of selected osmotic regulatory proteins in the gill during seawater acclimation of chum salmon (Oncorhynchus keta) fry. Sci. Rep. 2020, 10, 1987. [Google Scholar] [CrossRef] [PubMed]
- Kang, W.; Zhang, M.; Wang, Q.; Gu, D.; Huang, Z.; Wang, H.; Xiang, Y.; Xia, Q.; Cui, Z.; Jin, X. The SLC family are candidate diagnostic and prognostic biomarkers in clear cell renal cell carcinoma. Biomed. Res. Int. 2020, 2020, 1932948. [Google Scholar] [PubMed]
- Cutler, C.P.; Murray, D.; Ojo, T.; Harmon, S.; MacIver, B.; Cramb, G.; Zeidel, M.L. Aquaporin (AQP) channels in the spiny dogfish, Squalus acanthias I: Characterization of AQP3 and AQP15 function and expression, and localization of the proteins in gill and spiral valve intestine. Comp. Biochem. Physiol. Part B 2021, 258, 110702. [Google Scholar] [CrossRef]
- Hou, Z.-S.; Wen, H.-S. Neuropeptide Y and melanocortin receptors in fish: Regulators of energy homeostasis. Mar. Life Sci. Technol. 2021, 4, 42–51. [Google Scholar] [CrossRef]
- Shang, X.; Ma, A.; Wang, X.; Xia, D.; Zhuang, J. Isolation, characterization and expression analysis of TRPV4 in half-smooth tongue sole Cynoglossus semilaevis. J. Oceanol. Limnol. 2020, 38, 294–305. [Google Scholar] [CrossRef]
- Talbot, A.; Gargan, L.; Moran, G.; Prudent, L.; O’Connor, I.; Mirimin, L.; Carlsson, J.; MacCarthy, E. Investigation of the transcriptomic response in Atlantic salmon (Salmo salar) gill exposed to Paramoeba perurans during early onset of disease. Sci. Rep. 2021, 11, 20682. [Google Scholar] [CrossRef] [PubMed]
- Fakriadis, I.; Sigelaki, I.; Papadaki, M.; Papandroulakis, N.; Raftopoulos, A.; Tsakoniti, K.; Mylonas, C.C. Control of reproduction of greater amberjack Seriola dumerili reared in aquaculture facilities. Aquaculture 2020, 519, 734880. [Google Scholar] [CrossRef]
- Yokoyama, S.; Ishikawa, M.; Koshio, S. Dietary bovine lactoferrin enhances defense factors on body surface and anti-parasitic effects against Neobenedenia girellae infection, and mitigates low-salinity stress in amberjack (Seriola dumerili) juveniles. Aquaculture 2019, 504, 52–58. [Google Scholar] [CrossRef]
- Navarro-Guillén, C.; Conceição, L.E.C.; Pinto, W.; Siguero, I.; Urrutia, P.; Moyano, F.J.; Yúfera, M. Fast growing greater amberjack post-larvae require a high energy-high protein weaning diet. Aquaculture 2019, 499, 195–202. [Google Scholar] [CrossRef]
- Chen, C.; Ji, R.; Huang, J.; He, H.; Liao, Z. The relationship between the salinity and the embryonic, early larval development in Seriola dumerili. J. Shanghai Ocean Univ. 1997, 6, 1. [Google Scholar]
- Azodi, M.; Bahabadi, M.N.; Ghasemi, A.; Morshedi, V.; Mozanzadeh, M.T.; Shahraki, R.; Khademzadeh, O.; Hamedi, S.; Avizhgan, S. Effects of salinity on gills’ chloride cells, stress indices, and gene expression of Asian seabass (Lates calcarifer, Bloch, 1790). Fish Physiol. Biochem. 2021, 47, 2027–2039. [Google Scholar] [CrossRef] [PubMed]
- Lin, X.; Zhou, D.; Zhang, X.; Li, G.; Zhang, Y.; Huang, C.; Zhang, Z.; Tian, C. A First Insight into the Gonad Transcriptome of Hong Kong Catfish (Clarias fuscus). Animals 2021, 11, 1131. [Google Scholar] [CrossRef]
- Araki, K.; Aokic, J.-Y.; Kawase, J.; Hamada, K.; Ozaki, A.; Fujimoto, H.; Yamamoto, I.; Usuki, H. Whole genome sequencing of greater amberjack (Seriola dumerili) for SNP identification on aligned scaffolds and genome structural variation analysis using parallel resequencing. Int. J. Genom. 2018, 2018, 7984292. [Google Scholar] [CrossRef]
- Shi, H.; Ru, X.; Mustapha, U.F.; Jiang, D.; Huang, Y.; Pan, S.; Zhu, C.; Li, G. Characterization, expression, and regulatory effects of nr0b1a and nr0b1b in spotted scat (Scatophagus argus). Comp. Biochem. Physiol. Part B 2021, 256, 110644. [Google Scholar] [CrossRef] [PubMed]
- Zupa, R.; Rodríguez, C.; Mylonas, C.C.; Rosenfeld, H.; Fakriadis, I.; Papadaki, M.; Perez, J.A.; Pousis, C.; Basilone, G.; Corriero, A. Comparative study of reproductive development in wild and captive-reared greater amberjack Seriola dumerili (Risso, 1810). PLoS ONE 2017, 12, e0169645. [Google Scholar]
- Pousis, C.; Mylonas, C.C.; De Virgilio, C.; Santamaria, N.; Passantino, L.; Zupa, R.; Papadaki, M.; Fakriadis, I.; Ferreri, R.; Corriero, A. The observed oogenesis impairment in greater amberjack Seriola dumerili (Risso, 1810) reared in captivity is not related to aninsufficient liver transcription or oocyte uptake of vitellogenin. Aquac. Res. 2017, 49, 1–10. [Google Scholar]
- Pousis, C.; Rodríguez, C.; De Ruvo, P.; De Virgilio, C.; Pérez, J.A.; Mylonas, C.C.; Zupa, R.; Passantino, L.; Santamaria, N.; Valentini, L.; et al. Vitellogenin receptor and fatty acid profiles of individual lipid classes of oocytes from wild and captive-reared greater amberjack (Seriola dumerili) during the reproductive cycle. Theriogenology 2019, 140, 73–83. [Google Scholar] [CrossRef] [PubMed]
- Nie, X.; Zhang, C.; Jiang, C.; Li, S.; Hong, W.; Chen, S.; Zhang, Y. Characterizing transcriptome changes in gill tissue of turbot (Scophthalmus maximus) for waterless preservation. Aquaculture 2020, 518, 734830. [Google Scholar] [CrossRef]
- Kelly, S.P.; Wood, C.M. Cortisol stimulates calcium transport across cultured gill epithelia from freshwater rainbow trout. In Vitro Cell. Dev. Biol. Anim. 2008, 44, 96–104. [Google Scholar] [CrossRef]
- Mankiewicz, J.L.; Deck, C.A.; Taylor, J.D.; Douros, J.D.; Borski, R.J. Epinephrine and glucose regulation of leptin synthesis and secretion in a teleost fish, the tilapia (Oreochromis mossambicus). Gen. Comp. Endocrinol. 2021, 302, 113669. [Google Scholar] [CrossRef]
- Sandhu, N.; McGeer, J.C.; Vijayan, M.M. Exposure to environmental levels of waterborne cadmium impacts corticosteroidogenic and metabolic capacities, and compromises secondary stressor performance in rainbow trout. Aquat. Toxicol. 2014, 146, 20–27. [Google Scholar] [CrossRef]
- Hartill, V.L.; Tysoe, C.; Manning, N.; Dobbie, A.; Santra, S.; Walter, J.; Caswell, R.; Koster, J.; Waterham, H.; Hobson, E. An unusual phenotype of X-linked developmental delay and extreme behavioral difficulties associated with a mutation in the EBP gene. Am. J. Med. Genet. Part A 2014, 164, 907–914. [Google Scholar] [CrossRef]
- Juhl, A.D.; Heegaard, C.W.; Werner, S.; Schneider, G.; Krishnan, K.; Covey, D.F.; Wustner, D. Quantitative imaging of membrane contact sites for sterol transfer between endo-lysosomes and mitochondria in living cells. Sci. Rep. 2021, 11, 8927. [Google Scholar] [CrossRef]
- Yang, F.; Kou, J.; Liu, Z.; Li, W.; Du, W. MYC Enhances cholesterol biosynthesis and supports cell proliferation through SQLE. Front. Cell Dev. Biol. 2021, 9, 655889. [Google Scholar] [CrossRef] [PubMed]
- Furtado, L.V.; Kelley, R.I.; Opitz, J.M. Disorders of sterol biosynthesis. Transl. Sci. Rare Dis. 2016, 1, 145–182. [Google Scholar] [CrossRef]
- Liu, Y.; Li, E.; Xu, C.; Su, Y.; Qin, J.G.; Chen, L.; Wang, X. Brain transcriptome profiling analysis of nile tilapia (Oreochromis niloticus) under long-term hypersaline stress. Front. Physiol. 2018, 9, 219. [Google Scholar] [CrossRef] [PubMed]
- Bai, W.; Zhang, C.; Chen, H. Transcriptomic analysis of Momordica charantia polysaccharide on streptozotocin-induced diabetic rats. Gene 2018, 675, 208–216. [Google Scholar] [CrossRef] [PubMed]
- Xin, Y.; Li, C.; Guo, Y.; Xiao, R.; Zhang, H.; Zhou, G. RNA-Seq analysis reveals a negative role of MSMO1 with a synergized NSDHL expression during adipogenesis of 3T3-L1. Biosci. Biotechnol. Biochem. 2019, 83, 641–652. [Google Scholar] [CrossRef] [PubMed]
- Chen, M.; Zhao, Y.; Yang, X.; Zhao, Y.; Liu, Q.; Liu, Y.; Hou, Y.; Sun, H.; Jin, W. NSDHL promotes triple-negative breast cancer metastasis through the TGFbeta signaling pathway and cholesterol biosynthesis. Breast Cancer Res. Treat. 2021, 187, 349–362. [Google Scholar] [CrossRef]
- Bauer, A.; De Lucia, M.; Jagannathan, V.; Mezzalira, G.; Casal, M.L.; Welle, M.M.; Leeb, T. A large deletion in the NSDHL gene in labrador retrievers with a congenital cornification disorder. G3 Bethesda 2017, 7, 3115–3121. [Google Scholar] [CrossRef][Green Version]
- Hlusko, L.J.; Carlson, J.P.; Chaplin, G.; Elias, S.A.; Hoffecker, J.F.; Huffman, M.; Jablonski, N.G.; Monson, T.A.; O’Rourke, D.H.; Pilloud, M.A.; et al. Environmental selection during the last ice age on the mother-to-infant transmission of vitamin D and fatty acids through breast milk. Proc. Natl. Acad. Sci. USA 2018, 115, E4426–E4432. [Google Scholar] [CrossRef]
- Wang, B.; Liang, Y.; Chai, X.; Chen, S.; Ye, Z.; Li, R.; Li, X.; Kong, G.; Li, Y.; Zhang, X.; et al. Ectodysplasin A receptor (EDAR) promotes colorectal cancer cell proliferation via regulation of the Wnt/beta-catenin signaling pathway. Exp. Cell Res. 2020, 395, 112170. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Kong, X.; Ren, J.; Yuan, S.; Liu, C.; Hou, Y.; Liu, Y.; Meng, L.; Zhang, G.; Du, Q.; et al. A novel EDAR missense mutation identified by whole-exome sequencing with non-syndromic tooth agenesis in a Chinese family. Mol. Genet. Genom. Med. 2021, 9, e1684. [Google Scholar] [CrossRef]
- Li, M.; Bai, Y.T.; Han, K.; Li, X.D.; Meng, J. Knockdown of ectodysplasin-A receptor-associated adaptor protein exerts a tumor-suppressive effect in tongue squamous cell carcinoma cells. Exp. Ther. Med. 2020, 19, 3337–3347. [Google Scholar] [CrossRef] [PubMed]
- Hauner, H.; Link, D.; Wabitsch, M.; Brinkmann, U.; Keuper, M.; Skurk, T.; Finck, A.; Laumen, H.; Fischer-Posovszky, P.; Tiller, G. Effects of TWEAK (TNF Superfamily Member 12) on differentiation, metabolism, and secretory function of human primary preadipocytes and adipocytes. Endocrinology 2009, 150, 5373–5383. [Google Scholar]
- Nguyen, T.V.; Jung, H.; Nguyen, T.M.; Hurwood, D.; Mather, P. Evaluation of potential candidate genes involved in salinity tolerance in striped catfish (Pangasianodon hypophthalmus) using an RNA-Seq approach. Mar. Genom. 2016, 25, 75–88. [Google Scholar] [CrossRef] [PubMed]
- Mailloux, R.J.; Ayre, D.C.; Christian, S.L. Induction of mitochondrial reactive oxygen species production by GSH mediated S-glutathionylation of 2-oxoglutarate dehydrogenase. Redox Biol. 2016, 8, 285–297. [Google Scholar] [CrossRef] [PubMed]
- Mu, Y.; Li, W.; Wei, Z.; He, L.; Zhang, W.; Chen, X. Transcriptome analysis reveals molecular strategies in gills and heart of large yellow croaker (Larimichthys crocea) under hypoxia stress. Fish Shellfish Immunol. 2020, 104, 304–313. [Google Scholar] [CrossRef] [PubMed]
- Shmukler, B.E.; Reimold, F.R.; Heneghan, J.F.; Chen, C.; Zhao, T.; Paw, B.H.; Alper, S.L. Molecular cloning and functional characterization of zebrafish Slc4a3/Ae3 anion exchanger. Pflug. Arch. Eur. J. Physiol. 2014, 466, 1605–1618. [Google Scholar] [CrossRef]
- Liu, S.T.; Horng, J.L.; Chen, P.Y.; Hwang, P.P.; Lin, L.Y. Salt secretion is linked to acid-base regulation of ionocytes in seawater-acclimated medaka: New insights into the salt-secreting mechanism. Sci. Rep. 2016, 6, 31433. [Google Scholar] [CrossRef]
- Madsen, S.S.; Jensen, L.N.; Tipsmark, C.K.; Kiilerich, P.; Borski, R.J. Differential regulation of cystic fibrosis transmembrane conductance regulator and Na+,K+-ATPase in gills of striped bass, Morone saxatilis: Effect of salinity and hormones. J. Endocrinol. 2007, 192, 249–260. [Google Scholar] [CrossRef]
- Scott, G.R.; Schulte, P.M. Intraspecific variation in gene expression after seawater transfer in gills of the euryhaline killifish Fundulus heteroclitus. Comp. Biochem. Physiol. Part A 2005, 141, 176–182. [Google Scholar] [CrossRef]
- Singer, T.D.; Clements, K.M.; Semple, J.W.; Schulte, P.M.; Bystriansky, J.S.; Finstad, B.; Fleming, I.A.; McKinley, R.S. Seawater tolerance and gene expression in two strains of Atlantic salmon smolts. Can. J. Fish. Aquat. Sci. 2002, 59, 125–135. [Google Scholar] [CrossRef]
- Ma, Q.; Liu, X.; Li, A.; Liu, S.; Zhuang, Z. Effects of osmotic stress on the expression profiling of aquaporin genes in the roughskin sculpin (Trachidermus fasciatus). Acta Oceanol. Sin. 2020, 39, 19–25. [Google Scholar] [CrossRef]
- Cutler, C.P.; Maciver, B.; Cramb, G.; Zeidel, M. Aquaporin 4 is a ubiquitously expressed isoform in the dogfish (Squalus acanthias) shark. Front. Physiol. 2011, 2, 107. [Google Scholar] [CrossRef] [PubMed]
- Roy, J.; Terrier, F.; Marchand, M.; Herman, A.; Heraud, C.; Surget, A.; Lanuque, A.; Sandres, F.; Marandel, L. Effects of low stocking densities on zootechnical parameters and physiological responses of rainbow trout (Oncorhynchus mykiss) juveniles. Biology 2021, 10, 1040. [Google Scholar] [CrossRef] [PubMed]
- Zhang, S.; Li, C.Z.; Yang, Q.H.; Dong, X.H.; Chi, S.Y.; Liu, H.Y.; Shi, L.L.; Tan, B.P. Molecular cloning, characterization and expression analysis of Wnt4, Wnt5, Wnt6, Wnt7, Wnt10 and Wnt16 from Litopenaeus vannamei. Fish Shellfish Immunol. 2016, 54, 445–455. [Google Scholar] [CrossRef] [PubMed]
- Weng, S.; You, F.; Fan, Z.; Wang, L.; Wu, Z.; Zou, Y. Molecular cloning and sexually dimorphic expression of wnt4 in olive flounder (Paralichthys olivaceus). Fish Physiol. Biochem. 2016, 42, 1167–1176. [Google Scholar] [CrossRef] [PubMed]
- Schmitz, M.; Baekelandt, S.; Bequet, S.; Kestemont, P. Chronic hyperosmotic stress inhibits renal Toll-Like Receptors expression in striped catfish (Pangasianodon hypophthalmus, Sauvage) exposed or not to bacterial infection. Dev. Comp. Immunol. 2017, 73, 139–143. [Google Scholar] [CrossRef]
- Liu, F.; Su, B.; Fu, Q.; Shang, M.; Gao, C.; Tan, F.; Li, C. Identification, characterization and expression analysis of TLR5 in the mucosal tissues of turbot (Scophthalmus maximus L.) following bacterial challenge. Fish Shellfish Immunol. 2017, 68, 272–279. [Google Scholar] [CrossRef]
- Yang, J.; Yan, H. TLR5: Beyond the recognition of flagellin. Cell. Mol. Immunol. 2017, 14, 1017–1019. [Google Scholar] [CrossRef]
- Luo, S.; Deng, X.; Liu, Q.; Pan, Z.; Zhao, Z.; Zhou, L.; Luo, X. Emodin ameliorates ulcerative colitis by the flagellin-TLR5 dependent pathway in mice. Int. Immunopharmacol. 2018, 59, 269–275. [Google Scholar] [CrossRef]



| Samples a | Clean Reads | Clean Bases | GC Contents (%) | Clean Reads Q20 (%) b | Clean Reads Q30 (%) c |
|---|---|---|---|---|---|
| G20-1 | 33,618,558 | 10,069,190,306 | 49.64 | 98.66 | 95.70 |
| G20-2 | 30,542,362 | 9,149,565,768 | 49.59 | 98.63 | 95.63 |
| G20-3 | 30,626,059 | 9,174,115,482 | 49.82 | 98.59 | 95.51 |
| G30-1 | 29,118,900 | 8,723,118,946 | 49.98 | 98.65 | 95.68 |
| G30-2 | 30,326,490 | 9,084,622,756 | 50.10 | 98.64 | 95.62 |
| G30-3 | 34,209,823 | 10,247,870,004 | 49.86 | 98.61 | 95.59 |
| G40-1 | 26,628,061 | 7,974,715,550 | 50.08 | 98.49 | 95.35 |
| G40-2 | 29,931,391 | 8,964,260,212 | 49.86 | 98.63 | 95.72 |
| G40-3 | 22,273,974 | 6,672,988,308 | 49.68 | 98.60 | 95.63 |
| Category | Number | Percentage (%) |
|---|---|---|
| Total number of annotated genes | 23,972 | |
| Gene matches against GO | 14,525 | 60.59 |
| Gene matches against KEGG | 14,970 | 62.45 |
| Gene matches against KOG | 15,991 | 66.71 |
| Gene matches against Swiss-Prot | 16,505 | 68.85 |
| Gene matches against NR | 23,950 | 99.91 |
| Gene matches against Pfam | 21,503 | 89.70 |
| Gene matches against COG | 7382 | 30.79 |
| Gene matches against eggNOG | 22,838 | 95.27 |
| Gene Names | log2FC | Description |
|---|---|---|
| G30 vs. G20 | ||
| Seriola_dumerili_newGene_5930 | 2.80986 | protein NLRC3-like |
| Seriola_dumerili_newGene_10944 | 2.51744 | unnamed protein product |
| LOC111226299 | 2.19450 | C-type lectin domain family 4 member F-like |
| LOC111229131 | 2.14477 | growth/differentiation factor 8-like |
| frmd3 | 2.09078 | FERM domain-containing protein 3 |
| LOC111231723 | 1.99161 | uncharacterized protein LOC111231723 |
| lyve1 | 1.97051 | lymphatic vessel endothelial hyaluronic acid receptor 1 |
| LOC111239963 | 1.92871 | solute carrier family 12 member 3-like |
| slc5a6a | 1.81352 | solute carrier family 5 member 6a |
| Seriola_dumerili_newGene_13911 | 1.75102 | macrophage mannose receptor 1-like |
| LOC111240141 | −3.71184 | E3 ubiquitin-protein ligase TRIM21-like |
| Seriola_dumerili_newGene_14148 | −3.58766 | pol-like protein |
| LOC111223147 | −2.40705 | uncharacterized protein LOC111223147 |
| satb1a | −2.38908 | SATB homeobox 1a |
| ribc2 | −2.22092 | RIB43A domain with coiled-coils 2 |
| wnt7bb | −2.09191 | wingless-type MMTV integration site family, member 7Bb |
| LOC111228626 | −2.01800 | interferon-induced protein 44-like |
| trabd2a | −1.94610 | TraB domain containing 2A |
| Seriola_dumerili_newGene_12544 | −1.85641 | Retrotransposable element Tf2 protein type 1 |
| LOC111220383 | −1.70808 | gastrula zinc finger protein XlCGF57.1-like |
| G30 vs. G40 | ||
| Seriola_dumerili_newGene_10944 | 2.67769 | unnamed protein product |
| LOC111231293 | 1.50771 | C-X-C motif chemokine 10-like |
| LOC111219635 | 1.47708 | intraflagellar transport protein 140 homolog |
| slc4a4a | 1.24110 | solute carrier family 4 member 4a |
| cftr | 1.23478 | cystic fibrosis transmembrane conductance regulator |
| camk1a | 1.14853 | calcium/calmodulin-dependent protein kinase type 1-like |
| LOC111224523 | 1.01042 | kinesin-like protein KIF21A |
| map7d2b | 1.00381 | MAP7 domain containing 2b |
| LOC111235326 | −3.05319 | pleckstrin-like |
| LOC111220915 | −2.00153 | ladderlectin-like |
| LOC111235291 | −1.29715 | indoleamine 2,3-dioxygenase 2-like |
| LOC111240189 | −1.15190 | GTPase IMAP family member 7-like |
| sqle | −1.12522 | squalene monooxygenase |
| LOC111228808 | −1.08252 | C-X-C chemokine receptor type 4-like |
| Seriola_dumerili_newGene_4780 | −1.03479 | nuclear receptor subfamily 1 group D member 1-like |
| nr1d1 | −1.03175 | nuclear receptor subfamily 1, group d, member 1 |
| LOC111236503 | −1.01894 | granzyme A-like |
| Pathway ID | Pathway Term | Gene Name |
|---|---|---|
| G30 vs. G20 | ||
| ko04060 | Cytokine–cytokine receptor interaction | LOC111221262, LOC111222833, LOC111224301, LOC111225535, LOC111230808, LOC111231248, edar, tnfsf12 |
| ko04210 | Apoptosis | LOC111218420, LOC111225343, LOC111228846, LOC111230905, bcl2l11, pik3cb |
| ko00100 | Steroid biosynthesis | LOC111222971, ebp, lss, msmo1, nsdhl, sqle |
| ko04150 | mTOR signaling pathway | LOC111220460, LOC111228846, LOC111229726, LOC111238290, pik3cb, wnt4 |
| ko00860 | Porphyrin and chlorophyll metabolism | newGene_6606, LOC111217420, LOC111219041, LOC111226879, LOC111237667 |
| ko04020 | Calcium signaling pathway | newGene_6757, LOC111224354, LOC111230822, nos1, ptafr |
| ko04510 | Focal adhesion | LOC111228846, LOC111231768, itga9, pak5, pik3cb |
| ko04340 | Hedgehog signaling pathway | LOC111228096, evc, hhip, ptch1 |
| ko00240 | Pyrimidine metabolism | LOC111217958, LOC111221687, cda, cmpk2 |
| ko04060 | Cytokine–cytokine receptor interaction | LOC111221262, LOC111222833, LOC111224301, LOC111225535, LOC111230808, LOC111231248, edar, tnfsf12 |
| G30 vs. G40 | ||
| ko00380 | Tryptophan metabolism | LOC111235291 |
| ko04144 | Endocytosis | LOC111228808 |
| ko02010 | ABC transporters | cftr |
| ko04060 | Cytokine–cytokine receptor interaction | LOC111228808, LOC111231293 |
| ko04623 | Cytosolic DNA-sensing pathway | LOC111231293 |
| ko04672 | Intestinal immune network for IgA production | LOC111228808 |
| ko00100 | Steroid biosynthesis | sqle |
| ko04620 | Toll-like receptor signaling pathway | LOC111231293 |
| ko04622 | RIG-I-like receptor signaling pathway | LOC111231293 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Peng, Y.; Shi, H.; Liu, Y.; Huang, Y.; Zheng, R.; Jiang, D.; Jiang, M.; Zhu, C.; Li, G. RNA Sequencing Analysis Reveals Divergent Adaptive Response to Hypo- and Hyper-Salinity in Greater Amberjack (Seriola dumerili) Juveniles. Animals 2022, 12, 327. https://doi.org/10.3390/ani12030327
Peng Y, Shi H, Liu Y, Huang Y, Zheng R, Jiang D, Jiang M, Zhu C, Li G. RNA Sequencing Analysis Reveals Divergent Adaptive Response to Hypo- and Hyper-Salinity in Greater Amberjack (Seriola dumerili) Juveniles. Animals. 2022; 12(3):327. https://doi.org/10.3390/ani12030327
Chicago/Turabian StylePeng, Yuhao, Hongjuan Shi, Yuqi Liu, Yang Huang, Renchi Zheng, Dongneng Jiang, Mouyan Jiang, Chunhua Zhu, and Guangli Li. 2022. "RNA Sequencing Analysis Reveals Divergent Adaptive Response to Hypo- and Hyper-Salinity in Greater Amberjack (Seriola dumerili) Juveniles" Animals 12, no. 3: 327. https://doi.org/10.3390/ani12030327
APA StylePeng, Y., Shi, H., Liu, Y., Huang, Y., Zheng, R., Jiang, D., Jiang, M., Zhu, C., & Li, G. (2022). RNA Sequencing Analysis Reveals Divergent Adaptive Response to Hypo- and Hyper-Salinity in Greater Amberjack (Seriola dumerili) Juveniles. Animals, 12(3), 327. https://doi.org/10.3390/ani12030327






