Antimicrobial Resistance in Loggerhead Sea Turtles (Caretta caretta): A Comparison between Clinical and Commensal Bacterial Isolates
Abstract
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
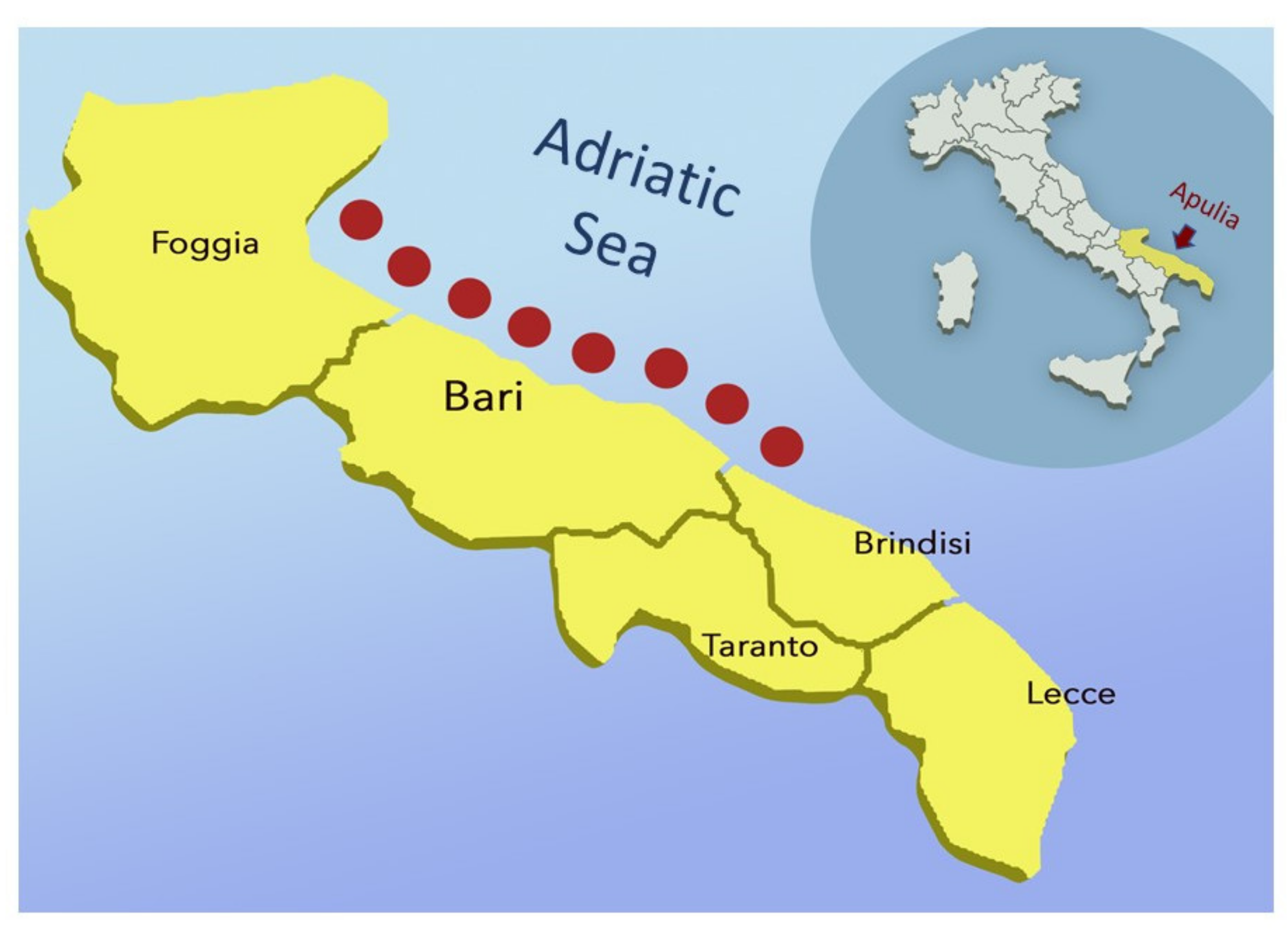
2.1. Turtles and Specimen Collection
2.2. Microbiological Analysis
2.3. Antimicrobial Susceptibility Test
2.4. Statistical Analysis
3. Results
3.1. Subsections
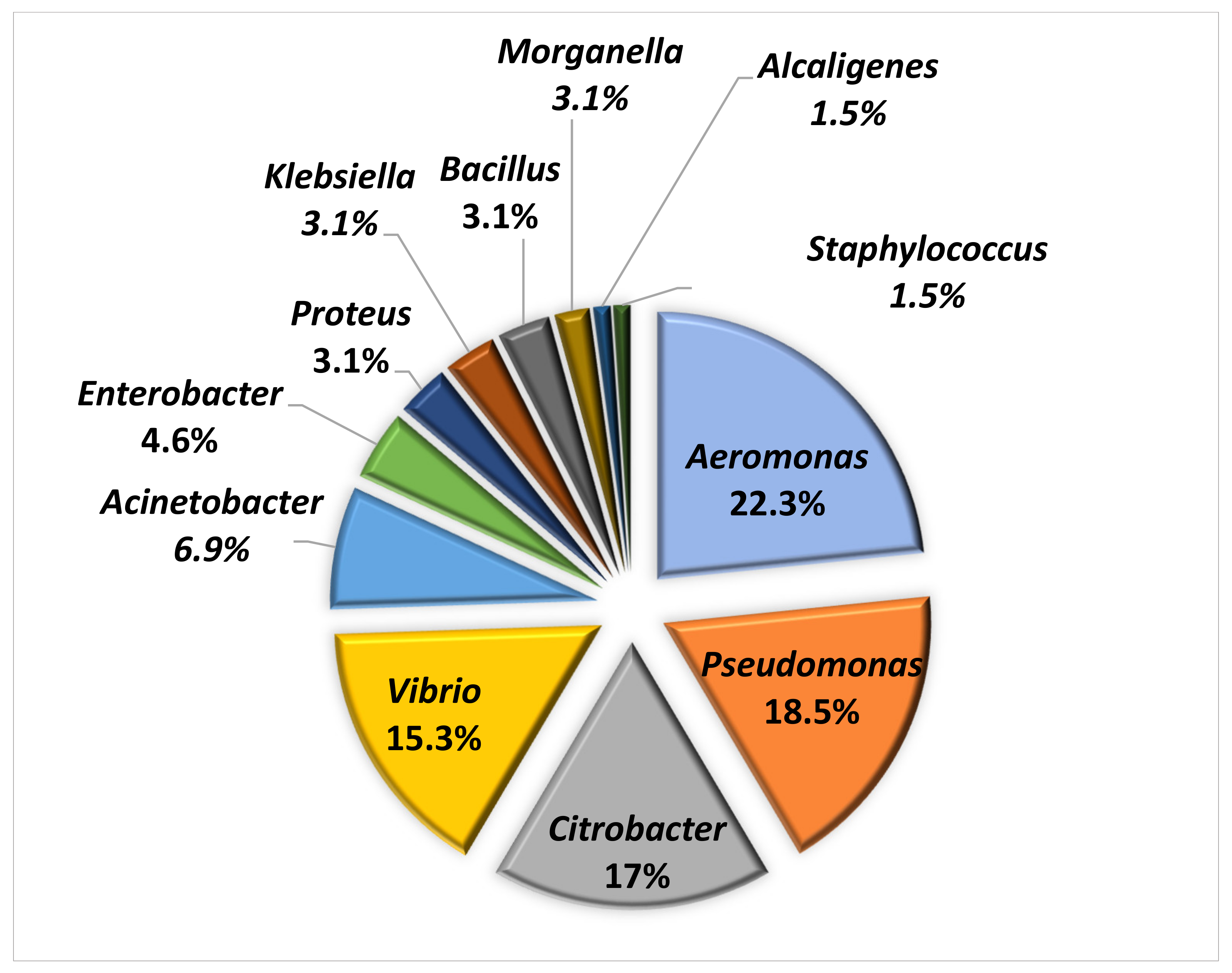
3.1.1. Identification of Bacterial Isolates from Oral and Cloacal Samples
3.1.2. Identification of Bacterial Isolates from Wound Samples
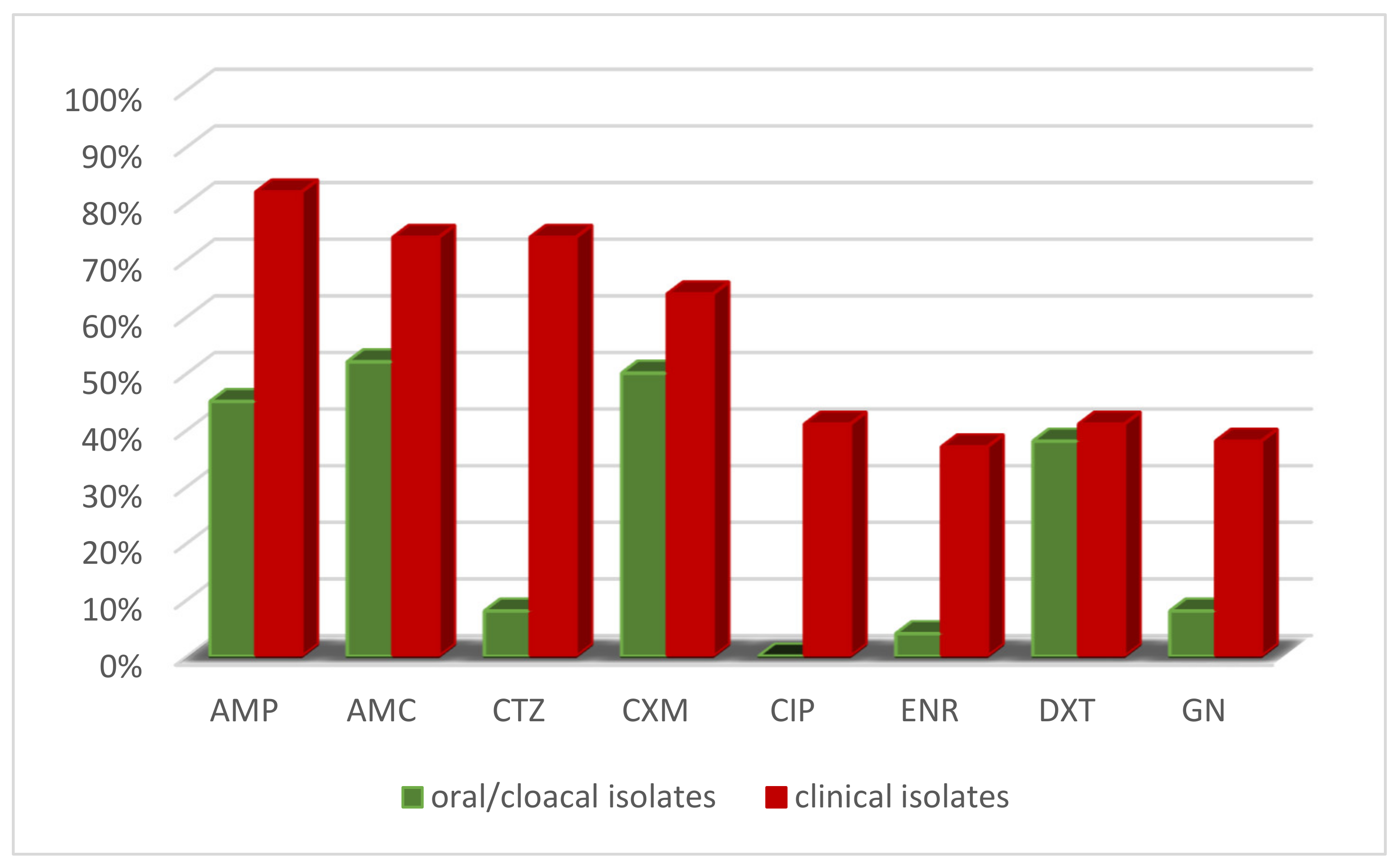
3.1.3. Summary of AMR Patterns
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Conflicts of Interest
References
- Holmes, A.H.; Moore, L.S.P.; Sundsfjord, A.; Steinbakk Sadie Regmi, M.; Karkey, A.; Guerin, P.J.; Piddock, L.J.V. Understanding the mechanisms and drivers of antimicrobial resistance. Lancet Infect. Dis. 2016, 387, 176–187. [Google Scholar] [CrossRef]
- Report World Health Organization. Antimicrobial Resistance: An Emerging Water, Sanitation and Hygiene Issue. 2014. Available online: https://www.who.int/water_sanitation_health/publications/antimicrobial-resistance/en/ (accessed on 13 May 2021).
- Corcoran, E.; Nellemann, C.; Baker, E.; Bos, R.; Osborn, D.; Savelli, H. Sick water? The Central Role of Wastewater Management in Sustainable Development. A rapid Response Assessment. United Nations Environment Programme, UN-Habitat, Grid-Arendal 2010. Available online: http://www.unwater.org/downloads/SickWater_unep_unh.pdf (accessed on 13 May 2020).
- Foti, M.; Giacopello, C.; Bottari, T.; Fisichella, V.; Rinaldo, D.; Mammina, C. Antibioitc resistance of gram negatives isoaltes from loggerhead sea turtles (Caretta caretta) in the Central Mediterrananean Sea. Mar. Pollut. Bull. 2009, 58, 1363–1366. [Google Scholar] [CrossRef]
- Di Cesare, A.; Vignaroli, C.; Luna, G.M.; Pasquaroli, S.; Biavasco, F. Antibiotic resistant enterococci in seawater and sediments from a costal fish farm. Microb. Drug Resist. 2012, 18, 502–509. [Google Scholar] [CrossRef] [PubMed]
- Stewart, J.R.; Townsend, E.; Lane, S.M.; Dyar, E.; Hohn, A.A.; Rowles, T.K.; Staggsd, L.A.; Wells, R.S.; Balmer, B.C.; Schwacje, L.H. Survey of antibiotic-resistant bacteria isolated from bottlenose dolphins Tursiops truncatus in the southeastern USA. Dis. Aquat. Organ. 2014, 108, 91–102. [Google Scholar] [CrossRef]
- Miranda, C.D.; Zemelman, R. Antibiotic resistant bacteria in fish from the Concepción Bay, Chile. Mar. Pollut. Bull. 2001, 42, 1006–1102. [Google Scholar] [CrossRef]
- Frazzon, A.P.J. Antibiotic resistant bacteria in free-living marine species. Vet. Record. 2017, 179, 648–649. [Google Scholar] [CrossRef]
- Brahmi, S.; Touati, A.; Dunyach-Remy, C.; Sotto, A.; Pantel, A.; Lavigne, J.P. High prevalence of extended-Spectrum β-lactamase-producing Enterobacteriaceae in wild fish from the Mediterranean Sea in Algeria. Microb. Drug Resist. 2018, 24, 290–298. [Google Scholar] [CrossRef] [PubMed]
- Candan, O.; Candan, E.D. Bacterial diversity of the green turtle (Chelonia mydas) nest environment. Sci. Total Environ. 2018, 10, 137717. [Google Scholar] [CrossRef]
- Senet, S. Mediterranean is Europe’s most waste-polluted sea. Journal de l’Environnement Reports 2019, 1–3. Available online: https://www.euractiv.com/section/energy-environment/news/mediterannean-is-europes-most-waste-polluted-sea-study-says/ (accessed on 15 February 2020).
- Maravić, M.; Skočibušić, M.; Cvjetan, S.; Šamanić, I.; Fredotović, Z.; Puizina, J. Prevalence and diversity of extended-spectrum-β-lactamase-producing Enterobacteriaceae from marine beach water. Mar. Pollut. Bull. 2015, 90, 60–67. [Google Scholar] [CrossRef] [PubMed]
- Maravić, M.; Skočibušić, M.; Šamanić, I.; Fredotović, Z.; Cvjetan, S.; Jutronić, M.; Puizina, J. Aeromonas spp. simultaneously harbouring blaCTX-M-15, blaSHV-12 and blaFOX-2, in wild-growing Mediterranean mussel (Mytilus galloprovincialis) from Adriatic Sea, Croatia. Int. J. Food Microbiol. 2013, 166, 301–308. [Google Scholar] [CrossRef]
- Gordon, L.; Giraud, E.; Ganière, J.P.; Armand, F.; Bouju-Albert, A.; de la Cotte, N.; Mangion, C.; Le Briùs, H. Antimicrobial resistance survey in a river receiving effluents from freshwater fish farms. J. Appl. Microbiol. 2007, 102, 1167–1176. [Google Scholar] [CrossRef] [PubMed]
- Galgani, F.; Claro, F.; Depledge, M.; Fossi, C. Monitoring the impact of litter in large vertebrates in the Mediterranean Sea within the European marine strategy framework directive (MSFD): Constraints, specificities and recommendations. Mar. Environ. Res. 2014, 100, 3–9. [Google Scholar] [CrossRef] [PubMed]
- Camacho, M.; Oros, J.; Boada, L.D.; Zaccaroni, A.; Silvi, M.; Formigaro, C.; López, P.; Zumbado, M.; Luzardo, O.P. Potential adverse effect of inorganic pollutants on clinical parameters of loggerhead sea turtles (Caretta caretta): Results from a nesting colony form Cape Verde, West Africa. Mar. Environ. Res. 2013, 92, 15–22. [Google Scholar] [CrossRef] [PubMed]
- Novillo, O.; Pertusa, J.F.; Tomás, J. Exploring the presence of pollutants of sea: Monitoring heavy metals in loggerhead turtles (Caretta caretta) from the Western Mediterranenan. Sci. Total Environ. 2017, 598, 1130–1139. [Google Scholar] [CrossRef]
- Casale, P.; Broderick, A.; Camiñas, J.; Cardona, L.; Carreras, C.; Demetropoulos, A.; Fuller, W.; Godley, B.; Hochscheid, S.; Kaska, Y.; et al. Mediterranean Sea turtles: Current knowledge and priorities for conservation and research. Endanger. Species Res. 2018, 36, 229–267. [Google Scholar] [CrossRef]
- Aguirre, A.A.; O’Hara, T.M.E.; Speaker, T.R.; Jessup, D.A. Monitoring the Health and Conservation of Marine Mammals, Sea Turtles, and Their Ecosystems In Conservation Medicine: Ecological Health in Practice; Aguirre, A.A., Ostfeld, R.S., Tabor, G.M., House, C., Pearl, M.C., Eds.; Oxford University Press: Oxford, UK, 2002; pp. 79–94. [Google Scholar]
- Caracappa, S.; Persichetti, M.F.; Piazza, A.; Caracappa, G.; Gentile, A.; Marineo, S.; Crucitti, D.; Arculeo, M. Incidental catch of loggerhead sea turtles (Caretta caretta) along the Sicilian coasts by longline fisheries. PeerJ 2018, 6, e5392. [Google Scholar] [CrossRef]
- Ciccarelli, S.; Valastro, C.; Di Bello, A.; Paci, S.; Caprio, F.; Corrente, M.L.; Trotta, A.; Franchini, D. Diagnosis and treatment of pulmonary disease in sea turtles (Caretta caretta). Animals 2020, 10, 1355. [Google Scholar] [CrossRef]
- Lazar, B.; Gračan, R. Ingestion of marine debris by loggerhead sea turtles, Caretta caretta, in the Adriatic Sea. Mar. Pollut. Bull. 2011, 62, 43–47. [Google Scholar] [CrossRef] [PubMed]
- Innis, C.; Nyaoke, A.C.; Williams, C.R., 3rd; Dunnigan, B.; Merigo, C.; Woodward, D.L.; Weber, E.S., 3rd; Frasca, S., Jr. Pathology and parasitology findings of cold-stunned Kemp’s ridley sea turtle (Lepidochelys kempii) stranded on Cape Cod, Massachusetts 2001–2006. J. Wildl. Dis. 2009, 45, 594–610. [Google Scholar] [CrossRef]
- Innis, C.J.; Frasca, S.J. Bacterial and fungal diseases. In Sea Turtles Health and Rehabilitation; Manire, C.A., Norton, T.N., Stacy, B.A., Innis, J.C., Harms, C.A., Eds.; J. Ross Publishing: Fort Lauderdale, FL, USA, 2017; pp. 779–790. [Google Scholar]
- Vega-Manriquez, D.X.; Dávila-Arellano, R.P.; Elsava-Campos, C.A.; Salazar Jiménez, E.; Negrete-Philipe, A.C.; Raigoza-figueras, R.; Muñoz-Tenería, F.A. Identification of bacteria present in ulcerative stomatitis lesions of captive sea turtles Chelonia mydas. Vet. Res. Commun. 2018, 42, 251–254. [Google Scholar] [CrossRef]
- Pace, A.; Dipineto, L.; Fioretti, A.; Hochscheid, S. Loggerhead Sea turtles as sentinel in the western Mediterranean: Antibiotic resistance and environment-related modifications of gram-negative bacteria. Mar. Pollut. Bull. 2019, 149, 110575. [Google Scholar] [CrossRef]
- Blasi, M.F.; Migliore, L.; Mattei, D.; Rotini, A.; Thaller, M.C.; Alduina, R. Antibiotic resistance of gram-negative bacteria from wild captured Loggerhead Sea turtles. Antibiotics 2020, 9, 162. [Google Scholar] [CrossRef] [PubMed]
- Al-Bahry, S.N.; Al-Zadjali, M.A.; Mahmoud, I.Y.; Elshafie, A.E. Biomonitoring marine habitats in reference to antibiotic resistant bacteria and ampicillin resistance determinants from oviductal fluid of the nesting green sea turtles, Chelonia mydas. Chemosphere 2012, 87, 1308–1315. [Google Scholar] [CrossRef] [PubMed]
- Ahasan, M.S.; Picard, J.; Elliott, L.; Kinobe, R.; Owens, L.; Ariel, E. Evidence of antibiotic resistance in Enterobacteriales isolated from green sea turtles, Chelonia mydas on the Great Barrier Reef. Mar. Pollut. Bull. 2017, 120, 18–27. [Google Scholar] [CrossRef] [PubMed]
- Trotta, A.; Cirilli, M.; Marinaro, M.; Bosak, S.; Diakoudi, G.; Ciccarelli, S.; Paci, S.; Buonavoglia, D.; Corrente, M. Detection of multi-drug resistance and AmpC β-lactamase/extended-spectrum β-lactamase genes in bacterial isolates of loggerhead sea turtles (Caretta caretta) from the Mediterranean Sea. Mar. Pollut. Bull. 2021, 164, 112015. [Google Scholar] [CrossRef] [PubMed]
- Harms, C.A.; Mihnovets, A.N.; McNeill, J.B.; Kelly, T.R.; Avens, L. Cloacal bacterial isolates and antimicrobial resistant patterns in juvenile loggerhead sea turtle (Caretta caretta) in Core Sound, North Carolina, USA. In Proceedings of the 26th International Symposium on Sea Turtles Conservation and Biology, Crete, Greece, 3–8 April 2006. [Google Scholar]
- Al-Bahry, S.N.; Mahmoud, I.Y.; Elshafie, A.E.; Al-Harthy, A.; Al-Ghafri, S.; Al-Amir, I.; Alkindi, A. Bacterial flora and antibiotic resistance from eggs of Green turtles Chelonia mydas: An indication of polluted effluents. Mar. Pollut. Bull. 2009, 58, 720–725. [Google Scholar] [CrossRef] [PubMed]
- Alduina, R.; Gambino, D.; Presentato, A.; Gentile, A.; Sucato, A.; Savoca, D.; Filippello, S.; Visconti, G.; Caracappa, G.; Vicari, D.; et al. Is Caretta Caretta a carrier of antibiotic resistance in the Mediterranean Sea? Antibiotics 2020, 9, 116. [Google Scholar] [CrossRef] [PubMed]
- Fichi, C.; Cardeti, G.; Cersini, A.; Mancusi, C.; Garducci, M.; Di Guardo, G.; Terracciano, G. Bacterial and viral pathogens detected in sea turtles stranded along the coast of Tuscany, Italy. Vet. Microbiol. 2016, 15, 56–61. [Google Scholar] [CrossRef]
- Casale, P.; Freggi, D.; Basso, R.; Argano, R. Size at male maturity, sexing methods and adult sex ratio in loggerhead turtles (Caretta caretta) from Italian waters investigated through tail measurements. Herpetol. J. 2005, 15, 145–148. [Google Scholar]
- Casale, P.; Lazar, B.; Pont, S.; Tomás, J.; Zizzo, N.; Alegre, F.; Badillo, J.; Di Summa, A.; Freggi, D.; Lackovic, G.; et al. Sex ratios of juvenile loggerhead sea turtles Caretta caretta in the Mediterranean Sea. Mar. Ecol. Prog. Ser. 2006, 324, 281–285. [Google Scholar] [CrossRef]
- Corrente, M.; Madio, A.; Friedrich, K.; Greco, G.; Desario, C.; Tagliabue, S.; D’Incau, M.; Campolo, M.; Buonavoglia, C. Isolation of Salmonella strains from reptile faeces and comparison of different culture media. J. Appl. Microbiol. 2004, 96, 709–715. [Google Scholar] [CrossRef]
- Khan, A.A.; Nawaz, M.S.; Khan, S.A.; Cerniglia, C.E. Detection of multidrug-resistant Salmonella typhimurium DT104 by multiplex polymerase chain reaction. FEMS Microbiol. Lett. 2000, 2, 355–360. [Google Scholar] [CrossRef]
- Stecher, B.; Chaffron, S.; Käppeli, R.; Hapfelmeier, S.; Freedrich, S.; Weber, T.C.; Kirundi, J.; Suar, M.; McCoy, K.D.; Von Mering, C.; et al. Like will to like: Abundance of closely related species can predict susceptibility to intestinal colonization by pathogenic and commensal bacteria. PLoS Pathog. 2010, 6, e1000711. [Google Scholar] [CrossRef] [PubMed]
- CLSI (Clinical and Laboratory Standards Institute). Performance Standards for Antimicrobial Disk and Dilution Susceptibility Tests for Bacteria Isolated from Animals, 4th ed.; CLSI document VET01-A4 and VET01-S2; CLSI: Wayne, PA, USA, 2018. [Google Scholar]
- Magiorakos, A.P.; Srinivasan, A.; Carey, R.B.; Carmeli, Y.; Falagas, M.E.; Giske, C.G.; Harbarth, S.; Hindler, J.F.; Kahlmeter, G.; Olsson-Liljequist, B.; et al. Multidrug-resistant, extensively drug-resistant and pandrug-resistant bacteria: An international expert proposal for interim standard definitions for acquired resistance. Clin. Microbiol. Infect. 2012, 18, 268–281. [Google Scholar] [CrossRef] [PubMed]
- Innis, C.J.; Braverman, J.M.; Cavin, M.L.; Ceresia, L.R.; Baden, L.R.; Kuhn, D.M.; Frasca, S., Jr.; McGowan, J.P.; Hirokawa, K.; Weber, E.S., 3rd; et al. Diagnosis and management of Enterococcus spp. infections during rehabilitation of cold-stunned Kemp’s ridley turtle (Lepidochelys kempii): 50 cases (2006–2012). J. Am. Vet. Med. Assoc. 2014, 245, 315–323. [Google Scholar] [CrossRef] [PubMed]
- Wyneken, J.; Burke, T.K.; Salmon, M.; Pederson, D.K. Egg failure in natural and relocated sea turtle nests. J. Herpetol. 1988, 22, 88–96. [Google Scholar] [CrossRef]
- Aguirre, A.A.; Balazs, G.H.; Zimmerman, B.; Spraker, T.R. Evaluation of Hawaiian green turtles (Chelonia mydas) for potential pathogens associated with fibropapillomas. J. Wildl. Dis. 1994, 30, 8–15. [Google Scholar] [CrossRef] [PubMed]
- Santoro, M.; Hernandez, G.; Caballer, M.; Garcia, F. Aerobic bacterial flora of nesting Green turtles (Chelonia mydas) from Tortuguero National Park, Costa Rica. J. Zoo Wildl. Med. 2006, 37, 549–552. [Google Scholar] [CrossRef] [PubMed]
- Innis, C.; Merigo, C.; Dodge, K.; Tlusty, M.; Dodge, M.; Sharp, B.; Myers, A.; McIntosh, A.; Wunn, D.; Perkins, C.; et al. Health evaluation of Leatherback turtles (Dermochelys coriacea) in the northwestern Adriatic during direct capturing and fisheries gear disentanglement. Chelonian Conserv. Biol. 2010, 9, 205–222. [Google Scholar] [CrossRef]
- Warwich, C.; Arena, P.C.; Steedman, C. Health implications associated with exposure to farmed and wild sea turtles. JRSM Short Rep. 2013, 4, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Sarmiento-Ramírez, J.M.; van der Voort, M.; Raaijmakers, J.M.; Diéguez-Uribeondo, J. Unravelling the microbiome of eggs of the endangered sea turtles Eretmochelys imbricata identifies bacteria with activity against the emerging pathogen Fusarium falciforme. PLoS ONE 2014, 9, e95206. [Google Scholar] [CrossRef]
- Santoro, H.; Hernandéz, G.; Caballero, M.; García, F. Potential bacterial pathogens carried by nesting leatherback turtles (Dermochelys coriacea) in Costa Rica. Chelonian Conserv. Biol. 2008, 7, 104–108. [Google Scholar] [CrossRef]
- Aguirre, A.A.; Luz, P. Marine turtles as sentinel of ecosystem health: Is fibropapillomatosis an indicator? Ecohealth 2004, 1, 275–283. [Google Scholar] [CrossRef]
- Raidal, S.R.; Ohara, M.; Hobbs, R.P.; Prince, R.I.T. Gram negative bacterial infections and cardiovascular parasitism in green sea turtles (Chelonia mydas). Aust. Vet. J. 1998, 76, 415–417. [Google Scholar] [CrossRef]
- Chuen-Im, T.M.; Areekijseree, M.; Chonthammakun, S.; Graham, S.V. Aerobic bacterial infections in captive juvenile green turtles (Chelonia mydas) and hawksbill turtles (Eretmochelys imbricata) from Thailand. Chelonian Conserv. Biol. 2010, 9, 135–142. [Google Scholar] [CrossRef]
- Pace, A.; Meomartino, L.; Affuso, A.; Mennonna, G.; Hochscheid, S.; Dipineto, L. Aeromonas indiced polyostotic osteomyelitis in a juvenile loggerhead sea turtle Caretta caretta. Dis. Aquat. Org. 2018, 132, 79–84. [Google Scholar] [CrossRef] [PubMed]
- Work, T.M.; Dagenais, J.; Stacy, B.A.; Ladner, J.T.; Lorch, J.M.; Balazs, G.H.; Barquero-Calvo, E.; Berlowski-Zier, B.M.; Breeden, R.; Corrales-Gómez, N.; et al. A novel host-adapted strain of Salmonella Thypimurium causes renal disease in olive ridley turtles (Lephdochelys olivacea) in the Pacific. Sci. Rep. 2019, 9, 9313. [Google Scholar] [CrossRef]
- Ferronato, B.O.; Marques, T.S.; Souza, F.L.; Verdade, L.M.; Matushima, E.R. Oral bacterial microbiota and traumatic injures of free-ranging Phrynops geoffroanus (Testudines, Chelide) in Southeastern Brazil. Phyllomedusa 2009, 8, 19–25. [Google Scholar] [CrossRef][Green Version]
- Glazebrook, J.S.; Campbell, R.S.F. A survey of the diseases of marine turtles in northern Australia. ΙΙ. Oceanarium-reared and wild turtles. Dis. Aquat. Org. 1990, 9, 97–104. [Google Scholar] [CrossRef]
- Fahlman, A.; Crespo-Picazo, J.L.; Sterba-Boatwright, B.; Stacy, B.A.; Garcia-Parraga, D. Defining risk variables causing gas embolism in loggerhead sea turtles Caretta caretta caught in trawls and gillnets. Sci. Rep. 2017, 7, 2739. [Google Scholar] [CrossRef] [PubMed]
- Parga, M.; Crespo-Picazo, J.L.; García-Párraga, D.; Stacy, B.A.; Harms, C.H. Fisheries and sea turtles. In Sea Turtles Health and Rehabilitation; Manire, C.A., Norton, T.N., Stacy, B.A., Innis, J.C., Harms, C.A., Eds.; J. Ross Publishing: Fort Lauderdale, FL, USA, 2017; Chapter XXXV; pp. 859–897. [Google Scholar]
- Rodríguez-Navarro, J.; Miró, E.; Brown-Jaque, M.; Hurtado, J.C.; Moreno, A.; Muniesa, M.; González-López, J.J.; Vila, J.; Espinal, P.; Navarro, F. Comparison of Commensal and Clinical Isolates for Diversity of Plasmids in Escherichia coli and Klebsiella pneumoniae. Antimicrob. Agents Chemother. 2020, 64, e02064-19. [Google Scholar] [CrossRef]
- ISPRA Guidelines (89/2013). Linee guida per il recupero, soccorso, affidamento e gestione delle tartarughe marine ai fine della riabilitazione e per la manipolazione a scopi scientifici. Available online: https://www.isprambiente.gov.it/en/publications/handbooks-and-guidelines/guidelines-for-the-recovery-rescue-and-management-of-sea-turtles-for-the-purposes-of-rehabilitation-and-for-scientific-purposes (accessed on 15 April 2021).
- Fernandes, M.; Grilo, M.L.; Carneiro, C.; Cunha, E.; Tavares, L.; Patino-Martinez, J.; Oliveira, M. Antibiotic Resistance and Virulence Profiles of Gram-Negative Bacteria Isolated from Loggerhead Sea Turtles (Caretta caretta) of the Island of Maio, Cape Verde. Antibiotics 2021, 10, 771. [Google Scholar] [CrossRef]
- Dandachi, I.; Chabo, S.; Daoud, Z.; Rolain, J.-M. Prevalence and Emergence of Extended-spectrum Cephalosporin-, Carbapenem- and Colistin resistant gram negative bacteria of animal origin in the Mediterranean Basin. Front. Microbiol. 2018, 9, 1–26. [Google Scholar] [CrossRef]
- Wang, J.; Ma, Z.-B.; Zeng, Z.-L.; Yand, X.-W.; Huang, Y.; Liu, J.-H. The role of wildlife (wild birds) in the global transmission of antimicrobial resistance gene. Zool. Res. 2017, 38, 55–80. [Google Scholar] [CrossRef]
- Maravić, M.; Skočibušić, M.; Fredotović, Z.; Šamanić, I.; Cvjetan, S.; Knezović, M.; Puizina, J. Urban River environment is a source of multi-drug resistant and ESBL-producing clinically important Acinetobacter spp. Environ. Sci. Pollut. Res. 2016, 23, 3525–3535. [Google Scholar] [CrossRef] [PubMed]
- Corum, O.; Altan, F.; Yildiz, R.; Ider, M.; Ok, M.; Uney, K. Pharmacokinetics of enrofloxacin and danofloxacin in premature calves. J. Vet. Pharmacol. Ther. 2019, 42, 624–631. [Google Scholar] [CrossRef]
- Sanders, C.C. Mechanisms responsible for cross-resistance and dichotomus resistance among quinolones. Clin. Infect. Dis. 2001, 32 (Suppl. S1), S1–S8. [Google Scholar] [CrossRef] [PubMed]
- Hopper, D.C.; Jacoby, G.A. Mechanisms of drug resistance: Quinolone resistance. Ann. N. Y. Acad. Sci. 2015, 1353, 12–31. [Google Scholar] [CrossRef] [PubMed]
- Chuen-Im, T.; Sawetsuwannakun, K.; Neesanant, P.; Kitkumthorn, N. Antibiotic-Resistant Bacteria in Green Turtle (Chelonia mydas) Rearing Seawater. Animals 2021, 11, 1841. [Google Scholar] [CrossRef]
- Drane, K.; Huerlimann, R.; Power, M.; Whelan, A.; Ariel, E.; Sheehan, M.; Kinobe, R. Testudines as Sentinels for Monitoring the Dissemination of Antibiotic Resistance in Marine Environments: An Integrative Review. Antibiotics 2021, 10, 775. [Google Scholar] [CrossRef] [PubMed]

| Juvenile | Subadult | Adult | |
|---|---|---|---|
| Number of subjects | 100 | 76 | 24 |
| CCL a | 31 ± 9.2 | 53 ± 2.3 | 74 ± 10.3 |
| Sex b | / | / | 11 males/13 females |
| Weight c | 22 ± 8.7 | 37 ± 6.3 | 45 ± 2.4 |
| Healthy animals | 70 | 23 | 5 |
| Injured animals | 30 | 53 | 19 |
| Antibiotics | Phenotypic Profiles in 410 Isolates | Percentage of Resistance in Oral/Cloacal and Clinical Isolates | ||
|---|---|---|---|---|
| R | R (%) | O/C Isolates (280) R (%) | Clinical Isolates (130) R (%) | |
| AMP | 233 * | 56.8 ** | 45 | 82 |
| AMC | 242 | 59 | 52 | 73.8 |
| CTZ | 300 | 73.1 | 7.8 | 73.8 |
| CXM | 223 | 54.3 | 50 | 63.8 |
| CIP | 53 | 12.9 | 0 | 40.7 |
| ENR | 59 | 14.3 | 3.9 | 36.9 |
| DXT | 161 | 39.2 | 38.5 | 40.7 |
| GN | 72 | 17.5 | 7.8 | 38.4 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Trotta, A.; Marinaro, M.; Sposato, A.; Galgano, M.; Ciccarelli, S.; Paci, S.; Corrente, M. Antimicrobial Resistance in Loggerhead Sea Turtles (Caretta caretta): A Comparison between Clinical and Commensal Bacterial Isolates. Animals 2021, 11, 2435. https://doi.org/10.3390/ani11082435
Trotta A, Marinaro M, Sposato A, Galgano M, Ciccarelli S, Paci S, Corrente M. Antimicrobial Resistance in Loggerhead Sea Turtles (Caretta caretta): A Comparison between Clinical and Commensal Bacterial Isolates. Animals. 2021; 11(8):2435. https://doi.org/10.3390/ani11082435
Chicago/Turabian StyleTrotta, Adriana, Mariarosaria Marinaro, Alessio Sposato, Michela Galgano, Stefano Ciccarelli, Serena Paci, and Marialaura Corrente. 2021. "Antimicrobial Resistance in Loggerhead Sea Turtles (Caretta caretta): A Comparison between Clinical and Commensal Bacterial Isolates" Animals 11, no. 8: 2435. https://doi.org/10.3390/ani11082435
APA StyleTrotta, A., Marinaro, M., Sposato, A., Galgano, M., Ciccarelli, S., Paci, S., & Corrente, M. (2021). Antimicrobial Resistance in Loggerhead Sea Turtles (Caretta caretta): A Comparison between Clinical and Commensal Bacterial Isolates. Animals, 11(8), 2435. https://doi.org/10.3390/ani11082435







