Proteomic Analysis of the Secretome and Exosomes of Feline Adipose-Derived Mesenchymal Stem Cells
Abstract
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Isolation, Expansion and Characterization of fAd-MSCs
2.2. Production of the fAd-MSC Secretome
2.3. Isolation of Exosomes from fAd-MSCs
2.4. Transmission Electron Microscopy
2.5. Size Distribution and Electronegativity of Exosomes
2.6. Expression of Exosomal Markers
2.7. Proteomic Analysis by UHPLC-HRMS
2.8. Bioinformatic Analysis
3. Results
3.1. Characterization of fAd-MSCs
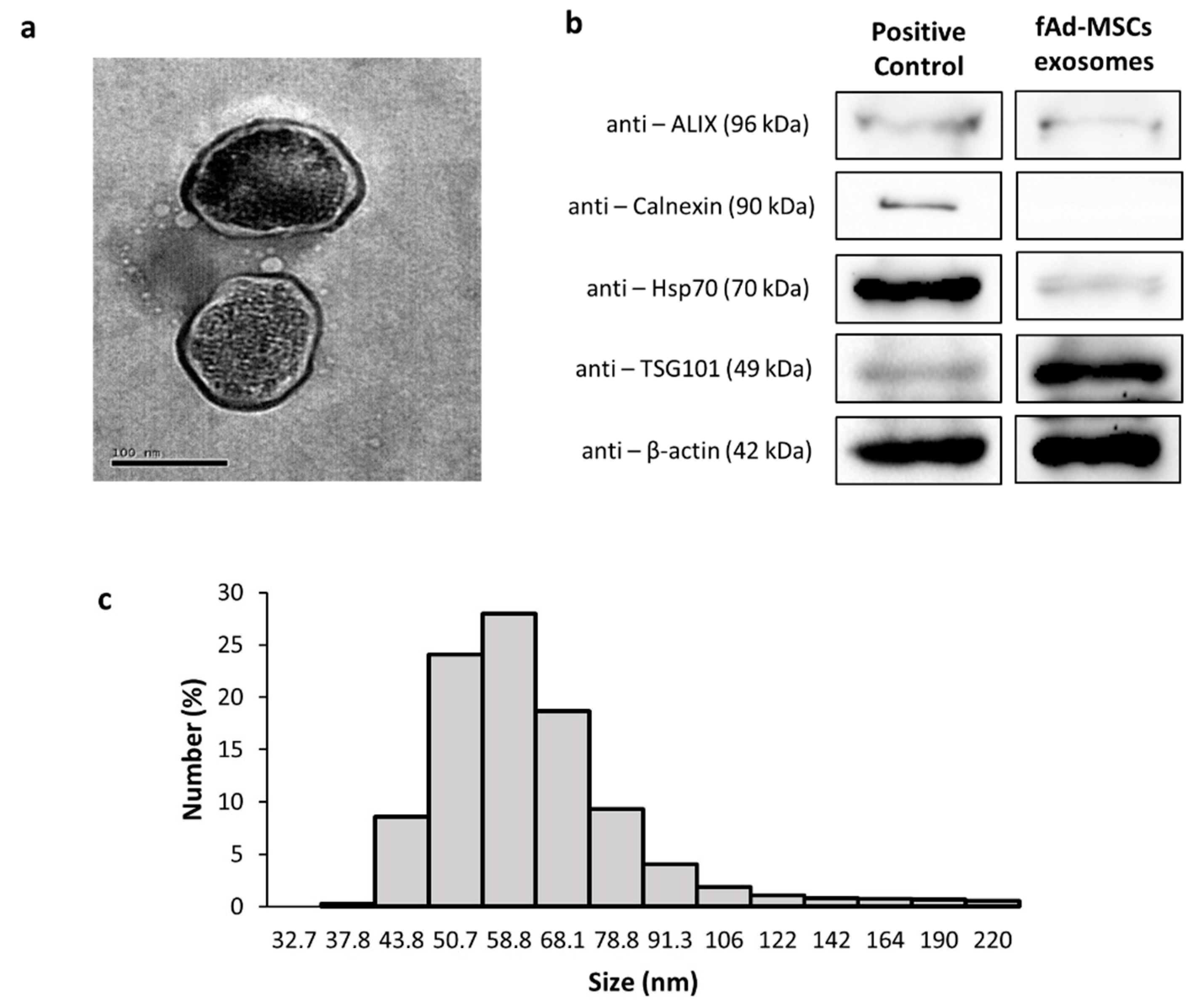
3.2. Characterization of the Exosomes of fAd-MSCs
3.3. Proteomic Analysis of the fAd-MSCs Secretome and fAd-MSCs Exosomes
3.4. Functional Annotation Based on Gene Ontology (GO)
3.5. Interactions between Differentially Expressed Proteins
3.6. KEGG Biological Pathway Analysis of Differentially Expressed Proteins
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Arzi, B.; Clark, K.C.; Sundaram, A.; Spriet, M.; Verstraete, F.J.M.; Walker, N.J.; Loscar, M.R.; Fazel, N.; Murphy, W.J.; Vapniarsky, N.; et al. Therapeutic Efficacy of Fresh, Allogeneic Mesenchymal Stem Cells for Severe Refractory Feline Chronic Gingivostomatitis. Stem. Cells Transl. Med. 2017, 6, 1710–1722. [Google Scholar] [CrossRef]
- Quimby, J.M.; Borjesson, D.L. Mesenchymal stem cell therapy in cats: Current knowledge and future potential. J. Feline Med. Surg. 2018, 20, 208–216. [Google Scholar] [CrossRef] [PubMed]
- Villatoro, A.J.; Claros, S.; Fernandez, V.; Alcoholado, C.; Farinas, F.; Moreno, A.; Becerra, J.; Andrades, J.A. Safety and efficacy of the Mesenchymal Stem Cell in Feline Eosinophilic Keratitis Treatment. BMC Vet. Res. 2018, 14, 116. [Google Scholar] [CrossRef] [PubMed]
- Martin, D.R.; Cox, N.R.; Hathcock, T.L.; Niemeyer, G.P.; Baker, H.J. Isolation and Characterization of Multipotential Mesenchymal Stem Cells from Feline Bone Marrow. Exp. Hematol. 2002, 30, 879–886. [Google Scholar] [CrossRef]
- Webb, T.L.; Quimby, J.M.; Dow, S.W. In Vitro Comparison of Feline Bone Marrow-Derived and Adipose Tissue-Derived Mesenchymal Stem Cells. J. Feline Med. Surg. 2012, 14, 165–168. [Google Scholar] [CrossRef] [PubMed]
- Sato, K.; Yamawaki-Ogata, A.; Kanemoto, I.; Usui, A.; Narita, Y. Isolation and Characterisation of Peripheral Blood-Derived Feline Mesenchymal Stem Cells. Vet. J. 2016, 216, 183–188. [Google Scholar] [CrossRef] [PubMed]
- Iacono, E.; Cunto, M.; Zambelli, D.; Ricci, F.; Tazzari, P.L.; Merlo, B. Could Fetal Fluid and Membranes Be an Alternative Source for Mesenchymal Stem Cells (MSCs) in the Feline Species? A Preliminary Study. Vet. Res. Commun. 2012, 36, 107–118. [Google Scholar] [CrossRef]
- Quimby, J.M.; Webb, T.L.; Habenicht, L.M.; Dow, S.W. Safety and Efficacy of Intravenous Infusion of Allogeneic Cryopreserved Mesenchymal Stem Cells for Treatment of Chronic Kidney Disease in Cats: Results of Three Sequential Pilot Studies. Stem. Cell Res. Ther. 2013, 4, 48. [Google Scholar] [CrossRef]
- Trzil, J.E.; Masseau, I.; Webb, T.L.; Chang, C.H.; Dodam, J.R.; Cohn, L.A.; Liu, H.; Quimby, J.M.; Dow, S.W.; Reinero, C.R. Long-Term Evaluation of Mesenchymal Stem Cell Therapy in a Feline Model of Chronic Allergic Asthma. Clin. Exp. Allergy 2014, 44, 1546–1557. [Google Scholar] [CrossRef]
- Webb, T.L.; Webb, C.B. Stem Cell Therapy in Cats with Chronic Enteropathy: A Proof-Of-Concept Study. J. Feline Med. Surg. 2015, 17, 901–908. [Google Scholar] [CrossRef]
- Carrade, D.D.; Borjesson, D.L. Immunomodulation by Mesenchymal Stem Cells in Veterinary Species. Comp. Med. 2013, 63, 207–217. [Google Scholar] [PubMed]
- Gómez, M.C.; Qin, Q.; Biancardi, M.N.; Galiguis, J.; Dumas, C.; Wang, G.; Pope, C.E. 340 Biological Characteristics and Functional Capability of Feline Adipose Tissue-Derived Mesenchymal Stem Cells. Reprod. Fertil. Dev. 2015, 27, 258. [Google Scholar] [CrossRef]
- Kono, S.; Kazama, T.; Kano, K.; Harada, K.; Uechi, M.; Matsumoto, T. Phenotypic and Functional Properties of Feline Dedifferentiated Fat Cells and Adipose-Derived Stem. Cells Vet. J. 2014, 199, 88–96. [Google Scholar] [CrossRef]
- An, J.H.; Song, W.J.; Li, Q.; Kim, S.M.; Yang, J.I.; Ryu, M.O.; Nam, A.R.; Bhang, D.H.; Jung, Y.C.; Youn, H.Y. Prostaglandin E2 Secreted from Feline Adipose Tissue-Derived Mesenchymal Stem Cells Alleviate DSS-Induced Colitis by Increasing Regulatory T Cells in Mice. BMC Vet. Res. 2018, 14, 354. [Google Scholar] [CrossRef] [PubMed]
- Taechangam, N.; Iyer, S.S.; Walker, N.J.; Arzi, B.; Borjesson, D.L. Mechanisms Utilized by Feline Adipose-Derived Mesenchymal Stem Cells to Inhibit T Lymphocyte Proliferation. Stem. Cell Res. Ther. 2019, 10. [Google Scholar] [CrossRef]
- Clark, K.C.; Fierro, F.A.; Ko, E.M.; Walker, N.J.; Arzi, B.; Tepper, C.G.; Dahlenburg, H.; Cicchetto, A.; Kol, A.; Marsh, L.; et al. Human and Feline Adipose-Derived Mesenchymal Stem Cells have Comparable Phenotype, Immunomodulatory Functions, and Transcriptome. Stem. Cell Res. Ther. 2017, 8, 69. [Google Scholar] [CrossRef]
- Villatoro, A.J.; Alcoholado, C.; Martin-Astorga, M.C.; Fernandez, V.; Cifuentes, M.; Becerra, J. Comparative Analysis and Characterization of Soluble Factors and Exosomes from Cultured Adipose Tissue and Bone Marrow Mesenchymal Stem Cells in Canine Species. Vet. Immunol. Immunopathol. 2019, 208, 6–15. [Google Scholar] [CrossRef]
- Bang, C.; Thum, T. Exosomes: New Players in Cell-Cell Communication. Int. J. Biochem. Cell Biol. 2012, 44, 2060–2064. [Google Scholar] [CrossRef]
- Lener, T.; Gimona, M.; Aigner, L.; Borger, V.; Buzas, E.; Camussi, G.; Chaput, N.; Chatterjee, D.; Court, F.A.; Del Portillo, H.A.; et al. Applying Extracellular Vesicles Based Therapeutics in Clinical Trials—An ISEV Position Paper. J. Extracell. Vesicles 2015, 4, 30087. [Google Scholar] [CrossRef]
- Konala, V.B.; Mamidi, M.K.; Bhonde, R.; Das, A.K.; Pochampally, R.; Pal, R. The Current Landscape of the Mesenchymal Stromal Cell Secretome: A New Paradigm for Cell-Free Regeneration. Cytotherapy 2016, 18, 13–24. [Google Scholar] [CrossRef]
- Bjorge, I.M.; Kim, S.Y.; Mano, J.F.; Kalionis, B.; Chrzanowski, W. Extracellular Vesicles, Exosomes and Shedding Vesicles in Regenerative Medicine—A New Paradigm For Tissue Repair. Biomater. Sci. 2017, 6, 60–78. [Google Scholar] [CrossRef] [PubMed]
- Phinney, D.G.; Pittenger, M.F. Concise Review: MSC-Derived Exosomes for Cell-Free Therapy. Stem. Cells 2017, 35, 851–858. [Google Scholar] [CrossRef] [PubMed]
- Harrell, C.; Fellabaum, C.; Arsenijevic, A.; Markovic, B.; Djonov, V.; Volarevic, V. Therapeutic Potential of Mesenchymal Stem Cells and Their Secretome in the Treatment of Glaucoma. Stem. Cells Int. 2019, 2019, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Fernandes-Cunha, G.M.; Na, K.S.; Putra, I.; Lee, H.J.; Hull, S.; Cheng, Y.C.; Blanco, I.J.; Eslani, M.; Djalilian, A.R.; Myung, D. Corneal Wound Healing Effects of Mesenchymal Stem Cell Secretome Delivered Within a Viscoelastic Gel Carrier. Stem. Cells Transl. Med. 2019, 8, 478–489. [Google Scholar] [CrossRef] [PubMed]
- Cho, B.S.; Kim, J.O.; Ha, D.H.; Yi, Y.W. Exosomes Derived from Human Adipose Tissue-Derived Mesenchymal Stem Cells Alleviate Atopic Dermatitis. Stem. Cell Res. Ther. 2018, 9, 187. [Google Scholar] [CrossRef] [PubMed]
- Elahi, F.M.; Farwell, D.G.; Nolta, J.A.; Anderson, J.D. Preclinical Translation of Exosomes Derived from Mesenchymal Stem/Stromal Cells. Stem. Cells 2020, 38, 15–21. [Google Scholar] [CrossRef] [PubMed]
- Mocchi, M.; Dotti, S.; Del Bue, M.; Villa, R.; Bari, E.; Perteghella, S.; Torre, M.L.; Grolli, S. Veterinary Regenerative Medicine for Musculoskeletal Disorders: Can Mesenchymal Stem/Stromal Cells and Their Secretome Be the New Frontier? Cells 2020, 9, 1453. [Google Scholar] [CrossRef]
- Lange-Consiglio, A.; Rossi, D.; Tassan, S.; Perego, R.; Cremonesi, F.; Parolini, O. Conditioned Medium from Horse Amniotic Membrane-Derived Multipotent Progenitor Cells: Immunomodulatory Activity in Vitro and First Clinical Application in Tendon and Ligament Injuries In Vivo. Stem. Cells Dev. 2013, 22, 3015–3024. [Google Scholar] [CrossRef]
- El-Tookhy, O.S.; Shamaa, A.A.; Shehab, G.G.; Abdallah, A.N.; Azzam, O.M. Histological Evaluation of Experimentally Induced Critical Size Defect Skin Wounds Using Exosomal Solution of Mesenchymal Stem Cells Derived Microvesicles. Int. J. Stem. Cells 2017, 10, 144–153. [Google Scholar] [CrossRef]
- Klymiuk, M.C.; Balz, N.; Elashry, M.I.; Heimann, M.; Wenisch, S.; Arnhold, S. Exosomes Isolation and Identification from Equine Mesenchymal Stem Cells. BMC Vet. Res. 2019, 15, 42. [Google Scholar] [CrossRef]
- Baglio, S.R.; Rooijers, K.; Koppers-Lalic, D.; Verweij, F.J.; Perez Lanzon, M.; Zini, N.; Naaijkens, B.; Perut, F.; Niessen, H.W.; Baldini, N.; et al. Human Bone Marrow- and Adipose-Mesenchymal Stem Cells Secrete Exosomes Enriched in Distinctive Mirna and t-RNA Species. Stem. Cell Res. Ther. 2015, 6, 127. [Google Scholar] [CrossRef] [PubMed]
- Villatoro, A.J.; Alcoholado, C.; Martín-Astorga, M.C.; Rico, G.; Fernández, V.; Becerra, J. Characterization of the Secretory Profile and Exosomes of Limbal Stem Cells in the Canine Species. PLoS ONE 2020, 15, e0244327. [Google Scholar] [CrossRef] [PubMed]
- Simpson, R.J.; Lim, J.W.; Moritz, R.L.; Mathivanan, S. Exosomes: Proteomic Insights and Diagnostic Potential. Expert Rev. Proteom. 2009, 6, 267–283. [Google Scholar] [CrossRef] [PubMed]
- Simpson, R.J.; Jensen, S.S.; Lim, J.W. Proteomic Profiling of Exosomes: Current Perspectives. Proteomics 2008, 8, 4083–4099. [Google Scholar] [CrossRef]
- Kupcova Skalnikova, H. Proteomic Techniques for Characterization of Mesenchymal Stem Cell Secretome. Biochimie 2013, 95, 2196–2211. [Google Scholar] [CrossRef]
- Hoffman, A.M.; Dow, S.W. Concise Review: Stem Cell Trials Using Companion Animal Disease Models. Stem. Cells 2016, 34, 1709–1729. [Google Scholar] [CrossRef]
- Galipeau, J.; Krampera, M.; Barrett, J.; Dazzi, F.; Deans, R.J.; DeBruijn, J.; Dominici, M.; Fibbe, W.E.; Gee, A.P.; Gimble, J.M.; et al. International Society for Cellular Therapy Perspective on Immune Functional Assays for Mesenchymal Stromal Cells as Potency Release Criterion for Advanced Phase Clinical Trials. Cytotherapy 2016, 18, 151–159. [Google Scholar] [CrossRef]
- Dominici, M.; Le Blanc, K.; Mueller, I.; Slaper-Cortenbach, I.; Marini, F.; Krause, D.; Deans, R.; Keating, A.; Prockop, D.; Horwitz, E. Minimal Criteria for Defining Multipotent Mesenchymal Stromal Cells. The International Society for Cellular Therapy Position Statement. Cytotherapy 2006, 8, 315–317. [Google Scholar] [CrossRef]
- Villatoro, A.; Martín-Astorga, M.; Alcoholado, C.; Becerra, J. Canine Colostrum Exosomes: Characterization and Influence on the Canine Mesenchymal Stem Cell Secretory Profile and Fibroblast Anti-Oxidative Capacity. BMC Vet. Res. 2020, 16. [Google Scholar] [CrossRef]
- Lotvall, J.; Hill, A.F.; Hochberg, F.; Buzas, E.I.; Di Vizio, D.; Gardiner, C.; Gho, Y.S.; Kurochkin, I.V.; Mathivanan, S.; Quesenberry, P.; et al. Minimal Experimental Requirements for Definition of Extracellular Vesicles and Their Functions: A Position Statement from The International Society For Extracellular Vesicles. J. Extracell. Vesicles 2014, 3, 26913. [Google Scholar] [CrossRef]
- Käll, L.; Canterbury, J.; Weston, J.; Noble, W.; MacCoss, M. Semi-Supervised Learning for Peptide Identification from Shotgun Proteomics Datasets. Nat. Methods 2007, 4, 923–925. [Google Scholar] [CrossRef] [PubMed]
- Haraszti, R.A.; Didiot, M.C.; Sapp, E.; Leszyk, J.; Shaffer, S.A.; Rockwell, H.E.; Gao, F.; Narain, N.R.; DiFiglia, M.; Kiebish, M.A.; et al. High-Resolution Proteomic and Lipidomic Analysis of Exosomes and Microvesicles from Different Cell Sources. J. Extracell. Vesicles 2016, 5, 32570. [Google Scholar] [CrossRef] [PubMed]
- Schey, K.L.; Luther, J.M.; Rose, K.L. Proteomics Characterization of Exosome Cargo. Methods 2015, 87, 75–82. [Google Scholar] [CrossRef] [PubMed]
- STRING: Functional Protein Association Networks. Available online: https://string-db.org (accessed on 13 May 2020).
- Bader, G.; Hogue, C. An Automated Method for Finding Molecular Complexes in Large Protein Interaction Networks. BMC Bioinform. 2003, 4, 2. [Google Scholar] [CrossRef]
- Bindea, G.; Mlecnik, B.; Hackl, H.; Charoentong, P.; Tosolini, M.; Kirilovsky, A.; Fridman, W.H.; Pagès, F.; Trajanoski, Z.; Galon, J. Cluego: A Cytoscape Plug-In to Decipher Functionally Grouped Gene Ontology and Pathway Annotation Networks. Bioinformatics 2009, 25, 1091–1093. [Google Scholar] [CrossRef] [PubMed]
- Tran, C.; Damaser, M.S. Stem Cells as Drug Delivery Methods: Application of Stem Cell Secretome for Regeneration. Adv. Drug Deliv. Rev. 2015, 82, 1–11. [Google Scholar] [CrossRef]
- Phan, J.; Kumar, P.; Hao, D.; Gao, K.; Farmer, D.; Wang, A. Engineering Mesenchymal Stem Cells to Improve Their Exosome Efficacy and Yield for Cell-Free Therapy. J. Extracell. Vesicles 2018, 7, 1522236. [Google Scholar] [CrossRef]
- Walker, L.C.; Overstreet, M.A.; Siddiqui, A.; De Paepe, A.; Ceylaner, G.; Malfait, F.; Symoens, S.; Atsawasuwan, P.; Yamauchi, M.; Ceylaner, S.; et al. A Novel Mutation in the Lysyl Hydroxylase 1 Gene Causes Decreased Lysyl Hydroxylase Activity in an Ehlers-Danlos VIA Patient. J. Invest. Dermatol. 2005, 124, 914–918. [Google Scholar] [CrossRef]
- Barski, O.A.; Tipparaju, S.M.; Bhatnagar, A. The Aldo-Keto Reductase Superfamily and its Role in Drug Metabolism and Detoxification. Drug Metab. Rev. 2008, 40, 553–624. [Google Scholar] [CrossRef]
- Osaki, M.; Oshimura, M.; Ito, H. PI3K-Akt Pathway: Its Functions and Alterations in Human Cancer. Apoptosis 2004, 9, 667–676. [Google Scholar] [CrossRef]
- Kanazu, T.; Shinoda, M.; Nakayama, T.; Deyashiki, Y.; Hara, A.; Sawada, H. Aldehyde Reductase is a Major Protein Associated with 3-Deoxyglucosone Reductase Activity in Rat; Pig and Human Livers. Biochem. J. 1991, 279, 903–906. [Google Scholar] [CrossRef] [PubMed]
- Stetler, R.A.; Gan, Y.; Zhang, W.; Liou, A.K.; Gao, Y.; Cao, G.; Chen, J. Heat Shock Proteins: Cellular and Molecular Mechanisms in the CNS. Prog. Neurobiol. 2010, 92, 184–211. [Google Scholar] [CrossRef] [PubMed]
- Drees, F.; Pokutta, S.; Yamada, S.; Nelson, W.J.; Weis, W.I. Alpha-Catenin is a Molecular Switch that Binds E-Cadherin-Beta-Catenin and Regulates Actin-Filament Assembly. Cell 2005, 123, 903–915. [Google Scholar] [CrossRef]
- Bathgate, R.; Halls, M.; van der Westhuizen, E.; Callander, G.; Kocan, M.; Summers, R. Relaxin Family Peptides and Their Receptors. Physiol. Rev. 2013, 93, 405–480. [Google Scholar] [CrossRef] [PubMed]
- Bajic, G.; Degn, S.; Thiel, S.; Andersen, G. Complement Activation; Regulation; and Molecular Basis for Complement-Related Diseases. EMBO J. 2015, 34, 2735–2757. [Google Scholar] [CrossRef]
- Karnoub, A.; Weinberg, R. Ras Oncogenes: Split Personalities. Nat. Rev. Mol. Cell Biol. 2008, 9, 517–531. [Google Scholar] [CrossRef]
- Engelman, J.; Luo, J.; Cantley, L. The Evolution of Phosphatidylinositol 3-Kinases as Regulators of Growth and Metabolism. Nat. Rev. Genet. 2006, 7, 606–619. [Google Scholar] [CrossRef]
- Zong, H.; Ward, M.; Stitt, A. AGEs, RAGE, and Diabetic Retinopathy. Curr. Diab. Rep. 2011, 11, 244–252. [Google Scholar] [CrossRef]
- Yamagishi, S. Role of Advanced Glycation End Products (Ages) and Receptor for Ages (RAGE) in Vascular Damage in Diabetes. Exp. Gerontol. 2011, 46, 217–224. [Google Scholar] [CrossRef]
- Tremblay, A.; Camargo, F. Hippo Signaling in Mammalian Stem Cells. Semin. Cell Dev. Biol. 2012, 23, 818–826. [Google Scholar] [CrossRef]
- Nikitovic, D.; Katonis, P.; Tsatsakis, A.; Karamanos, N.; Tzanakakis, G. Lumican; A Small Leucine-Rich Proteoglycan. IUBMB Life 2008, 60, 818–823. [Google Scholar] [CrossRef] [PubMed]
- van Buul, J.; Hordijk, P. Signaling in Leukocyte Transendothelial Migration. Arterioscler. Thromb. Vasc. Biol. 2004, 24, 824–833. [Google Scholar] [CrossRef] [PubMed]
- Määttänen, P.; Gehring, K.; Bergeron, J.; Thomas, D. Protein Quality Control in the ER: The Recognition of Misfolded Proteins. Semin. Cell Dev. Biol. 2010, 21, 500–511. [Google Scholar] [CrossRef] [PubMed]
- Stuart, L.; Ezekowitz, R. Phagocytosis: Elegant complexity. Immunity 2005, 22, 539–550. [Google Scholar] [CrossRef] [PubMed]
- Available online: https://www.clinicaltrials.gov (accessed on 20 December 2020).
- Whitford, W.; Guterstam, P. Exosome Manufacturing Status. Future Med. Chem. 2019, 11, 1225–1236. [Google Scholar] [CrossRef] [PubMed]
- Labriola, N.R.; Azagury, A.; Gutierrez, R.; Mathiowitz, E.; Darling, E.M. Concise Review: Fabrication, Customization, and Application of Cell Mimicking Microparticles in Stem Cell Science. Stem. Cells Transl. Med. 2018, 7, 232–240. [Google Scholar] [CrossRef]
- Bari, E.; Perteghella, S.; Di Silvestre, D.; Sorlini, M.; Catenacci, L.; Sorrenti, M.; Marrubini, G.; Rossi, R.; Tripodo, G.; Mauri, P.; et al. Pilot Production of Mesenchymal Stem/Stromal Freeze-Dried Secretome for Cell-Free Regenerative Nanomedicine: A Validated GMP-Compliant Process. Cells 2018, 7, 190. [Google Scholar] [CrossRef]
- Madrigal, M.; Rao, K.S.; Riordan, N.H. A Review of Therapeutic Effects of Mesenchymal Stem Cell Secretions and Induction of Secretory Modification by Different Culture Methods. J. Transl. Med. 2014, 12, 260. [Google Scholar] [CrossRef]
- Jeon, Y.J.; Kim, J.; Cho, J.H.; Chung, H.M.; Chae, J.I. Comparative Analysis of Human Mesenchymal Stem Cells Derived from Bone Marrow, Placenta, and Adipose Tissue as Sources of Cell Therapy. J. Cell Biochem. 2016, 117, 1112–1125. [Google Scholar] [CrossRef]
- Pires, A.O.; Mendes-Pinheiro, B.; Teixeira, F.G.; Anjo, S.I.; Ribeiro-Samy, S.; Gomes, E.D.; Serra, S.C.; Silva, N.A.; Manadas, B.; Sousa, N.; et al. Unveiling the Differences of Secretome of Human Bone Marrow Mesenchymal Stem Cells, Adipose Tissue-Derived Stem Cells, and Human Umbilical Cord Perivascular Cells: A Proteomic Analysis. Stem. Cells Dev. 2016, 25, 1073–1083. [Google Scholar] [CrossRef]
- Wang, Z.G.; He, Z.Y.; Liang, S.; Yang, Q.; Cheng, P.; Chen, A.M. Comprehensive Proteomic Analysis of Exosomes Derived from Human Bone Marrow, Adipose Tissue, and Umbilical Cord Mesenchymal Stem Cells. Stem. Cell Res. Ther. 2020, 11, 511. [Google Scholar] [CrossRef] [PubMed]
- Amable, P.R.; Teixeira, M.V.; Carias, R.B.; Granjeiro, J.M.; Borojevic, R. Protein synthesis and secretion in human mesenchymal cells derived from bone marrow, adipose tissue and Wharton’s jelly. Stem. Cell Res. Ther. 2014, 5, 53. [Google Scholar] [CrossRef] [PubMed]




Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Villatoro, A.J.; Martín-Astorga, M.d.C.; Alcoholado, C.; Sánchez-Martín, M.d.M.; Becerra, J. Proteomic Analysis of the Secretome and Exosomes of Feline Adipose-Derived Mesenchymal Stem Cells. Animals 2021, 11, 295. https://doi.org/10.3390/ani11020295
Villatoro AJ, Martín-Astorga MdC, Alcoholado C, Sánchez-Martín MdM, Becerra J. Proteomic Analysis of the Secretome and Exosomes of Feline Adipose-Derived Mesenchymal Stem Cells. Animals. 2021; 11(2):295. https://doi.org/10.3390/ani11020295
Chicago/Turabian StyleVillatoro, Antonio J., María del Carmen Martín-Astorga, Cristina Alcoholado, María del Mar Sánchez-Martín, and José Becerra. 2021. "Proteomic Analysis of the Secretome and Exosomes of Feline Adipose-Derived Mesenchymal Stem Cells" Animals 11, no. 2: 295. https://doi.org/10.3390/ani11020295
APA StyleVillatoro, A. J., Martín-Astorga, M. d. C., Alcoholado, C., Sánchez-Martín, M. d. M., & Becerra, J. (2021). Proteomic Analysis of the Secretome and Exosomes of Feline Adipose-Derived Mesenchymal Stem Cells. Animals, 11(2), 295. https://doi.org/10.3390/ani11020295






