Prediction and Experimental Verification of a Hierarchical Transcription Factor Regulatory Network of Porcine Myoglobin (Mb)
Abstract
:Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Animal Collection and RNA Extraction
2.2. In Silico Analysis of the Porcine Mb Promoter to Identify Transcription Factors Binding Sites
2.3. Identification of Differentially Expressed Genes (DEGs)
2.4. Functional Enrichment Analysis of DEGs
2.5. Mb-TFs Interaction Network (TTI) Construction
2.6. Meat Color Apparent Index Determination
2.7. Western Blotting
2.8. Quantitative Real-Time PCR
2.9. Statistical Analysis
3. Results
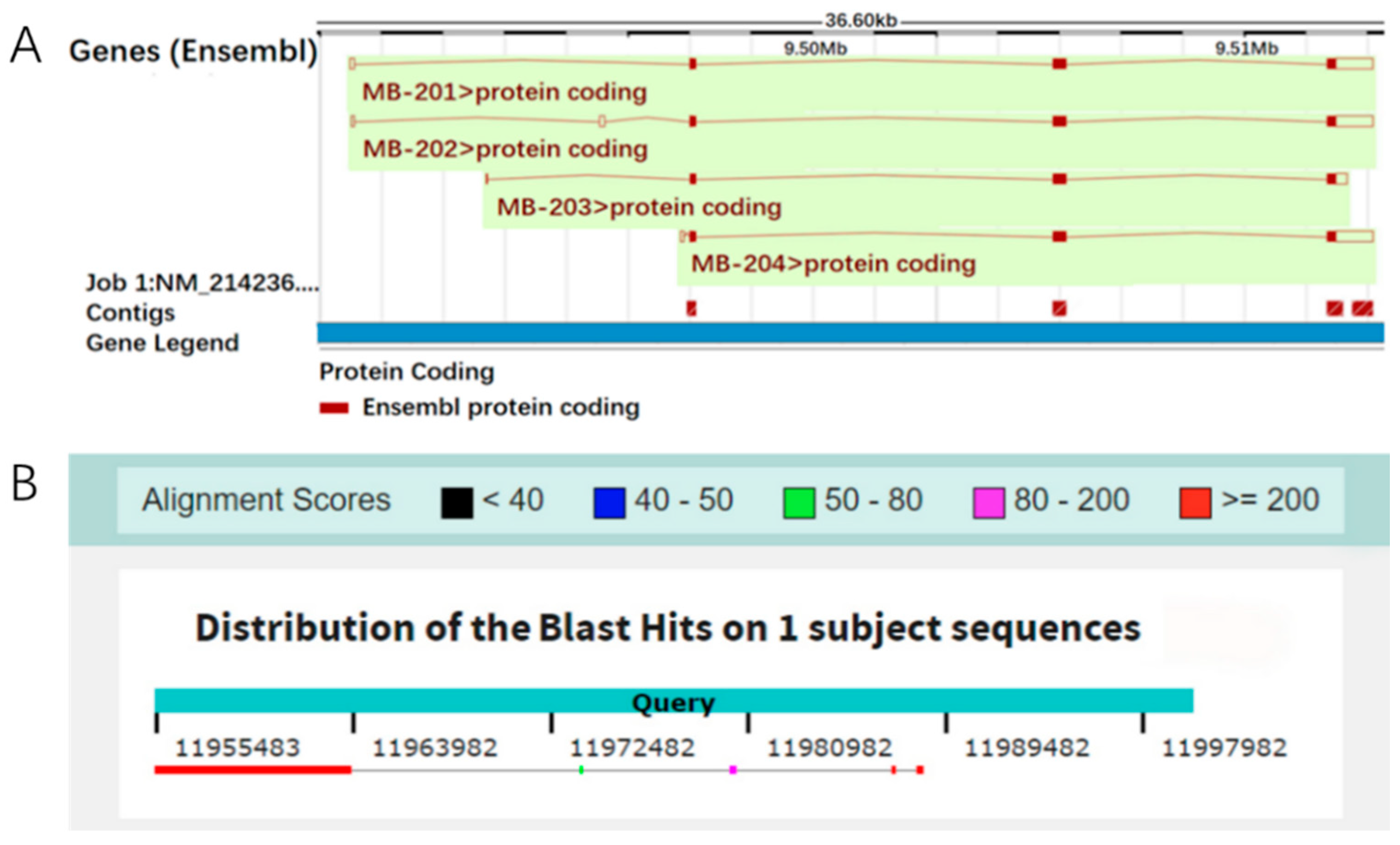
3.1. The Localization of the Mb Gene on the Porcine Reference Genome
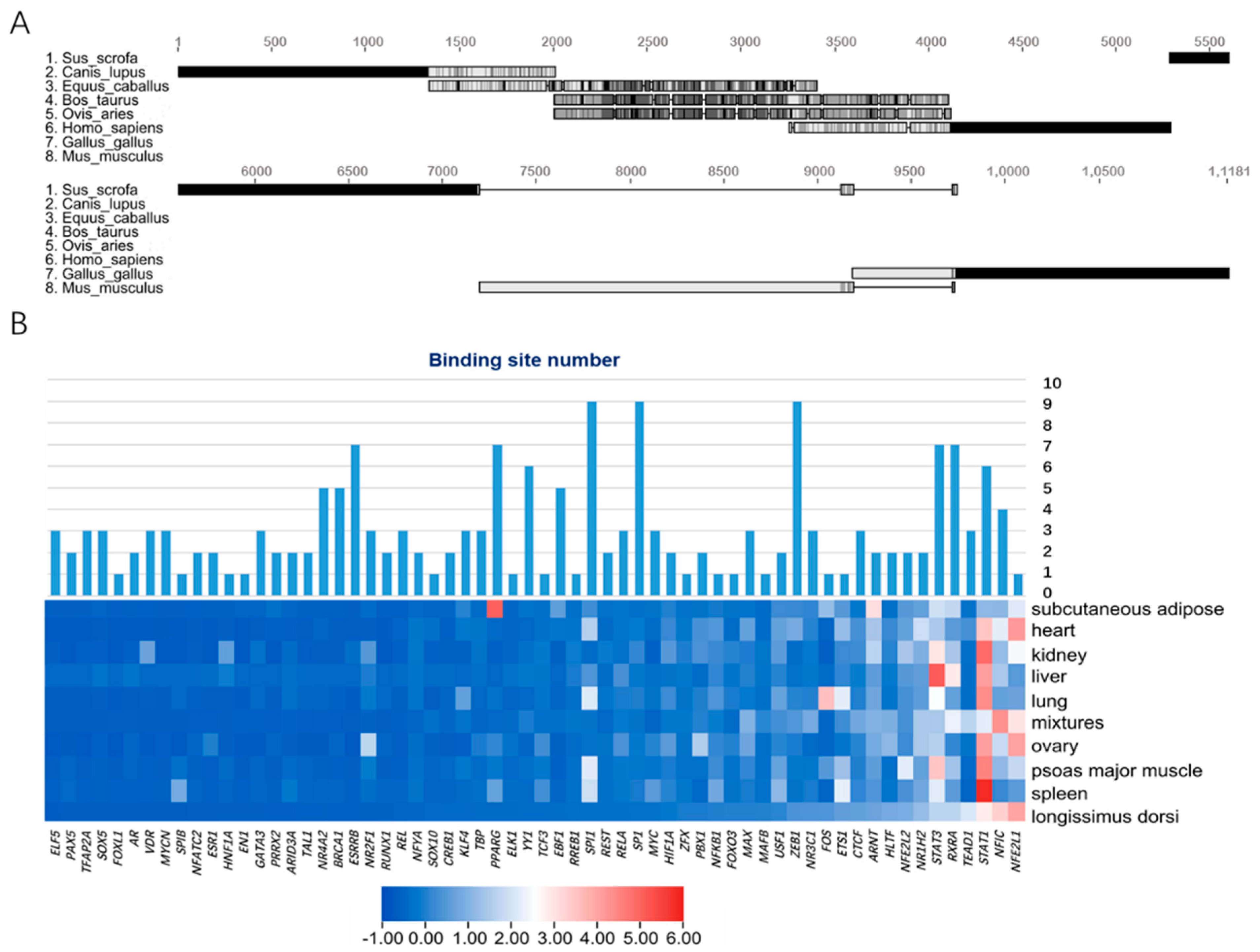
3.2. Prediction and Enrichment of Transcription Factors of Porcine Mb Gene
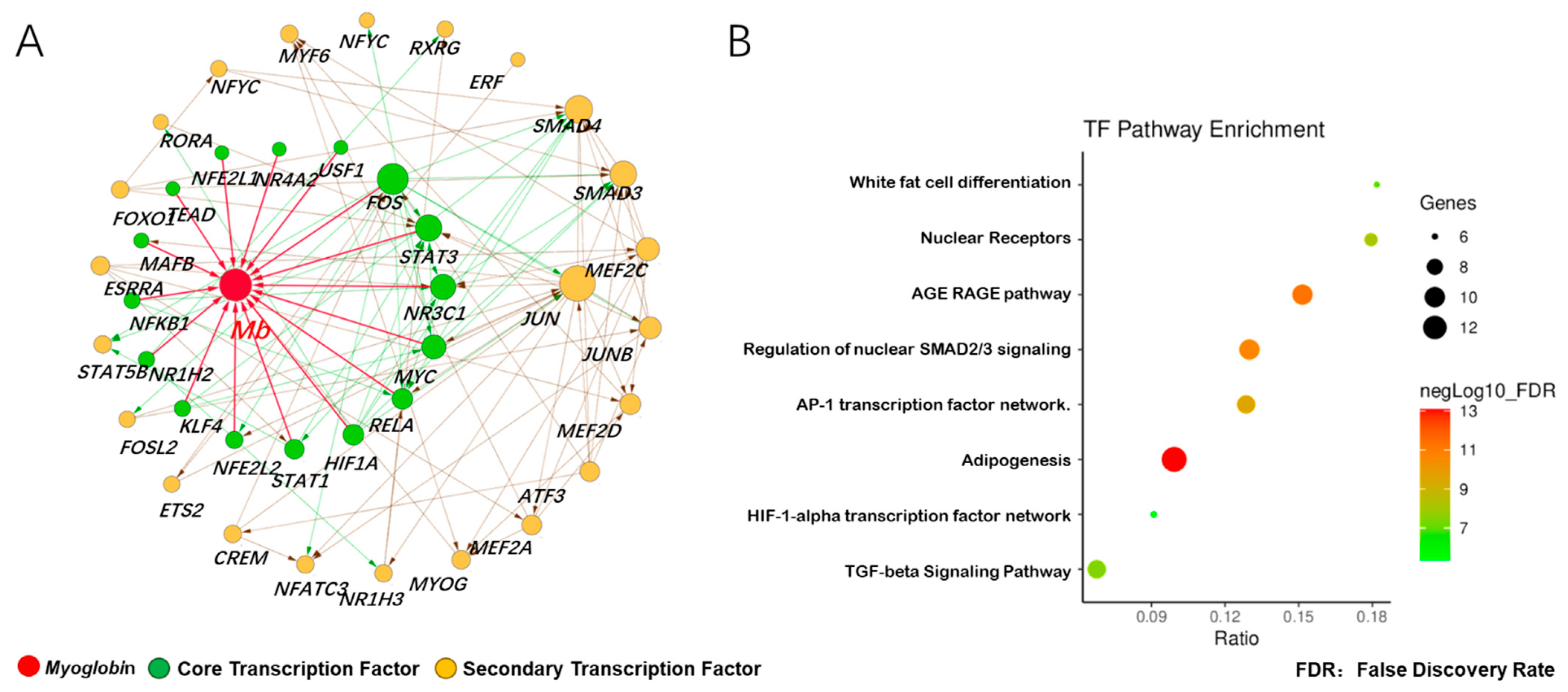
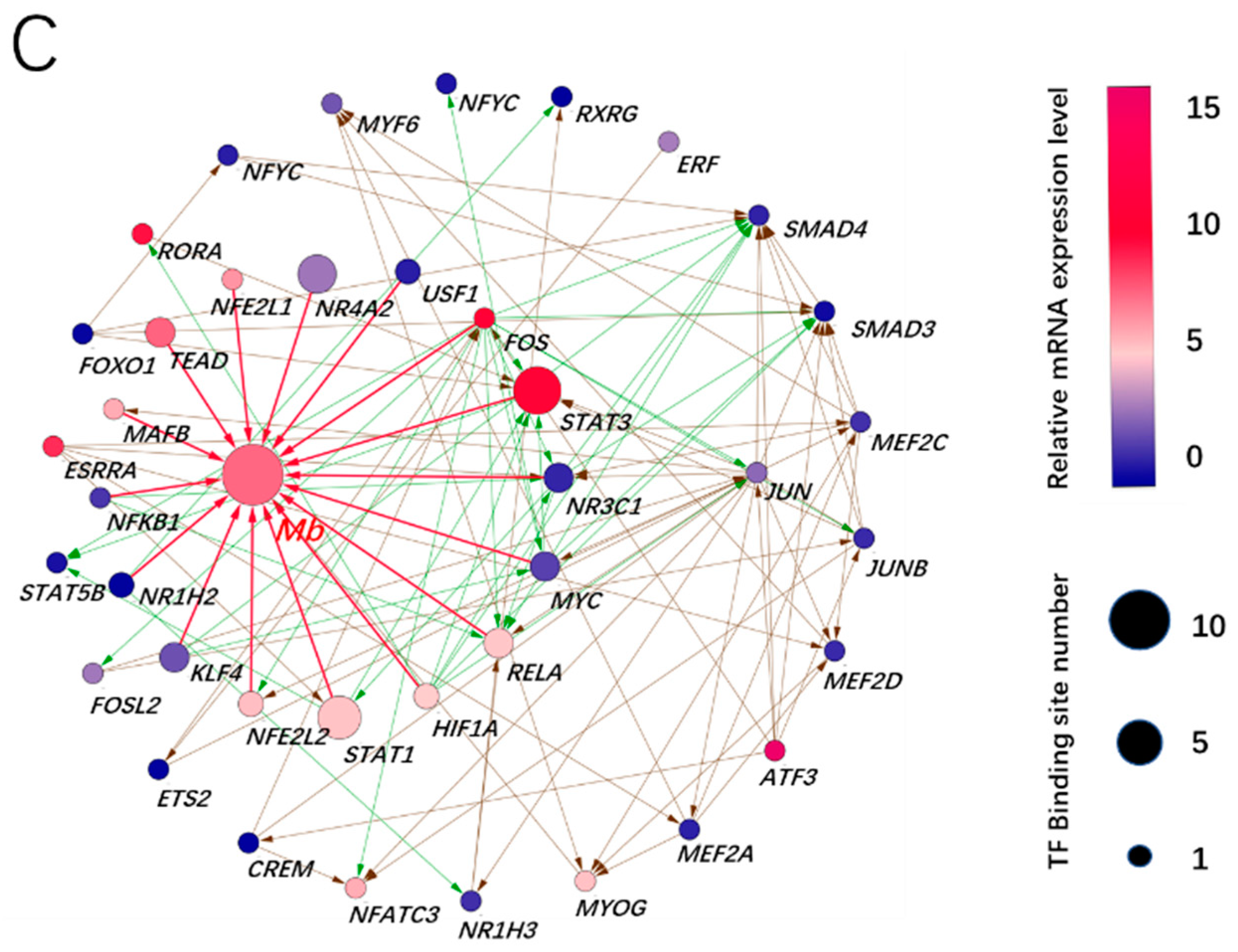
3.3. Prediction of Adaptive Transcription Factors of Porcine Mb Gene and Construction of Its Interaction Network
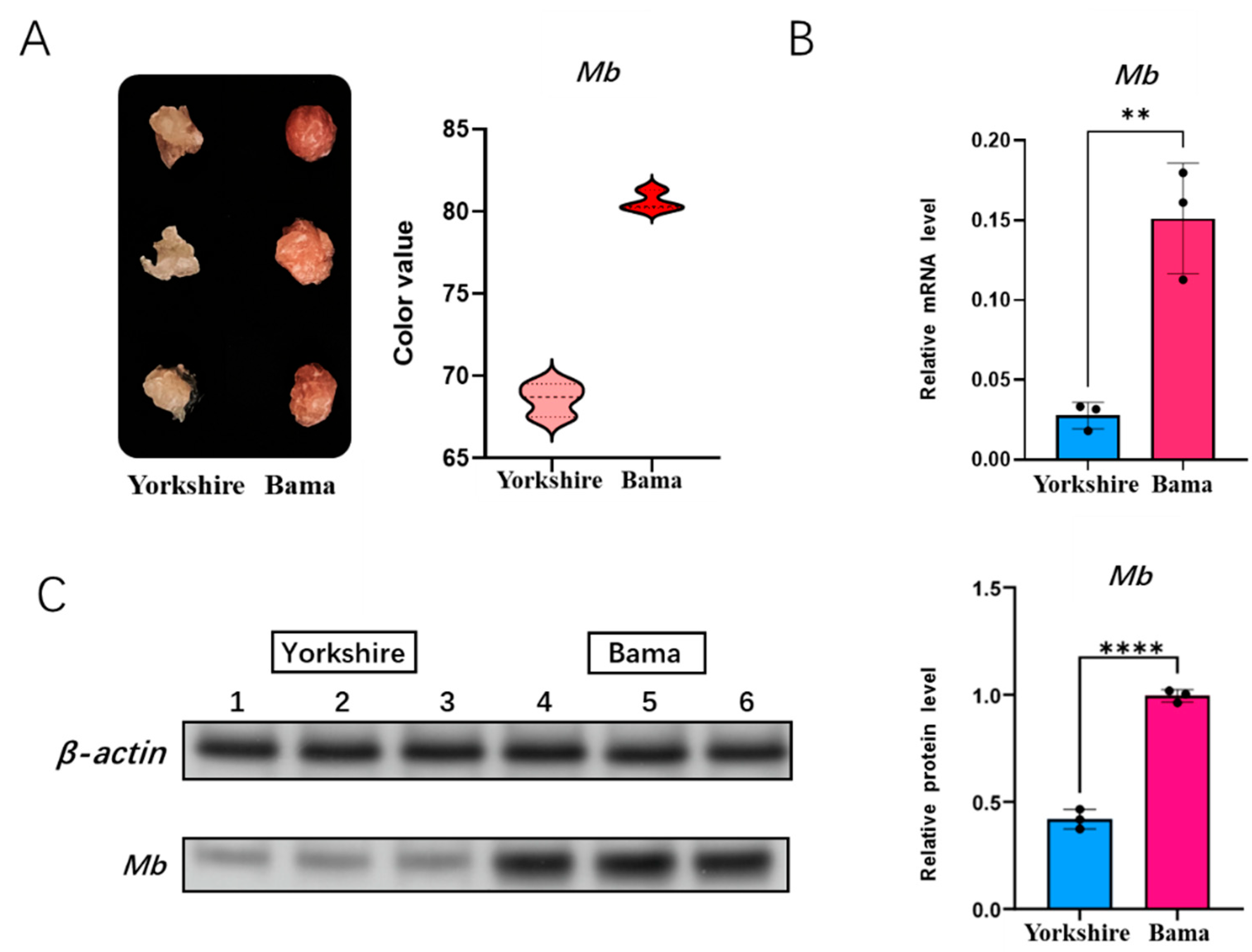
3.4. Detection of Meat Color and Mb Gene Expression
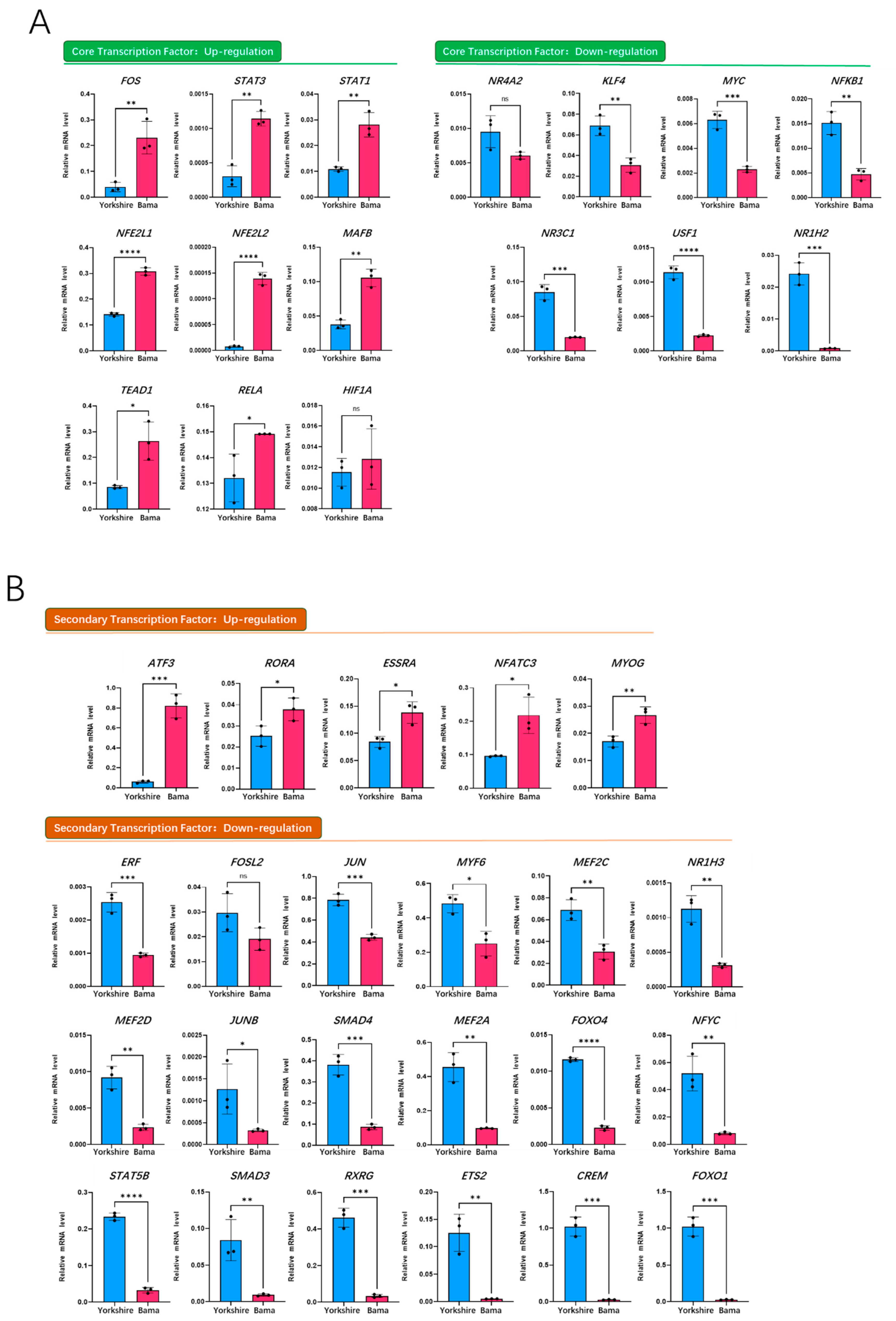
3.5. qRT-PCR Validation of Transcription Factors on Porcine Mb Gene Regulatory Network of the Bama Breed
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Henchion, M.; McCarthy, M.; Resconi, V.C.; Troy, D. Meat consumption: Trends and quality matters. Meat Sci. 2014, 98, 561–568. [Google Scholar] [CrossRef] [Green Version]
- Liu, X.; Xiong, X.; Yang, J.; Zhou, L.; Yang, B.; Ai, H.; Ma, H.; Xie, X.; Huang, Y.; Fang, S.; et al. Genome-wide association analyses for meat quality traits in Chinese Erhualian pigs and a Western Duroc × (Landrace × Yorkshire) commercial population. Genet. Sel. Evol. 2015, 47, 44. [Google Scholar] [CrossRef] [Green Version]
- Faucitano, L.; Ielo, M.; Ster, C.; Fiego, D.P.L.; Methot, S.; Saucier, L. Shelf life of pork from five different quality classes. Meat Sci. 2010, 84, 466–469. [Google Scholar] [CrossRef] [PubMed]
- Font-I-Furnols, M.; Guerrero, L. Consumer preference, behavior and perception about meat and meat products: An overview. Meat Sci. 2014, 98, 361–371. [Google Scholar] [CrossRef]
- Mancini, R.A.; Hunt, M.C. Current research in meat color. Meat Sci. 2005, 71, 100–121. [Google Scholar] [CrossRef]
- Faustman, C.; Cassens, R. The biochemical basis for discoloration in fresh meat: A review. J. Muscle Foods 1990, 1, 217–243. [Google Scholar] [CrossRef]
- Smith, G.; Belk, K.E.; Sofos, J.N.; Tatum, J.D.; Williams, S.N. Economic implications of improved color stability in beef. In Antioxidants in Muscle Foods: Nutritional Strategies to Improve Quality; John and Wiley and Sons: Hoboken, NJ, USA, 2000; pp. 397–426. [Google Scholar]
- Dagevos, H.; Voordouw, J. Sustainability and meat consumption: Is reduction realistic? Sustain. Sci. Pract. Policy 2013, 9, 60–69. [Google Scholar] [CrossRef]
- Jeong, J.; Hur, S.; Yang, H.; Moon, S.; Hwang, Y.; Park, G.; Joo, S. Discoloration Characteristics of 3 Major Muscles From Cattle During Cold Storage. J. Food Sci. 2009, 74, C1–C5. [Google Scholar] [CrossRef] [PubMed]
- Karamucki, T.; Jakubowska, M.; Rybarczyk, A.; Gardzielewska, J. The influence of myoglobin on the colour of minced pork loin. Meat Sci. 2013, 94, 234–238. [Google Scholar] [CrossRef]
- Nerín, C.; Tovar, L.; Djenane, D.; Camo, J.; Salafranca, J.; Beltrán, J.A.; Roncalés, P. Stabilization of Beef Meat by a New Active Packaging Containing Natural Antioxidants. J. Agric. Food Chem. 2006, 54, 7840–7846. [Google Scholar] [CrossRef]
- Alexandratos, N.; Bruinsma, J. World Agriculture towards 2030/2050: The 2012 Revision; FAO: Rome, Italy, 2012. [Google Scholar]
- Herron, S.E. 2018 Outlook for the US Livestock and Poultry Sectors; USDA: Arlington, VA, USA, 2018.
- Gustavsson, J.; Cederberg, C.; Sonesson, U.; Van Otterdijk, R.; Meybeck, A. Global Food Losses and Food Waste-FAO Report; Food and Agriculture Organization (FAO) of the United Nations: Rome, Italy, 2011; pp. 1–37. [Google Scholar]
- Morrissey, P.A.; Sheehy, P.J.; Galvin, K.; Kerry, J.P.; Buckley, D.J. Lipid stability in meat and meat products. Meat Sci. 1998, 49, S73–S86. [Google Scholar] [CrossRef]
- Faustman, C.; Liebler, D.C.; McClure, T.D.; Sun, Q. α, β-Unsaturated aldehydes accelerate oxymyoglobin oxidation. J. Agric. Food Chem. 1999, 47, 3140–3144. [Google Scholar] [CrossRef]
- Suman, S.P.; Faustman, C.; Stamer, S.L.; Liebler, D.C. Proteomics of lipid oxidation-induced oxidation of porcine and bovine oxymyoglobins. Proteomics 2007, 7, 628–640. [Google Scholar] [CrossRef] [PubMed]
- Grimsrud, P.A.; Xie, H.; Griffin, T.; Bernlohr, D.A. Oxidative Stress and Covalent Modification of Protein with Bioactive Aldehydes. J. Biol. Chem. 2008, 283, 21837–21841. [Google Scholar] [CrossRef] [Green Version]
- Newcom, D.W.; Stalder, K.J.; Baas, T.J.; Goodwin, R.N.; Parrish, F.C.; Wiegand, B.R. Breed differences and genetic parameters of myoglobin concentration in porcine longissimus muscle. J. Anim. Sci. 2004, 82, 2264–2268. [Google Scholar] [CrossRef] [Green Version]
- Devlin, B.H.; Wefald, F.C.; Kraus, W.E.; Bernard, T.S.; Williams, R.S. Identification of a Muscle-specific Enhancer within the 5′-Flanking Region of the Human Myoglobin Gene. J. Biol. Chem. 1989, 264, 13896–13901. [Google Scholar] [CrossRef]
- Kanatous, S.B.; Mammen, P.P.A. Regulation of myoglobin expression. J. Exp. Biol. 2010, 213, 2741–2747. [Google Scholar] [CrossRef] [Green Version]
- Gentleman, R.C. Bioconductor: Open software development for computational biology and bioinformatics. Genome Biol. 2004, 5, R80. [Google Scholar] [CrossRef] [Green Version]
- Yu, G. Using ClusterProfiler to Identify and Compare Functional Profiles of Gene Lists. 2013. Available online: https://www.bioconductor.org/packages//2.13/bioc/vignettes/clusterProfiler/inst/doc/clusterProfiler.pdf (accessed on 17 July 2021).
- Han, H.; Cho, J.-W.; Lee, S.-Y.; Yun, A.; Kim, H.; Bae, D.; Yang, S.; Kim, C.Y.; Lee, M.; Kim, E.; et al. TRRUST v2: An expanded reference database of human and mouse transcriptional regulatory interactions. Nucleic Acids Res. 2018, 46, D380–D386. [Google Scholar] [CrossRef] [PubMed]
- Mahmood, T.; Yang, P.-C. Western blot: Technique, theory, and trouble shooting. N. Am. J. Med. Sci. 2012, 4, 429. [Google Scholar]
- Schmittgen, T.D.; Livak, K.J. Analyzing real-time PCR data by the comparative CT method. Nat. Protoc. 2008, 3, 1101–1108. [Google Scholar] [CrossRef]
- Hruz, T.; Wyss, M.; Docquier, M.; Pfaffl, M.W.; Masanetz, S.; Borghi, L.; Verbrugghe, P.; Kalaydjieva, L.; Bleuler, S.; Laule, O.; et al. RefGenes: Identification of reliable and condition specific reference genes for RT-qPCR data normalization. BMC Genom. 2011, 12, 156. [Google Scholar] [CrossRef] [Green Version]
- Troy, D.; Kerry, J. Consumer perception and the role of science in the meat industry. Meat Sci. 2010, 86, 214–226. [Google Scholar] [CrossRef] [PubMed]
- Livingston, D.; Brown, W. The chemistry of myoglobin and its reactions. Food Technol. 1981, v.35, 244–252, (RESEARCH). [Google Scholar]
- Cornforth, D.P.; Jayasingh, P. Chemical and physical characteristics of meat/Color and Pigment. In Encyclopedia of Meat Sciences; Jensen, W.K., Devine C. and Dikeman, M., Eds.; Elsevier Science Ltd.: Oxford, UK, 2004; Volume 1, pp. 249–256. [Google Scholar]
- Pinter, T.B.J.; Dodd, E.L.; Bohle, D.S.; Stillman, M.J. Spectroscopic and Theoretical Studies of Ga(III)protoporphyrin-IX and Its Reactions with Myoglobin. Inorg. Chem. 2012, 51, 3743–3753. [Google Scholar] [CrossRef] [PubMed]
- Wittenberg, J.B.; Wittenberg, B.A. Myoglobin function reassessed. J. Exp. Biol. 2003, 206, 2011–2020. [Google Scholar] [CrossRef] [Green Version]
- Salathe, E.P.; Chen, C. The role of myoglobin in retarding oxygen depletion in skeletal muscle. Math. Biosci. 1993, 116, 1–20. [Google Scholar] [CrossRef]
- Kanatous, S.B.; Garry, D.J. Gene deletional strategies reveal novel physiological roles for myoglobin in striated muscle. Respir. Physiol. Neurobiol. 2006, 151, 151–158. [Google Scholar] [CrossRef]
- Buckley, D.J.; Morrissey, P.A.; Gray, J.I. Influence of dietary vitamin E on the oxidative stability and quality of pig meat. J. Anim. Sci. 1995, 73, 3122–3130. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Milde-Langosch, K. The Fos family of transcription factors and their role in tumourigenesis. Eur. J. Cancer 2005, 41, 2449–2461. [Google Scholar] [CrossRef]
- Pedraza-Alva, G.; Zingg, J.; Jost, J. AP-1 binds to a putative cAMP response element of the MyoD1 promoter and negatively modulates MyoD1 expression in dividing myoblasts. J. Biol. Chem. 1994, 269, 6978–6985. [Google Scholar] [CrossRef]
- Chang, Y.-H.; Huang, C.-N.; Shiau, M.-Y. Association of IL-4 receptor gene polymorphisms with high density lipoprotein cholesterol. Cytokine 2012, 59, 309–312. [Google Scholar] [CrossRef] [PubMed]
- Sun, L.; Ma, K.; Wang, H.; Xiao, F.; Gao, Y.; Zhang, W.; Wang, K.; Gao, X.; Ip, N.; Wu, Z. JAK1–STAT1–STAT3, a key pathway promoting proliferation and preventing premature differentiation of myoblasts. J. Cell Biol. 2007, 179, 129–138. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Butturini, E.; De Prati, A.C.; Mariotto, S. Redox Regulation of STAT1 and STAT3 Signaling. Int. J. Mol. Sci. 2020, 21, 7034. [Google Scholar] [CrossRef] [PubMed]
- Grohmann, M.; Wiede, F.; Dodd, G.; Gurzov, E.; Ooi, G.J.; Butt, T.; Rasmiena, A.A.; Kaur, S.; Gulati, T.; Goh, P.K.; et al. Obesity Drives STAT-1-Dependent NASH and STAT-3-Dependent HCC. Cell 2018, 175, 1289–1306.e20. [Google Scholar] [CrossRef] [Green Version]
- Guillet-Deniau, I.; Burnol, A.-F.; Girard, J. Identification and Localization of a Skeletal Muscle Secrotonin 5-HT2A Receptor Coupled to the Jak/STAT Pathway. J. Biol. Chem. 1997, 272, 14825–14829. [Google Scholar] [CrossRef] [Green Version]
- Glal, D.; Sudhakar, J.N.; Lu, H.H.; Liu, M.-C.; Chiang, H.-Y.; Liu, Y.-C.; Cheng, C.-F.; Shui, J.-W. ATF3 Sustains IL-22-Induced STAT3 Phosphorylation to Maintain Mucosal Immunity Through Inhibiting Phosphatases. Front. Immunol. 2018, 9, 2522. [Google Scholar] [CrossRef] [Green Version]
- Hou, Y.; Liu, Z.; Zuo, Z.; Gao, T.; Fu, J.; Wang, H.; Xu, Y.; Liu, D.; Yamamoto, M.; Zhu, B.; et al. Adipocyte-specific deficiency of Nfe2l1 disrupts plasticity of white adipose tissues and metabolic homeostasis in mice. Biochem. Biophys. Res. Commun. 2018, 503, 264–270. [Google Scholar] [CrossRef]
- Zhang, Z.-Y.; Bao, X.-L.; Cong, Y.-Y.; Fan, B.; Li, G.-Y. Autophagy in Age-Related Macular Degeneration: A Regulatory Mechanism of Oxidative Stress. Oxidative Med. Cell. Longev. 2020, 2020, 2896036. [Google Scholar] [CrossRef]
- Kataoka, K.; Noda, M.; Nishizawa, M. Maf nuclear oncoprotein recognizes sequences related to an AP-1 site and forms heterodimers with both Fos and Jun. Mol. Cell. Biol. 1994, 14, 700–712. [Google Scholar] [PubMed] [Green Version]
- Hamada, M.; Nakamura, M.; Tran, M.T.N.; Moriguchi, T.; Hong, C.; Ohsumi, T.; Dinh, T.T.H.; Kusakabe, M.; Hattori, M.; Katsumata, T.; et al. MafB promotes atherosclerosis by inhibiting foam-cell apoptosis. Nat. Commun. 2014, 5, 3147. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tran, M.T.N.; Hamada, M.; Nakamura, M.; Jeon, H.; Kamei, R.; Tsunakawa, Y.; Kulathunga, K.; Lin, Y.; Fujisawa, K.; Kudo, T.; et al. MafB deficiency accelerates the development of obesity in mice. FEBS Open Bio 2016, 6, 540–547. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mahmassani, Z.S.; Reidy, P.T.; McKenzie, A.I.; Stubben, C.; Howard, M.T.; Drummond, M.J. Age-dependent skeletal muscle transcriptome response to bed rest-induced atrophy. J. Appl. Physiol. 2019, 126, 894–902. [Google Scholar] [CrossRef]
- Tong, X.; Hou, L.; He, W.; Mei, C.; Huang, B.; Zhang, C.; Hu, C.; Wang, C. Whole genome sequence analysis reveals genetic structure and X-chromosome haplotype structure in indigenous Chinese pigs. Sci. Rep. 2020, 10, 9433. [Google Scholar] [CrossRef] [PubMed]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yuan, D.; Yu, H.; Liu, S.; Hao, L.; Zhang, J. Prediction and Experimental Verification of a Hierarchical Transcription Factor Regulatory Network of Porcine Myoglobin (Mb). Animals 2021, 11, 3599. https://doi.org/10.3390/ani11123599
Yuan D, Yu H, Liu S, Hao L, Zhang J. Prediction and Experimental Verification of a Hierarchical Transcription Factor Regulatory Network of Porcine Myoglobin (Mb). Animals. 2021; 11(12):3599. https://doi.org/10.3390/ani11123599
Chicago/Turabian StyleYuan, Di, Hao Yu, Songcai Liu, Linlin Hao, and Jing Zhang. 2021. "Prediction and Experimental Verification of a Hierarchical Transcription Factor Regulatory Network of Porcine Myoglobin (Mb)" Animals 11, no. 12: 3599. https://doi.org/10.3390/ani11123599
APA StyleYuan, D., Yu, H., Liu, S., Hao, L., & Zhang, J. (2021). Prediction and Experimental Verification of a Hierarchical Transcription Factor Regulatory Network of Porcine Myoglobin (Mb). Animals, 11(12), 3599. https://doi.org/10.3390/ani11123599






