The German Shorthair Pointer Dog Breed (Canis lupus familiaris): Genomic Inbreeding and Variability
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
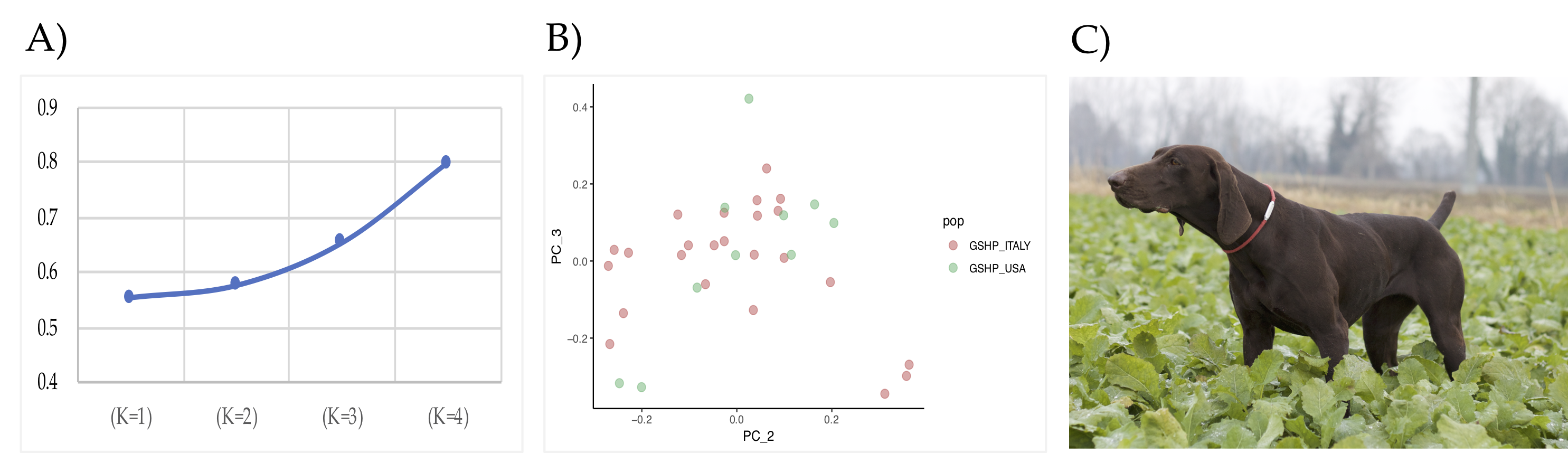
2.1. Sampling and Genotyping
2.2. Runs of homozygosity (ROH)
2.3. Inbreeding Coefficients
3. Results
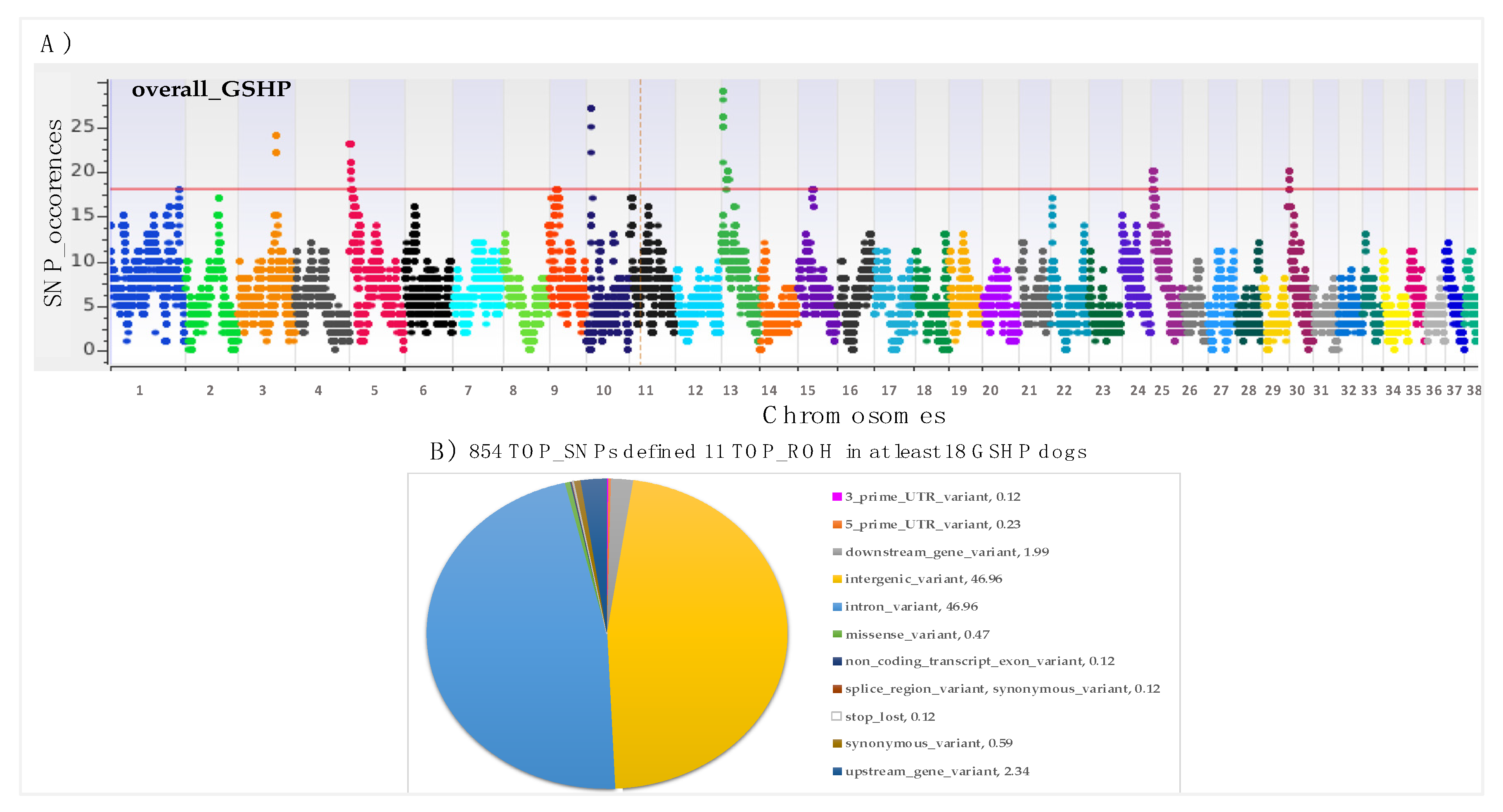
3.1. Runs of Homozygosity (ROH)
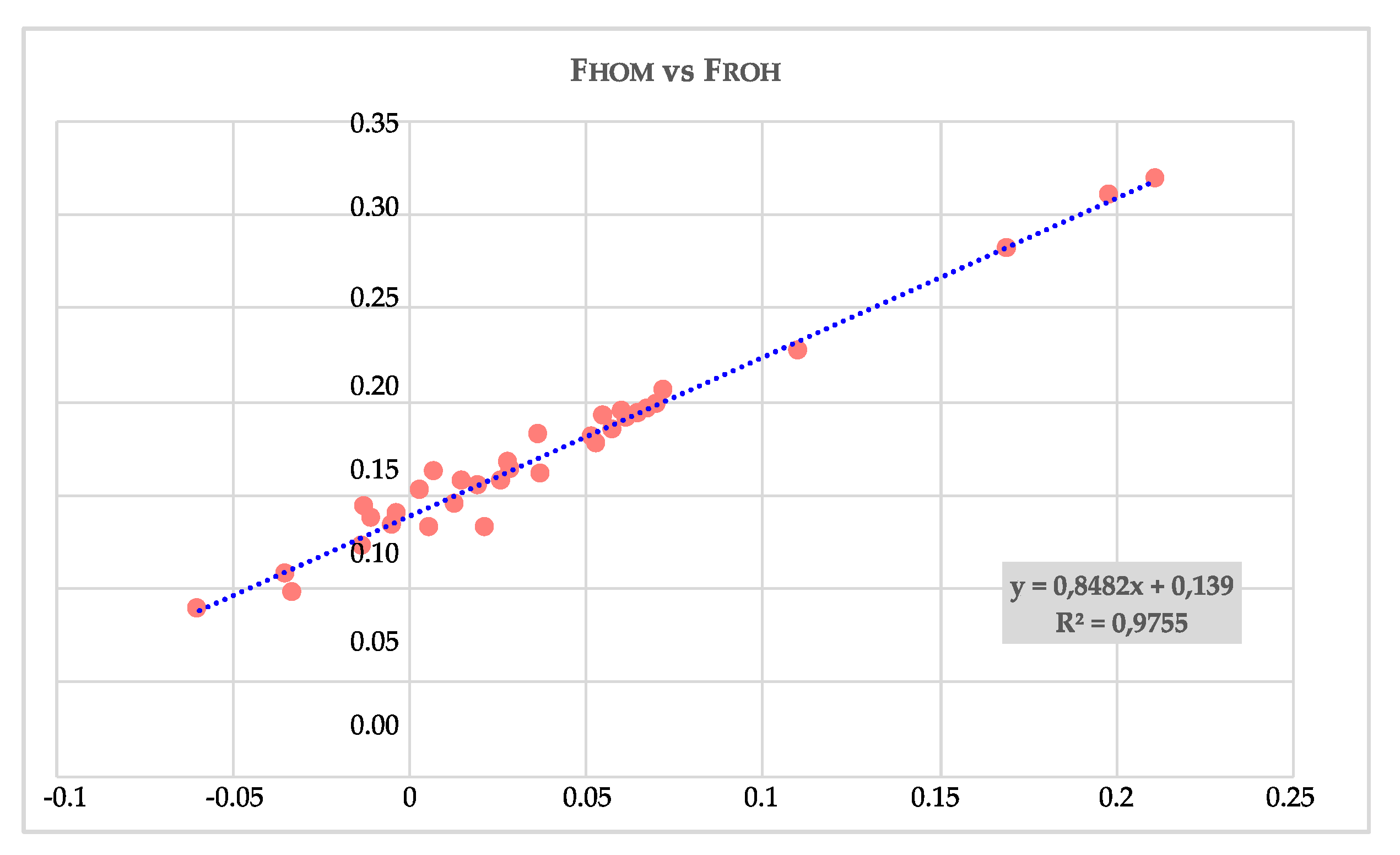
3.2. Inbreeding Coefficients
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Asher, L.; Diesel, G.; Summers, J.F.; McGreevy, P.D.; Collins, L.M. Inherited defects in pedigree dogs. Part 1: Disorders related to breed standards. Vet. J. 2009, 182, 402–411. [Google Scholar] [CrossRef]
- Parker, H.G.; Kim, L.V.; Sutter, N.B.; Carlson, S.; Lorentzen, T.D.; Malek, T.B.; Johnson, G.S.; DeFrance, H.B.; Ostrander, E.A.; Kruglyak, L. Genetic structure of the purebred domestic dog. Science 2004, 304, 1160–1164. [Google Scholar] [CrossRef] [PubMed]
- Lindblad-Toh, K.; Wade, C.M.; Mikkelsen, T.S.; Karlsson, E.K.; Jaffe, D.B.; Kamal, M.; Clamp, M.; Chang, J.L.; Kulbokas, E.J., III; Zody, M.C. Genome sequence, comparative analysis and haplotype structure of the domestic dog. Nature 2005, 438, 803–819. [Google Scholar] [CrossRef] [PubMed]
- Schoenebeck, J.J.; Ostrander, E.A. Insights into morphology and disease from the dog genome project. Ann. Rev. Cell Dev. Biol. 2014, 30, 535–560. [Google Scholar] [CrossRef] [PubMed]
- Pollinger, J.P.; Lohmueller, K.E.; Han, E.; Parker, H.G.; Quignon, P.; Degenhardt, J.D.; Boyko, A.R.; Earl, D.A.; Auton, A.; Reynolds, A. Genome-wide SNP and haplotype analyses reveal a rich history underlying dog domestication. Nature 2010, 464, 898–902. [Google Scholar]
- Vilà, C.; Leonard, J.A. Canid phylogeny and origin of the domestic dog. In The Genetics of the Dog; CABI: Wallingford, UK, 2012. [Google Scholar]
- Dreger, D.L.; Davis, B.W.; Cocco, R.; Sechi, S.; Di Cerbo, A.; Parker, H.G.; Polli, M.; Marelli, S.P.; Crepaldi, P.; Ostrander, E.A. Commonalities in development of pure breeds and population isolates revealed in the genome of the sardinian Fonni’s Dog. Genetics 2016, 204, 737–755. [Google Scholar] [CrossRef] [PubMed]
- Dreger, D.L.; Rimbault, M.; Davis, B.W.; Bhatnagar, A.; Parker, H.G.; Ostrander, E.A. Whole-genome sequence, SNP chips and pedigree structure: Building demographic profiles in domestic dog breeds to optimize genetic-trait mapping. Dis. Model. Mech. 2016, 9, 1445–1460. [Google Scholar] [CrossRef]
- Chu, E.T.; Simpson, M.J.; Diehl, K.; Page, R.L.; Sams, A.J.; Boyko, A.R. Inbreeding depression causes reduced fecundity in Golden Retrievers. Mamm. Genome 2019, 30, 166–172. [Google Scholar] [CrossRef]
- Mortlock, S.-A.; Khatkar, M.S.; Williamson, P. Comparative analysis of genome diversity in bullmastiff dogs. PLoS ONE 2016, 11, e0147941. [Google Scholar] [CrossRef]
- Pertoldi, C.; Kristensen, T.N.; Loeschcke, V.; Berg, P.; Praebel, A.; Stronen, A.V.; Proschowsky, H.F.; Fredholm, M. Characterization of the genetic profile of five Danish dog breeds. J. Anim. Sci. 2013, 91, 5122–5127. [Google Scholar] [CrossRef]
- Pfahler, S.; Distl, O. Effective population size, extended linkage disequilibrium and signatures of selection in the rare dog breed Lundehund. PLoS ONE 2015, 10, e0122680. [Google Scholar] [CrossRef] [PubMed]
- Pedersen, N.C.; Pooch, A.S.; Liu, H. A genetic assessment of the English bulldog. Canine Genet. Epidemiol. 2016, 3, 6. [Google Scholar] [CrossRef] [PubMed]
- Pedersen, N.; Liu, H.; Theilen, G.; Sacks, B. The effects of dog breed development on genetic diversity and the relative influences of performance and conformation breeding. J. Anim. Breed. Genet. 2013, 130, 236–248. [Google Scholar] [CrossRef]
- Keijser, S.F.A.; Fieten, H.; Vos-Loohuis, M.; Piek, C.J.; Anderson, H.; Donner, J.; Scholten, I.; Nielen, M.; Hesselink, J.W.; van Steenbeek, F.G. Heterozygosity testing and multiplex DNA panel screening as a potential tool to monitor health and inbreeding in a small, closed dog population. Canine Genet. Epidemiol. 2018, 5, 12. [Google Scholar] [CrossRef] [PubMed]
- Wiener, P.; Sánchez-Molano, E.; Clements, D.N.; Woolliams, J.A.; Haskell, M.J.; Blott, S.C. Genomic data illuminates demography, genetic structure and selection of a popular dog breed. BMC Genom. 2017, 18, 609. [Google Scholar] [CrossRef] [PubMed]
- Bigi, D.; Marelli, S.P.; Liotta, L.; Frattini, S.; Talenti, A.; Pagnacco, G.; Polli, M.; Crepaldi, P. Investigating the population structure and genetic differentiation of livestock guard dog breeds. Animal 2018, 12, 2009–2016. [Google Scholar] [CrossRef] [PubMed]
- Mastrangelo, S.; Biscarini, F.; Auzino, B.; Ragatzu, M.; Spaterna, A.; Ciampolini, R. Genome-wide diversity and runs of homozygosity in the “Braque Français, type Pyrénées” dog breed. BMC Res. Notes 2018, 11, 13. [Google Scholar] [CrossRef]
- Kardos, M.; Luikart, G.; Allendorf, F.W. Measuring individual inbreeding in the age of genomics: Marker-based measures are better than pedigrees. Heredity (Edinb) 2015, 115, 63. [Google Scholar] [CrossRef]
- Mastrangelo, S.; Biscarini, F.; Tolone, M.; Auzino, B.; Ragatzu, M.; Spaterna, A.; Ciampolini, R. Genomic characterization of the Braque Français type Pyrénées dog and relationship with other breeds. PLoS ONE 2018, 13, e0208548. [Google Scholar] [CrossRef]
- Mortlock, S.-A.; Booth, R.; Mazrier, H.; Khatkar, M.S.; Williamson, P. Visualization of genome diversity in german shepherd dogs. Bioinform. Biol. Insights 2015, 9, BBI-S30524. [Google Scholar] [CrossRef]
- Wright, S. Genetics of populations. Encycl. Br. 1948, 10, 111-AD. [Google Scholar]
- Clutton-Brock, J. A Natural History of Domesticated Mammals; Cambridge University Press: Cambridge, UK, 1999; ISBN 0521634954. [Google Scholar]
- Parra, D.; Méndez, S.; Canon, J.; Dunner, S. Genetic differentiation in pointing dog breeds inferred from microsatellites and mitochondrial DNA sequence. Anim. Genet. 2008, 39, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Udell, M.A.R.; Ewald, M.; Dorey, N.R.; Wynne, C.D.L. Exploring breed differences in dogs (Canis familiaris): Does exaggeration or inhibition of predatory response predict performance on human-guided tasks? Anim. Behav. 2014, 89, 99–105. [Google Scholar] [CrossRef]
- Turcsán, B.; Kubinyi, E.; Miklósi, Á. Trainability and boldness traits differ between dog breed clusters based on conventional breed categories and genetic relatedness. Appl. Anim. Behav. Sci. 2011, 132, 61–70. [Google Scholar] [CrossRef]
- Bell, J.; Cavanagh, K.; Tilley, L.; Smith, F.W.K. Veterinary Medical Guide to Dog and Cat Breeds; CRC Press: Boca Raton, FL, USA, 2012; ISBN 1482241412. [Google Scholar]
- Alexander, D.H.; Novembre, J.; Lange, K. Fast model-based estimation of ancestry in unrelated individuals. Genome Res. 2009, 19, 1655–1664. [Google Scholar] [CrossRef] [PubMed]
- ENSEMBL Genome Browser. Available online: https://www.ensembl.org/index.html (accessed on 16 March 2020).
- DAVID 6.8 Data Base. Available online: https://david.ncifcrf.gov (accessed on 16 March 2020).
- Biscarini, F.; Cozzi, P.; Gaspa, G.; Marras, G. detectRUNS: Detect Runs of Homozygosity and Runs of Heterozygosity in Diploid Genomes. 2018. Available online: http://orca.cf.ac.uk/id/eprint/108906 (accessed on 16 March 2020).
- Kinghorn, B.P.; Kinghorn, A.J. Pedigree Viewer 6.5; University of New England: Armidale, Australia, 2010. [Google Scholar]
- Mastrangelo, S.; Portolano, B.; Di Gerlando, R.; Ciampolini, R.; Tolone, M.; Sardina, M.T.; Consortium, I.S.G. Genome-wide analysis in endangered populations: A case study in Barbaresca sheep. Animal 2017, 11, 1107–1116. [Google Scholar] [CrossRef]
- Lamac, N.; Sölkner, J.; Mészáros, G. Analysis of Excessive Homozygous Regions in Rhodesian Ridgeback Dogs. Agric. Conspec. Sci. 2017, 82, 277–280. [Google Scholar]
- MacLean, E.; Snyder-Mackler, N.; Serpell, J. Highly heritable and functionally relevant breed differences in dog behavior. BioRxiv 2019, 286, 509315. [Google Scholar]
- Vaysse, A.; Ratnakumar, A.; Derrien, T.; Axelsson, E.; Pielberg, G.R.; Sigurdsson, S.; Fall, T.; Seppälä, E.H.; Hansen, M.S.T.; Lawley, C.T. Identification of genomic regions associated with phenotypic variation between dog breeds using selection mapping. PLoS Genet. 2011, 7, e1002316. [Google Scholar] [CrossRef]
- Gurgul, A.; Jasielczuk, I.; Semik-Gurgul, E.; Pawlina-Tyszko, K.; Stefaniuk-Szmukier, M.; Szmatoła, T.; Polak, G.; Tomczyk-Wrona, I.; Bugno-Poniewierska, M. A genome-wide scan for diversifying selection signatures in selected horse breeds. PLoS ONE 2019, 14, e0210751. [Google Scholar] [CrossRef]
- Kim, J.; Williams, F.J.; Dreger, D.L.; Plassais, J.; Davis, B.W.; Parker, H.G.; Ostrander, E.A. Genetic selection of athletic success in sport-hunting dogs. Proc. Natl. Acad. Sci. USA 2018, 115, E7212–E7221. [Google Scholar] [CrossRef] [PubMed]
- Maciejewska-Skrendo, A.; Cięszczyk, P.; Chycki, J.; Sawczuk, M.; Smółka, W. Genetic Markers Associated with Power Athlete Status. J. Hum. Kinet. 2019, 68, 17. [Google Scholar] [CrossRef] [PubMed]
- Du, W.; Hu, H.; Zhang, J.; Bao, G.; Chen, R.; Quan, R. The Mechanism of MAPK Signal Transduction Pathway Involved with Electroacupuncture Treatment for Different Diseases. Evidence-Based Complement. Altern. Med. 2019, 2019, 8138017. [Google Scholar] [CrossRef] [PubMed]
- Kettunen, A.; Daverdin, M.; Helfjord, T.; Berg, P. Cross-breeding is inevitable to conserve the highly inbred population of puffin hunter: The Norwegian Lundehund. PLoS ONE 2017, 12, e0170039. [Google Scholar] [CrossRef]
- Leroy, G. Genetic diversity, inbreeding and breeding practices in dogs: Results from pedigree analyses. Vet. J. 2011, 189, 177–182. [Google Scholar] [CrossRef] [PubMed]
- Leroy, G.; Mary-Huard, T.; Verrier, E.; Danvy, S.; Charvolin, E.; Danchin-Burge, C. Methods to estimate effective population size using pedigree data: Examples in dog, sheep, cattle and horse. Genet. Sel. Evol. 2013, 45, 1. [Google Scholar] [CrossRef]
- Jansson, M.; Laikre, L. Pedigree data indicate rapid inbreeding and loss of genetic diversity within populations of native, traditional dog breeds of conservation concern. PLoS ONE 2018, 13, e0202849. [Google Scholar] [CrossRef]
- Talenti, A.; Dreger, D.L.; Frattini, S.; Polli, M.; Marelli, S.; Harris, A.C.; Liotta, L.; Cocco, R.; Hogan, A.N.; Bigi, D. Studies of modern Italian dog populations reveal multiple patterns for domestic breed evolution. Ecol. Evol. 2018, 8, 2911–2925. [Google Scholar] [CrossRef]
- Cecchi, F.; Paci, G.; Spaterna, A.; Ciampolini, R. Genetic variability in Bracco Italiano dog breed assessed by pedigree data. Ital. J. Anim. Sci. 2013, 12, e54. [Google Scholar] [CrossRef]
- Cecchi, F.; Paci, G.; Spaterna, A.; Ragatzu, M.; Ciampolini, R. Demographic approach on the study of genetic parameters in the dog Braque Français type Pyrénées italian population. Ital. J. Anim. Sci. 2016, 15, 30–36. [Google Scholar] [CrossRef]
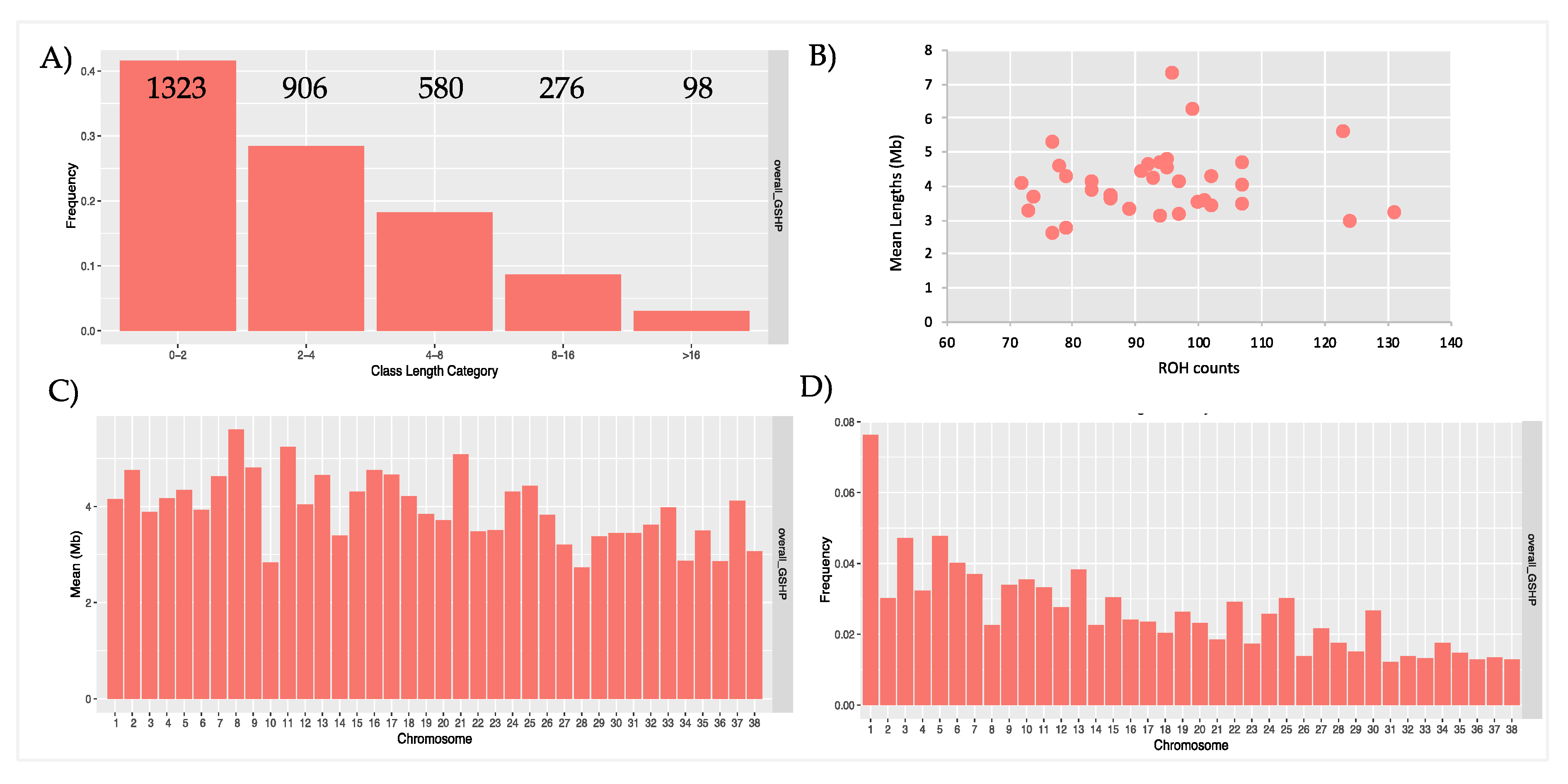
| POP | N. ROHs | Min - Max (Mean) of ROH Per Samples | Min ROH Length | Max ROH Length | Mean ROH Length | (%) Total Coverage |
|---|---|---|---|---|---|---|
| overall_GSHP | 3183 | 71-131 (93.6) | 1000717 | 56893025 | 4072407 | 5.9 |
| TOP_ROH_id | Chr | Start Position | End Position | *N. Samples | Genes |
|---|---|---|---|---|---|
| TOP_ROH_1 | 1 | 112009648 | 112324183 | 18 | CNFN, MEGF8, TMEM145, PRR19, PAFAH1B3, CIC, ERF, GSK3A, ZNF526, DEDD2, POU2F2, ZNF574, GRIK5 |
| TOP_ROH_2 | 3 | 60892911 | 62824148 | 22-24 | DOK7, HGFAC, RGS12, MSANTD1, HTT, GRK4, NOP14, MFSD10, ADD1, SH3BP2, TNIP2, FAM193A, RNF4, CFAP99, ZFYVE28, MXD4, HAUS3, POLN, NAT8L, C3H4orf48, NELFA, SCARNA22, NSD2, LETM1, FGFR3, TACC3, TMEM129, SLBP, FAM53A, NKX1-1, UVSSA, MAEA, FAM184B |
| TOP_ROH_3 | 5 | 247752 | 3480976 | 18-23 | B3GAT1, GLB1L2 (LOC606786), GLB1L3, ACAD8, THYN1, cfa-mir-8868, VPS26B, NCAPD3, JAM3, IGSF9B, SPATA19, OPCML, NTM |
| TOP_ROH_4 | 5 | 4900070 | 4979148 | 18 | PRDM10, NFRKB |
| TOP_ROH_5 | 9 | 12539669 | 14668121 | 18 | KPNA2, C9H17orf58, BPTF, NOL11, PITPNC1, PSMD12, HELZ, CACNG1, CACNG4, CACNG5, PRKCA, APOH, CEP112, AXIN2 |
| TOP_ROH_6 | 10 | 7278956 | 8418771 | 22-27 | RASSF3, GNS, TBC1D30, WIF1, LEMD3, MSRB3, HMGA2 |
| TOP_ROH_7 | 13 | 2962719 | 4505605 | 21-29 | ZNF706, GRHL2, NCALD, RRM2B, UBR5, ODF1, KLF10, AZIN1 |
| TOP_ROH_8 | 13 | 10151357 | 13036989 | 18-20 | SYBU, KCNV1, CSMD3 |
| TOP_ROH_9 | 15 | 24204830 | 25089306 | 18 | CCDC59, METTL25, TMTC2 |
| TOP_ROH_10 | 25 | 2268042 | 3887386 | 18-20 | UFM1, TRPC4, POSTN, SUPT20H, EXOSC8, ALG5, SMAD9, RFXAP |
| TOP_ROH_11 | 30 | 927947 | 2775667 | 18-20 | EMC7, CHRM5, AVEN, RYR3, FMN1, GREM1, SCG5, ARHGAP11A, GJD2, ACTC1, AQR, ZNF770 |
| Category | Term (Nominal p-Value) | Genes |
|---|---|---|
| GO:0019226~transmission of nerve impulse (0.005) | CHRM5, CACNG5, JAM3 | |
| GOTERM (BP) | GO:0071407~cellular response to organic cyclic compound (0.02) | SMAD9, GSK3A, AXIN2 |
| GO:0043161~proteasome-mediated ubiquitin-dependent protein catabolic process (0.035) | MAEA, RNF4, PSMD12, GSK3A | |
| GOTERM (CC) | GO:0005654~nucleoplasm (0.01) | HAUS3, SMAD9, HTT, NFRKB, RRM2B, GRHL2, BPTF, UBR5, SYBU, POLN, PITPNC1, AXIN2, TNIP2, KPNA2, DEDD2, ADD1 |
| KEGG | cfa04260: Cardiac muscle contraction (0.005); cfa05410: Hypertrophic cardiomyopathy (HCM) (0.007); cfa05414: Dilated cardiomyopathy (0.008); cfa04261: Adrenergic signaling in cardiomyocytes (0.028). | ACTC1, CACNG5, CACNG4, CACNG1 |
| cfa04921: Oxytocin signaling pathway (0.005); cfa04010: MAPK signaling pathway (0.032). | PRKCA, RYR3, CACNG5, CACNG4, CACNG1 | |
| cfa05412: Arrhythmogenic right ventricular cardiomyopathy (ARVC) (0.042) | CACNG5, CACNG4, CACNG1 |
| TOP_ROH_id | Gene Name | Behavior Traits [35] | Morphological Traits [36] |
|---|---|---|---|
| TOP_ROH_2 | FAM193A | Chasing, Dog Aggression | |
| TOP_ROH_2 | HTT | Chasing | |
| TOP_ROH_2 | RNF4 | Chasing, Dog Aggression | |
| TOP_ROH_3 | NCAPD3 | Separation Problems | |
| TOP_ROH_3 | SPATA19 | Chasing | |
| TOP_ROH_5 | NOL11 | Energy | |
| TOP_ROH_5 | OPCML | Dog Aggression | |
| TOP_ROH_6 | HMGA2 | Body size | |
| TOP_ROH_6 | MSRB3 | Dog Fear | Ear morphology |
| TOP_ROH_6 | TBC1D30 | Attachment/attention-seeking, Dog Fear | |
| TOP_ROH_6 | WIF1 | Ear morphology | |
| TOP_ROH_8 | CSMD3 | Excitability | |
| TOP_ROH_9 | TMTC2 | Attachment/attention-seeking, Excitability | |
| TOP_ROH_11 | FMN1 | Energy, Trainability | |
| TOP_ROH_11 | RYR3 | attachment/attention-seeking, Energy |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Boccardo, A.; Marelli, S.P.; Pravettoni, D.; Bagnato, A.; Busca, G.A.; Strillacci, M.G. The German Shorthair Pointer Dog Breed (Canis lupus familiaris): Genomic Inbreeding and Variability. Animals 2020, 10, 498. https://doi.org/10.3390/ani10030498
Boccardo A, Marelli SP, Pravettoni D, Bagnato A, Busca GA, Strillacci MG. The German Shorthair Pointer Dog Breed (Canis lupus familiaris): Genomic Inbreeding and Variability. Animals. 2020; 10(3):498. https://doi.org/10.3390/ani10030498
Chicago/Turabian StyleBoccardo, Antonio, Stefano Paolo Marelli, Davide Pravettoni, Alessandro Bagnato, Giuseppe Achille Busca, and Maria Giuseppina Strillacci. 2020. "The German Shorthair Pointer Dog Breed (Canis lupus familiaris): Genomic Inbreeding and Variability" Animals 10, no. 3: 498. https://doi.org/10.3390/ani10030498
APA StyleBoccardo, A., Marelli, S. P., Pravettoni, D., Bagnato, A., Busca, G. A., & Strillacci, M. G. (2020). The German Shorthair Pointer Dog Breed (Canis lupus familiaris): Genomic Inbreeding and Variability. Animals, 10(3), 498. https://doi.org/10.3390/ani10030498





