New Insights into Modelling Bacterial Growth with Reference to the Fish Pathogen Flavobacterium psychrophilum
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Datasets
2.2. Mathematical Considerations
2.2.1. Potential Growth
2.2.2. Actual Growth
Rectangular Hyperbola
Simple Exponential
2.3. Model Fitting
2.4. Statistical Analysis
2.5. Model Validation
3. Results
3.1. Paramater Estimates
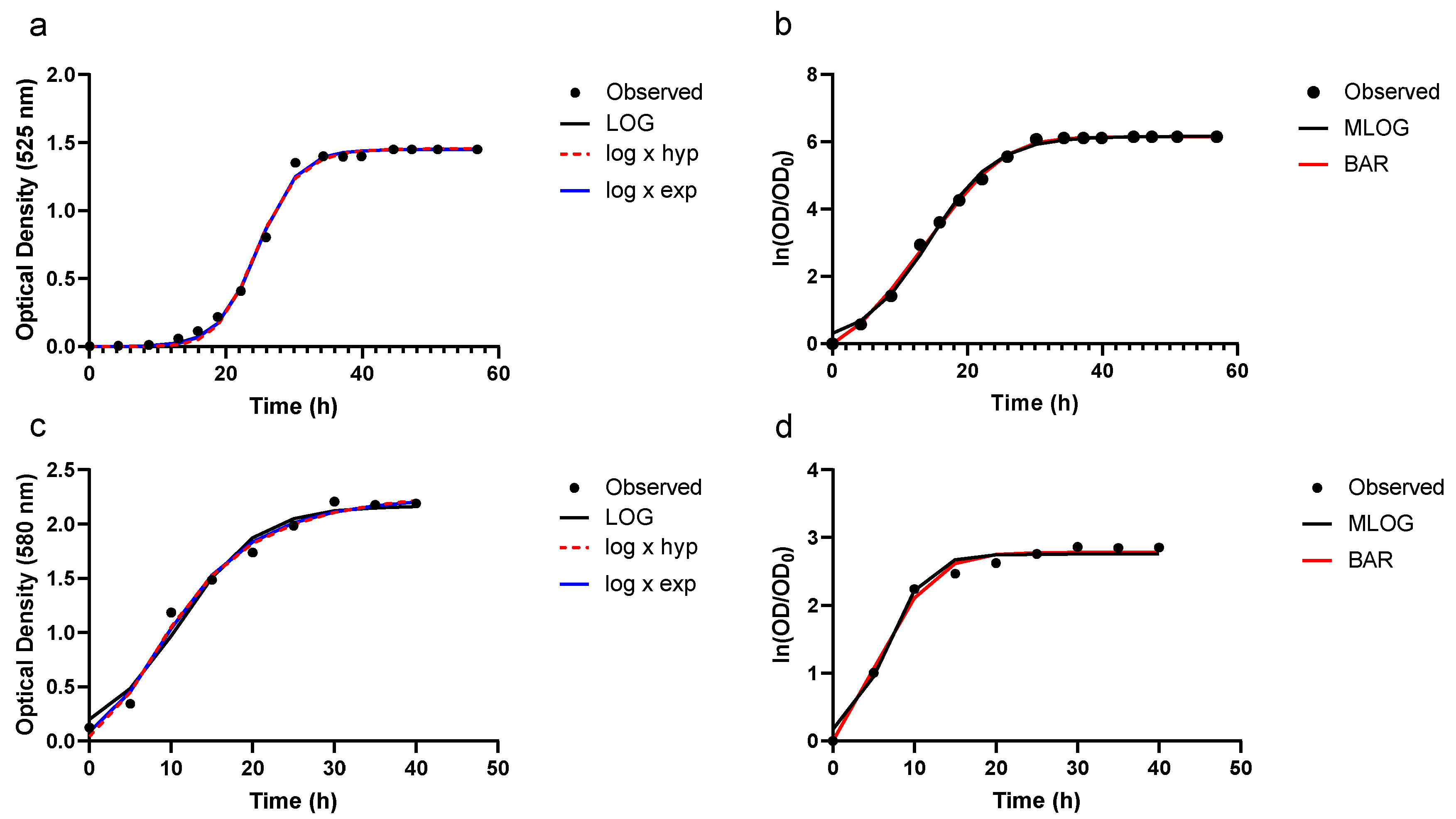
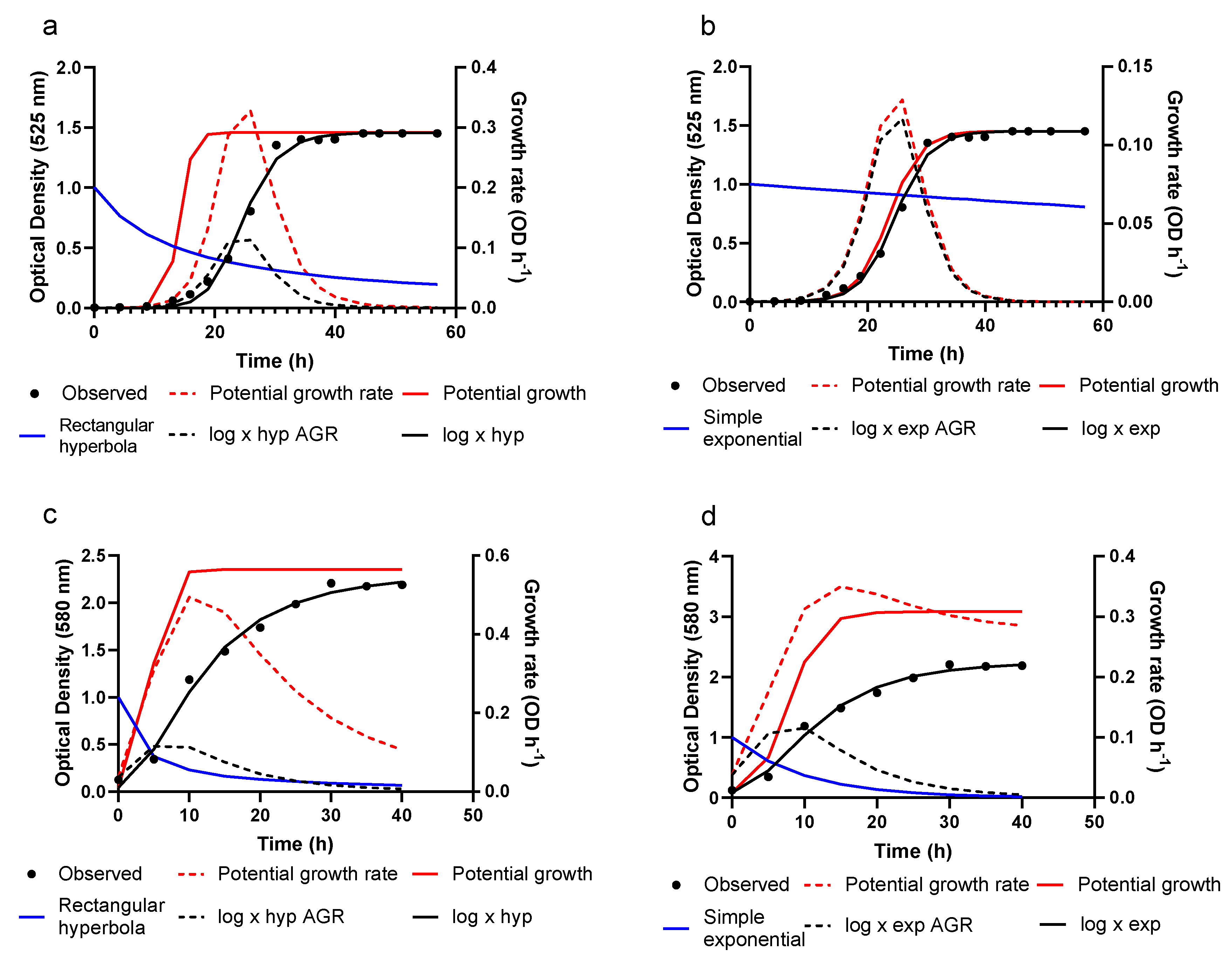
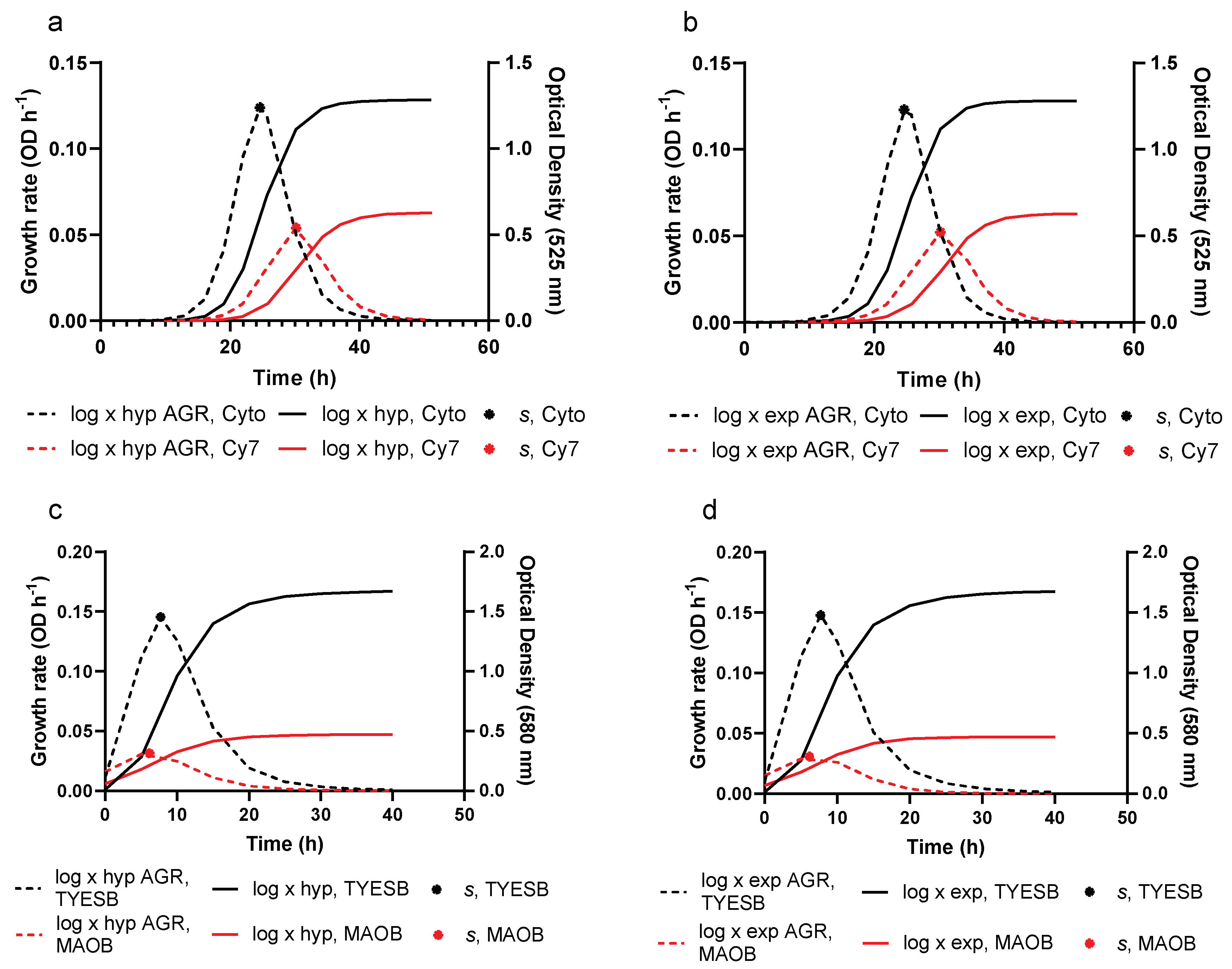
3.2. Growth Prediction
3.3. Model Evaluation
3.4. Model Validation
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Whiting, R.C. Notes on reparameterization of bacterial growth curves—A reply to J. Baranyi and W. E. Garthright. Food Microbiol. 1992, 9, 173–174. [Google Scholar] [CrossRef]
- Baranyi, J.; Roberts, T.A. A dynamic approach to predicting bacterial growth in food. Int. J. Food Microbiol. 1994, 23, 277–294. [Google Scholar] [CrossRef]
- López, S.; Prieto, M.; Dijkstra, J.; Dhanoa, M.S.; France, J. Statistical evaluation of mathematical models for microbial growth. Int. J. Food Microbiol. 2004, 96, 289–300. [Google Scholar] [CrossRef]
- Dalgaard, P.; Koutsoumanis, K. Comparison of maximum specific growth rates and lag times estimated from absorbance and viable count data by different mathematical models. J. Microbiol. Methods 2001, 43, 183–196. [Google Scholar] [CrossRef]
- Vine, N.; Leukes, W.; Kaiser, H. In vitro growth characteristics of five candidate aquaculture probiotics and two fish pathogens grown in fish intestinal mucus. FEMS Microbiol. Lett. 2004, 231, 145–152. [Google Scholar] [CrossRef]
- Dalgaard, P.; Ross, T.; Kamperman, L.; Neumeyer, K.; McMeekin, T.A. Estimation of bacterial growth rates from turbidimetric and viable count data. Int. J. Food Microbiol. 1994, 23, 391–404. [Google Scholar] [CrossRef]
- Perni, S.; Andrew, P.W.; Shama, G. Estimating the maximum growth rate from microbial growth curves: Definition is everything. Food Microbiol. 2005, 22, 491–495. [Google Scholar] [CrossRef]
- Campana, R.; Favi, G.; Baffone, W.; Lucarini, S. Marine alkaloid 2,2-bis(6-bromo-3-indolyl) ethylamine and its synthetic derivatives inhibit microbial biofilms formation and disaggregate developed biofilms. Microorganisms 2019, 7, 28. [Google Scholar] [CrossRef]
- Farber, R.; Rosenberg, A.; Rozenfeld, S.; Benet, G.; Cahan, R. Bioremediation of artificial diesel-contaminated soil using bacterial consortium immobilized to plasma-pretreated wood waste. Microorganisms 2019, 7, 497. [Google Scholar] [CrossRef]
- Deeb, J.G.; Smith, J.; Belvin, B.R.; Grzech-Leśniak, K.; Lewis, J. Er:YAG laser irradiation reduces microbial viability when used in combination with irrigation with sodium hypochlorite, chlorhexidine, and hydrogen peroxide. Microorganisms 2019, 7, 612. [Google Scholar] [CrossRef]
- Mauerhofer, L.M.; Pappenreiter, P.; Paulik, C.; Seifert, A.H.; Bernacchi, S.; Rittmann, S.K.M.R. Methods for quantification of growth and productivity in anaerobic microbiology and biotechnology. Folia Microbiol. (Praha). 2019, 321–360. [Google Scholar] [CrossRef]
- Zwietering, M.H.; Jongenburger, I.; Rombouts, F.M.; van ’T Riet, K. Modeling of the bacterial growth curve. Appl. Environ. Microbiol. 1990, 56, 1875–1881. [Google Scholar] [CrossRef]
- Buchanan, R.L.; Phillips, J.G. Response surface model for predicting the effects of temperature pH, sodium chloride content, sodium nitrite concentration and atmosphere on the growth of Listeria monocytogenes. J. Food Prot. 1990, 53, 370–376. [Google Scholar] [CrossRef]
- Garthright, W.E. Refinements in the prediction of microbial growth curves. Food Microbiol. 1991, 8, 239–248. [Google Scholar] [CrossRef]
- Baranyi, J.; Robinson, T.P.; Kaloti, A.; Mackey, B.M. Predicting growth of Brochothrix thermosphacta at changing temperature. Int. J. Food Microbiol. 1995, 27, 61–75. [Google Scholar] [CrossRef]
- Peleg, M. A model of microbial growth and decay in a closed habitat based on combined Fermi’s and the logistic equations. J. Sci. Food Agric. 1996, 71, 225–230. [Google Scholar] [CrossRef]
- Fujikawa, H.; Kai, A.; Morozumi, S. A new logistic model for bacterial growth. J. Food Hyg. Soc. Japan 2003, 44, 155–160. [Google Scholar] [CrossRef]
- Huang, L. Growth kinetics of Listeria monocytogenes in broth and beef frankfurters - Determination of lag phase duration and exponential growth rate under isothermal conditions. J. Food Sci. 2008, 73, 235–242. [Google Scholar] [CrossRef]
- Huang, L. A new mechanistic growth model for simultaneous determination of lag phase duration and exponential growth rate and a new Bělehdrádek-type model for evaluating the effect of temperature on growth rate. Food Microbiol. 2011, 28, 770–776. [Google Scholar] [CrossRef]
- Gompertz, B. On the nature of the function expressive of the law of human mortality, and on a new mode of determining the value of life contingencies. Philos. Trans. R. Soc. London 1825, 115, 513–583. [Google Scholar]
- Verhulst, P.F. Notice sur la loi que la population suit dans son accroissement. Corresp. Mathématique Phys. 1838, 10, 113–121. [Google Scholar]
- McMeekin, T.A. Predictive Microbiology: Theory and Application; Innovation in Microbiology Series 5; Research Studies Press Ltd.: Taunton, Somerset, UK, 1993; ISBN 0863801323. [Google Scholar]
- Baranyi, J.; Roberts, T.A.; McClure, P. A non-autonomous differential equation to model bacterial growth. Food Microbiol. 1993, 10, 43–59. [Google Scholar] [CrossRef]
- Baranyi, J. Simple is good as long as it is enough. Food Microbiol. 1997, 14, 189–192. [Google Scholar] [CrossRef]
- Buchanan, R.L.; Whiting, R.C.; Damert, W.C. When is simple good enough: A comparison of the Gompertz, Baranyi, and three-phase linear models for fitting bacterial growth curves. Food Microbiol. 1997, 14, 313–326. [Google Scholar] [CrossRef]
- Lorenzen, E.; Dalsgaard, I.; Bernardet, J. Characterization of isolates of Flavobacterium psychrophilum associated with coldwater disease or rainbow trout fry syndrome: Phenotypic and genomic studies. Dis. Aquat. Organ. 1997, 31, 197–208. [Google Scholar] [CrossRef]
- Lafrentz, B.R.; Lapatra, S.E.; Jones, G.R.; Cain, K.D. Passive immunization of rainbow trout, Oncorhynchus mykiss (Walbaum), against Flavobacterium psychrophilum, the causative agent of bacterial coldwater disease and rainbow trout fry syndrome. J. Fish Dis. 2003, 26, 377–384. [Google Scholar] [CrossRef]
- Starliper, C.E. Bacterial coldwater disease of fishes caused by Flavobacterium psychrophilum. J. Adv. Res. 2011, 2, 97–108. [Google Scholar] [CrossRef]
- Holt, R.A. Cytophaga psychrophila, the Causative Agent of Bacterial Cold-Water Disease in Salmonid Fish. Ph.D. Thesis, Oregon State University, Corvallis, OR, USA, 1987. [Google Scholar]
- Wiens, G.D.; LaPatra, S.E.; Welch, T.J.; Evenhuis, J.P.; Rexroad III, C.E.; Leeds, T.D. On-farm performance of rainbow trout (Oncorhynchus mykiss) selectively bred for resistance to bacterial cold water disease: Effect of rearing environment on survival phenotype. Aquaculture 2013, 388, 128–136. [Google Scholar] [CrossRef]
- Bruun, M.S.; Schmidt, A.; Madsen, L.; Dalsgaard, I. Antimicrobial resistance patterns in Danish isolates of Flavobacterium psychrophilum. Aquaculture 2000, 187, 201–212. [Google Scholar] [CrossRef]
- Shieh, H.S. Studies on the nutrition of a fish pathogen, Flexibacter columnaris. Microbios Lett. 1980, 13, 129–133. [Google Scholar]
- Wakabayashi, H. Characteristics of myxobacteria associated with some freshwater fish diseases in Japan. Bull. Japanese Soc. Sci. Fish 1974, 40, 751–757. [Google Scholar] [CrossRef]
- Reichenbach, H.; Dworkin, M. Introduction to the gliding bacteria. In The Prokaryotes; Starr, M.P., Stolp, H., Trüper, H.G., Balows, A., Schlegel, H.G., Eds.; Springer: Berlin, 1981; pp. 315–327. [Google Scholar]
- Cepeda, C.; García-Márquez, S.; Santos, Y. Improved growth of Flavobacterium psychrophilum using a new culture medium. Aquaculture 2004, 238, 75–82. [Google Scholar] [CrossRef]
- Segel, I.H. Enzyme Kinetics: Behavior and Analysis of Rapid Equilibrium and Steady-State Enzyme Systems; Wiley Classics Library; Wiley: New York, NY, USA, 1993; ISBN 0471303097. [Google Scholar]
- Miller, G. Numerical Analysis for Engineers and Scientists; Cambridge University Press: Cambridge, UK, 2014; ISBN 9781107021082. [Google Scholar]
- SAS Institute Inc. SAS/STAT® 9.2 User’s Guide, 2nd ed.; SAS Institute Inc.: Cary, NC, USA, 2009; ISBN 978-1607644392. [Google Scholar]
- Bibby, J.; Toutenburg, H. Prediction and Improved Estimation in Linear Models; John Wiley and Sons: Chichester, UK, 1977; ISBN 0-471-01656-X. [Google Scholar]
- Lin, L.I.-K. A concordance correlation coefficient to evaluate reproducibility. Biometrics 1989, 45, 255–268. [Google Scholar] [CrossRef]
- Akaike, H. A new look at the statistical model identification. IEEE Trans. Automat. Contr. 1974, 19, 716–723. [Google Scholar] [CrossRef]
- Ross, T. Indices for performance evaluation of predictive models in food microbiology. J. Appl. Bacteriol. 1996, 81, 501–508. [Google Scholar] [CrossRef]
- Baranyi, J.; Pin, C.; Ross, T. Validating and comparing predictive models. Int. J. Food Microbiol. 1999, 48, 159–166. [Google Scholar] [CrossRef]
- Draper, N.R.; Smith, H. Applied Regression Analysis, 3rd ed.; Wiley Series in Probability and Statistics; John Wiley & Sons: New York, NY, YSA, 1998; ISBN 9780471170822. [Google Scholar]
- Allen, D.M. The relationship between variable selection and data augmentation and a method for prediction. Technometrics 1974, 16, 125–127. [Google Scholar] [CrossRef]
- Tarpey, T. A note on the prediction sum of squares statistic for restricted least squares. Am. Stat. 2000, 54, 116–118. [Google Scholar] [CrossRef]
- Holiday, D.B.; Ballard, J.E.; McKeown, B.C. PRESS-related statistics: Regression tools for cross-validation and case diagnostics. Med. Sci. Sports Exerc. 1995, 27, 612–620. [Google Scholar] [CrossRef]
- Richards, F.J. A flexible growth function for empirical use. J. Exp. Bot. 1959, 10, 290–301. [Google Scholar] [CrossRef]
- Dalgaard, P. Modelling of microbial activity and prediction of shelf life for packed fresh fish. Int. J. Food Microbiol. 1995, 305–317. [Google Scholar] [CrossRef]
- Stenholm, A.R.; Dalsgaard, I.; Middelboe, M. Isolation and characterization of bacteriophages infecting the fish pathogen Flavobacterium psychrophilum. Appl. Environ. Microbiol. 2008, 74, 4070–4078. [Google Scholar] [CrossRef] [PubMed]
- Aoki, M.; Kondo, M.; Kawai, K.; Oshima, S.I. Experimental bath infection with Flavobacterium psychrophilum, inducing typical signs of rainbow trout Oncorhynchus mykiss fry syndrome. Dis. Aquat. Organ. 2005, 67, 73–79. [Google Scholar] [CrossRef] [PubMed]
| Functional Form | |
|---|---|
| Simple logistic (LOG) | |
| Modified logistic (MLOG) | |
| Baranyi four parameter (BAR) | where µmaxT = ln[1 + |
| log × hyp | |
| log × exp |
| LOG | log × hyp | log × exp | MLOG | BAR | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Dataset | s | s/x* | T | s | s/x* | T | s | s/x* | T | µmax | T | µmax | T |
| 1 | 0.122 | 0.168 | 18.8 | 0.122 | 0.182 | 18.7 | 0.122 | 0.170 | 18.3 | 0.314 | 4.6 | 0.292 | 3.7 |
| 2 | 0.096 | 0.138 | 16.8 | 0.095 | 0.151 | 16.6 | 0.097 | 0.163 | 17.1 | 0.362 | 5.6 | 0.348 | 5.3 |
| 3 | 0.122 | 0.191 | 19.8 | 0.124 | 0.207 | 19.8 | 0.123 | 0.197 | 19.5 | 0.302 | 5.5 | 0.291 | 5.1 |
| 4 | 0.052 | 0.165 | 24.5 | 0.053 | 0.181 | 24.7 | 0.052 | 0.168 | 24.2 | 0.219 | 7.8 | 0.221 | 7.7 |
| 5 | 0.111 | 0.103 | 3.1 | 0.124 | 0.174 | 1.7 | 0.137 | 0.161 | 1.6 | 0.285 | 1.7 | 0.334 | 1.8 |
| 6 | 0.133 | 0.161 | 3.7 | 0.145 | 0.225 | 3.5 | 0.148 | 0.237 | 3.6 | 0.309 | 2.9 | 0.506 | 4.4 |
| 7 | 0.120 | 0.206 | 4.1 | 0.119 | 0.217 | 4.0 | 0.107 | 0.279 | 4.8 | 0.847 | 7.6 | 1.656 | 8.3 |
| 8 | 0.031 | 0.130 | 1.4 | 0.031 | 0.146 | 1.2 | 0.030 | 0.130 | 0.7 | 0.457 | 7.7 | 1.452 | 8.7 |
| Measure of Goodness-of-Fit | LOG | log × hyp | log × exp | MLOG | BAR |
|---|---|---|---|---|---|
| AIC | |||||
| Study 1-Average (±SE) | −117.1 (11.5) | −110.0 (11.5) | −111.6 (11.3) | −58.8 (3.1) | −67.5 (5.5) |
| SStudy 2-Average (±SE) | −44.6 (3.9) | −43.9 (2.7) | −45.0 (2.9) | −44.4 (3.6) | −44.0 (2.9) |
| SOverall-Average (±SE) | −80.8 (15.1) | −77.0 (14.0) | −78.3 (14.0) | −51.6 (3.7) | −55.7 (5.6) |
| MSPE | |||||
| SStudy 1-Average (±SE) | 0.001 (0.0002) | 0.001 (0.0003) | 0.001 (0.0002) | 0.024 (0.003) | 0.014 (0.004) |
| SStudy 2-Average (±SE) | 0.005 (0.002) | 0.004 (0.001) | 0.003 (0.001) | 0.005 (0.002) | 0.004 (0.001) |
| SOverall-Average (±SE) | 0.003 (0.001) | 0.002 (0.001) | 0.002 (0.001) | 0.013 (0.004) | 0.009 (0.003) |
| CCC | |||||
| SStudy 1-Average (±SE) | 0.998 (0.001) | 0.998 (0.001) | 0.998 (0.001) | 0.997 (0.001) | 0.998 (0.001) |
| SStudy 2-Average (±SE) | 0.987 (0.004) | 0.988 (0.005) | 0.989 (0.005) | 0.997 (0.001) | 0.998 (0.001) |
| SOverall-Average (±SE) | 0.992 (0.003) | 0.993 (0.003) | 0.993 (0.003) | 0.997 (0.001) | 0.998 (0.001) |
| AF | |||||
| SStudy 1-Average (±SE) | 1.013 (0.002) | 1.016 (0.003) | 1.015 (0.002) | 1.039 (0.005) | 1.024 (0.005) |
| SStudy 2-Average (±SE) | 1.033 (0.005) | 1.030 (0.003) | 1.027 (0.002) | 1.022 (0.007) | 1.016 (0.003) |
| SOverall-Average (±SE) | 1.023 (0.005) | 1.023 (0.003) | 1.021 (0.002) | 1.031 (0.005) | 1.020 (0.003) |
| Test for Examination of Residuals | LOG | log × hyp | log × exp | MLOG | BAR |
|---|---|---|---|---|---|
| Runs test | |||||
| Runs were random | 6 | 5 | 6 | 8 | 8 |
| Too few runs | 2 | 3 | 2 | 0 | 0 |
| No. curves exhibiting serial correlation determined by DW statistic (α = 0.01) | |||||
| No serial correlation | 7 | 7 | 7 | 8 | 8 |
| Positive Correlation | 1 | 1 | 1 | 0 | 1 |
| BF | |||||
| SStudy 1-Average (±SE) | 0.996 (0.002) | 0.993 (0.001) | 0.994 (0.000) | 1.017 (0.001) | 1.003 (0.001) |
| SStudy 2-Average (±SE) | 1.001 (0.003) | 0.994 (0.002) | 0.994 (0.003) | 1.006 (0.003) | 1.001 (0.000) |
| SOverall-Average (±SE) | 0.999 (0.002) | 0.994 (0.001) | 0.994 (0.001) | 1.011 (0.003) | 1.002 (0.001) |
| Cross-Validation Test | Model | ||
|---|---|---|---|
| LOG | log × hyp | log × exp | |
| PRESS | |||
| Dataset 1 | 0.0717 | 0.0718 | 0.0599 * |
| Dataset 2 | 0.0110 * | 0.0125 | 0.0134 |
| Dataset 5 | 0.2640 | 0.2135 | 0.1858 * |
| Dataset 6 | 0.1184 | 0.1153 | 0.1149 * |
| CCC | |||
| Dataset 1 | 0.9945 | 0.9945 | 0.9953 * |
| Dataset 2 | 0.9990 * | 0.9989 | 0.9988 |
| Dataset 5 | 0.9726 | 0.9782 | 0.9814 * |
| Dataset 6 | 0.9806 | 0.9807 | 0.9808 * |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Powell, C.D.; López, S.; France, J. New Insights into Modelling Bacterial Growth with Reference to the Fish Pathogen Flavobacterium psychrophilum. Animals 2020, 10, 435. https://doi.org/10.3390/ani10030435
Powell CD, López S, France J. New Insights into Modelling Bacterial Growth with Reference to the Fish Pathogen Flavobacterium psychrophilum. Animals. 2020; 10(3):435. https://doi.org/10.3390/ani10030435
Chicago/Turabian StylePowell, Christopher D., Secundino López, and James France. 2020. "New Insights into Modelling Bacterial Growth with Reference to the Fish Pathogen Flavobacterium psychrophilum" Animals 10, no. 3: 435. https://doi.org/10.3390/ani10030435
APA StylePowell, C. D., López, S., & France, J. (2020). New Insights into Modelling Bacterial Growth with Reference to the Fish Pathogen Flavobacterium psychrophilum. Animals, 10(3), 435. https://doi.org/10.3390/ani10030435






