Molecular Typing and Antimicrobial Susceptibility Profile of Staphylococcus aureus Isolates Recovered from Bovine Mastitis and Nasal Samples
Abstract
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. S. aureus Isolates from Milk Samples
2.2. S. aureus Isolates from Muzzle/Nare Samples
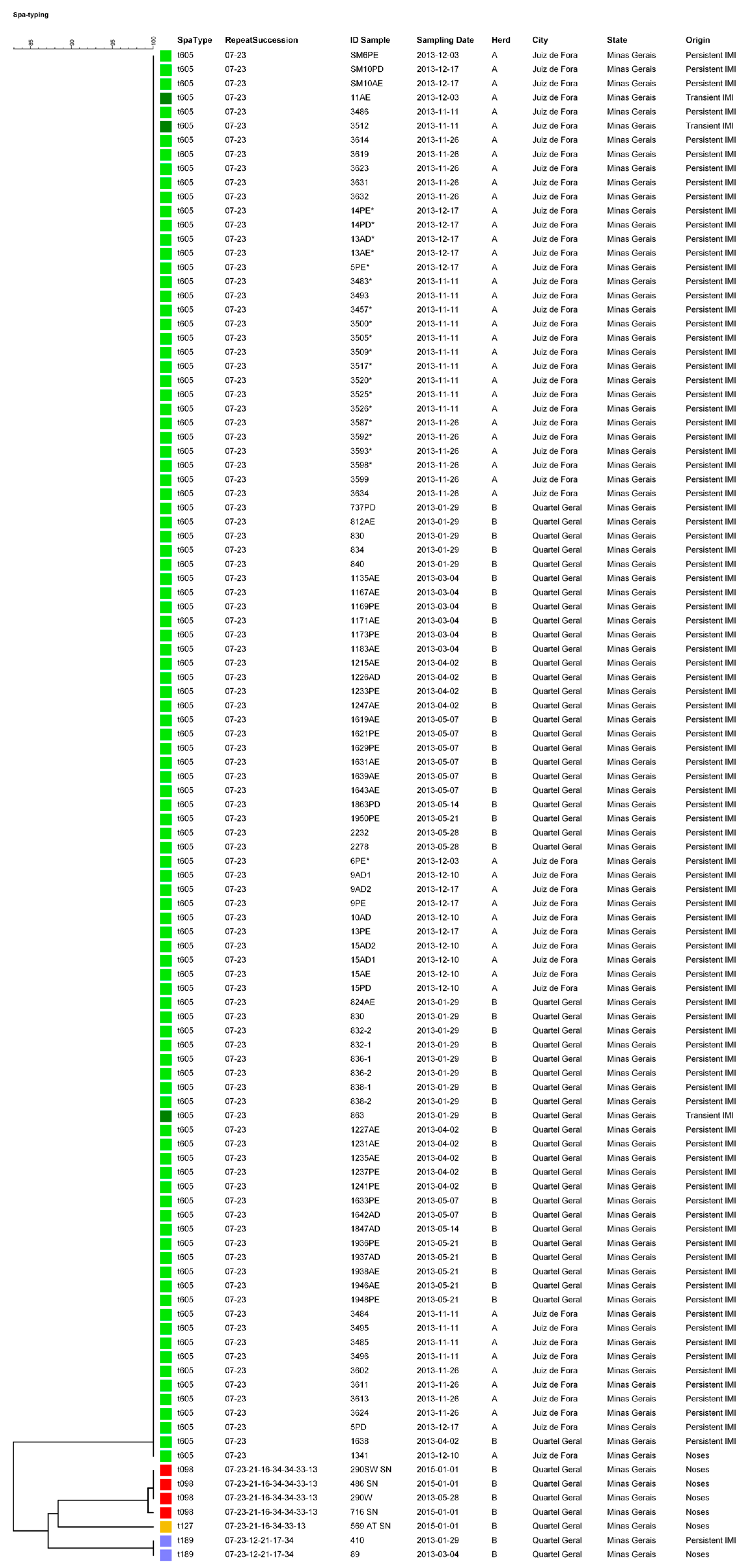
2.3. Spa Typing
2.4. Antimicrobial Susceptibility Tests
3. Results
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Cunha, A.; Andrade, H.; Souza, F.; Júnior, L.F.; Rosa, D.; Sanchez, E.R.; Gidlund, M.; Goto, H.; Brito, M.; Guimarães, A.; et al. Comparison of antibody repertories against Staphylococcus aureus in healthy and infected dairy cows with a distinct mastitis history and vaccinated with a polyvalent mastitis vaccine. J. Dairy Sci. 2020, 103, 4588–4605. [Google Scholar] [CrossRef]
- Richardson, E.J.; Bacigalupe, R.; Harrison, E.M.; Weinert, L.A.; Lycett, S.; Vrieling, M.; Robb, K.; Hoskisson, P.A.; Holden, M.T.; Feil, E.J.; et al. Gene exchange drives the ecological success of a multi-host bacterial pathogen. Nat. Ecol. Evol. 2018, 2, 1468–1478. [Google Scholar] [CrossRef] [PubMed]
- Štěpán, J.; Pantůček, R.; Doškař, J. Molecular diagnostics of clinically important staphylococci. Folia Microbiol. 2004, 49, 353–386. [Google Scholar] [CrossRef]
- Harmsen, D.; Claus, H.; Witte, W.; Rothgänger, J.; Turnwald, D.; Vogel, U. Typing of Methicillin-Resistant Staphylococcus aureus in a University Hospital Setting by Using Novel Software for spa Repeat Determination and Database Management. J. Clin. Microbiol. 2003, 41, 5442–5448. [Google Scholar] [CrossRef] [PubMed]
- Souza, F.N.; Cunha, A.F.; Rosa, D.L.; Brito, M.A.V.; Guimarães, A.S.; Mendonça, L.C.; Souza, G.N.; Lage, A.P.; Blagitz, M.G.; Della Libera, A.M.; et al. Somatic cell count and mastitis pathogen detection in composite and single or duplicate quarter milk samples. Pesquisa Veterinária Brasileira 2016, 36, 811–818. [Google Scholar] [CrossRef]
- Nonnemann, B.; Lyhs, U.; Svennesen, L.; Kristensen, K.A.; Klaas, I.C.; Pedersen, K. Bovine mastitis bacteria resolved by MALDI-TOF mass spectrometry. J. Dairy Sci. 2019, 102, 2515–2524. [Google Scholar] [CrossRef]
- Sasaki, T.; Tsubakishita, S.; Tanaka, Y.; Sakusabe, A.; Ohtsuka, M.; Hirotaki, S.; Kawakami, T.; Fukata, T.; Hiramatsu, K. Multiplex-PCR Method for Species Identification of Coagulase-Positive Staphylococci. J. Clin. Microbiol. 2010, 48, 765–769. [Google Scholar] [CrossRef]
- Roberson, J.R.; Fox, L.K.; Hancock, D.D.; Gay, J.M.; Besser, T.E. Ecology of Staphylcoccus aureus isolated from various sites on dairy farms. J. Dairy Sci. 1994, 77, 3354–3364. [Google Scholar]
- Fan, H.H.; Kleven, S.H.; Jackwood, M.W. Application of Polymerase Chain Reaction with Arbitrary Primers to Strain Identification of Mycoplasma gallisepticum. Avian Dis. 1995, 39, 729. [Google Scholar] [CrossRef]
- Magiorakos, A.-P.; Srinivasan, A.; Carey, R.; Carmeli, Y.; Falagas, M.; Giske, C.; Harbarth, S.; Hindler, J.; Kahlmeter, G.; Olsson-Liljequist, B.; et al. Multidrug-resistant, extensively drug-resistant and pandrug-resistant bacteria: An international expert proposal for interim standard definitions for acquired resistance. Clin. Microbiol. Infect. 2012, 18, 268–281. [Google Scholar] [CrossRef]
- CLSI. Performance Standards for Antimicrobial Disk and Dilution Susceptibility Test for Bacteria Isolated from Animals, 5th ed.; Clinical and Laboratory Standards Institute: Wayne, PA, USA, 2018; ISBN 978-1-68440-008-9. [Google Scholar]
- CLSI. Performance Standards for Antimicrobial Disk and Dilution Susceptibility Test for Bacteria Isolated from Animals, 4th ed.; Clinical and Laboratory Standards Institute: Wayne, PA, USA, 2018; ISBN 978-1-68440-010-2. [Google Scholar]
- EUCAST. The European Committee on Antimicrobial Susceptibility Testing. Breakpoints Tables for Interpretation of MICs and Zone Diameters–Version 9.0. 2019. Available online: http://www.eucast.org/fileadmin/src/media/PDFs/EUCAST_files/Breakpoint_tables/v_9.0_Breakpoint_Tables.pdf (accessed on 13 June 2020).
- Mehrotra, M.; Wang, G.; Johnson, W.M. Multiplex PCR for Detection of Genes forStaphylococcus aureus Enterotoxins, Exfoliative Toxins, Toxic Shock Syndrome Toxin 1, and Methicillin Resistance. J. Clin. Microbiol. 2000, 38, 1032–1035. [Google Scholar] [CrossRef] [PubMed]
- Paterson, G.K.; Larsen, A.R.; Robb, A.; Edwards, G.E.; Pennycott, T.W.; Foster, G.; Mot, D.; Hermans, K.; Baert, K.; Peacock, S.J.; et al. The newly described mecA homologue, mecALGA251, is present in methicillin-resistant Staphylococcus aureus isolates from a diverse range of host species. J. Antimicrob. Chemother. 2012, 67, 2809–2813. [Google Scholar] [CrossRef] [PubMed]
- Capurro, A.; Aspán, A.; Unnerstad, H.E.; Waller, K.P.; Artursson, K. Identification of potential sources of Staphylococcus aureus in herds with mastitis problems. J. Dairy Sci. 2010, 93, 180–191. [Google Scholar] [CrossRef]
- Adkins, P.R.F.; Dufour, S.; Spain, J.; Calcutt, M.; Reilly, T.; Stewart, G.; Middleton, J. Molecular characterization of non-aureus Staphylococcus spp. from heifer intramammary infections and body sites. J. Dairy Sci. 2018, 101, 5388–5403. [Google Scholar] [CrossRef]
- Leuenberger, A.; Sartori, C.; Boss, R.; Resch, G.; Oechslin, F.; Steiner, A.; Moreillon, P.; Graber, H. Genotypes of Staphylococcus aureus: On-farm epidemiology and the consequences for prevention of intramammary infections. J. Dairy Sci. 2019, 102, 3295–3309. [Google Scholar] [CrossRef]
- Paduch, J.-H.; Krömker, V. Colonization of the teat skin and the teat canal of lactating dairy cattle by mastitis pathogens. Tierärztl. Prax. Ausg. G: Großtiere/Nutztiere 2011, 39, 71–76. [Google Scholar]
- Srednik, M.E.; Usongo, V.; Lépine, S.; Janvier, X.; Archambault, M.; Gentilini, E.R. Characterization of Staphylococcus aureus strains isolated from mastitis bovine milk in Argentina. J. Dairy Res. 2018, 85, 57–63. [Google Scholar]
- Silva, N.; Guimarães, F.; Manzi, M.; Budri, P.; Gómez-Sanz, E.; Benito, D.; Langoni, H.; Rall, V.; Torres, C. Molecular characterization and clonal diversity of methicillin-susceptible Staphylococcus aureus in milk of cows with mastitis in Brazil. J. Dairy Sci. 2013, 96, 6856–6862. [Google Scholar] [CrossRef]
- Mesquita, A.A.; Rocha, C.M.; Bruhn, F.R.; Custódio, D.A.; Braz, M.S.; Pinto, S.M.; Silva, D.B.; Da Costa, G.M. Staphylococcus aureus and Streptococcus agalactiae: Prevalence, resistance to antimicrobials, and their relationship with the milk quality of dairy cattle herds in Minas Gerais state, Brazil. J. Pesq. Vet. Bras. 2019, 39, 308–316. [Google Scholar] [CrossRef]
- Rabello, R.F.; Bonelli, R.R.; Penna, B.A.; Albuquerque, J.P.; Souza, R.M.; Cerqueira, A.M.F. Antimicrobial Resistance in Farm Animals in Brazil: An Update Overview. Animals 2020, 10, 552. [Google Scholar] [CrossRef]
- Barkema, H.W.; Schukken, Y.; Zadoks, R. Invited Review: The Role of Cow, Pathogen, and Treatment Regimen in the Therapeutic Success of Bovine Staphylococcus aureus Mastitis. J. Dairy Sci. 2006, 89, 1877–1895. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Santos, R.P.; Souza, F.N.; Oliveira, A.C.D.; de Souza Filho, A.F.; Aizawa, J.; Moreno, L.Z.; da Cunha, A.F.; Cortez, A.; Della Libera, A.M.M.P.; Heinemann, M.B.; et al. Molecular Typing and Antimicrobial Susceptibility Profile of Staphylococcus aureus Isolates Recovered from Bovine Mastitis and Nasal Samples. Animals 2020, 10, 2143. https://doi.org/10.3390/ani10112143
Santos RP, Souza FN, Oliveira ACD, de Souza Filho AF, Aizawa J, Moreno LZ, da Cunha AF, Cortez A, Della Libera AMMP, Heinemann MB, et al. Molecular Typing and Antimicrobial Susceptibility Profile of Staphylococcus aureus Isolates Recovered from Bovine Mastitis and Nasal Samples. Animals. 2020; 10(11):2143. https://doi.org/10.3390/ani10112143
Chicago/Turabian StyleSantos, Renata P., Fernando N. Souza, Ana Claudia D. Oliveira, Antônio F. de Souza Filho, Juliana Aizawa, Luisa Z. Moreno, Adriano F. da Cunha, Adriana Cortez, Alice M.M.P. Della Libera, Marcos B. Heinemann, and et al. 2020. "Molecular Typing and Antimicrobial Susceptibility Profile of Staphylococcus aureus Isolates Recovered from Bovine Mastitis and Nasal Samples" Animals 10, no. 11: 2143. https://doi.org/10.3390/ani10112143
APA StyleSantos, R. P., Souza, F. N., Oliveira, A. C. D., de Souza Filho, A. F., Aizawa, J., Moreno, L. Z., da Cunha, A. F., Cortez, A., Della Libera, A. M. M. P., Heinemann, M. B., & Cerqueira, M. M. O. P. (2020). Molecular Typing and Antimicrobial Susceptibility Profile of Staphylococcus aureus Isolates Recovered from Bovine Mastitis and Nasal Samples. Animals, 10(11), 2143. https://doi.org/10.3390/ani10112143






