Biodiversity of Phages Infecting the Dairy Bacterium Streptococcus thermophilus
Abstract
:1. Introduction
2. Current Taxonomy of Dairy Streptococcal Phages
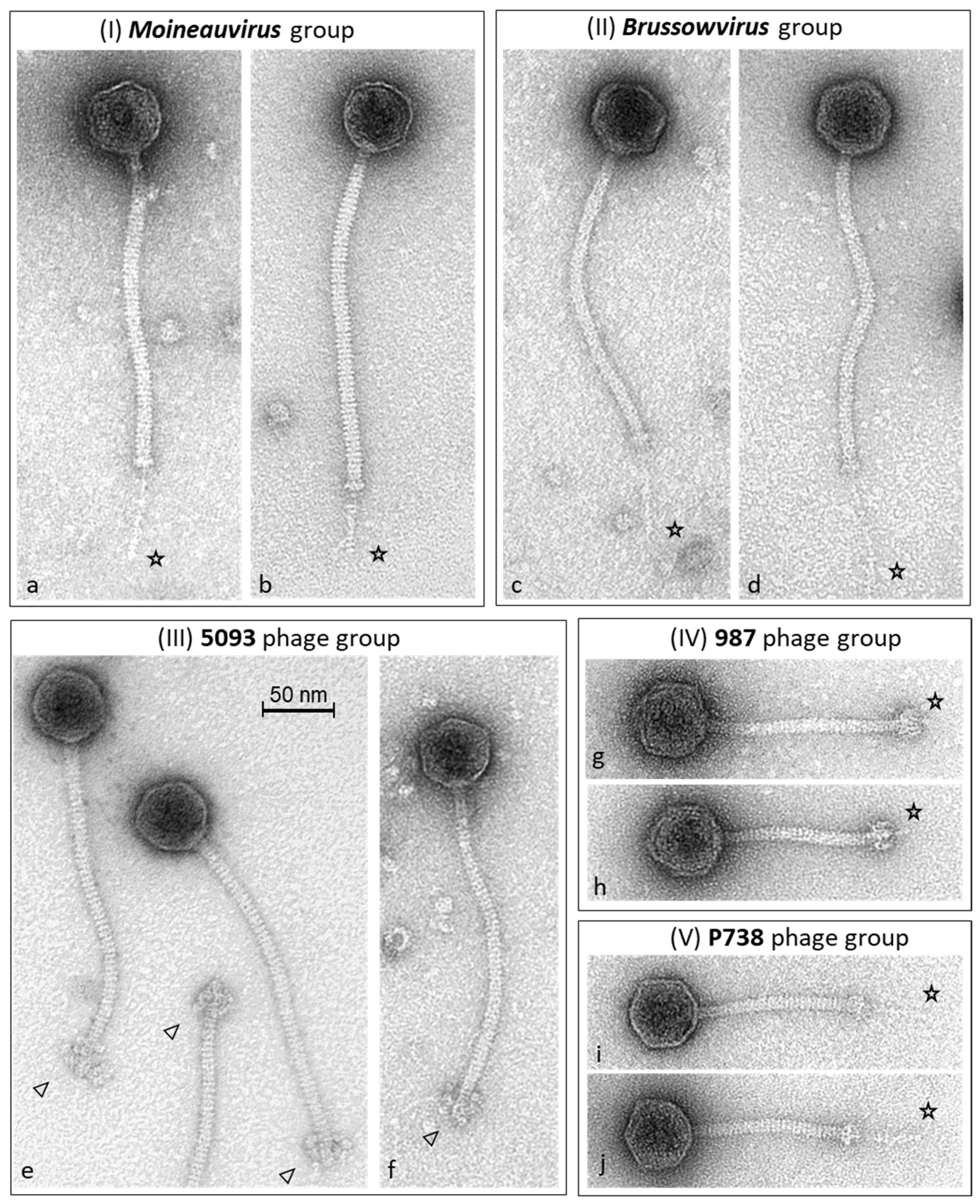
2.1. Biodiversity and Morphological Characteristics of Dairy Streptococcal Phages
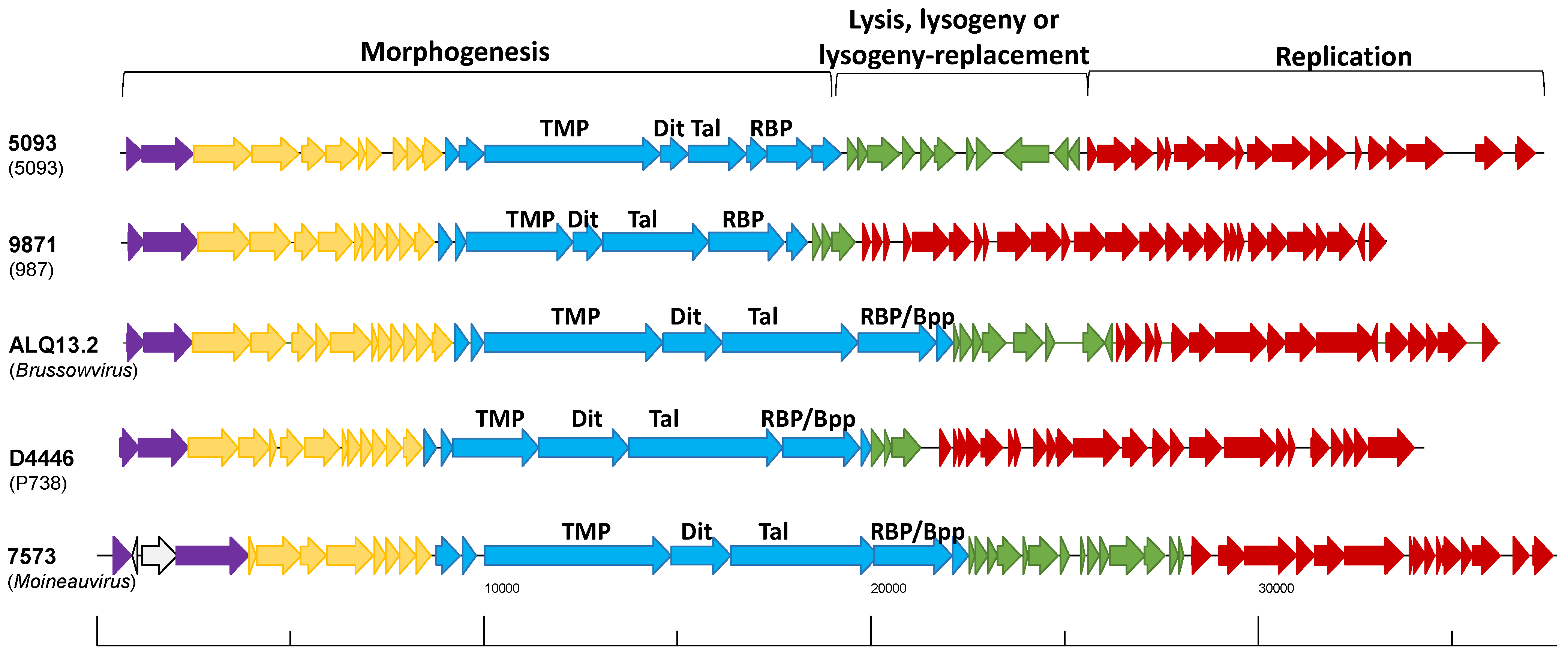
2.2. Genetic Diversity of Dairy Streptococcal Phages
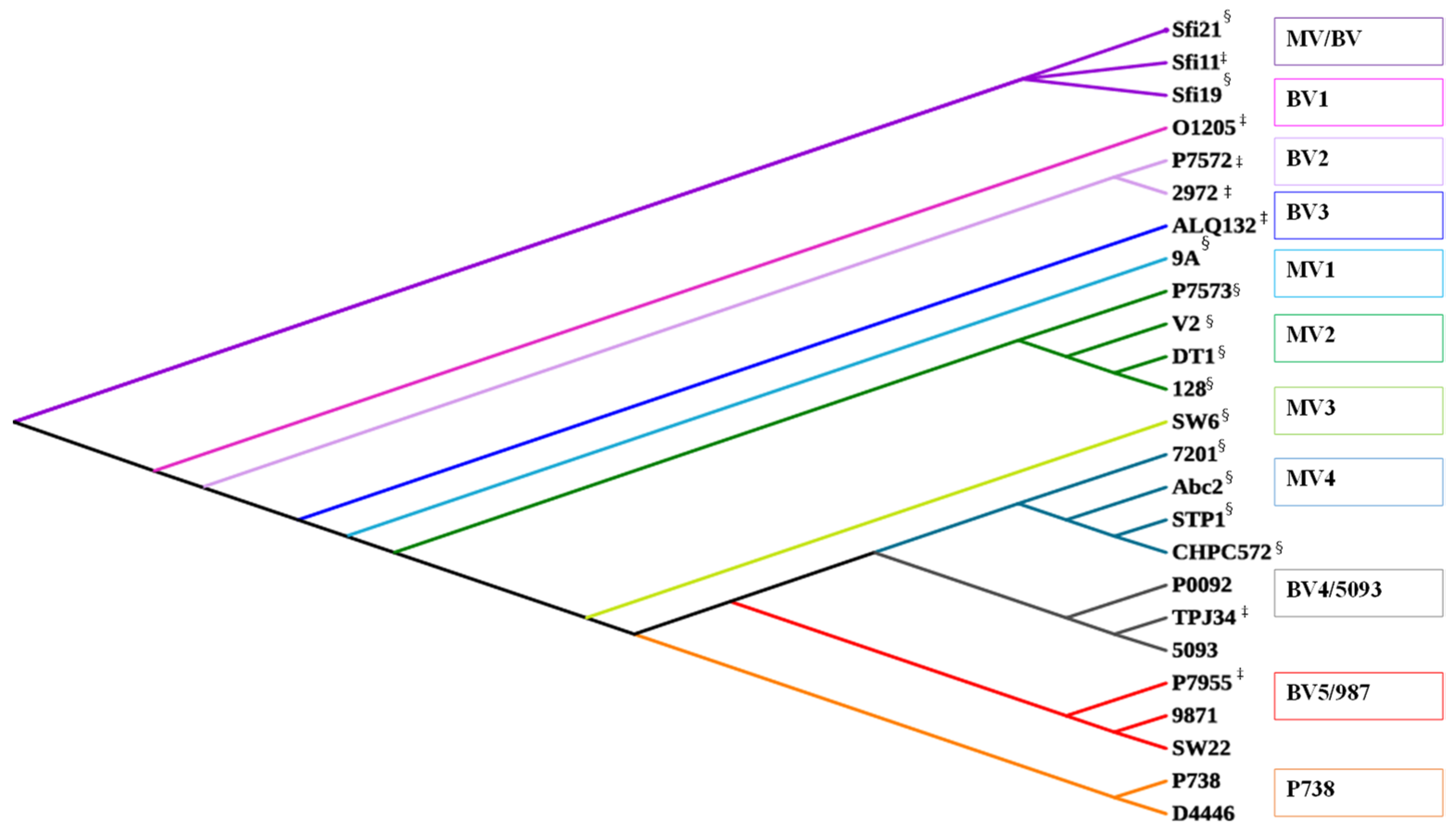
2.2.1. Replication Modules Display Inter-Group Relatedness
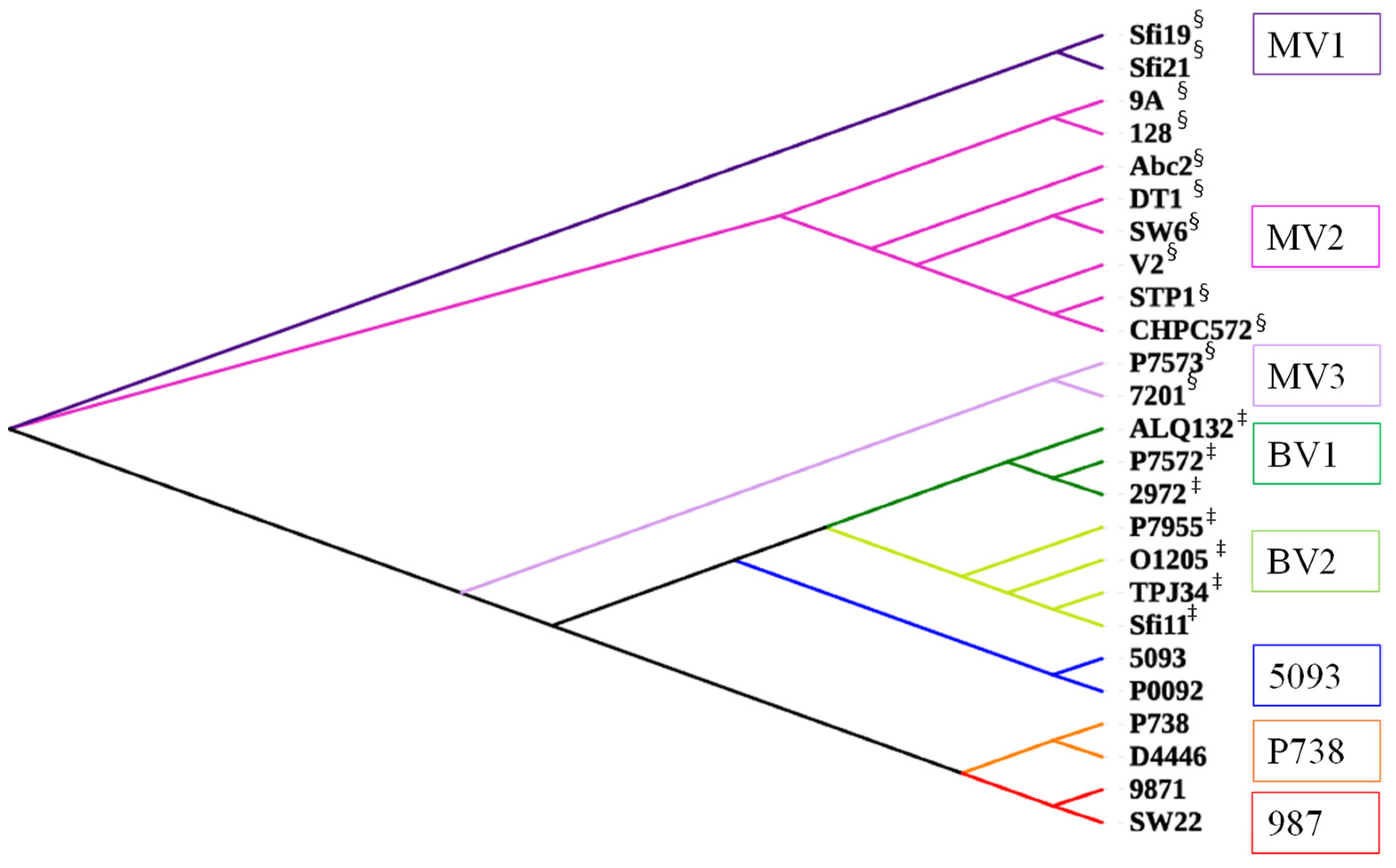
2.2.2. Morphogenesis Modules Are Highly Group-Specific
2.3. Evolutionary Pathways of Dairy Streptococcal Phages
3. Interactions of Dairy Streptococcal Phages and Their Hosts
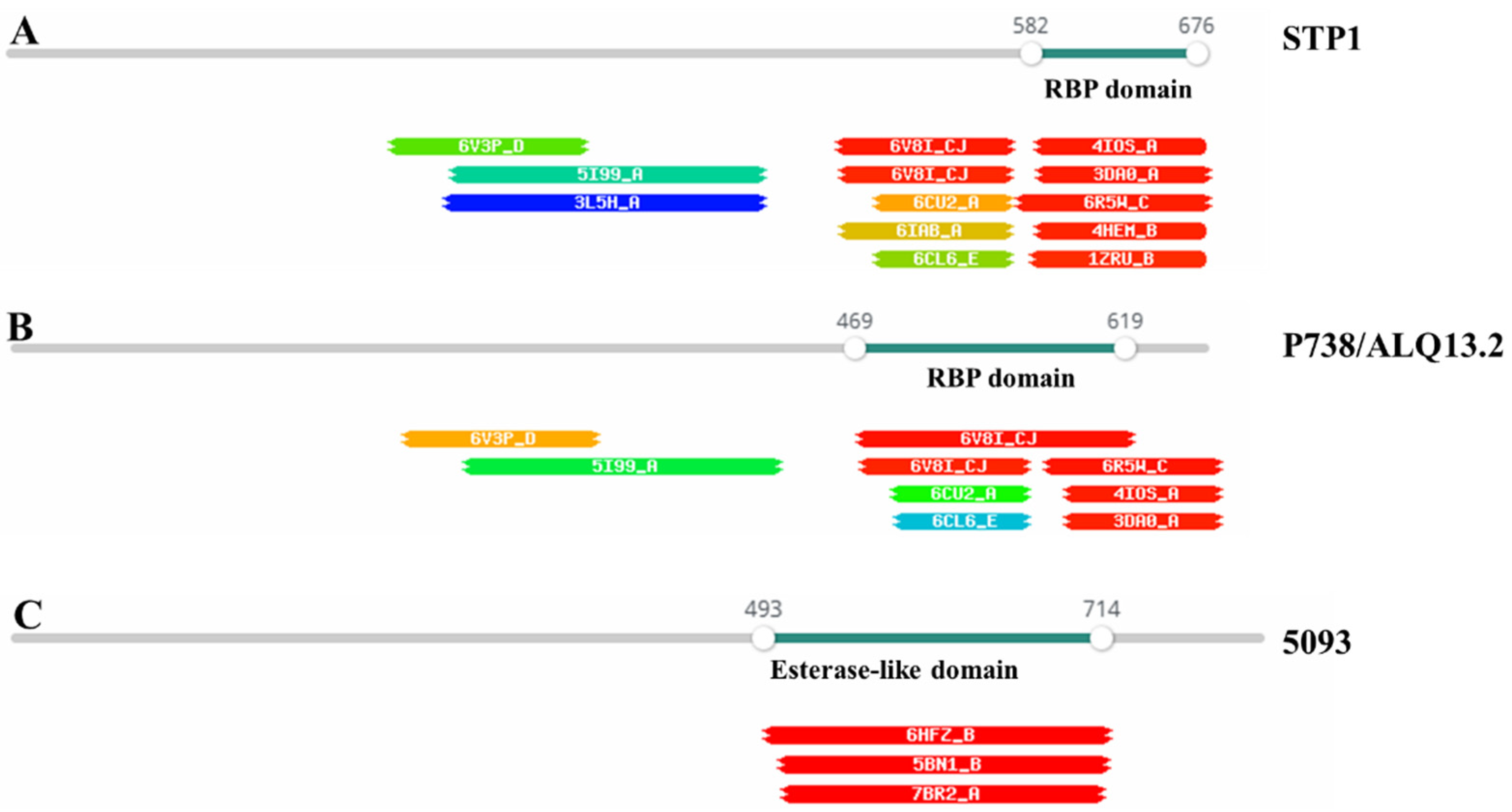
3.1. Phage Adhesion Device
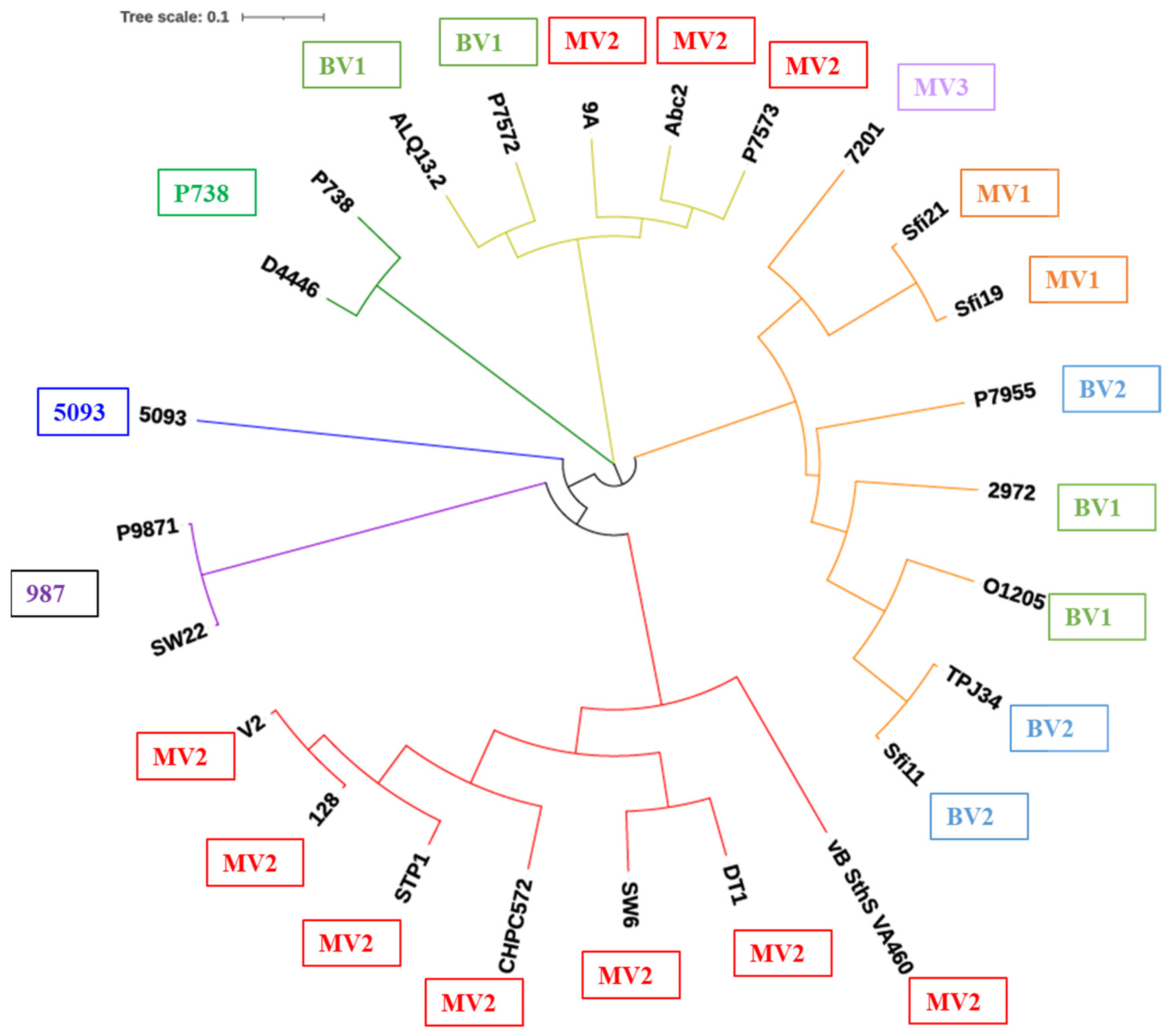
3.2. RBP Phylogeny Is Not Necessarily Linked to That of the Overall Morphogenesis Module
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Alexandraki, V.; Kazou, M.; Blom, J.; Pot, B.; Papadimitriou, K.; Tsakalidou, E. Comparative genomics of Stretococcus thermophilus support important traits concerning the evolution, biology and technological properties of the species. Front. Microbiol. 2019, 10, 2916. [Google Scholar] [CrossRef]
- Deveau, H.; Garneau, J.E.; Moineau, S. CRISPR/Cas system and its role in phage-bacteria interactions. Annu. Rev. Microbiol. 2010, 64, 475–493. [Google Scholar] [CrossRef]
- Garneau, J.E.; Dupuis, M.E.; Villion, M.; Romero, D.A.; Barrangou, R.; Boyaval, P.; Fremaux, C.; Horvath, P.; Magadan, A.H.; Moineau, S. The CRISPR/Cas bacterial immune system cleaves bacteriophage and plasmid DNA. Nature 2010, 468, 67–71. [Google Scholar] [CrossRef] [PubMed]
- Philippe, C.; Moineau, S. The endless battle between phages and CRISPR-Cas systems in Streptococcus thermophilus. Biochem. Cell. Biol. 2021, 99. [Google Scholar] [CrossRef] [PubMed]
- Deveau, H.; Barrangou, R.; Garneau, J.E.; Labonte, J.; Fremaux, C.; Boyaval, P.; Romero, D.A.; Horvath, P.; Moineau, S. Phage response to CRISPR-encoded resistance in Streptococcus thermophilus. J. Bacteriol. 2008, 190, 1390–1400. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hynes, A.P.; Rousseau, G.M.; Agudelo, D.; Goulet, A.; Amigues, B.; Loehr, J.; Romero, D.A.; Fremaux, C.; Horvath, P.; Doyon, Y.; et al. Widespread anti-CRISPR proteins in virulent bacteriophages inhibit a range of Cas9 proteins. Nat. Commun. 2018, 9, 2919. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Brussow, H.; Fremont, M.; Bruttin, A.; Sidoti, J.; Constable, A.; Fryder, V. Detection and classification of Streptococcus thermophilus bacteriophages isolated from industrial milk fermentation. Appl. Environ. Microbiol. 1994, 60, 4537–4543. [Google Scholar] [CrossRef] [Green Version]
- Carminati, D.; Giraffa, G. Evidence and characterization of temperate bacteriophage in Streptococcus salivarius subsp. thermophilus St18. J. Dairy Res. 1992, 59, 71–79. [Google Scholar] [CrossRef] [PubMed]
- Le Marrec, C.; van Sinderen, D.; Walsh, L.; Stanley, E.; Vlegels, E.; Moineau, S.; Heinze, P.; Fitzgerald, G.; Fayard, B. Two groups of bacteriophages infecting Streptococcus thermophilus can be distinguished on the basis of mode of packaging and genetic determinants for major structural proteins. Appl. Environ. Microbiol. 1997, 63, 3246–3253. [Google Scholar] [CrossRef] [Green Version]
- Adriaenssens, E.M.; Wittmann, J.; Kuhn, J.H.; Turner, D.; Sullivan, M.B.; Dutilh, B.E.; Jang, H.B.; van Zyl, L.J.; Klumpp, J.; Lobocka, M.; et al. Taxonomy of prokaryotic viruses: 2017 update from the ICTV Bacterial and Archaeal Viruses subcommittee. Arch. Virol. 2018, 163, 1125–1129. [Google Scholar] [CrossRef] [Green Version]
- Mills, S.; Griffin, C.; O’Sullivan, O.; Coffey, A.; McAuliffe, O.E.; Meijer, W.C.; Serrano, L.M.; Ross, R.P. A new phage on the ‘Mozzarella’ block: Bacteriophage 5093 shares a low level of homology with other Streptococcus thermophilus phages. Int. Dairy J. 2011, 21, 963–969. [Google Scholar] [CrossRef]
- Szymczak, P.; Janzen, T.; Neves, A.R.; Kot, W.; Hansen, L.H.; Lametsch, R.; Neve, H.; Franz, C.; Vogensen, F.K. Novel Variants of Streptococcus thermophilus Bacteriophages Are Indicative of Genetic Recombination among Phages from Different Bacterial Species. Appl. Environ. Microbiol. 2017, 83, e02748-16. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- McDonnell, B.; Mahony, J.; Neve, H.; Hanemaaijer, L.; Noben, J.P.; Kouwen, T.; van Sinderen, D. Identification and analysis of a novel group of bacteriophages infecting the lactic acid bacterium Streptococcus thermophilus. Appl. Environ. Microbiol. 2016, 82, 5153–5165. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Philippe, C.; Levesque, S.; Dion, M.B.; Tremblay, D.M.; Horvath, P.; Luth, N.; Cambillau, C.; Franz, C.; Neve, H.; Fremaux, C.; et al. Novel Genus of Phages Infecting Streptococcus thermophilus: Genomic and Morphological Characterization. Appl. Environ. Microbiol. 2020, 86, e00227-20. [Google Scholar] [CrossRef]
- Mahony, J.; Van Sinderen, D. Current taxonomy of phages infecting lactic acid bacteria. Front. Microbiol. 2014, 5, 7. [Google Scholar] [CrossRef] [Green Version]
- Lavelle, K.; Martinez, I.; Neve, H.; Lugli, G.A.; Franz, C.M.A.P.; Ventura, M.; Bello, F.D.; Sinderen, D.v.; Mahony, J. Biodiversity of Streptococcus thermophilus phages in global dairy fermentations. Viruses 2018, 10, 577. [Google Scholar] [CrossRef] [Green Version]
- Lavelle, K.; Murphy, J.; Fitzgerald, B.; Lugli, G.A.; Zomer, A.; Neve, H.; Ventura, M.; Franz, C.M.; Cambillau, C.; van Sinderen, D.; et al. A decade of Streptococcus thermophilus phage evolution in an Irish dairy plant. Appl. Environ. Microbiol. 2018, 84, e02855-17. [Google Scholar] [CrossRef] [Green Version]
- Szymczak, P.; Rau, M.H.; Monteiro, J.M.; Pinho, M.G.; Filipe, S.R.; Vogensen, F.K.; Zeidan, A.A.; Janzen, T. A comparative genomics approach for identifying host–range determinants in Streptococcus thermophilus bacteriophages. Sci. Rep. 2019, 9, 7991. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Romero, D.A.; Magill, D.; Millen, A.; Horvath, P.; Fremaux, C. Dairy lactococcal and streptococcal phage-host interactions: An industrial perspective in an evolving phage landscape. FEMS Microbiol. Rev. 2020, 44, 909–932. [Google Scholar] [CrossRef] [PubMed]
- McDonnell, B.; Mahony, J.; Hanemaaijer, L.; Neve, H.; Noben, J.-P.; Lugli, G.A.; Ventura, M.; Kouwen, T.R.; van Sinderen, D. Global survey and genome exploration of bacteriophages infecting the lactic acid bacterium Streptococcus thermophilus. Front. Microbiol. 2017, 8, 1754. [Google Scholar] [CrossRef] [PubMed]
- Lucchini, S.; Desiere, F.; Brüssow, H. Comparative genomics of Streptococcus thermophilus phage species supports a modular evolution theory. J. Virol. 1999, 73, 8647–8656. [Google Scholar] [CrossRef] [Green Version]
- Stanley, E.; Walsh, L.; van der Zwet, A.; Fitzgerald, G.F.; van Sinderen, D. Identification of four loci isolated from two Streptococcus thermophilus phage genomes responsible for mediating bacteriophage resistance. FEMS Microbiol. Lett. 2000, 182, 271–277. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Duplessis, M.; Moineau, S. Identification of a genetic determinant responsible for host specificity in Streptococcus thermophilus bacteriophages. Mol. Microbiol. 2001, 41, 325–336. [Google Scholar] [CrossRef] [PubMed]
- Duplessis, M.; Levesque, C.M.; Moineau, S. Characterization of Streptococcus thermophilus host range phage mutants. Appl. Environ. Microbiol. 2006, 72, 3036–3041. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lavelle, K.; Goulet, A.; McDonnell, B.; Spinelli, S.; van Sinderen, D.; Mahony, J.; Cambillau, C. Revisiting the host adhesion determinants of Streptococcus thermophilus siphophages. Microb. Biotechnol. 2020, 13, 1765–1779. [Google Scholar] [CrossRef]
- Zimmermann, L.; Stephens, A.; Nam, S.-Z.; Rau, D.; Kübler, J.; Lozajic, M.; Gabler, F.; Söding, J.; Lupas, A.N.; Alva, V. A completely reimplemented MPI bioinformatics toolkit with a new HHpred server at its core. J. Mol. Biol. 2018, 430, 2237–2243. [Google Scholar] [CrossRef]
- Sadovskaya, I.; Vinogradov, E.; Courtin, P.; Armalyte, J.; Meyrand, M.; Giaouris, E.; Palussiere, S.; Furlan, S.; Pechoux, C.; Ainsworth, S.; et al. Another brick in the wall: A rhamnan polysaccharide trapped inside peptidoglycan of Lactococcus lactis. Mbio 2017, 8. [Google Scholar] [CrossRef] [Green Version]
- McDonnell, B.; Hanemaaijer, L.; Bottacini, F.; Kelleher, P.; Lavelle, K.; Sadovskaya, I.; Vinogradov, E.; Ver Loren van Themaat, E.; Kouwen, T.; Mahony, J.; et al. A cell wall-associated polysaccharide is required for bacteriophage adsorption to the Streptococcus thermophilus cell surface. Mol. Microbiol. 2020, 114, 31–45. [Google Scholar] [CrossRef] [PubMed]
- Bourgoin, F.; Pluvinet, A.; Gintz, B.; Decaris, B.; Guedon, G. Are horizontal transfers involved in the evolution of the Streptococcus thermophilus exopolysaccharide synthesis loci? Gene 1999, 233, 151–161. [Google Scholar] [CrossRef]
- Knirel, Y.A.; Van Calsteren, M.-R. Bacterial exopolysaccharides. In Reference Module in Chemistry, Molecular Sciences and Chemical Engineering; Elsevier Inc.: Amsterdam, The Netherlands, 2020. [Google Scholar] [CrossRef]
- Brown, S.; Santa Maria, J.P., Jr.; Walker, S. Wall teichoic acids of Gram-positive bacteria. Annu. Rev. Microbiol. 2013, 67, 313–336. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Siponen, M.; Spinelli, S.; Blangy, S.; Moineau, S.; Cambillau, C.; Campanacci, V. Crystal structure of a chimeric receptor binding protein constructed from two lactococcal phages. J. Bacteriol. 2009, 191, 3220–3225. [Google Scholar] [CrossRef] [Green Version]
- Dunne, M.; Hupfeld, M.; Klumpp, J.; Loessner, M.J. Molecular basis of bacterial host interactions by Gram-positive targeting bacteriophages. Viruses 2018, 10, 397. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Szymczak, P.; Filipe, S.R.; Covas, G.; Vogensen, F.K.; Neves, A.R.; Janzen, T. Cell Wall Glycans Mediate Recognition of the Dairy Bacterium Streptococcus thermophilus by Bacteriophages. Appl. Environ. Microbiol. 2018, 84, e01847-18. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Vale, P.F.; Lafforgue, G.; Gatchitch, F.; Gardan, R.; Moineau, S.; Gandon, S. Costs of CRISPR-Cas-mediated resistance in Streptococcus thermophilus. Proc. Biol. Sci. 2015, 282, 20151270. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Horvath, P.; Romero, D.A.; Coute-Monvoisin, A.C.; Richards, M.; Deveau, H.; Moineau, S.; Boyaval, P.; Fremaux, C.; Barrangou, R. Diversity, activity, and evolution of CRISPR loci in Streptococcus thermophilus. J. Bacteriol. 2008, 190, 1401–1412. [Google Scholar] [CrossRef] [Green Version]
- Achigar, R.; Magadan, A.H.; Tremblay, D.M.; Julia Pianzzola, M.; Moineau, S. Phage-host interactions in Streptococcus thermophilus: Genome analysis of phages isolated in Uruguay and ectopic spacer acquisition in CRISPR array. Sci. Rep. 2017, 7, 43438. [Google Scholar] [CrossRef]

| Phage | Phage Group | Replication Module Type | Morphogenesis Module Type |
|---|---|---|---|
| 9A | Moineauvirus | MV1 | MV2 |
| V2, DT1, 128 | Moineauvirus | MV2 | MV2 |
| P7573 | Moineauvirus | MV2 | MV3 |
| SW6 | Moineauvirus | MV3 | MV2 |
| STP1, CHPC572, Abc2 | Moineauvirus | MV4 | MV2 |
| 7201 | Moineauvirus | MV4 | MV3 |
| Sfi19, Sfi21 | Moineauvirus | MV/BV | MV1 |
| Sfi11 | Brussowvirus | MV/BV | BV2 |
| O1205 | Brussowvirus | BV1 | BV2 |
| P7572, 2972 | Brussowvirus | BV2 | BV1 |
| ALQ13.2 | Brussowvirus | BV3 | BV1 |
| TP-J34 | Brussowvirus | BV4/5093 | BV2 |
| P7955 | Brussowvirus | BV5/987 | BV2 |
| 9871, SW22 | 987 | BV5/987 | 987 |
| 5093, P0092 | 5093 | BV4/5093 | 5093 |
| P738, D4446 | P738 | P738 (S. pyogenes phage T12) | P738 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hanemaaijer, L.; Kelleher, P.; Neve, H.; Franz, C.M.A.P.; de Waal, P.P.; van Peij, N.N.M.E.; van Sinderen, D.; Mahony, J. Biodiversity of Phages Infecting the Dairy Bacterium Streptococcus thermophilus. Microorganisms 2021, 9, 1822. https://doi.org/10.3390/microorganisms9091822
Hanemaaijer L, Kelleher P, Neve H, Franz CMAP, de Waal PP, van Peij NNME, van Sinderen D, Mahony J. Biodiversity of Phages Infecting the Dairy Bacterium Streptococcus thermophilus. Microorganisms. 2021; 9(9):1822. https://doi.org/10.3390/microorganisms9091822
Chicago/Turabian StyleHanemaaijer, Laurens, Philip Kelleher, Horst Neve, Charles M. A. P. Franz, Paul P. de Waal, Noël N. M. E. van Peij, Douwe van Sinderen, and Jennifer Mahony. 2021. "Biodiversity of Phages Infecting the Dairy Bacterium Streptococcus thermophilus" Microorganisms 9, no. 9: 1822. https://doi.org/10.3390/microorganisms9091822
APA StyleHanemaaijer, L., Kelleher, P., Neve, H., Franz, C. M. A. P., de Waal, P. P., van Peij, N. N. M. E., van Sinderen, D., & Mahony, J. (2021). Biodiversity of Phages Infecting the Dairy Bacterium Streptococcus thermophilus. Microorganisms, 9(9), 1822. https://doi.org/10.3390/microorganisms9091822







