Abstract
Escherichia (E.) coli is the main causative pathogen of neonatal and post-weaning diarrhea and edema disease in swine production. There is a significant health concern due to an increasing number of human infections associated with food and/or environmental-borne pathogenic and multidrug-resistant E. coli worldwide. Monitoring the presence of pathogenic and antimicrobial-resistant E. coli isolates is essential for sustainable disease management in livestock and human medicine. A total of 102 E. coli isolates of diseased pigs were characterized by antimicrobial and biocide susceptibility testing. Antimicrobial resistance genes, including mobile colistin resistance genes, were analyzed by PCR and DNA sequencing. The quinolone resistance-determining regions of gyrA and parC in ciprofloxacin-resistant isolates were analyzed. Clonal relatedness was investigated by two-locus sequence typing (CH clonotyping). Phylotyping was performed by the Clermont multiplex PCR method. Virulence determinants were analyzed by customized DNA-based microarray technology developed in this study for fast and economic molecular multiplex typing. Thirty-five isolates were selected for whole-genome sequence-based analysis. Most isolates were resistant to ampicillin and tetracycline. Twenty-one isolates displayed an ESBL phenotype and one isolate an AmpC β-lactamase-producing phenotype. Three isolates had elevated colistin minimal inhibitory concentrations and carried the mcr-1 gene. Thirty-seven isolates displayed a multi-drug resistance phenotype. The most predominant β-lactamase gene classes were blaTEM-1 (56%) and blaCTX-M-1 (13.71%). Mutations in QRDR were observed in 14 ciprofloxacin-resistant isolates. CH clonotyping divided all isolates into 51 CH clonotypes. The majority of isolates belonged to phylogroup A. Sixty-four isolates could be assigned to defined pathotypes wherefrom UPEC was predominant. WGS revealed that the most predominant sequence type was ST100, followed by ST10. ST131 was detected twice in our analysis. This study highlights the importance of monitoring antimicrobial resistance and virulence properties of porcine E. coli isolates. This can be achieved by applying reliable, fast, economic and easy to perform technologies such as DNA-based microarray typing. The presence of high-risk pathogenic multi-drug resistant zoonotic clones, as well as those that are resistant to critically important antibiotics for humans, can pose a risk to public health. Improved protocols may be developed in swine farms for preventing infections, as well as the maintenance and distribution of the causative isolates.
Keywords:
antimicrobial resistance; pig; E. coli; molecular characterization; microarray; colistin; WGS 1. Introduction
Escherichia (E.) coli is a facultatively anaerobic Gram-negative rod with many facets. The majority of E. coli strains inhabit the intestinal tract of humans and warm-blooded animals as commensal bacteria in a mutually beneficial association with its hosts [1,2,3]. However, some strains of E. coli have acquired virulence-associated genes (VAGs), rendering them pathogenic and empowering them to play an important role as pathogens in humans and animals [3]. E. coli is a prominent cause for a wide range of bacterial infections in swine but might also play a role as a bacterial foodborne pathogen. In particular, VAGs enable E. coli to cause enteritis, urinary tract infections, peritonitis, meningitis, and septicemia in humans. In swine, E. coli is more prominently associated with diarrhea [4]. Depending on their VAGs, their patho-mechanisms and their clinical symptoms, E. coli strains are classified into numerous pathotypes. Diarrhea-associated strains include enterotoxigenic E. coli (ETEC), enteropathogenic E. coli (EPEC), enterohemorrhagic E. coli (EHEC), enteroaggregative E. coli (EAEC), and enteroinvasive E. coli (EIEC). Extraintestinal infections are caused by extraintestinal pathogenic E. coli strains (EXPEC). EXPEC are mostly innocuous gut commensals that are harmful only if they reach other body sites. They include uropathogenic strains (UPEC), or strains that are involved in septicemia in humans and animals (SEPEC), as well as E. coli that are involved in neonatal meningitis of humans (MENEC) [4,5,6].
E. coli represents a versatile and diverse enterobacterial species with a broad genetic flexibility and adaptability to constantly changing environments [7]. E. coli has acquired antimicrobial resistance mechanisms [8]. The genetic adaptation of E. coli to antibiotic exposure may select for decreased susceptibility to several antimicrobial agents [9]. Antimicrobial resistance (AMR) is recognized as a global problem in human and veterinary medicine. The high prevalence of multidrug-resistant (MDR) bacteria causes a significant concern in public health [10]. The extended use of critically important antibiotics in livestock also affects the emergence, prevalence, and dissemination of AMR [11]. VAGs and antimicrobial resistance genes are often carried on mobile genetic elements that might enable zoo-anthropogenic transfer. Therefore, monitoring the presence of pathogenic and drug-resistant E. coli isolates is essential for sustainable disease management in livestock and human medicine [12].
The testing and screening of virulence genes of porcine E. coli by single and/or multiplex PCRs is an economic factor in the frame of routine microbiological diagnostics [13]. There are numerous VAGs, but a limited number of them are usually examined by a combination of single or multiplex PCRs [13]. Accurate and time saving determination of a wide variety of genes can be accomplished using DNA microarray-based assays [14]. In the present study, we developed a microarray-based diagnostic tool combining oligonucleotides designed to detect a customized set of VAGs for use in routine diagnostics.
In Austria, there is a limited body of data describing the genomic epidemiology of E. coli from swine. Therefore, the objective of the present study was to characterize porcine E. coli, isolated during routine diagnostics, by a polyphasic approach including pheno- and genotypic susceptibility testing and whole-genome sequencing of selected isolates. For the rapid identification of virulence genes in E. coli, customized DNA microarray assay were developed within this study.
2. Materials and Methods
2.1. E. coli Isolates
A total of 102 E. coli isolates of suckling and weaning pigs were included in the present study. All isolates were gut-associated and were obtained during routine bacteriological diagnostics at the Institute of Microbiology of the University of Veterinary Medicine Vienna, Austria and from BS-Immun GmbH Vienna, Austria. All isolates originated from clinical samples received from third parties and therefore were not subject to reporting obligations of the Ethics and Animal Welfare Commission of the University of Veterinary Medicine Vienna. Isolates were stored at −80 °C until further examination.
2.2. Antimicrobial Susceptibility Testing
Antimicrobial susceptibility testing was performed by agar disk diffusion according to the CLSI [15]. Escherichia coli ATCC® 25,922 served as the quality control strain. The following antimicrobials were used: ampicillin (10 µg), piperacillin (10 µg), cefotaxime (30 µg), ceftazidime (30 µg), cefepime (30 µg), aztreonam (30 µg), meropenem (10 µg), imipenem (10 µg), gentamicin (10 µg), amikacin (30 µg), tobramycin (10 µg), ciprofloxacin (5 µg), trimethoprim–sulfamethoxazole (1.25/23.75 µg), tetracycline (30 µg), fosfomycin (200 µg), and chloramphenicol (30 µg) (Becton Dickinson, Heidelberg, Germany). Isolates were further examined for extended-spectrum β-lactamase (ESBL) production by combination disk tests using cefotaxime and ceftazidime with and without clavulanic acid (Becton Dickinson, Heidelberg, Germany) [15]. Furthermore, cefoxitin (30 μg) (BD, Heidelberg, Germany) was utilized to detect AmpC β-lactamase-producing (AmpC) phenotypes. Minimal inhibitory concentration of isolates mobile colistin resistance (mcr) determinants were screened by broth microdilution testing method in accordance with the CLSI document VET01-A4 [16]. Colistin susceptibility testing was interpreted according to the CLSI document MR01 [17]. Escherichia coli ATCC® 25,922 served as quality control strain. E. coli isolates displaying the AmpC phenotype were analyzed for mutations in the chromosomal ampC promoter/attenuator region as described previously [18]. The following resistance genes were screened via PCRs: blaCMY, blaCTX, blaOXA-1, blaOXA-2, blaSHV, blaTEM, sul1, sul2, sul3, dfrA1, dfrA12, dfrA14, dfrA17, dfrA19, strA, strB, aadA1, aadA2, aadA4, aadA5, aadB, qepA, qnrA, qnrB, qnrC, qnrD, qnrS, aac(6′)-Ib-cr, catA1, cfr, cmlA1, floR, tet(A), tet(B), tet(C), tet(D), tet(E), tet(G) as described elsewhere [19,20]. In addition, the genes blaCMY, blaCTX-M, blaSHV, and blaTEM were sequenced after PCR amplification. All amplicons in the present study were sequenced at LGC Genomics, Berlin, Germany. Sequences were aligned with BLAST (Basic Local Alignment Search Tool. Available online: https://blast.ncbi.nlm.nih.gov/Blast.cgi, accessed on 29 July 2021) and compared with reference sequences available in GenBank and the National Center for Biotechnology Information (NCBI) database (Beta Lactamase Data Resources. Available online: http://www.ncbi.nlm.nih.gov/pathogens/beta-lactamase-data-resources/, accessed on 29 July 2021). PCR for plasmid-mediated colistin resistance genes, mcr-1, mcr-2, mcr-3, mcr-4, mcr-5, was performed according to the protocol of European Union Reference Laboratory for Antimicrobial Resistance [21]. The quinolone resistance-determining regions (QRDR) of gyrA and parC in ciprofloxacin-resistant isolates were amplified by PCR and sequenced [22].
2.3. Bicocide Susceptibility Testing
Biocide susceptibility testing was performed according to the previously established protocol by Schug et al. [23]. Established minimal inhibitory concentration (MIC) values of investigated biocides on reference strains are shown in Supplementary Materials Table S2. Benzalkonium chloride (Acros Organics, Geel, Belgium, 21541), as a representative of the quaternary ammonium compounds, was tested at concentration ranges 0.000015–0.016%; chlorhexidine (Sigma-Aldrich, Schnelldorf, Germany, 55-56-1), as a representative of cationic compounds, was tested at concentration ranges 0.000015–0.002%; glutardialdehyde (Chempur, Piekary Slaskie, Poland, 424610240), as a representative of aldehydes, was tested at concentration ranges 0.0075–1%; and isopropanol (99.9%, PHPU Eurochem BGD, Tarnow, Poland), as a representative of alcohols, was tested at concentration ranges 1–14%. The method was performed in 96-well polystyrene microtiter plates with U bottom (Sarstedt, Numbrecht, Germany, 82.1582.001). The bacterial inoculum was prepared according to the CLSI standard (Clinical and Laboratory Standards Institute, 2020), using Trypticasein soy broth (BioMaxima, Lublin, Poland, PS 23-500). The final concentration of bacteria inoculated into the wells was 2.5–5 × 105 CFU/mL.
2.4. Clonal Relatedness of E. coli and Whole-Genome Sequencing
E. coli DNA was extracted as previously described [24]. Isolates were phylotyped using the quadruplex assignment method [25]. Clonal relatedness of E. coli isolates was assessed by two-locus sequence typing, or “CH-clonotyping”, using combined data of fumC and fimH sequences as described by Weissman et al. [26]. Allele and CH clonotype numbers were used for goeBURST analysis using PHYLOViZ [27]. Thirty-five selected E. coli isolates were analyzed by whole-genome sequencing (WGS), which was performed by isolating bacterial DNA using the MagAttract HMW DNA Kit (Qiagen, Hilden, Germany). Ready-to-sequence libraries were prepared using Nextera XT DNA Library Preparation Kit (Illumina, San Diego, United States). Sequencing was performed on the Illumina MiSeq platform [28]. De novo assembly of the 300 bp paired-end reads was conducted using SPAdes 3.9.0 [29]. WGS data analysis was performed with SeqSphere+ software (Ridom, Münster, Germany). To assess the genetic relatedness between the E. coli isolates, multi-locus sequence typing (MLST) and core genome multi-locus sequence-based typing (cgMLST) were performed as previously described [30]. To identify acquired resistance genes or chromosomal mutations, Comprehensive Antibiotic Resistance Database [31] as well as ResFinder 4.1 [32,33] were used. Genes associated with biocide resistance were compared with BacMet database (Antibacterial Biocide and Metal Resistance Genes Database. Available online: http://bacmet.biomedicine.gu.se/, accessed on 29 July 2021) [34]. Virulence genes were identified using VirulenceFinder [35,36]. CH types were characterized as mentioned above. Serogenotypes were analyzed by SerotypeFinder [37]. E. coli phylotypes were extracted from WGS by Clermont typing [38]. The presence of plasmids was determined using PlasmidFinder [39]. Probability prediction of the location of a given virulence or antibiotic resistance gene was achieved by applying mlplasmids trained on E. coli [40]. Posterior plasmid probability (ppp) scores ≥0.7 at a minimum contig length of 700 bp indicate that a given contig sequence is plasmid-derived. For selected contigs with lower ppp scores, BLAST analyses against the Enterobacterales nucleotide collection at NCBI were performed. Plasmid probability was assumed for mlplasmid scores > 0.699 or if BLAST analyses identified E. coli plasmids for at least 90% of contig length with >90% identity. The genomes of WGS isolates were deposited under PRJNA728557 in the NCBI BioProject database.
2.5. Microarray-Based Detection of Virulence-Associated Genes
A set of virulence genes was determined for all isolates using a DNA microarray-based technology developed in the present study frame. The technology is based on methods as described previously [41], and custom-made microarrays from INTER-ARRAY (INTER-ARRAY by fzmb GmbH, Bad Langensalza, Germany) were used according to manufacturer’s instructions. The complete list of virulence-associated genes can be found at INTER-ARRAY website (Virulence Genes for Manuscript. Available online: https://www.inter-array.com/porcineEcoli/VirulenceGenesformanuscript_supplementary_material.xlsx, accessed on 29 July 2021). A split network tree was used to visualize similarities between hybridization patterns as described previously [13].
3. Results
3.1. Antimicrobial Susceptibility Testing
All isolates were susceptible to amikacin and carbapenems. Out of the 102 E. coli strains, 79.41% were resistant to at least one of the remaining antimicrobial agents tested. Twenty-one isolates displayed an extended-spectrum β-lactamase (ESBL) phenotype, whereas a single isolate displayed an AmpC phenotype. In total, 36.27% of the isolates exhibited an MDR phenotype [10]. The majority of isolates were resistant to ampicillin (61.75%) and/or tetracycline (58.81%). Further resistance rates were found to piperacillin (26.46%), sulfamethoxazole–trimethoprim (23.53%), cefotaxime (13.71%), chloramphenicol (11.75%), ceftazidime (8.81%), cefepime (7.83%), gentamicin (6.85%), fluoroquinolone (5.87%), aztreonam (4.90%), tobramycin (3.91%), and fosfomycin (1.95%). A total of 2.94% of all investigated isolates exhibited elevated colistin MICs of ≥4 µg/mL. All results of antimicrobial susceptibility testing are summarized in Table 1 and Table 2.

Table 1.
Pheno- and genotypic characterization of E. coli isolated from porcine sources.

Table 2.
Characterization of whole-genome-sequenced porcine E. coli.
3.2. Characterization of Genotypic Antibiotic Resistance
In 13.71% of the isolates, genes from the blaCTX family were detected alone or combined with other bla genes. One of the isolates displayed an AmpC phenotype and carried a blaCMY-2 gene. The most prevalent β-lactamase genes detected were blaTEM-1 (56.00%) followed by blaCTX M-1 (13.71%). Three isolates carried the mobile colistin resistance gene mcr-1.1.
The gyrA and parC sequences of 13.72% ciprofloxacin-resistant isolates were analyzed and revealed mutations that resulted in the following amino acid substitutions: 10.78% of the isolates had a Ser83Leu, one isolate a Ser83Ala, and another 10.78% of isolates an Asp87Asn substitution in gyrA, while in parC 11 isolates displayed a Ser80Ile, 1.96% of the isolates showed Glu84Gly mutation while one isolate revealed a Cys56Thr substitution. A total of 1.96% of all isolates had an Ile355Thr mutation in parE. Results are listed in Table 1.
3.3. Biocide Susceptibility Testing
The obtained MIC values of all tested biocides against ATCC strains, including E. coli ATCC 10,536, were in the acceptable susceptibility. MIC values of benzalkonium chloride (BAC) for all clinical E. coli isolates ranged from 0.0005% to 0.002%. The obtained BAC MIC values were 0.0005% for 1.9% of isolates (2/104), 0.001% for 54.7% (59/104) of isolates, and 0.002% for 41.3% (43/104) of isolates. Chlorhexidine (CHX) MIC values comprised seven dilutions steps from 0.00003% to 0.002%. In comparison to BAC with a unimodal distribution, a bimodal MIC distribution was seen for CHX. This bimodal distribution might point towards a possibly acquired resistance property for the isolates with CHX MICs of 0.00025%. For glutaraldehyde (GLU), unimodal MIC distribution comprising five dilution steps (0.03% to 0.5%) was observed. Except for one isolate with an MIC of <1%, the remaining isolates had isopropanol (ISO) MICs from 2% to 10%. The results of the biocide susceptibility testing of E. coli are shown in Supplementary Materials Tables S1 and S2.
3.4. E. coli Phylotyping
Among all E. coli isolates, the most dominant phylogenetic group was A (50.98%), followed by B1 (25.48%), while the remaining belonged to C (8.81%), D (5.87%), B2 (3.91%), F (1.95%), E, G, and clade 1 (each 0.97%). Results of E. coli phylotyping are shown in Table 1 and Table 2 and Supplementary Materials Table S3.
3.5. E. coli Clonotyping
The fumC and fimH (CH) typing divided all isolates into 51 distinct CH clonotypes and revealed the clonal relatedness of 12 isolates (CH27-0), 9 isolates (CH11-54) and 8 isolates (CH11-23). E. coli-predicted CH clonotype CH40-24 was clearly determined in isolates 24_8 and 99_74. The relatedness of isolates is visualized in Figure 1.
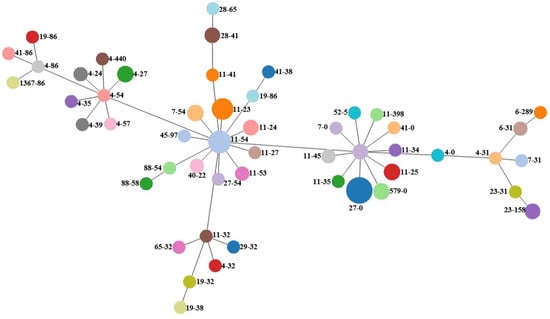
Figure 1.
goeBURST diagram for the CH clonotyping dataset of E. coli isolates. An eBURST diagram was calculated using PHYLOViZ with the goeBURST algorithm. E. coli isolates were grouped according to their CH profiles.
3.6. Whole-Genome Sequencing (WGS) of Selected E. coli Isolates
In our study, 35 isolates were analyzed by whole-genome sequencing (WGS). WGS revealed a total of 16 distinct STs. The most common sequence type was ST10 (n = 6), which clustered together by cgMLST. Further sequence types were ST100 (n = 5), ST354, ST131 (n = 2 each), and singletons ST6404, ST6365, ST1112, ST1079, ST760, ST744, ST641, ST117, ST101, ST56, ST42, and ST23. New sequence types could be obtained in three isolates: ST12008 (37_21), ST12009 (46_30), ST12010 (98_73).
The WGS analysis revealed 12 different serogenotypes (WGS-predicted serotypes). The remaining 23 strains were O-non-typeable with 11 different H types. Three isolates could not be assigned to a known serotype. Isolates belonging to O25:H4 were detected in two cases. Three isolates carrying the gene stx2e could be assigned to serotype O138:H14. Another stx2-carrying isolate belonged to serotype O121:H10.
In total, 6 out of 35 Shiga toxin-producing E. coli were detected in the present study carrying the genes stx2, stx2e, stx2A and stx2B.
Two out of 35 isolates belonged to the successful evolutionary line ST131 and could be assigned to phylogroup B2 (fimH22). Isolate 24_8 was a blaTEM-1C and blaCTX-M-1-producing ESBL E. coli whereas isolate 99_74 produced only blaTEM-1C. Both strains revealed mutations in the QRDR of gyrA and parC and were multidrug-resistant. Virulence potential for both strains was inferred by the detection of multiple VAGs determining the UPEC pathotype. Virulence profile similarity among the two isolates was high and the types of virulence genes presented in these strains were coding for adhesins, toxins, siderophores, hemolysins, and protectins.
PlasmidFinder was used for the analysis of WGS data and revealed the presence of the plasmid replicons IncFIB(AP001918), IncFIC(FII), IncHI2, IncHI2A, IncX1, IncFII, IncN, IncY, IncI1-I(Alpha), IncFIA, IncQ1, p0111, IncFII(pHN7A8), Col(MG828), IncR, IncFIB(H89-PhagePlasmid), IncFII(29), IncFII(pCoo), Col156, IncFII(pSE11), IncI2(Delta), IncI2, IncFII(pRSB107), Col440II, ColpVC, IncX4 and IncB/O/K/Z. IncX4 was identified as the replicon of all mcr-1-carrying plasmids. IncFIB(AP001918) plasmids were predominant (27 of 35) and carried the VAGs ompT, hlyF, cia, and etsC, followed by IncI1-I(Alpha) plasmids carrying cia and blaCTX-M-1 and the IncX1 plasmids carrying blaTEM-1B. IncFII and IncFII(pCoo) carried traT. The full list of VAG and AMR genes and their predicted plasmid probability are shown in Table 1, Supplementary Materials Tables S3–S6.
3.7. E. coli Pathotyping
All isolates carrying VAGs and VAGs related to pathogenic E. coli subtypes were frequently detected. A total of 30 genes were screened by using microarray-based diagnostics. The adhesion gene fimH was present in all but one isolate and therefore was the most frequent gene of the adhesins category. The iron acquisition gene iucD was found in 24 isolates and was always represented together with the fimbrial gene papC. Among toxin-encoding genes, astA was the most predominant (n = 27) gene, followed by itcA (n = 13). The shigatoxin stx2e gene was detected in five isolates and the gene hlyA (n = 9) occurred more often than the cnf1 gene (n = 3). WGS detected the toxin-associated gene sta1 in four isolates. Of all analyzed isolates, the combination of the VAGs fimH, papC and iucD characterizing the UPEC pathotype was the most frequent one (23.52%), followed by the combination of a fimbrial gene/adhesion gene and a toxin gene characterizing the ETEC pathotype (22.54%). Further pathotypes were EDEC (4.90%), atypical ETEC and EPEC (each 3.92%), STEC (0.98%) and UPEC with enterotoxin (2.94%). In total, 40.19% of all E. coli isolates could not be assigned to a specific pathotype.
4. Discussion
This study aimed to characterize E. coli isolates from pig farms in Austria by using pheno- and genotyping methods as well as WGS. Resistance to antimicrobial agents was found in 81 (79%) isolates and 37 isolates met the MDR definition of Sweeney and colleagues [10]. Twenty-one isolates were susceptible to all antimicrobial agents tested. Resistance rates to penicillins (61.73%) and tetracyclines (58.81%) were similar to results of previous studies where penicillins and tetracyclines were the most common antibiotics with AMR in global pig production [42]. The distribution of resistance rates is similar to that in other European studies. Especially, an increased resistance to ampicillin was already reported in the EFSA surveillance program [43]. The variation in resistance in pathogenic E. coli was broad. This emphasizes the importance of performing antimicrobial susceptibility testing after pathotype identification for determining prognosis and guiding clinical management [44].
Colistin is considered by the WHO as a last-resort agent in the treatment of severe bacterial infections caused by multi-drug resistant Gram-negative bacteria [11]. Different genetic mechanisms are known to lead to colistin resistance. In particular, for isolates showing reduced susceptibility to colistin, this may be conferred by chromosomal alterations in pmrAB genes, which encode a two-component signal transduction system regulating the endogenous LPS modification system [45,46,47]. In 2015, the emergence and also the spread of mobile colistin resistance (mcr) genes were detected [48]. Although only three isolates in this study carried a MCR resistance gene, namely mcr-1.1, there is a scarcity of surveillance studies focusing on MCR genes in both human and veterinary medicine in Austria. Indeed, Austrian surveillance programs until now have not mentioned the presence of any colistin-resistant E. coli [49]. Only single reports from human medicine [50] and a study on the Austrian pig population reported the presence of MCR genes [50] previously. Regarding co-resistance, the fact that two of the mcr-1.1-positive isolates showed MDR to penicillins, tetracyclines and trimethoprim–sulfamethoxazole highlights the threat of these clones to therapeutic choices [45]. In animal production, colistin is extensively used for metaphylactic and therapeutic purposes, which may contribute to increasing levels of colistin resistance [45]. For this reason, the European Medicine Agency has raised serious concerns in regard to the use of colistin in animals and the increasing risk for humans that this antimicrobial resistance poses [43].
In addition to colistin, fluoroquinolones are critically important antimicrobials and sometimes they are the sole or one of limited available therapies to treat serious bacterial infections in people (EARS Net Reports. Available online: https://www.ecdc.europa.eu, accessed on 29 July 2021). Resistance to fluoroquinolones among the investigated E. coli isolates was observed in 14/102 isolates (13.7%). Although results must be compared with caution because of the different methodologies performed, the proportion of samples with resistance to fluoroquinolones was lower than in other studies performed on humans, which was revealed to be 18.2% on average [51].
Different E. coli lineages are responsible for animal as well as for human E. coli infections, with previous studies having identified food and food animal reservoirs as sources for zoo-anthropogenic E. coli clones [52]. A study conducted on ESBL-positive E. coli isolates of human and animal origin in the Netherlands, the UK and Germany revealed that human E. coli isolates in the three countries were more closely related to one another than to isolates from animals [53]. In our study, we found isolates of distinct E. coli clonal lineages, including the specific international high-risk clone O25:H4-ST131-H22, which emphasizes its wide distribution and would be the first report of ST131 in pigs of Austrian origin. In addition, recent studies demonstrated the potential of E. coli O25:H4-ST131 to serve as a foodborne UPEC [54] and revealed the close relationship of human and porcine ST131 strains [55]. Indeed, enhanced virulence and antimicrobial resistance were compared with other E. coli ST131 strains from our recent work [56]. Interestingly, a number of virulence genes, encoding colonization, iron uptake, and biofilm formation, which are key enabling factors for the clinical success of ST131 [54,56,57,58], were present in both isolate types (24 VAGs in 24_8, 26 VAGs in 99_74).
Concerning E. coli ST10, an ancestral and ubiquitously occurring lineage comprising both commensal and pathogenic strains, it was detected in six out of 35 sequenced isolates. All but one isolate showed MDR, including a plasmid-predicted carriage of the mcr-1.1 gene (IncX4) in two isolates and blaCTX-M-1 (IncI1-I(Alpha)) in one isolate. Previous studies confirmed ST10 as the dominant ST from swine in Northern Europe with a broad host range and association with hospital- and community-acquired infections [59]. Shepard et al. [60] found that ST10 is one of the main E. coli clonal complexes associated with porcine ETEC, and Garcia et al. identified ST10 as primarily responsible for mcr-4 spread [61]. Nevertheless, more investigations are necessary to verify if E. coli from porcine sources may be derived from the same bacterial lineages or share common evolutionary roots with human isolates.
The reporting of STEC O26 infections has been steadily increasing in the EU due to improved diagnostics of non-O157 sero-pathotypes (EARS Net Reports. Available online: https://www.ecdc.europa.eu, accessed on 29 July 2021). Among characterized E. coli strains, an atypical enteropathogenic E. coli (aEPEC), O26:H11_ST88, was detected. Besides the intimin (eae), which confers the ability to cause attaching and effacing (AE) lesions, the strain harbored heat-stable toxin gene astA and a further 20 VAGs. Previous studies described aEPEC as a possible progenitor of stx-producing O26:H11 STEC that is a major pathogen by causing severe gastrointestinal infections in animals and humans [62] and hemolytic–uremic syndrome (HUS) in humans [63]. Further studies indicated that aEPEC isolates may be able to acquire stx by integrating the stx-prophage into their genome and further function as STEC [64]. In addition, the isolate in our study was MDR and harbored a plasmid-predicted mcr-1.1 gene.
ETEC strains are recognized as the most common cause of porcine neonatal diarrhea (ND) and PWD in pigs [44], and were found in 23 of the investigated isolates. Interestingly, the pathotype UPEC was found to be the most common (24 isolates), although collected samples were mainly associated with ND and PWD. In total, 41 isolates could not be assigned to a specific pathotype because of lacking a specific combination of VAGs, or because of harboring VAGs that are specific for more than one pathotype. This circumstance may confirm expectations of Robins-Browne et al. and Müller et al. [3,65] that some of the typing schemes in current use will eventually be replaced, allowing more pathotypes to be identified (2016).
Phylogenetic analyses found groups A and B1 to be the most common, which corresponded to the results of similar studies [66]. Phylogroup B2 was represented by 4/102 isolates, all of which represented the UPEC pathotype, including both ST131 isolates, as previously confirmed by Nicolas-Chanoine et al. [67].
The plasmid types IncF, IncI and IncX, carrying VAGs and AMR genes, were found. These findings are a cause for concern, as these elements can easily be transferred from animal host pathogens to human pathogens, increasing their AMR and virulence [8]. lncF is the most frequently described plasmid type found in E. coli of human and animal sources. Interestingly, our investigation revealed that the traT gene, which codes for surface exclusion, was IncFI1-associated [68]. In a single isolate, blaCTX-M-1 was predicted to be on an lncl1 plasmid. Such plasmids are predominantly described as blaCTX-M-1 carriers in E. coli of European poultry and are further considered as a possible source for human infections [69]. In our study, three of 102 isolates carried the mcr-1.1 gene on an IncX4 plasmid, which is in agreement with other works on Salmonella and E. coli isolates obtained from human and animal sources where IncX plasmids are also shown to carry mcr genes [69].
Biocides are applied as an integral part of infection control in pig production and slaughterhouses. The selection of bacteria with reduced susceptibility to disinfectants has already been confirmed [70]. In our study, we investigated biocide susceptibility and revealed unimodal MIC distributions for benzalkonium chloride, glutardialdehyde and isopropanol. In comparison, a bimodal MIC distribution was observed for chlorhexidine, which might point towards the acquisition of the respective resistance properties. Previous studies confirmed that biocide-like disinfectants and surfactants are effective to select for AMR [71].
In our study, the newly developed oligonucleotide microarray offered an accurate and rapid solution to detect a large set of E. coli VAGs. Previous studies compared the accuracy and time needed to perform a microarray-based method with conventional multiplex PCR [72], and showed that microarray-based diagnostics was less labor-intensive and, therefore, more cost-effective. In addition, the error rates occurring in the amplification process during multiplex PCR do not exist when using microarrays [73]. Therefore, in our study, microarray technology offered an accurate and rapid tool to detect a large set of VAGs in parallel.
5. Conclusions
In this study, we have found porcine high-risk zoonotic E. coli clones that are both pathogenic and multi-drug resistant. The threat that these clones can pose to public health is derived from their AMR to critically important antibiotics for humans. Therefore, our work highlights the importance of monitoring AMR and VAGs in porcine E. coli isolates. This can be achieved by applying reliable, fast, economical, and easy to perform technologies such as DNA-based microarray typing. Nevertheless, preventive measures in swine farms in addition to surveillance must be applied to avoid infection of the pigs with resistant and pathogenic E. coli strains and to avoid their spread.
6. Limitations of Our Study
Data on prevalence, serotypes, and pathotypes of porcine E. coli in Austria and other countries were scarce, which made comparisons difficult. In our study, we were not able to compare our data on the national level because resistance in swine is not monitored yet in a harmonized way in Austria.
Supplementary Materials
The following are available online at https://www.mdpi.com/article/10.3390/microorganisms9081676/s1, Tables S1 and S2: Biocide susceptibility, Table S3: Plasmid presence_virulence genes_sorted, Table S4: Plasmid probability prediction—AMR genes, Table S5: Plasmid probability prediction_virulence genes, Table S6: Overview plasmid presence AMR genes. Figure S1: Splitstree.
Author Contributions
Conceptualization, K.F., A.L. and I.L.; methodology, T.B.-H., E.M., A.C.-R., D.M., A.T.F., M.P.S., W.R., S.D.B., S.M., R.E., S.S. and I.L.; validation, A.C.-R., A.T.F., M.P.S., W.R., S.D.B., S.M., R.E., S.S. and I.L.; formal analysis, A.C.-R., M.K., D.M. and I.L.; investigation, T.B.-H., A.C.-R., M.K., D.M., K.A., O.G., M.P.S., W.R., R.E. and I.L.; resources, L.S., D.M., A.W., F.A., A.T.F., A.L., J.S. and R.E.; data curation, A.C.-R., M.K., D.M., K.F., A.T.F., M.P.S., W.R., S.D.B. and I.L.; writing—original draft preparation, T.B.-H. and I.L.; writing—review and editing, A.C.-R., D.M., A.T.F., F.A., M.P.S., W.R., S.D.B., S.M., R.E., S.S. and I.L.; visualization, A.T.F.; supervision, I.L.; project administration, L.S., I.L.; funding acquisition, R.E. All authors have read and agreed to the published version of the manuscript.
Funding
The part of the research that was performed by Leibniz Institute of Photonic Technology (IPHT), Jena, Germany, was funded by Project DRESI: Rapid and accurate diagnosis and resistance testing of sepsis pathogens in the intensive care unit (13GW0423C, Start: 03/2020; BMBF).
Institutional Review Board Statement
All clinical samples were obtained within the scope of routine diagnostic health assessments and only subsequently evaluated within the scope of this study and, therefore, not subject to reporting obligations of the Ethics and Animal Welfare Commission of the University of Veterinary Medicine in Vienna. For this reason, ethical review and approval were waived for this study.
Informed Consent Statement
Not applicable.
Data Availability Statement
All data are contained within the article or Supplementary Materials.
Acknowledgments
We thank Spencer Sawyer for proofreading and Rita Bernreiter, Anna Korath, Barbara Tischler and Martina Zimmermann for their technical assistance. Open Access Funding by the University of Veterinary Medicine Vienna.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Janda, J.M.; Abbott, S.L. The Changing Face of the Family Enterobacteriaceae (Order: “Enterobacterales”): New Members, Taxonomic Issues, Geographic Expansion, and New Diseases and Disease Syndromes. Clin. Microbiol. Rev. 2021, 34, e00174-20. [Google Scholar] [CrossRef]
- Kaper, J.B.; Nataro, J.P.; Mobley, H.L. Pathogenic Escherichia coli. Nat. Rev. Genet. 2004, 2, 123–140. [Google Scholar] [CrossRef] [PubMed]
- Robins-Browne, R.M.; Holt, K.E.; Ingle, D.J.; Hocking, D.M.; Yang, J.; Tauschek, M. Are Escherichia coli Pathotypes Still Relevant in the Era of Whole-Genome Sequencing? Front. Cell. Infect. Microbiol. 2016, 6, 141. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Poolman, J.T.; Wacker, M. Extraintestinal Pathogenic Escherichia coli, a Common Human Pathogen: Challenges for Vaccine Development and Progress in the Field. J. Infect. Dis. 2016, 213, 6–13. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Basmaci, R.; Bonacorsi, S.; Bidet, P.; Biran, V.; Aujard, Y.; Bingen, E.; Béchet, S.; Cohen, R.; Levy, C. Escherichia ColiMeningitis Features in 325 Children From 2001 to 2013 in France. Clin. Infect. Dis. 2015, 61, 779–786. [Google Scholar] [CrossRef] [Green Version]
- Sarowska, J.; Futoma-Koloch, B.; Jama-Kmiecik, A.; Frej-Madrzak, M.; Ksiazczyk, M.; Bugla-Ploskonska, G.; Choroszy-Krol, I. Virulence factors, prevalence and potential transmission of extraintestinal pathogenic Escherichia coli isolated from different sources: Recent reports. Gut Pathog. 2019, 11, 10. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pitout, J.D.D. Extraintestinal Pathogenic Escherichia coli: A Combination of Virulence with Antibiotic Resistance. Front. Microbiol. 2012, 3, 9. [Google Scholar] [CrossRef] [Green Version]
- Partridge, S.R.; Kwong, S.M.; Firth, N.; Jensen, S.O. Mobile Genetic Elements Associated with Antimicrobial Resistance. Clin. Microbiol. Rev. 2018, 31, e00088-17. [Google Scholar] [CrossRef] [Green Version]
- Knöppel, A.; Näsvall, J.; Andersson, D.I. Evolution of Antibiotic Resistance without Antibiotic Exposure. Antimicrob. Agents Chemother. 2017, 61, 61. [Google Scholar] [CrossRef] [Green Version]
- Sweeney, M.T.; Lubbers, B.V.; Schwarz, S.; Watts, J.L. Applying definitions for multidrug resistance, extensive drug resistance and pandrug resistance to clinically significant livestock and companion animal bacterial pathogens. J. Antimicrob. Chemother. 2018, 73, 1460–1463. [Google Scholar] [CrossRef]
- World Health Organization. WHO Guidelines on Use of Medically Important Antimicrobials in Food-Producing Animals; World Health Organization: Geneva, Switzerland, 2017; ISBN 9241550139.
- Boerlin, P.; Travis, R.; Gyles, C.L.; Reid-Smith, R.; Lim, N.J.H.; Nicholson, V.; McEwen, S.A.; Friendship, R.; Archambault, M. Antimicrobial Resistance and Virulence Genes of Escherichia coli Isolates from Swine in Ontario. Appl. Environ. Microbiol. 2005, 71, 6753–6761. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gwida, M.; Awad, A.; El-Ashker, M.; Hotzel, H.; Monecke, S.; Ehricht, R.; Müller, E.; Reißig, A.; Barth, S.; Berens, C.; et al. Microarray-based detection of resistance and virulence factors in commensal Escherichia coli from livestock and farmers in Egypt. Veter Microbiol. 2020, 240, 108539. [Google Scholar] [CrossRef]
- Anjum, M.F.; Mafura, M.; Slickers, P.; Ballmer, K.; Kuhnert, P.; Woodward, M.J.; Ehricht, R. Pathotyping Escherichia coli by Using Miniaturized DNA Microarrays. Appl. Environ. Microbiol. 2007, 73, 5692–5697. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Weinstein, M.P. Performance Standards for Antimicrobial Susceptibility Testing: Supplement M100, 30th ed.; Clinical and Laboratory Standards Institute: Wayne, PA, USA, 2020; ISBN 9781684400669. [Google Scholar]
- Watts, J.L. Performance Standards for Antimicrobial Disk and Dilution Susceptibility Tests for Bacteria Isolated from Animals: Approved Standard; Documents/Clinical and Laboratory Standards Institute VET01-A4: Wayne, PA, USA, 2013. [Google Scholar]
- Humphries, R.M. Polymyxin Breakpoints for Enterobacterales, Pseudomonas aeruginosa, and Acinetobacter spp. 2020. Available online: https://clsi.org/standards/products/microbiology/companion/mr01/ (accessed on 21 July 2021).
- Caroff, N.; Espaze, E.; Bérard, I.; Richet, H.; Reynaud, A. Mutations in the ampC promoter of Escherichia coli isolates resistant to oxyiminocephalosporins without extended spectrum β-lactamase production. FEMS Microbiol. Lett. 1999, 173, 459–465. [Google Scholar] [CrossRef]
- Kehrenberg, C.; Schwarz, S. Distribution of Florfenicol Resistance Genes fexA and cfr among Chloramphenicol-Resistant Staphylococcus Isolates. Antimicrob. Agents Chemother. 2006, 50, 1156–1163. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dolejska, M.; Frolkova, P.; Florek, M.; Jamborova, I.; Purgertova, M.; Kutilova, I.; Cizek, A.; Guenther, S.; Literak, I. CTX-M-15-producing Escherichia coli clone B2-O25b-ST131 and Klebsiella spp. isolates in municipal wastewater treatment plant effluents. J. Antimicrob. Chemother. 2011, 66, 2784–2790. [Google Scholar] [CrossRef]
- Rebelo, A.R.; Bortolaia, V.; Kjeldgaard, J.S.; Pedersen, S.K.; Leekitcharoenphon, P.; Hansen, I.M.; Guerra, B.; Malorny, B.; Borowiak, M.; Hammerl, J.A.; et al. Multiplex PCR for detection of plasmid-mediated colistin resistance determinants, mcr-1, mcr-2, mcr-3, mcr-4 and mcr-5 for surveillance purposes. Eurosurveillance 2018, 23, 17-00672. [Google Scholar] [CrossRef] [PubMed]
- Everett, M.J.; Jin, Y.F.; Ricci, V.; Piddock, L.J. Contributions of individual mechanisms to fluoroquinolone resistance in 36 Escherichia coli strains isolated from humans and animals. Antimicrob. Agents Chemother. 1996, 40, 2380–2386. [Google Scholar] [CrossRef] [Green Version]
- Schug, A.R.; Bartel, A.; Scholtzek, A.D.; Meurer, M.; Brombach, J.; Hensel, V.; Fanning, S.; Schwarz, S.; Feßler, A.T. Biocide susceptibility testing of bacteria: Development of a broth microdilution method. Veter Microbiol. 2020, 248, 108791. [Google Scholar] [CrossRef]
- Loncaric, I.; Misic, D.; Szostak, M.P.; Künzel, F.; Schäfer-Somi, S.; Spergser, J. Broad-Spectrum Cephalosporin-Resistant and/or Fluoroquinolone-Resistant Enterobacterales Associated with Canine and Feline Urogenital Infections. Antibiotics 2020, 9, 387. [Google Scholar] [CrossRef]
- Clermont, O.; Christenson, J.K.; Denamur, E.; Gordon, D.M. The Clermont Escherichia coli phylo-typing method revisited: Improvement of specificity and detection of new phylo-groups. Environ. Microbiol. Rep. 2013, 5, 58–65. [Google Scholar] [CrossRef]
- Weissman, S.J.; Johnson, J.R.; Tchesnokova, V.; Billig, M.; Dykhuizen, D.; Riddell, K.; Rogers, P.; Qin, X.; Butler-Wu, S.; Cookson, B.T.; et al. High-Resolution Two-Locus Clonal Typing of Extraintestinal Pathogenic Escherichia coli. Appl. Environ. Microbiol. 2012, 78, 1353–1360. [Google Scholar] [CrossRef] [Green Version]
- Francisco, A.P.; Bugalho, M.; Ramirez, M.; Carriço, J.A. Global optimal eBURST analysis of multilocus typing data using a graphic matroid approach. BMC Bioinform. 2009, 10, 152. [Google Scholar] [CrossRef] [Green Version]
- Lepuschitz, S.; Huhulescu, S.; Hyden, P.; Springer, B.; Rattei, T.; Allerberger, F.; Mach, R.; Ruppitsch, W. Characterization of a community-acquired-MRSA USA300 isolate from a river sample in Austria and whole genome sequence based comparison to a diverse collection of USA300 isolates. Sci. Rep. 2018, 8, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Bankevich, A.; Nurk, S.; Antipov, D.; Gurevich, A.A.; Dvorkin, M.; Kulikov, A.S.; Lesin, V.M.; Nikolenko, S.I.; Pham, S.; Prjibelski, A.D.; et al. SPAdes: A New Genome Assembly Algorithm and Its Applications to Single-Cell Sequencing. J. Comput. Biol. 2012, 19, 455–477. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wirth, T.; Falush, D.; Lan, R.; Colles, F.; Mensa, P.; Wieler, L.H.; Karch, H.; Reeves, P.; Maiden, M.; Ochman, H.; et al. Sex and virulence in Escherichia coli: An evolutionary perspective. Mol. Microbiol. 2006, 60, 1136–1151. [Google Scholar] [CrossRef] [Green Version]
- Alcock, B.P.; Raphenya, A.R.; Lau, T.T.Y.; Tsang, K.K.; Bouchard, M.; Edalatmand, A.; Huynh, W.; Nguyen, A.-L.V.; Cheng, A.A.; Liu, S.; et al. CARD 2020: Antibiotic resistome surveillance with the comprehensive antibiotic resistance database. Nucleic Acids Res. 2020, 48, D517–D525. [Google Scholar] [CrossRef] [PubMed]
- Camacho, C.; Coulouris, G.; Avagyan, V.; Ma, N.; Papadopoulos, J.S.; Bealer, K.; Madden, T.L. BLAST+: Architecture and applications. BMC Bioinform. 2009, 10, 421. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Clausen, P.T.L.C.; Aarestrup, F.M.; Lund, O. Rapid and precise alignment of raw reads against redundant databases with KMA. BMC Bioinform. 2018, 19, 1–8. [Google Scholar] [CrossRef]
- Pal, C.; Bengtsson-Palme, J.; Rensing, C.; Kristiansson, E.; Larsson, D.G.J. BacMet: Antibacterial biocide and metal resistance genes database. Nucleic Acids Res. 2014, 42, D737–D743. [Google Scholar] [CrossRef] [Green Version]
- Joensen, K.G.; Scheutz, F.; Lund, O.; Hasman, H.; Kaas, R.S.; Nielsen, E.M.; Aarestrup, F. Real-Time Whole-Genome Sequencing for Routine Typing, Surveillance, and Outbreak Detection of Verotoxigenic Escherichia coli. J. Clin. Microbiol. 2014, 52, 1501–1510. [Google Scholar] [CrossRef] [Green Version]
- Tetzschner, A.M.M.; Johnson, J.R.; Johnston, B.D.; Lund, O.; Scheutz, F. In Silico Genotyping of Escherichia coli Isolates for Extraintestinal Virulence Genes by Use of Whole-Genome Sequencing Data. J. Clin. Microbiol. 2020, 58, 58. [Google Scholar] [CrossRef]
- Joensen, K.; Tetzschner, A.M.M.; Iguchi, A.; Aarestrup, F.M.; Scheutz, F. Rapid and Easy In Silico Serotyping of Escherichia coli Isolates by Use of Whole-Genome Sequencing Data. J. Clin. Microbiol. 2015, 53, 2410–2426. [Google Scholar] [CrossRef] [Green Version]
- Beghain, J.; Bridier-Nahmias, A.; Le Nagard, H.; Denamur, E.; Clermont, O. ClermonTyping: An easy-to-use and accurate in silico method for Escherichia genus strain phylotyping. Microb. Genom. 2018, 4, e000192. [Google Scholar] [CrossRef] [PubMed]
- Carattoli, A.; Zankari, E.; García-Fernández, A.; Larsen, M.V.; Lund, O.; Villa, L.; Aarestrup, F.; Hasman, H. In SilicoDetection and Typing of Plasmids using Plasmid Finder and Plasmid Multilocus Sequence Typing. Antimicrob. Agents Chemother. 2014, 58, 3895–3903. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Arredondo-Alonso, S.; Rogers, M.R.C.; Braat, J.C.; Verschuuren, T.D.; Top, J.; Corander, J.; Willems, R.J.L.; Schürch, A.C. mlplasmids: A user-friendly tool to predict plasmid- and chromosome-derived sequences for single species. Microb. Genom. 2018, 4, e000224. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Monecke, S.; Berger-Bächi, B.; Coombs, G.; Holmes, A.; Kay, I.; Kearns, A.; Linde, H.-J.; O’Brien, F.; Slickers, P.; Ehricht, R. Comparative genomics and DNA array-based genotyping of pandemic Staphylococcus aureus strains encoding Panton-Valentine leukocidin. Clin. Microbiol. Infect. 2007, 13, 236–249. [Google Scholar] [CrossRef] [Green Version]
- Lekagul, A.; Tangcharoensathien, V.; Yeung, S. Patterns of antibiotic use in global pig production: A systematic review. Veter Anim. Sci. 2019, 7, 100058. [Google Scholar] [CrossRef]
- European Centre for Disease Prevention and Control (ECDC). The European Union Summary Report on Antimicrobial Resistance in zoonotic and indicator bacteria from humans, animals and food in 2017/2018. EFSA J. 2020, 18, e06007. [Google Scholar] [CrossRef] [Green Version]
- Luppi, A. Swine enteric colibacillosis: Diagnosis, therapy and antimicrobial resistance. Porc. Health Manag. 2017, 3, 1–18. [Google Scholar] [CrossRef]
- Rhouma, M.; Beaudry, F.; Thériault, W.; Bergeron, N.; Beauchamp, G.; Laurent-Lewandowski, S.; Fairbrother, J.M.; Letellier, A. In vivo therapeutic efficacy and pharmacokinetics of colistin sulfate in an experimental model of enterotoxigenic Escherichia coli infection in weaned pigs. Vet. Res. 2016, 47, 58. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cannatelli, A.; Giani, T.; Aiezza, N.; Di Pilato, V.; Principe, L.; Luzzaro, F.; Galeotti, C.L.; Rossolini, G.M. An allelic variant of the PmrB sensor kinase responsible for colistin resistance in an Escherichia coli strain of clinical origin. Sci. Rep. 2017, 7, 1–6. [Google Scholar] [CrossRef]
- Elbediwi, M.; Li, X.; Paudyal, N.; Pan, H.; Xie, S.; Rajkovic, A.; Feng, Y.; Fang, W.; Rankin, S.C.; Yue, M. Global Burden of Colistin-Resistant Bacteria: Mobilized Colistin Resistance Genes Study (1980–2018). Microorganisms 2019, 7, 461. [Google Scholar] [CrossRef] [Green Version]
- Liu, Y.-Y.; Wang, Y.; Walsh, T.; Yi, L.-X.; Zhang, R.; Spencer, J.; Doi, Y.; Tian, G.; Dong, B.; Huang, X.; et al. Emergence of plasmid-mediated colistin resistance mechanism MCR-1 in animals and human beings in China: A microbiological and molecular biological study. Lancet Infect. Dis. 2016, 16, 161–168. [Google Scholar] [CrossRef]
- European Centre for Disease Prevention and Control. Antimicrobial resistance in the EU/EEA (EARS-Net)—AER for 2019: Annual Epidemiological Report for 2019. 2020. Available online: https://www.ecdc.europa.eu/en/publications-data/surveillance-antimicrobial-resistance-europe-2019 (accessed on 20 July 2021).
- Hartl, R.; Kerschner, H.; Lepuschitz, S.; Ruppitsch, W.; Allerberger, F.; Apfalter, P. Detection of the mcr-1 Gene in a Multidrug-Resistant Escherichia coli Isolate from an Austrian Patient. Antimicrob. Agents Chemother. 2017, 61, e02623-16. [Google Scholar] [CrossRef] [Green Version]
- Federal Ministry of Social Affairs, Health, Care and Consumer Protection. Resistenzbericht Österreich AURES. 2017. Available online: https://www.sozialministerium.at/dam/jcr:9f7a835d-7975-4cd5-9732-a706bf2a8203/AURES_2017.pdf (accessed on 20 July 2021).
- Manges, A.R.; Johnson, J.R. Food-Borne Origins of Escherichia coli Causing Extraintestinal Infections. Clin. Infect. Dis. 2012, 55, 712–719. [Google Scholar] [CrossRef] [Green Version]
- Wu, G.; Day, M.J.; Mafura, M.T.; Nunez-Garcia, J.; Fenner, J.J.; Sharma, M.; Van Essen-Zandbergen, A.; Rodríguez, I.; Dierikx, C.; Kadlec, K.; et al. Comparative Analysis of ESBL-Positive Escherichia coli Isolates from Animals and Humans from the UK, The Netherlands and Germany. PLoS ONE 2013, 8, e75392. [Google Scholar] [CrossRef] [Green Version]
- Liu, C.M.; Stegger, M.; Aziz, M.; Johnson, T.J.; Waits, K.; Nordstrom, L.; Gauld, L.; Weaver, B.; Rolland, D.; Statham, S.; et al. Escherichia coli ST131- H 22 as a Foodborne Uropathogen. mBio 2018, 9, e00470-18. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Reid, C.J.; Blau, K.; Jechalke, S.; Smalla, K.; Djordjevic, S.P. Whole Genome Sequencing of Escherichia coli From Store-Bought Produce. Front. Microbiol. 2020, 10, 3050. [Google Scholar] [CrossRef] [PubMed]
- Mathers, A.J.; Peirano, G.; Pitout, J.D.D. Escherichia coli ST131: The Quintessential Example of an International Multiresistant High-Risk Clone. Adv. Clin. Chem. 2015, 90, 109–154. [Google Scholar] [CrossRef]
- Nicolas-Chanoine, M.-H.; Blanco, J.; Leflon-Guibout, V.; Demarty, R.; Alonso, M.P.; Caniça, M.; Park, Y.-J.; Lavigne, J.-P.; Pitout, J.; Johnson, J.R. Intercontinental emergence of Escherichia coli clone O25:H4-ST131 producing CTX-M-15. J. Antimicrob. Chemother. 2008, 61, 273–281. [Google Scholar] [CrossRef] [Green Version]
- Pitout, J.D.J.; De Vinney, R. Escherichia coli ST131: A multidrug-resistant clone primed for global domination. F1000Research 2017, 6, 195. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Manges, A. Escherichia coli and urinary tract infections: The role of poultry-meat. Clin. Microbiol. Infect. 2016, 22, 122–129. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shepard, S.M.; Danzeisen, J.L.; Isaacson, R.E.; Seemann, T.; Achtman, M.; Johnson, T.J. Genome Sequences and Phylogenetic Analysis of K88- and F18-Positive Porcine Enterotoxigenic Escherichia coli. J. Bacteriol. 2012, 194, 395–405. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- García, V.; Meniño, I.G.; Mora, A.; Simon, S.C.F.; Jiménez, D.D.; Blanco, J.E.; Alonso, M.P.; Blanco, J. Co-occurrence of mcr-1, mcr-4 and mcr-5 genes in multidrug-resistant ST10 Enterotoxigenic and Shiga toxin-producing Escherichia coli in Spain (2006-2017). Int. J. Antimicrob. Agents 2018, 52, 104–108. [Google Scholar] [CrossRef] [PubMed]
- Watson, V.E.; Jacob, M.E.; Flowers, J.R.; Strong, S.J.; DebRoy, C.; Gookin, J.L. Association of Atypical Enteropathogenic Escherichia coli with Diarrhea and Related Mortality in Kittens. J. Clin. Microbiol. 2017, 55, 2719–2735. [Google Scholar] [CrossRef] [Green Version]
- Afset, J.E.; Bruant, G.; Brousseau, R.; Harel, J.; Anderssen, E.; Bevanger, L.; Bergh, K. Identification of Virulence Genes Linked with Diarrhea Due to Atypical Enteropathogenic Escherichia coli by DNA Microarray Analysis and PCR. J. Clin. Microbiol. 2006, 44, 3703–3711. [Google Scholar] [CrossRef] [Green Version]
- Brandal, L.T.; Sekse, C.; Lindstedt, B.-A.; Sunde, M.; Løbersli, I.; Urdahl, A.M.; Kapperud, G. Norwegian Sheep Are an Important Reservoir for Human-Pathogenic Escherichia coli O26:H11. Appl. Environ. Microbiol. 2012, 78, 4083–4091. [Google Scholar] [CrossRef] [Green Version]
- Müller, D.; Greune, L.; Heusipp, G.; Karch, H.; Fruth, A.; Tschäpe, H.; Schmidt, M.A. Identification of Unconventional Intestinal Pathogenic Escherichia coli Isolates Expressing Intermediate Virulence Factor Profiles by Using a Novel Single-Step Multiplex PCR. Appl. Environ. Microbiol. 2007, 73, 3380–3390. [Google Scholar] [CrossRef] [Green Version]
- Bok, E.; Kożańska, A.; Mazurek-Popczyk, J.; Wojciech, M.; Baldy-Chudzik, K. Extended Phylogeny and Extraintestinal Virulence Potential of Commensal Escherichia coli from Piglets and Sows. Int. J. Environ. Res. Public Health 2020, 17, 366. [Google Scholar] [CrossRef] [Green Version]
- Nicolas-Chanoine, M.-H.; Bertrand, X.; Madec, J.-Y. Escherichia coli ST131, an Intriguing Clonal Group. Clin. Microbiol. Rev. 2014, 27, 543–574. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Virolle, C.; Goldlust, K.; Djermoun, S.; Bigot, S.; Lesterlin, C. Plasmid Transfer by Conjugation in Gram-Negative Bacteria: From the Cellular to the Community Level. Genes 2020, 11, 1239. [Google Scholar] [CrossRef] [PubMed]
- Rozwandowicz, M.; Brouwer, M.S.M.; Fischer, J.; Wagenaar, J.A.; Gonzalez-Zorn, B.; Guerra, B.; Mevius, D.J.; Hordijk, J. Plasmids carrying antimicrobial resistance genes in Enterobacteriaceae. J. Antimicrob. Chemother. 2018, 73, 1121–1137. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Maertens, H.; De Reu, K.; Meyer, E.; Van Coillie, E.; Dewulf, J. Limited association between disinfectant use and either antibiotic or disinfectant susceptibility of Escherichia coli in both poultry and pig husbandry. BMC Veter Res. 2019, 15, 310. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Singer, A.C.; Shaw, H.; Rhodes, V.; Hart, A. Review of Antimicrobial Resistance in the Environment and Its Relevance to Environmental Regulators. Front. Microbiol. 2016, 7, 1728. [Google Scholar] [CrossRef] [Green Version]
- Bekal, S.; Brousseau, R.; Masson, L.; Prefontaine, G.; Fairbrother, J.; Harel, J. Rapid identification of Escherichia coli pathotypes by virulence gene detection with DNA microarrays. J. Clin. Microbiol. 2003, 41, 2113–2125. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Potapov, V.; Ong, J.L. Examining Sources of Error in PCR by Single-Molecule Sequencing. PLoS ONE 2017, 12, e0169774. [Google Scholar] [CrossRef] [PubMed]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).