Differential Exoproteome and Biochemical Characterisation of Neoparamoeba perurans
Abstract
1. Introduction
2. Materials and Methods
2.1. The Neoparamoeba perurans Culture
2.2. Extraction of Extracellular Proteome
2.3. In-Solution Trypsin Digestion
2.4. Liquid Chromatography Tandem Mass Spectrometry
2.5. Raw Data Processing and Label Free Quantification (LFQ)
2.6. Enzymatic Characterization
2.7. Enzyme Inhibition Assay
2.8. RTgill W1 Culture and Assay Preparation
2.9. Cytotoxicity Alamar Blue Preparation
3. Results
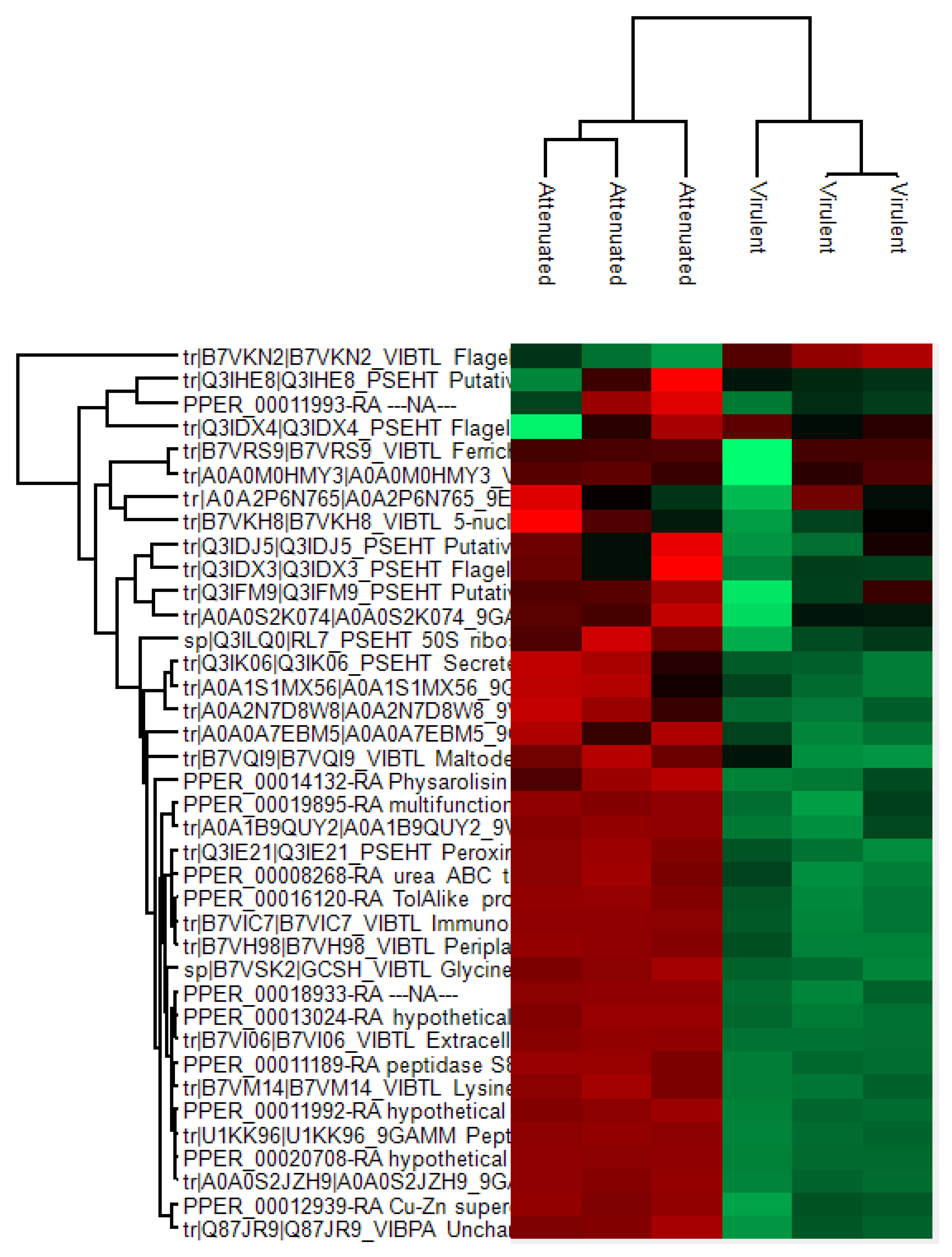
3.1. Exoproteome Identification Using Differential Proteomics
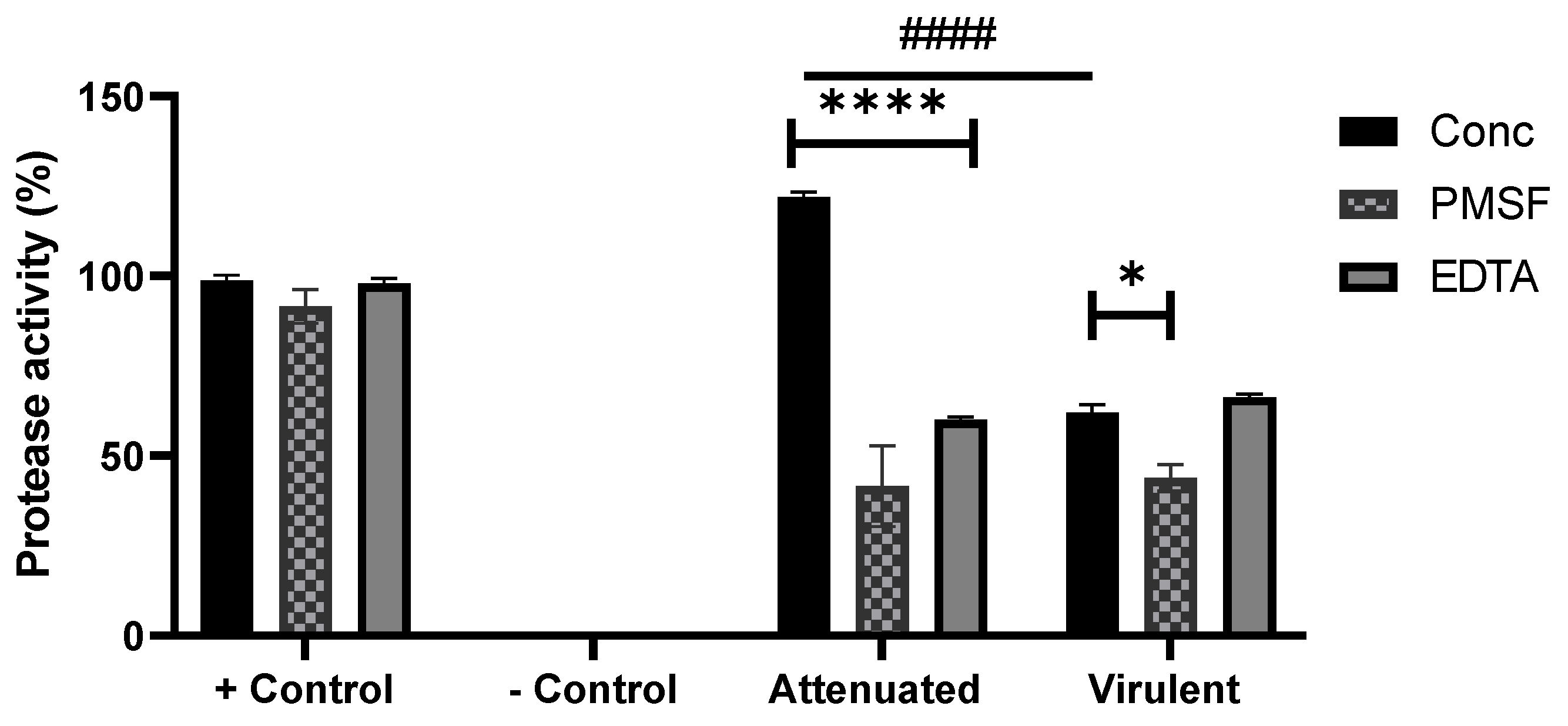
3.2. Enzymatic Characterisation and Inhibition Assay
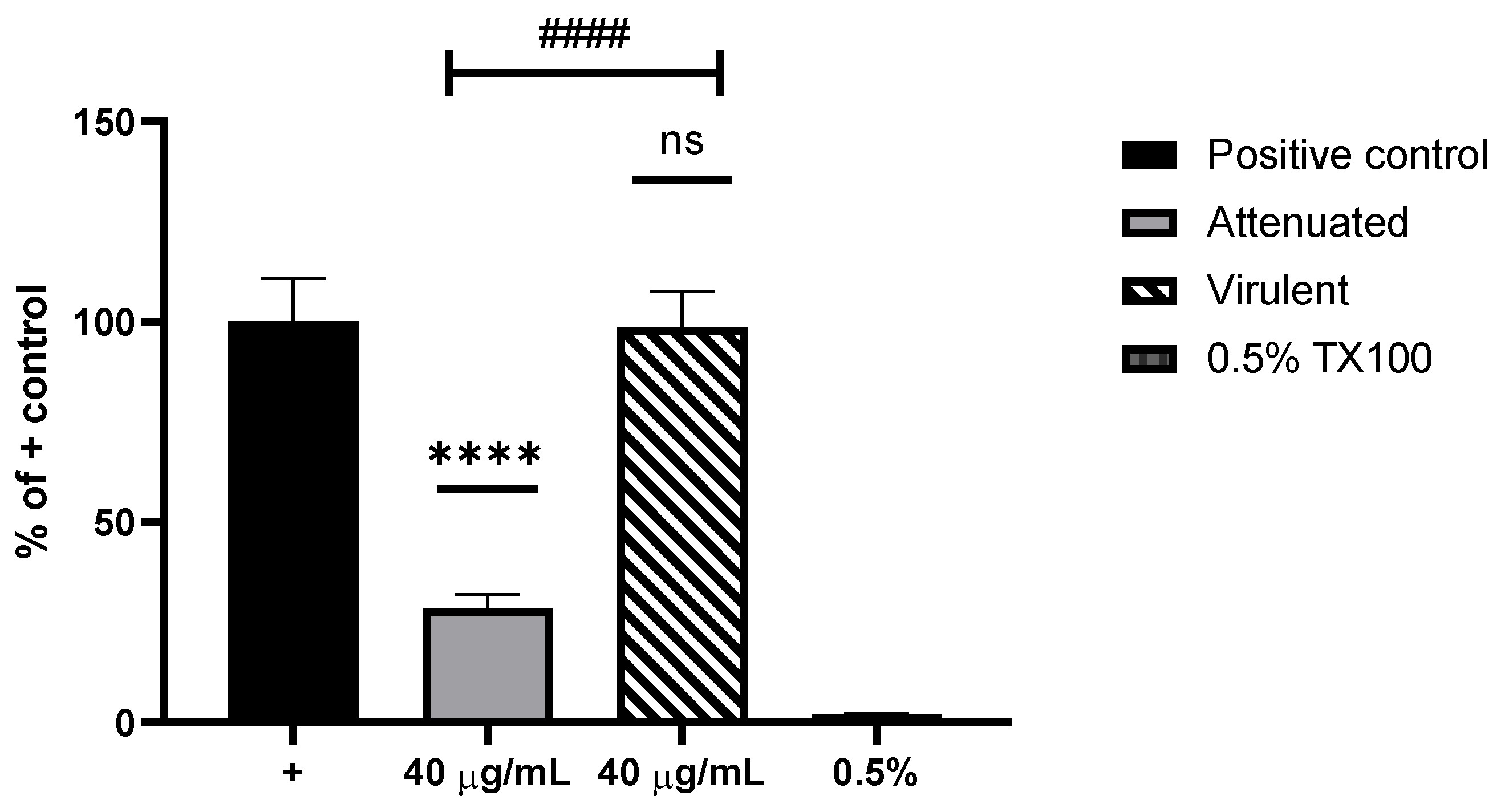
3.3. Cytotoxicity Assay
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Feehan, C.J.; Johnson-Mackinnon, J.; Scheibling, R.E.; Lauzon-Guay, J.-S.; Simpson, A.G. Validating the identity of Paramoeba invadens, the causative agent of recurrent mass mortality of sea urchins in Nova Scotia, Canada. Dis. Aquat. Org. 2013, 103, 209–227. [Google Scholar] [CrossRef] [PubMed]
- Young, N.D.; Crosbie, P.B.; Adams, M.B.; Nowak, B.F.; Morrison, R.N. Neoparamoeba perurans n. sp., an agent of amoebic gill disease of Atlantic salmon (Salmo salar). Int. J. Parasitol. 2007, 37, 1469–1481. [Google Scholar] [CrossRef] [PubMed]
- Munday, B.L. Disease of Salmonids. In Workshop on Diseases of Austrailian Fish & Shellfish; Humphrey, J.D., Langdon, J.S., Eds.; Department of Agriculture and Rural Affairs: Benalla, VIC, Australia, 1985; pp. 127–141. [Google Scholar]
- Steinum, T.; Kvellestad, A.; Rønneberg, L.; Nilsen, H.; Asheim, A.; Fjell, K.; Nygård, S.; Olsen, A.; Dale, O. First cases of amoebic gill disease (AGD) in Norwegian seawater farmed Atlantic salmon, Salmo salar L., and phylogeny of the causative amoeba using 18S cDNA sequences. J. Fish Dis. 2008, 31, 205–214. [Google Scholar] [CrossRef] [PubMed]
- Bustos, P.A.; Young, N.D.; Rozas, M.A.; Bohle, H.M.; Ildefonso, R.S.; Morrison, R.N.; Nowak, B.F. Amoebic gill disease (AGD) in Atlantic salmon (Salmo salar) farmed in Chile. Aquaculture 2011, 310, 281–288. [Google Scholar] [CrossRef]
- Rodger, H. Amoebic gill disease (AGD) in farmed salmon (Salmo salar) in Europe. Fish Vet. J. 2014, 14, 16–26. [Google Scholar]
- Rodger, H.D.; McArdle, J.F. An outbreak of amoebic gill disease in Ireland. Vet. Rec. 1996, 139, 348–349. [Google Scholar] [CrossRef] [PubMed]
- Oldham, T.; Rodger, H.; Nowak, B.F. Incidence and distribution of amoebic gill disease (AGD)—An epidemiological review. Aquaculture 2016, 457, 35–42. [Google Scholar] [CrossRef]
- Peyghan, R.; Powell, M. Histopathological study of gills in experimentally amoebic gill disease (AGD) infected Atlantic salmon, Salmo salar, L. Iran. J. Vet. Res. 2006, 7, 8–13. [Google Scholar]
- Adams, M.; Nowak, B. Sequential pathology after initial freshwater bath treatment for amoebic gill disease in cultured Atlantic salmon, Salmo salar L. J. Fish Dis. 2004, 27, 163–173. [Google Scholar] [CrossRef] [PubMed]
- Adams, M.; Nowak, B. Distribution and structure of lesions in the gills of Atlantic salmon, Salmo salar L., affected with amoebic gill disease. J. Fish Dis. 2001, 24, 535–542. [Google Scholar] [CrossRef]
- Ranganathan, S.; Garg, G. Secretome: Clues into pathogen infection and clinical applications. Genome Med. 2009, 1, 113. [Google Scholar] [CrossRef] [PubMed]
- Hiery, E.; Poetsch, A.; Moosbauer, T.; Amin, B.; Hofmann, J.; Burkovski, A. A Proteomic Study of Clavibacter Michiganensis Subsp. Michiganensis Culture Supernatants. Proteomes 2015, 3, 411–423. [Google Scholar] [CrossRef] [PubMed]
- Serrano-Luna, J.; Piña-Vázquez, C.; Reyes-López, M.; Ortiz-Estrada, G.; de la Garza, M. Proteases from Entamoeba spp. and pathogenic free-living amoebae as virulence factors. J. Trop. Med. 2013, 2013. [Google Scholar] [CrossRef]
- Gonçalves, D.d.S.F.; Ferreira, M.D.S.; Liedke, S.C.; Gomes, K.X.; de Oliveira, G.A.; Leão, P.E.L.; Cesar, G.V.; Seabra, S.H.; Cortines, J.R.; Casadevall, A.; et al. Extracellular vesicles and vesicle-free secretome of the protozoa Acanthamoeba castellanii under homeostasis and nutritional stress and their damaging potential to host cells. Virulence 2018, 9, 818–836. [Google Scholar] [CrossRef] [PubMed]
- Ralston, K.S.; Petri, W.A., Jr. Tissue destruction and invasion by Entamoeba histolytica. Trends Parasitol. 2011, 27, 254–263. [Google Scholar] [CrossRef]
- Siddiqui, R.; Khan, N.A. Primary amoebic meningoencephalitis caused by Naegleria fowleri: An old enemy presenting new challenges. PLoS Negl. Trop. Dis. 2014, 8, e3017. [Google Scholar] [CrossRef]
- Butler, R.; Nowak, B. In vitro interactions between Neoparamoeba sp. and Atlantic salmon epithelial cells. J. Fish Dis. 2004, 27, 343–349. [Google Scholar] [CrossRef]
- Bridle, A.R.; Davenport, D.L.; Crosbie, P.B.; Polinski, M.; Nowak, B.F. Neoparamoeba perurans loses virulence during clonal culture. Int. J. Parasitol. 2015, 45, 575–578. [Google Scholar] [CrossRef]
- Dhufaigh, K.N.; Dillon, E.; Botwright, N.; Talbot, A.; O’Connor, I.; MacCarthy, E.; Slattery, O. Comparative proteomic profiling of newly acquired, virulent and attenuated Neoparamoeba perurans proteins associated with amoebic gill disease. Sci. Rep. Rev. 2021, 11, 1–12. [Google Scholar] [CrossRef]
- Dayeh, V.R.; Schirmer, K.; Lee, L.E.J.; Bols, N.C. Rainbow Trout Gill Cell Line Microplate Cytotoxicity Test. In Small-Scale Freshwater Toxicity Investigations: Toxicity Test Methods; Blaise, C., Férard, J.-F., Eds.; Springer: Dordrecht, The Netherlands, 2005; pp. 473–503. [Google Scholar] [CrossRef]
- Bayer-Santos, E.; Aguilar-Bonavides, C.; Rodrigues, S.P.; Cordero, E.M.; Marques, A.F.; Varela-Ramirez, A.; Choi, H.; Yoshida, N.; da Silveira, J.F.; Almeida, I.C. Proteomic analysis of Trypanosoma cruzi secretome: Characterization of two populations of extracellular vesicles and soluble proteins. J. Proteome Res. 2013, 12, 883–897. [Google Scholar] [CrossRef]
- Rubin, E.; Pales Espinosa, E.; Koller, A.; Allam, B. Characterisation of the secretome of the clam parasite, QPX. Int. J. Parasitol. 2015, 45, 187–196. [Google Scholar] [CrossRef] [PubMed]
- Taylor, R.S.; Muller, W.J.; Cook, M.T.; Kube, P.D.; Elliott, N.G. Gill observations in Atlantic salmon (Salmo salar, L.) during repeated amoebic gill disease (AGD) field exposure and survival challenge. Aquaculture 2009, 290, 1–8. [Google Scholar] [CrossRef]
- Crosbie, P.; Bridle, A.; Cadoret, K.; Nowak, B. In vitro cultured Neoparamoeba perurans causes amoebic gill disease in Atlantic salmon and fulfils Koch’s postulates. Int. J. Parasitol. 2012, 42, 511–515. [Google Scholar] [CrossRef] [PubMed]
- Downes, J.; Henshilwood, K.; Collins, E.; Ryan, A.; O’Connor, I.; Rodger, H.; MacCarthy, E.; Ruane, N. A longitudinal study of amoebic gill disease on a marine Atlantic salmon farm utilising a real-time PCR assay for the detection of Neoparamoeba perurans. Aquac. Environ. Interact. 2015, 7, 239–251. [Google Scholar] [CrossRef]
- Kulak, N.A.; Pichler, G.; Paron, I.; Nagaraj, N.; Mann, M. Minimal, encapsulated proteomic-sample processing applied to copy-number estimation in eukaryotic cells. Nat. Methods 2014, 11, 319–324. [Google Scholar] [CrossRef]
- Cox, J.; Mann, M. MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat. Biotechnol. 2008, 26, 1367. Available online: https://www.nature.com/articles/nbt.1511#supplementary-information (accessed on 15 May 2020). [CrossRef]
- Cox, J.; Neuhauser, N.; Michalski, A.; Scheltema, R.A.; Olsen, J.V.; Mann, M. Andromeda: A peptide search engine integrated into the MaxQuant environment. J. Proteome Res. 2011, 10, 1794–1805. [Google Scholar] [CrossRef]
- Cox, J.; Hein, M.Y.; Luber, C.A.; Paron, I.; Nagaraj, N.; Mann, M. Accurate proteome-wide label-free quantification by delayed normalization and maximal peptide ratio extraction, termed MaxLFQ. Mol. Cell. Proteom. 2014, 13, 2513–2526. [Google Scholar] [CrossRef] [PubMed]
- Tyanova, S.; Temu, T.; Sinitcyn, P.; Carlson, A.; Hein, M.Y.; Geiger, T.; Mann, M.; Cox, J. The Perseus computational platform for comprehensive analysis of (prote)omics data. Nat. Methods 2016, 13, 731. [Google Scholar] [CrossRef] [PubMed]
- Marcos-López, M.; Ruiz, C.E.; Rodger, H.D.; O’Connor, I.; MacCarthy, E.; Esteban, M.Á. Local and systemic humoral immune response in farmed Atlantic salmon (Salmo salar L.) under a natural amoebic gill disease outbreak. Fish Shellfish Immunol. 2017, 66, 207–216. [Google Scholar] [CrossRef] [PubMed]
- Ross, N.W.; Firth, K.J.; Wang, A.; Burka, J.F.; Johnson, S.C. Changes in hydrolytic enzyme activities of naive Atlantic salmon Salmo salar skin mucus due to infection with the salmon louse Lepeophtheirus salmonis and cortisol implantation. Dis. Aquat. Org. 2000, 41, 43–51. [Google Scholar] [CrossRef]
- Bols, N.C.; Barlian, A.; Chirino-Trejo, M.; Caldwell, S.J.; Goegan, P.; Lee, L.E.J. Development of a cell line from primary cultures of rainbow trout, Oncorhynchus mykiss (Walbaum), gills. J. Fish Dis. 1994, 17, 601–611. [Google Scholar] [CrossRef]
- Kim, J.H.; Kim, D.; Shin, H.J. Contact-independent cell death of human microglial cells due to pathogenic Naegleria fowleri trophozoites. Korean J. Parasitol. 2008, 46, 217–221. [Google Scholar] [CrossRef] [PubMed]
- Lun, J.; Xia, C.; Yuan, C.; Zhang, Y.; Zhong, M.; Huang, T.; Hu, Z. The outer membrane protein, LamB (maltoporin), is a versatile vaccine candidate among the Vibrio species. Vaccine 2014, 32, 809–815. [Google Scholar] [CrossRef] [PubMed]
- Payne, S.M.; Mey, A.R.; Wyckoff, E.E. Vibrio Iron Transport: Evolutionary Adaptation to Life in Multiple Environments. Microbiol. Mol. Biol. Rev. 2016, 80, 69–90. [Google Scholar] [CrossRef]
- Christie-Oleza, J.A.; Armengaud, J.; Guerin, P.; Scanlan, D.J. Functional distinctness in the exoproteomes of marine Synechococcus. Environ. Microbiol. 2015, 17, 3781–3794. [Google Scholar] [CrossRef] [PubMed]
- Christie-Oleza, J.A.; Armengaud, J. In-Depth Analysis of Exoproteomes from Marine Bacteria by Shotgun Liquid Chromatography-Tandem Mass Spectrometry: The Ruegeria pomeroyi DSS-3 Case-Study. Mar. Drugs 2010, 8, 2223–2239. [Google Scholar] [CrossRef]
- Shabardina, V.; Kischka, T.; Kmita, H.; Suzuki, Y.; Makalowski, W. Environmental adaptation of Acanthamoeba castellanii and Entamoeba histolytica at genome level as seen by comparative genomic analysis. Int. J. Biol. Sci. 2018, 14, 306–320. [Google Scholar] [CrossRef] [PubMed]
- Takahashi, K. Structure and function studies on enzymes with a catalytic carboxyl group(s): From ribonuclease T1 to carboxyl peptidases. Proc. Jpn. Acad. Ser. B Phys. Biol. Sci. 2013, 89, 201–225. [Google Scholar] [CrossRef] [PubMed]
- Buchner, J. Molecular chaperones and protein quality control: An introduction to the JBC Reviews thematic series. J. Biol. Chem. 2019, 294, 2074–2075. [Google Scholar] [CrossRef] [PubMed]
- Hurt, M.; Neelam, S.; Niederkorn, J.; Alizadeh, H. Pathogenic Acanthamoeba spp secrete a mannose-induced cytolytic protein that correlates with the ability to cause disease. Infect. Immun. 2003, 71, 6243–6255. [Google Scholar] [CrossRef] [PubMed]
- Franco, E.; Vazquez-Prado, J.; Meza, I. Fibronectin-derived fragments as inducers of adhesion and chemotaxis of Entamoeba histolytica trophozoites. J. Infect. Dis. 1997, 176, 1597–1602. [Google Scholar] [CrossRef][Green Version]
- Menanteau-Ledouble, S.; Lawrence, M.L.; El-Matbouli, M. Invasion and replication of Yersinia ruckeri in fish cell cultures. BMC Vet. Res. 2018, 14, 81. [Google Scholar] [CrossRef] [PubMed]
- Tobback, E.; Decostere, A.; Hermans, K.; Van den Broeck, W.; Haesebrouck, F.; Chiers, K. In vitro markers for virulence in Yersinia ruckeri. J. Fish Dis. 2010, 33, 197–209. [Google Scholar] [CrossRef] [PubMed]
- Lee, L.E.; Dayeh Vr Fau-Schirmer, K.; Schirmer, K.; Fau-Bols, N.C.; Bols, N.C. Applications and potential uses of fish gill cell lines: Examples with RTgill-W1. Vitr. Cell. Dev. Biol. Anim. 2009, 45, 127–134. [Google Scholar] [CrossRef] [PubMed]
- Gjessing, M.C.; Aamelfot, M.; Batts, W.N.; Benestad, S.L.; Dale, O.B.; Thoen, E.; Weli, S.C.; Winton, J.R. Development and characterization of two cell lines from gills of Atlantic salmon. PLoS ONE 2018, 13, e0191792. [Google Scholar] [CrossRef] [PubMed]
- Perez-Riverol, Y.; Csordas, A.; Bai, J.; Bernal-Llinares, M.; Hewapathirana, S.; Kundu, D.J.; Inuganti, A.; Griss, J.; Mayer, G.; Eisenacher, M.; et al. The PRIDE database and related tools and resources in 2019: Improving support for quantification data. Nucleic Acids Res. 2019, 47, D442–D450. [Google Scholar] [CrossRef]
| Gene Bank IDs | Protein IDs | Fasta Headers | Mol. Weight (kDa) | Fold Change 1 | Atten Peptides 2 | Vir Peptides 2 | t-Test 3 |
|---|---|---|---|---|---|---|---|
| PRP79790.1 | A0A2P6N765 | PPM-type phosphatase domain-containing protein (Planoprotostelium fungivorum) | 190.44 | −0.144883 | 14 | 19 | NS |
| MW534129 | PPER_00018933-RA | NA | 281.31 | −4.72831 | 40 | 17 | + |
| MW534130 | PPER_00016120-RA | TolAlike protein | 36.42 | −3.89208 | 11 | 0 | + |
| MW534131 | PPER_00020708-RA | Hypothetical protein | 64.3 | −5.01281 | 10 | 0 | + |
| MW534132 | PPER_00011993-RA | NA | 35.25 | −1.15085 | 8 | 6 | + |
| MW534133 | PPER_00013024-RA | Hypothetical protein | 13.01 | −4.76979 | 8 | 3 | + |
| MW534135 | PPER_00019895-RA | Multifunctional chaperone | 28.08 | −3.49453 | 9 | 2 | + |
| MW534136 | PPER_00008268-RA | Urea ABC transporter substrate-binding protein Branched-chain amino acid | 47.26 | −3.18528 | 6 | 0 | + |
| MW534137 | PPER_00011992-RA | Hypothetical protein | 39.76 | −2.86775 | 6 | 6 | + |
| MT419758 | PPER_00012939-RA | Cu-Zn superoxide dismutase | 15.50 | −3.72917 | 6 | 2 | + |
| MW534138 | PPER_00014132-RA | Physarolisin | 49.49 | −4.06437 | 5 | 1 | + |
| MW534139 | PPER_00011189-RA | Peptidase S8 and S53 domain-containing protein | 58.69 | −2.78376 | 3 | 0 | + |
| Gene | Protein IDs | Fasta Headers | Organism | Mol. Weight (kDa) | Fold Change 1 | Att. Peptides 2 | Vir Peptides 2 | t-Test 3 |
|---|---|---|---|---|---|---|---|---|
| ST37_14230 | A0A1X1MR06 | ABC transporter substrate-binding protein | Vibrio sp. qd031 | 47.26 | −3.18528 | 6 | 0 | + |
| VS_II0220 | B7VQI9 | Maltodextrin-binding protein | Vibrio tasmaniensis (strain LGP32 | 42.39 | −4.90439 | 13 | 4 | + |
| VS_0355 | B7VIC7 | Immunogenic protein | Vibrio tasmaniensis (strain LGP32) | 34.44 | −5.20998 | 11 | 1 | + |
| ahpCB | Q3IE21 | Peroxiredoxin 2 (TSA) (PRP) | Pseudoalteromonas haloplanktis (strain TAC 125) | 22.12 | −4.75659 | 9 | 0 | + |
| PSHAa297; B1199_19990 | Q3IK06; A0A244CKW5 | Secreted alkaline phosphatase; Alkaline phosphatase | Pseudoalteromonas haloplanktis (strain TAC 125); Pseudoalteromonas ulvae | 56.1 | −4.1791 | 10 | 2 | + |
| PCIT_22080 | U1KK96 | Peptidase S8/S53 subtilisin kexin sedolisin | Pseudoalteromonas citrea DSM 8771 | 127.37 | −3.70559 | 6 | 3 | + |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ní Dhufaigh, K.; Botwright, N.; Dillon, E.; O’Connor, I.; MacCarthy, E.; Slattery, O. Differential Exoproteome and Biochemical Characterisation of Neoparamoeba perurans. Microorganisms 2021, 9, 1258. https://doi.org/10.3390/microorganisms9061258
Ní Dhufaigh K, Botwright N, Dillon E, O’Connor I, MacCarthy E, Slattery O. Differential Exoproteome and Biochemical Characterisation of Neoparamoeba perurans. Microorganisms. 2021; 9(6):1258. https://doi.org/10.3390/microorganisms9061258
Chicago/Turabian StyleNí Dhufaigh, Kerrie, Natasha Botwright, Eugene Dillon, Ian O’Connor, Eugene MacCarthy, and Orla Slattery. 2021. "Differential Exoproteome and Biochemical Characterisation of Neoparamoeba perurans" Microorganisms 9, no. 6: 1258. https://doi.org/10.3390/microorganisms9061258
APA StyleNí Dhufaigh, K., Botwright, N., Dillon, E., O’Connor, I., MacCarthy, E., & Slattery, O. (2021). Differential Exoproteome and Biochemical Characterisation of Neoparamoeba perurans. Microorganisms, 9(6), 1258. https://doi.org/10.3390/microorganisms9061258






