Targeted Molecular Detection of Nosocomial Carbapenemase-Producing Gram-Negative Bacteria—On Near- and Distant-Patient Surfaces
Abstract
1. Introduction
2. Materials and Methods
2.1. Sampling Strategy in the Patient Area
2.2. Sampling on Surfaces
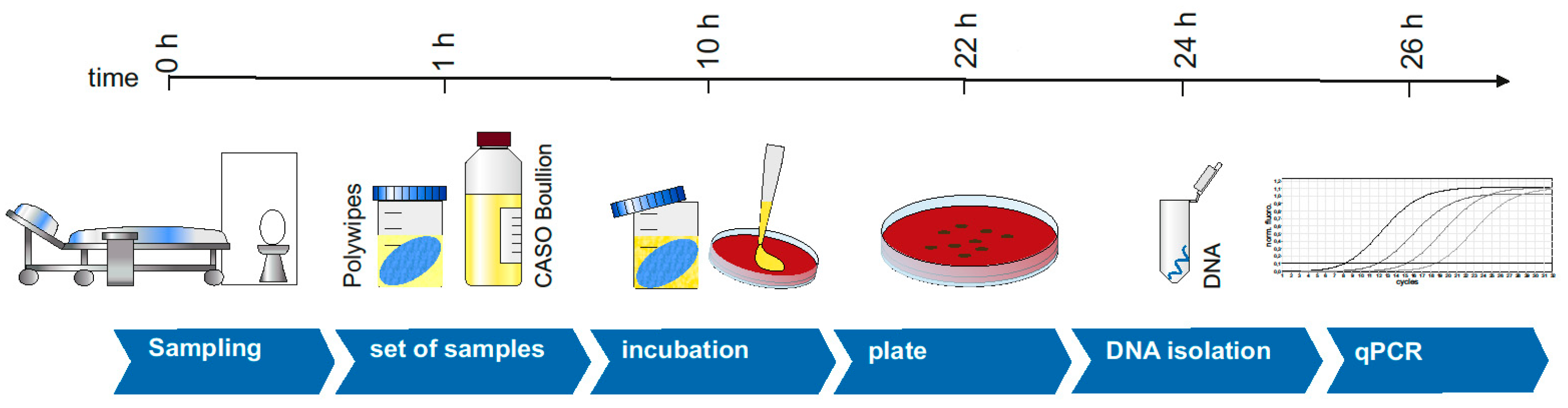
2.3. Enrichment and Incubation
2.4. DNA Isolation and PCR
3. Results
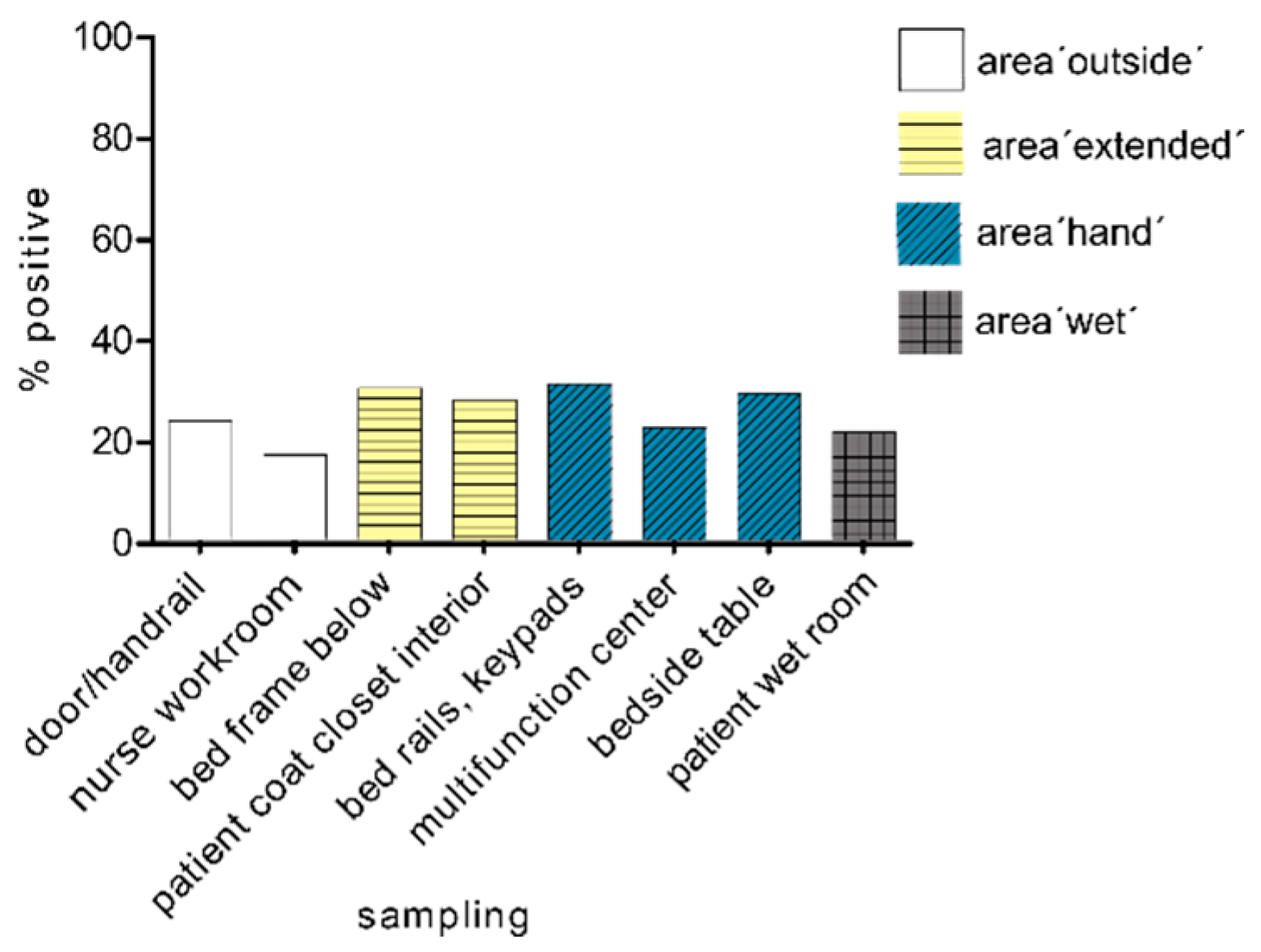
3.1. Sampling in the Patient Area
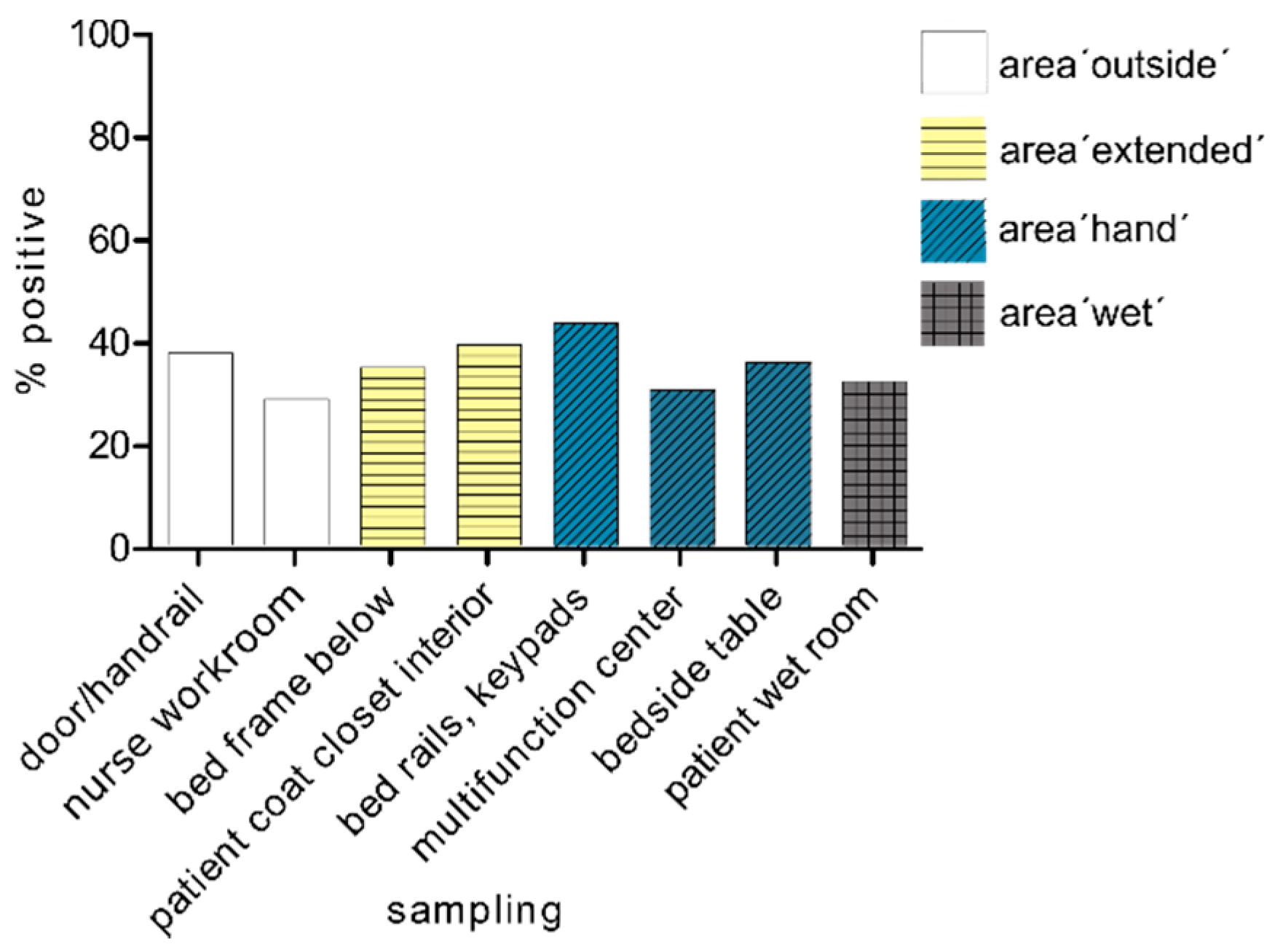
3.2. Spread of Multidrug-Resistant Gram-Negative Pathogens with Carbapenemase after 24 h
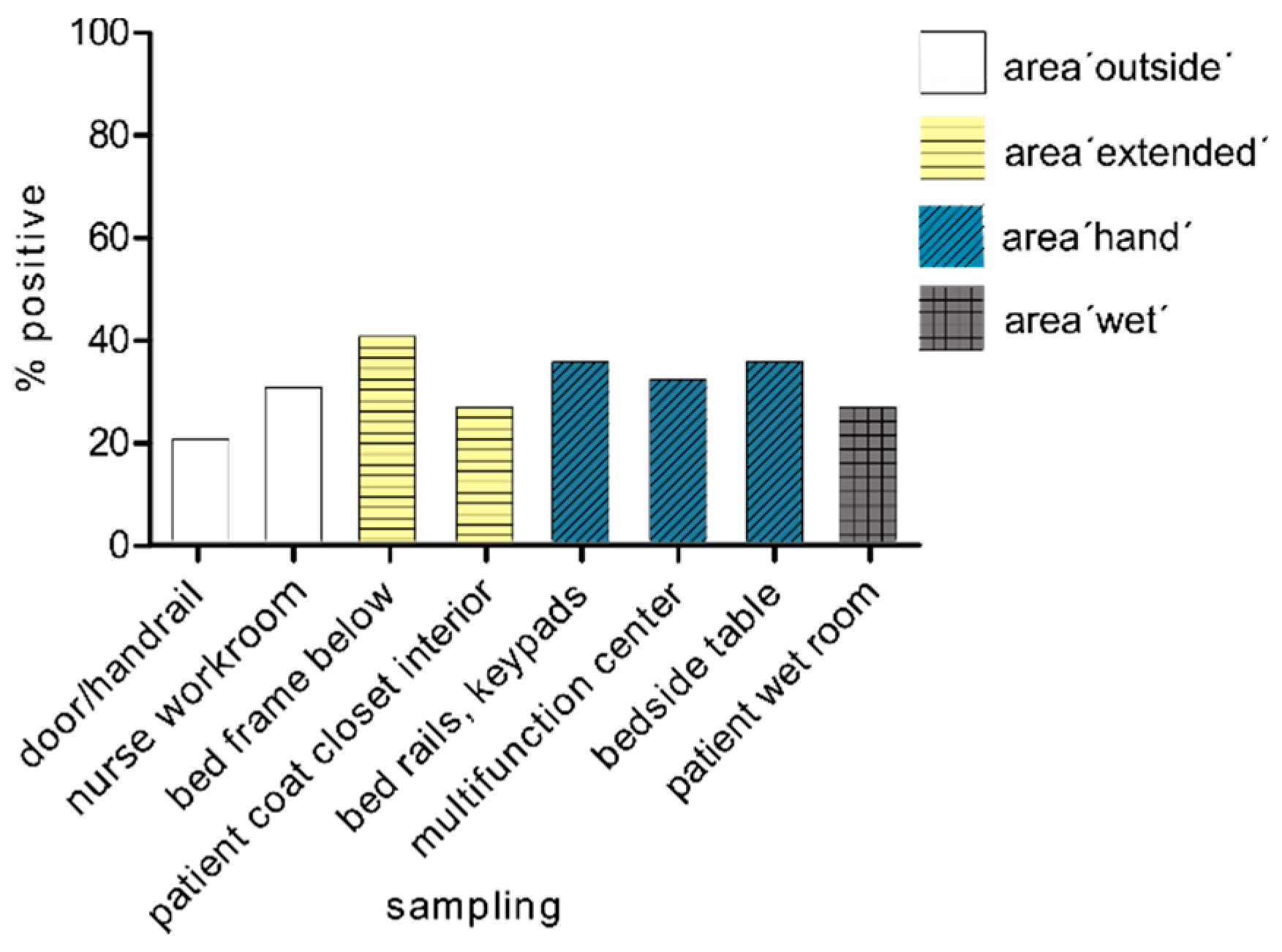
3.3. Spread of Multi-Resistant Gram-Negative Pathogens with Carbapenemase after 72 h
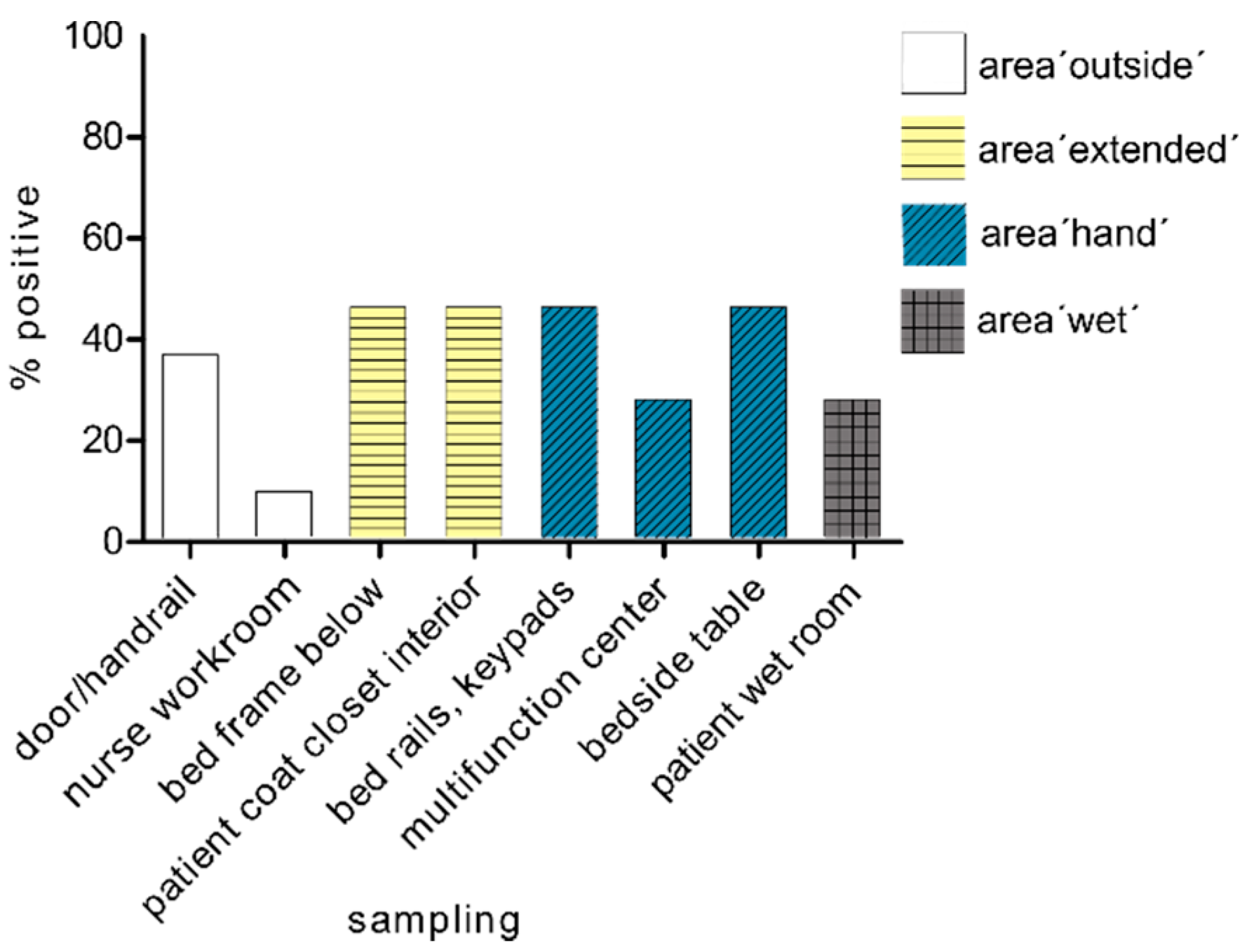
3.4. Spread of Multi-Resistant Gram-Negative Pathogens with Carbapenemase after One Week
3.5. Spread of Multi-Resistant Gram-Negative Pathogens (Independent of Time)
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Suleyman, G.; Alangaden, G.; Bardossy, A.C. The Role of Environmental Contamination in the Transmission of Nosocomial Pathogens and Healthcare-Associated Infections. Curr. Infect. Dis Rep. 2018, 20, 12. [Google Scholar] [CrossRef] [PubMed]
- Weber, D.J.; Rutala, W.A.; Anderson, D.J.; Chen, L.F.; Sickbert-Bennett, E.E.; Boyce, J.M. Effectiveness of ultraviolet devices and hydrogen peroxide systems for terminal room decontamination: Focus on clinical trials. Am. J. Infect. Control 2016, 44, e77–e84. [Google Scholar] [CrossRef] [PubMed]
- Rutala, W.A.; Weber, D.J. Are room decontamination units needed to prevent transmission of environmental pathogens? Infect. Control Hosp. Epidemiol. 2011, 32, 743–747. [Google Scholar] [CrossRef] [PubMed]
- Rutala, W.A.; Kanamori, H.; Gergen, M.F.; Sickbert-Bennett, E.E.; Anderson, D.J.; Sexton, D.J.; Weber, D.J.; CDC Prevention Epicenters Program. Evaluation of dilute hydrogen peroxide technology for continuous room decontamination of multidrug-resistant organisms. Infect. Control Hosp. Epidemiol. 2019, 40, 1438–1439. [Google Scholar] [CrossRef]
- Russotto, V.; Cortegiani, A.; Fasciana, T.; Iozzo, P.; Raineri, S.M.; Gregoretti, C.; Giammanco, A.; Giarratano, A. What Healthcare Workers Should Know about Environmental Bacterial Contamination in the Intensive Care Unit. BioMed Res. Int. 2017, 2017, 6905450. [Google Scholar] [CrossRef]
- Huslage, K.; Rutala, W.A.; Gergen, M.F.; Sickbert-Bennett, E.E.; Weber, D.J. Microbial assessment of high-, medium-, and low-touch hospital room surfaces. Infect. Control Hosp. Epidemiol. 2013, 34, 211–212. [Google Scholar] [CrossRef]
- Rutala, W.A.; Weber, D.J. Outbreaks of carbapenem-resistant Enterobacteriaceae infections associated with duodenoscopes: What can we do to prevent infections? Am. J. Infect. Control 2016, 44, e47–e51. [Google Scholar] [CrossRef]
- Kommission für Krankenhaushygiene und Infektionsprävention. Hygiene measures for infection or colonization with multidrug-resistant gram-negative bacilli. Commission recommendation for hospital hygiene and infection prevention (KRINKO) at the Robert Koch Institute (RKI). Bundesgesundheitsblatt Gesundheitsforschung Gesundheitsschutz 2012, 55, 1311–1354. (In German) [Google Scholar] [CrossRef]
- Bonomo, R.A.; Burd, E.M.; Conly, J.; Limbago, B.M.; Poirel, L.; Segre, J.A.; Westblade, L.F. Carbapenemase-Producing Organisms: A Global Scourge. Clin. Infect. Dis. 2018, 66, 1290–1297. [Google Scholar] [CrossRef]
- Park, J.W.; Kwak, S.H.; Jung, J.; Lee, J.Y.; Lim, Y.J.; Choi, H.S.; Hong, M.J.; Choi, S.H.; Kim, M.N.; Kim, S.H. The Rate of Acquisition of Carbapenemase-Producing Enterobacteriaceae among Close Contact Patients Depending on Carbapenemase Enzymes. Infect. Chemother. 2020, 52, 39–47. [Google Scholar] [CrossRef]
- Weber, D.J.; Rutala, W.A.; Kanamori, H.; Gergen, M.F.; Sickbert-Bennett, E.E. Carbapenem-resistant Enterobacteriaceae: Frequency of hospital room contamination and survival on various inoculated surfaces. Infect. Control Hosp. Epidemiol. 2015, 36, 590–593. [Google Scholar] [CrossRef]
- Decraene, V.; Phan, H.T.T.; George, R.; Wyllie, D.H.; Akinremi, O.; Aiken, Z.; Cleary, P.; Dodgson, A.; Pankhurst, L.; Crook, D.W.; et al. A Large, Refractory Nosocomial Outbreak of Klebsiella pneumoniae Carbapenemase-Producing Escherichia coli Demonstrates Carbapenemase Gene Outbreaks Involving Sink Sites Require Novel Approaches to Infection Control. Antimicrob. Agents Chemother. 2018, 62. [Google Scholar] [CrossRef] [PubMed]
- Turner, C.; Mosby, D.; Partridge, D.; Mason, C.; Parsons, H. A patient sink tap facilitating carbapenemase-producing enterobacteriales transmission. J. Hosp. Infect. 2019. [Google Scholar] [CrossRef]
- Clarivet, B.; Grau, D.; Jumas-Bilak, E.; Jean-Pierre, H.; Pantel, A.; Parer, S.; Lotthe, A. Persisting transmission of carbapenemase-producing Klebsiella pneumoniae due to an environmental reservoir in a university hospital, France, 2012 to 2014. Eurosurveillance 2016, 21. [Google Scholar] [CrossRef] [PubMed]
- Koklic, T.; Urbancic, I.; Zdovc, I.; Golob, M.; Umek, P.; Arsov, Z.; Drazic, G.; Pintaric, S.; Dobeic, M.; Strancar, J. Surface deposited one-dimensional copper-doped TiO2 nanomaterials for prevention of health care acquired infections. PLoS ONE 2018, 13, e0201490. [Google Scholar] [CrossRef] [PubMed]
- Boyle, M.A.; Kearney, A.; Carling, P.C.; Humphreys, H. ‘Off the rails’: Hospital bed rail design, contamination, and the evaluation of their microbial ecology. J. Hosp. Infect. 2019, 103, e16–e22. [Google Scholar] [CrossRef] [PubMed]
- Caselli, E.; D’Accolti, M.; Soffritti, I.; Lanzoni, L.; Bisi, M.; Volta, A.; Berloco, F.; Mazzacane, S. An Innovative Strategy for the Effective Reduction of MDR Pathogens from the Nosocomial Environment. Adv. Exp. Med. Biol. 2019, 1214, 79–91. [Google Scholar] [CrossRef] [PubMed]
- Anderson, D.J.; Moehring, R.W.; Weber, D.J.; Lewis, S.S.; Chen, L.F.; Schwab, J.C.; Becherer, P.; Blocker, M.; Triplett, P.F.; Knelson, L.P.; et al. Effectiveness of targeted enhanced terminal room disinfection on hospital-wide acquisition and infection with multidrug-resistant organisms and Clostridium difficile: A secondary analysis of a multicentre cluster randomised controlled trial with crossover design (BETR Disinfection). Lancet Infect. Dis. 2018, 18, 845–853. [Google Scholar] [CrossRef] [PubMed]
- Weber, D.J.; Kanamori, H.; Rutala, W.A. ‘No touch’ technologies for environmental decontamination: Focus on ultraviolet devices and hydrogen peroxide systems. Curr. Opin. Infect. Dis. 2016, 29, 424–431. [Google Scholar] [CrossRef]
- Eichner, A.; Holzmann, T.; Eckl, D.B.; Zeman, F.; Koller, M.; Huber, M.; Pemmerl, S.; Schneider-Brachert, W.; Baumler, W. Novel photodynamic coating reduces the bioburden on near-patient surfaces thereby reducing the risk for onward pathogen transmission: A field study in two hospitals. J. Hosp. Infect. 2020, 104, 85–91. [Google Scholar] [CrossRef]
- Weiss, D.; Engelmann, I.; Braun, S.D.; Monecke, S.; Ehricht, R. A multiplex real-time PCR for the direct, fast, economic and simultaneous detection of the carbapenemase genes blaKPC, blaNDM, blaVIM and blaOXA-48. J. Microbiol. Methods 2017, 142, 20–26. [Google Scholar] [CrossRef]
- Brandt, C.; Zander, E.; Pfeifer, Y.; Braun, S.D.; Ehricht, R.; Makarewicz, O.; Pletz, M.W. Development of a rapid diagnostic assay based on magnetic bead purification of OXA-beta-lactamase mRNA. Future Microbiol. 2016, 11, 617–629. [Google Scholar] [CrossRef]
- Aubron, C.; Poirel, L.; Ash, R.J.; Nordmann, P. Carbapenemase-producing Enterobacteriaceae, U.S. rivers. Emerg. Infect. Dis. 2005, 11, 260–264. [Google Scholar] [CrossRef] [PubMed]
- Poirel, L.; Walsh, T.R.; Cuvillier, V.; Nordmann, P. Multiplex PCR for detection of acquired carbapenemase genes. Diagn. Microbiol. Infect. Dis. 2011, 70, 119–123. [Google Scholar] [CrossRef] [PubMed]
- Garcia-Fernandez, A.; Miriagou, V.; Papagiannitsis, C.C.; Giordano, A.; Venditti, M.; Mancini, C.; Carattoli, A. An ertapenem-resistant extended-spectrum-beta-lactamase-producing Klebsiella pneumoniae clone carries a novel OmpK36 porin variant. Antimicrob. Agents Chemother. 2010, 54, 4178–4184. [Google Scholar] [CrossRef]
- Novais, A.; Rodrigues, C.; Branquinho, R.; Antunes, P.; Grosso, F.; Boaventura, L.; Ribeiro, G.; Peixe, L. Spread of an OmpK36-modified ST15 Klebsiella pneumoniae variant during an outbreak involving multiple carbapenem-resistant Enterobacteriaceae species and clones. Eur. J. Clin. Microbiol. Infect. Dis. 2012, 31, 3057–3063. [Google Scholar] [CrossRef] [PubMed]
- Fernandez, L.; Hancock, R.E. Adaptive and mutational resistance: Role of porins and efflux pumps in drug resistance. Clin. Microbiol. Rev. 2012, 25, 661–681. [Google Scholar] [CrossRef] [PubMed]
- De Geyter, D.; Blommaert, L.; Verbraeken, N.; Sevenois, M.; Huyghens, L.; Martini, H.; Covens, L.; Pierard, D.; Wybo, I. The sink as a potential source of transmission of carbapenemase-producing Enterobacteriaceae in the intensive care unit. Antimicrob. Resist. Infect. Control 2017, 6, 24. [Google Scholar] [CrossRef]
- Song, X.; Vossebein, L.; Zille, A. Efficacy of disinfectant-impregnated wipes used for surface disinfection in hospitals: A review. Antimicrob. Resist. Infect. Control 2019, 8, 139. [Google Scholar] [CrossRef] [PubMed]
- Weber, D.J.; Anderson, D.; Rutala, W.A. The role of the surface environment in healthcare-associated infections. Curr. Opin. Infect. Dis. 2013, 26, 338–344. [Google Scholar] [CrossRef] [PubMed]
- Rutala, W.A.; Kanamori, H.; Gergen, M.F.; Knelson, L.P.; Sickbert-Bennett, E.E.; Chen, L.F.; Anderson, D.J.; Sexton, D.J.; Weber, D.J.; CDC Prevention Epicenters Program (CDCPEP). Enhanced disinfection leads to reduction of microbial contamination and a decrease in patient colonization and infection. Infect. Control Hosp. Epidemiol. 2018, 39, 1118–1121. [Google Scholar] [CrossRef] [PubMed]
- Schneider, A.; Coope, C.; Michie, S.; Puleston, R.; Hopkins, S.; Oliver, I. Implementing a toolkit for the prevention, management and control of carbapenemase-producing Enterobacteriaceae in English acute hospitals trusts: A qualitative evaluation. BMC Health Serv. Res. 2019, 19, 689. [Google Scholar] [CrossRef]
- Shams, A.M.; Rose, L.J.; Edwards, J.R.; Cali, S.; Harris, A.D.; Jacob, J.T.; LaFae, A.; Pineles, L.L.; Thom, K.A.; McDonald, L.C.; et al. Assessment of the Overall and Multidrug-Resistant Organism Bioburden on Environmental Surfaces in Healthcare Facilities. Infect. Control Hosp. Epidemiol. 2016, 37, 1426–1432. [Google Scholar] [CrossRef]
- O’Fallon, E.; Schreiber, R.; Kandel, R.; D’Agata, E.M. Multidrug-resistant gram-negative bacteria at a long-term care facility: Assessment of residents, healthcare workers, and inanimate surfaces. Infect. Control Hosp. Epidemiol. 2009, 30, 1172–1179. [Google Scholar] [CrossRef]
- Rock, C.; Anderson, M.; Lewis, S.; Scheeler, V.; Nowakowski, E.; Hsu, Y.J.; Milstone, A.M.; Carroll, K.C.; Maragakis, L.L.; Simner, P.J.; et al. Comparison of nylon-flocked swab and cellulose sponge methods for carbapenem-resistant Enterobacteriaceae and gram-negative organism recovery from high-touch surfaces in patient rooms. Infect. Control Hosp. Epidemiol. 2018, 39, 1257–1261. [Google Scholar] [CrossRef] [PubMed]
- Schmidt, M.G.; Anderson, T.; Attaway, H.H., 3rd; Fairey, S.; Kennedy, C.; Salgado, C.D. Patient environment microbial burden reduction: A pilot study comparison of 2 terminal cleaning methods. Am. J. Infect. Control 2012, 40, 559–561. [Google Scholar] [CrossRef] [PubMed]
| Carbapenemase | Forward Primer | Reverse Primer | Annealing Temperature | Reference |
|---|---|---|---|---|
| VIM | TGGCAACGTACGCATCACC | CGCAGCACCGGGATAGAA | 61 °C | [21] |
| OXA-48 | GCGTGTATTAGCCTTATCGGCTG | GCGGGTAAAAATGCTTGGTTCGC | 60 °C | [22] |
| IMI | ATAGCCATCCTTGTTTAGCTC | TCTGCGATTACTTTATCCTC | 62 °C | [23] |
| NDM | GGTTTGGCGATCTGGTTTTC | CGGAATGGCTCATCACGATC | 68 °C | [24] |
| KPC | CGTCTAGTTCTGCTGTCTTG | CTTGTCATCCTTGTTAGGCG | 68 °C | [24] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Stein, C.; Lange, I.; Rödel, J.; Pletz, M.W.; Kipp, F. Targeted Molecular Detection of Nosocomial Carbapenemase-Producing Gram-Negative Bacteria—On Near- and Distant-Patient Surfaces. Microorganisms 2021, 9, 1190. https://doi.org/10.3390/microorganisms9061190
Stein C, Lange I, Rödel J, Pletz MW, Kipp F. Targeted Molecular Detection of Nosocomial Carbapenemase-Producing Gram-Negative Bacteria—On Near- and Distant-Patient Surfaces. Microorganisms. 2021; 9(6):1190. https://doi.org/10.3390/microorganisms9061190
Chicago/Turabian StyleStein, Claudia, Isabel Lange, Jürgen Rödel, Mathias W. Pletz, and Frank Kipp. 2021. "Targeted Molecular Detection of Nosocomial Carbapenemase-Producing Gram-Negative Bacteria—On Near- and Distant-Patient Surfaces" Microorganisms 9, no. 6: 1190. https://doi.org/10.3390/microorganisms9061190
APA StyleStein, C., Lange, I., Rödel, J., Pletz, M. W., & Kipp, F. (2021). Targeted Molecular Detection of Nosocomial Carbapenemase-Producing Gram-Negative Bacteria—On Near- and Distant-Patient Surfaces. Microorganisms, 9(6), 1190. https://doi.org/10.3390/microorganisms9061190







