Elevated Incidences of Antimicrobial Resistance and Multidrug Resistance in the Maumee River (Ohio, USA), a Major Tributary of Lake Erie
Abstract
1. Introduction
2. Materials and Methods
2.1. Sampling Sites, Study Duration and Sampling Methods
2.2. E. coli Strains Archival and Enumeration
2.3. Antimicrobial Resitance Analysis
2.4. Statistical Analysis
3. Results
3.1. E. coli Population
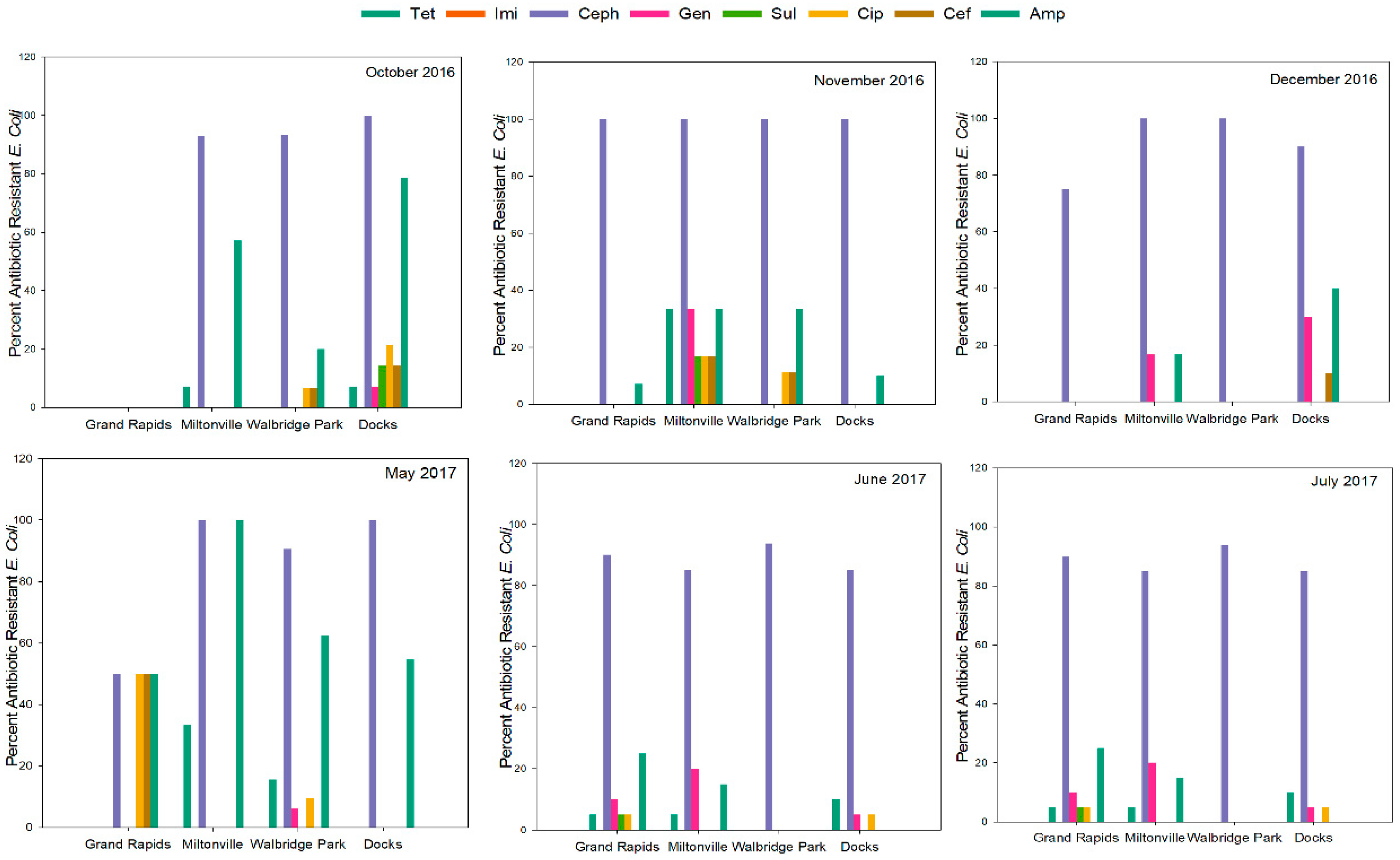
3.2. Antimicrobial Resistance Patterns
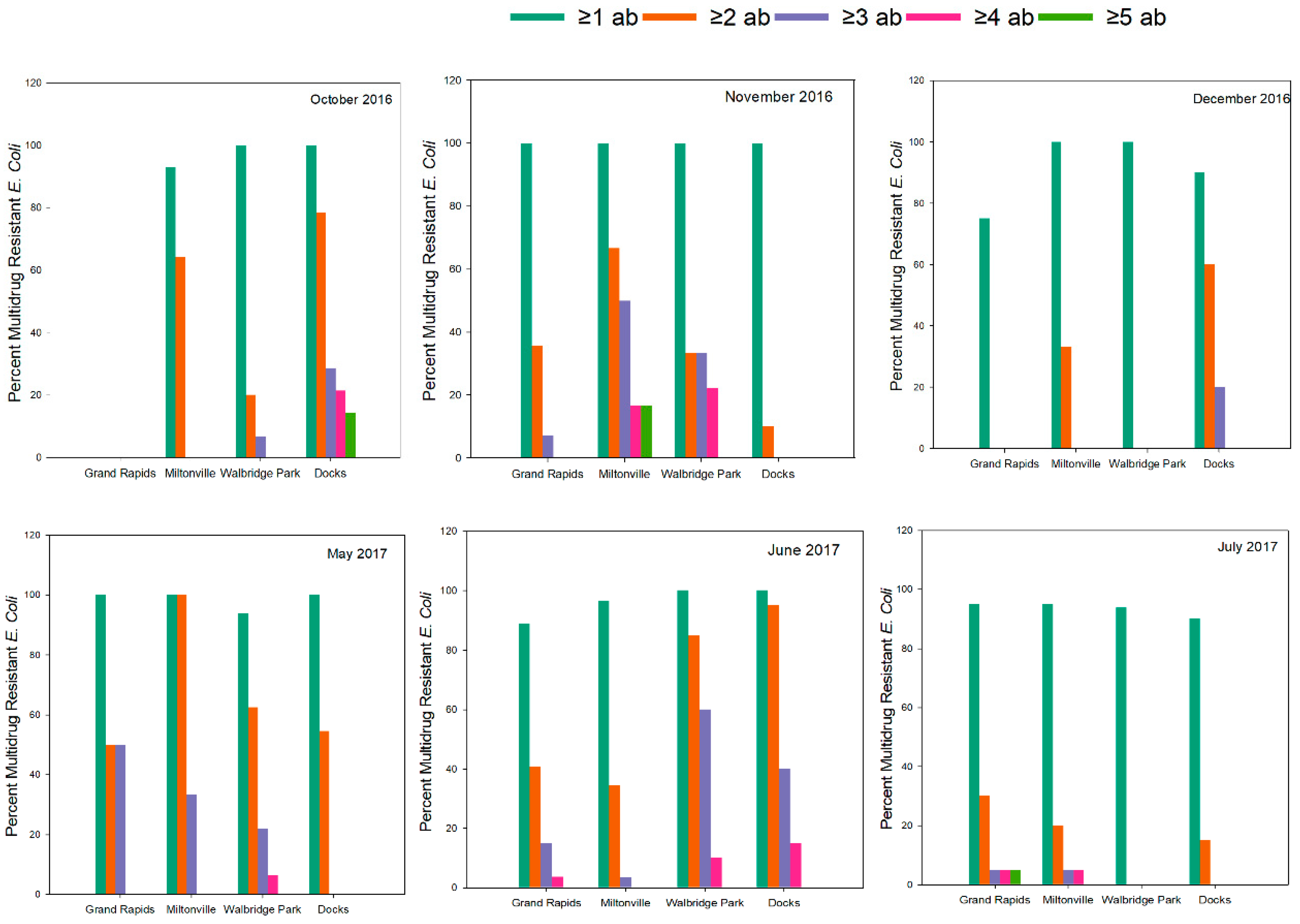
3.3. Multidrug Resistance Patterns
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Davison, J. Genetic Exchange between Bacteria in the Environment. Plasmid 1999, 42, 73–91. [Google Scholar] [CrossRef] [PubMed]
- Pepper, I.L.; Gentry, T.J. Microorganisms Found in the Environment. In Environmental Microbiology; Elsevier: Amsterdam, The Netherlands, 2015; pp. 9–36. [Google Scholar]
- Thomas, S.G.; Rieu, I.; Steber, C.M. Gibberellin Metabolism and Signaling. In Vitamins & Hormones; Litwack, G., Ed.; Academic Press: Cambridge, MA, USA, 2005; pp. 289–338. [Google Scholar]
- Pepper, I.L.; Gerba, C.P.; Gentry, T.J. Global Emerging Microbial Issues in the Anthropocene Era. In Environmental Microbiology; Elsevier: Amsterdam, The Netherlands, 2015; pp. 677–688. [Google Scholar]
- Bush, K.; Courvalin, P.; Dantas, G.; Davies, J.; Eisenstein, B.; Huovinen, P.; Jacoby, G.A.; Kishony, R.; Kreiswirth, B.N.; Kutter, E.; et al. Tackling Antibiotic Resistance. Nat. Rev. Microbiol. 2011, 9, 894–896. [Google Scholar] [CrossRef] [PubMed]
- Pruden, A. Balancing Water Sustainability and Public Health Goals in the Face of Growing Concerns about Antibiotic Resistance. Environ. Sci. Technol. 2014, 48, 5–14. [Google Scholar] [CrossRef]
- Allen, H.K.; Donato, J.; Wang, H.H.; Cloud-Hansen, K.A.; Davies, J.; Handelsman, J. Call of the Wild: Antibiotic Resistance Genes in Natural Environments. Nat. Rev. Microbiol. 2010, 8, 251–259. [Google Scholar] [CrossRef]
- Spellberg, B.; Guidos, R.; Gilbert, D.; Bradley, J.; Boucher, H.W.; Scheld, W.M.; Bartlett, J.G.; Edwards, J.; Infectious Diseases Society of America. The Epidemic of Antibiotic-Resistant Infections: A Call to Action for the Medical Community from the Infectious Diseases Society of America. Clin. Infect. Dis. 2008, 46, 155–164. [Google Scholar] [CrossRef] [PubMed]
- Freire-Moran, L.; Aronsson, B.; Manz, C.; Gyssens, I.C.; So, A.D.; Monnet, D.L.; Cars, O. Critical Shortage of New Antibiotics in Development against Multidrug-Resistant Bacteria—Time to React Is Now. Drug Resist. Updates 2011, 14, 118–124. [Google Scholar] [CrossRef]
- Kemper, N. Veterinary Antibiotics in the Aquatic and Terrestrial Environment. Ecol. Indic. 2008, 8, 1–13. [Google Scholar] [CrossRef]
- Zhang, X.-X.; Zhang, T.; Fang, H.H.P. Antibiotic resistance genes in water environment. Appl. Microbiol. Biotechnol. 2009, 82, 397–414. [Google Scholar] [CrossRef]
- Goossens, H.; Ferech, M.; Vander Stichele, R.; Elseviers, M.; ESAC Project Group. Outpatient Antibiotic Use in Europe and Association with Resistance: A Cross-National Database Study. Lancet 2005, 365, 579–587. [Google Scholar] [CrossRef]
- Lim, S.-M.; Webb, S.A.R. Nosocomial Bacterial Infections in Intensive Care Units. I: Organisms and Mechanisms of Antibiotic Resistance. Anaesthesia 2005, 60, 887–902. [Google Scholar] [CrossRef]
- Blot, S.; Vandijck, D.; Lizy, C.; Annemans, L.; Vogelaers, D.; Mauldin, P.D.; Salgado, C.D.; Bosso, J.A. Estimating the Length of Hospitalization Attributable to Multidrug Antibiotic Resistance. Antimicrob. Agents Chemother. 2010, 54, 4046–4047. [Google Scholar] [CrossRef]
- Lye, D.C.; Earnest, A.; Ling, M.L.; Lee, T.-E.; Yong, H.-C.; Fisher, D.A.; Krishnan, P.; Hsu, L.-Y. The Impact of Multidrug Resistance in Healthcare-Associated and Nosocomial Gram-Negative Bacteraemia on Mortality and Length of Stay: Cohort Study. Clin. Microbiol. Infect. 2012, 18, 502–508. [Google Scholar] [CrossRef]
- Chopra, I.; Schofield, C.; Everett, M.; O’Neill, A.; Miller, K.; Wilcox, M.; Frère, J.-M.; Dawson, M.; Czaplewski, L.; Urleb, U.; et al. Treatment of Health-Care-Associated Infections Caused by Gram-Negative Bacteria: A Consensus Statement. Lancet Infect. Dis. 2008, 8, 133–139. [Google Scholar] [CrossRef]
- Nikaido, H.; Pagès, J.-M. Broad-Specificity Efflux Pumps and Their Role in Multidrug Resistance of Gram-Negative Bacteria. FEMS Microbiol. Rev. 2012, 36, 340–363. [Google Scholar] [CrossRef]
- Naqvi, S.Z.H.; Kiran, U.; Ali, M.I.; Jamal, A.; Hameed, A.; Ahmed, S.; Ali, N. Combined efficacy of biologically synthesized silver nanoparticles and different antibiotics against multidrug-resistant bacteria. Int. J. Nanomed. 2013, 8, 3187–3195. [Google Scholar] [CrossRef]
- Worthington, R.J.; Melander, C. Combination Approaches to Combat Multidrug-Resistant Bacteria. Trends Biotechnol. 2013, 31, 177–184. [Google Scholar] [CrossRef] [PubMed]
- Wright, G.D. Antibiotic Resistance in the Environment: A Link to the Clinic? Curr. Opin. Microbiol. 2010, 13, 589–594. [Google Scholar] [CrossRef] [PubMed]
- Rizzo, L.; Manaia, C.; Merlin, C.; Schwartz, T.; Dagot, C.; Ploy, M.C.; Michael, I.; Fatta-Kassinos, D. Urban Wastewater Treatment Plants as Hotspots for Antibiotic Resistant Bacteria and Genes Spread into the Environment: A Review. Sci. Total Environ. 2013, 447, 345–360. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.; Jensen, J.N.; Aga, D.S.; Weber, A.S. Tetracycline as a Selector for Resistant Bacteria in Activated Sludge. Chemosphere 2007, 66, 1643–1651. [Google Scholar] [CrossRef]
- Schlüter, A.; Szczepanowski, R.; Pühler, A.; Top, E.M. Genomics of IncP-1 Antibiotic Resistance Plasmids Isolated from Wastewater Treatment Plants Provides Evidence for a Widely Accessible Drug Resistance Gene Pool. FEMS Microbiol. Rev. 2007, 31, 449–477. [Google Scholar] [CrossRef] [PubMed]
- Szczepanowski, R.; Linke, B.; Krahn, I.; Gartemann, K.-H.; Gützkow, T.; Eichler, W.; Pühler, A.; Schlüter, A. Detection of 140 Clinically Relevant Antibiotic-Resistance Genes in the Plasmid Metagenome of Wastewater Treatment Plant Bacteria Showing Reduced Susceptibility to Selected Antibiotics. Microbiology 2009, 155, 2306–2319. [Google Scholar] [CrossRef]
- Harwood, J.J. Molecular Markers for Identifying Municipal, Domestic and Agricultural Sources of Organic Matter in Natural Waters. Chemosphere 2014, 95, 3–8. [Google Scholar] [CrossRef]
- Bernhard, A.E.; Field, K.G. A PCR Assay To Discriminate Human and Ruminant Feces on the Basis of Host Differences in Bacteroides-Prevotella Genes Encoding 16S RRNA. Appl. Environ. Microbiol. 2000, 66, 4571–4574. [Google Scholar] [CrossRef]
- Shanks, O.C.; Atikovic, E.; Blackwood, A.D.; Lu, J.; Noble, R.T.; Domingo, J.S.; Seifring, S.; Sivaganesan, M.; Haugland, R.A. Quantitative PCR for Detection and Enumeration of Genetic Markers of Bovine Fecal Pollution. Appl. Environ. Microbiol. 2008, 74, 745–752. [Google Scholar] [CrossRef]
- Ervin, J.S.; Van De Werfhorst, L.C.; Murray, J.L.S.; Holden, P.A. Microbial Source Tracking in a Coastal California Watershed Reveals Canines as Controllable Sources of Fecal Contamination. Environ. Sci. Technol. 2014, 48, 9043–9052. [Google Scholar] [CrossRef]
- Guber, A.K.; Fry, J.; Ives, R.L.; Rose, J.B. Escherichia Coli Survival in, and Release from, White-Tailed Deer Feces. Appl. Environ. Microbiol. 2015, 81, 1168–1176. [Google Scholar] [CrossRef] [PubMed]
- Mukherjee, M.; Gentry, T.; Mjelde, H.; Brooks, J.; Harmel, D.; Gregory, L.; Wagner, K. Escherichia Coli Antimicrobial Resistance Variability in Water Runoff and Soil from a Remnant Native Prairie, an Improved Pasture, and a Cultivated Agricultural Watershed. Water 2020, 12, 1251. [Google Scholar] [CrossRef]
- Price, R.G.; Wildeboer, D.E. coli as an Indicator of Contamination and Health Risk in Environmental Waters. In Escherichia coli—Recent Advances on Physiology, Pathogenesis and Biotechnological Applications; InTech: London, UK, 2017. [Google Scholar]
- Ibekwe, A.M.; Murinda, S.E.; Graves, A.K. Genetic Diversity and Antimicrobial Resistance of Escherichia Coli from Human and Animal Sources Uncovers Multiple Resistances from Human Sources. PLoS ONE 2011, 6, e20819. [Google Scholar] [CrossRef]
- Blaak, H.; Lynch, G.; Italiaander, R.; Hamidjaja, R.A.; Schets, F.M.; de Roda Husman, A.M. Multidrug-Resistant and Extended Spectrum Beta-Lactamase-Producing Escherichia Coli in Dutch Surface Water and Wastewater. PLoS ONE 2015, 10, e0127752. [Google Scholar] [CrossRef] [PubMed]
- Ramírez Castillo, F.Y.; Avelar González, F.J.; Garneau, P.; Márquez Díaz, F.; Guerrero Barrera, A.L.; Harel, J. Presence of Multi-Drug Resistant Pathogenic Escherichia Coli in the San Pedro River Located in the State of Aguascalientes, Mexico. Front. Microbiol. 2013, 4, 147. [Google Scholar] [CrossRef] [PubMed]
- Sayah, R.S.; Kaneene, J.B.; Johnson, Y.; Miller, R. Patterns of Antimicrobial Resistance Observed in Escherichia Coli Isolates Obtained from Domestic- and Wild-Animal Fecal Samples, Human Septage, and Surface Water. Appl. Environ. Microbiol. 2005, 71, 1394–1404. [Google Scholar] [CrossRef]
- Servais, P.; Passerat, J. Antimicrobial Resistance of Fecal Bacteria in Waters of the Seine River Watershed (France). Sci. Total Environ. 2009, 408, 365–372. [Google Scholar] [CrossRef] [PubMed]
- Edge, S.E.; Morgan, M.B.; Gleason, D.F.; Snell, T.W. Development of a Coral CDNA Array to Examine Gene Expression Profiles in Montastraea Faveolata Exposed to Environmental Stress. Mar. Pollut. Bull. 2005, 51, 507–523. [Google Scholar] [CrossRef] [PubMed]
- Kappell, A.D.; DeNies, M.S.; Ahuja, N.H.; Ledeboer, N.A.; Newton, R.J.; Hristova, K.R. Detection of Multi-Drug Resistant Escherichia Coli in the Urban Waterways of Milwaukee, WI. Front. Microbiol. 2015, 6, 336. [Google Scholar] [CrossRef] [PubMed]
- Bullerjahn, G.S.; McKay, R.M.; Davis, T.W.; Baker, D.B.; Boyer, G.L.; D’Anglada, L.V.; Doucette, G.J.; Ho, J.C.; Irwin, E.G.; Kling, C.L.; et al. Global Solutions to Regional Problems: Collecting Global Expertise to Address the Problem of Harmful Cyanobacterial Blooms. A Lake Erie Case Study. Harmful Algae 2016, 54, 223–238. [Google Scholar] [CrossRef] [PubMed]
- Scavia, D.; Kalcic, M.; Muenich, R.L.; Read, J.; Aloysius, N.; Bertani, I.; Boles, C.; Confesor, R.; DePinto, J.; Gildow, M.; et al. Multiple Models Guide Strategies for Agricultural Nutrient Reductions. Front. Ecol. Environ. 2017, 15, 126–132. [Google Scholar] [CrossRef]
- Environmental Working Group. Manure from Unregulated Factory Farms Fuels Lake Erie’s Toxic Algae Blooms. AgMag. 2019. Available online: https://www.ewg.org/agmag/2019/03/manure-unregulated-factory-farms-fuels-lake-erie-s-toxic-algae-blooms (accessed on 23 April 2021).
- Environmental Working Group. Explosion of Unregulated Factory Farms in Maumee Watershed Fuels Lake Erie’s Toxic Blooms; Environmental Working Group: Washington, DC, USA, 2019; Available online: https://www.ewg.org/interactive-maps/2019_maumee/map/ (accessed on 23 April 2021).
- Bauer, A.W.; Kirby, W.M.; Sherris, J.C.; Turck, M. Antibiotic Susceptibility Testing by a Standardized Single Disk Method. Am. J. Clin. Pathol. 1966, 45, 493–496. [Google Scholar] [CrossRef]
- Schneider, C.A.; Rasband, W.S.; Eliceiri, K.W. NIH Image to ImageJ: 25 Years of Image Analysis. Nat. Methods 2012, 9, 671–675. [Google Scholar] [CrossRef]
- Laird, E.D. Characterization of Antibiotic Resistance Profiles of Surface Water Bacteria in an Urbanizing Watershed. Ph.D. Thesis, Texas A&M University, College Station, TX, USA, 2016. [Google Scholar]
- Janezic, K.J.; Ferry, B.; Hendricks, E.W.; Janiga, B.A.; Johnson, T.; Murphy, S.; Roberts, M.E.; Scott, S.M.; Theisen, A.N.; Hung, K.F.; et al. Phenotypic and Genotypic Characterization of Escherichia Coli Isolated from Untreated Surface Waters. Open Microbiol. J. 2013, 7, 9–19. [Google Scholar] [CrossRef]
- Sullivan, B.A.; Gentry, T.; Karthikeyan, R. Characterization of Tetracycline-Resistant Bacteria in an Urbanizing Subtropical Watershed. J. Appl. Microbiol. 2013, 115, 774–785. [Google Scholar] [CrossRef]
- Guyomard-Rabenirina, S.; Dartron, C.; Falord, M.; Sadikalay, S.; Ducat, C.; Richard, V.; Breurec, S.; Gros, O.; Talarmin, A. Resistance to Antimicrobial Drugs in Different Surface Waters and Wastewaters of Guadeloupe. PLoS ONE 2017, 12, e0173155. [Google Scholar] [CrossRef]
- Ash, R.J.; Mauck, B.; Morgan, M. Antibiotic Resistance of Gram-Negative Bacteria in Rivers, United States. Emerg. Infect Dis. 2002, 8, 713–715. [Google Scholar] [CrossRef] [PubMed]
- Nicolau, D.P.; Carmeli, Y.; Crank, C.W.; Goff, D.A.; Graber, C.J.; Lima, A.L.L.; Goldstein, E.J.C. Carbapenem Stewardship: Does Ertapenem Affect Pseudomonas Susceptibility to Other Carbapenems? A Review of the Evidence. Int. J. Antimicrob. Agents 2012, 39, 11–15. [Google Scholar] [CrossRef] [PubMed]
- Gupta, N.; Limbago, B.M.; Patel, J.B.; Kallen, A.J. Carbapenem-Resistant Enterobacteriaceae: Epidemiology and Prevention. Clin. Infect. Dis. 2011, 53, 60–67. [Google Scholar] [CrossRef] [PubMed]
- Kittinger, C.; Lipp, M.; Folli, B.; Kirschner, A.; Baumert, R.; Galler, H.; Grisold, A.J.; Luxner, J.; Weissenbacher, M.; Farnleitner, A.H.; et al. Enterobacteriaceae Isolated from the River Danube: Antibiotic Resistances, with a Focus on the Presence of ESBL and Carbapenemases. PLoS ONE 2016, 11, 1–17. [Google Scholar]
- Roe, M.T.; Vega, E.; Pillai, S.D. Antimicrobial Resistance Markers of Class 1 and Class 2 Integron-Bearing Escherichia Coli from Irrigation Water and Sediments. Emerg. Infect. Dis. 2003, 9, 822–826. [Google Scholar] [CrossRef]
- Bouki, C.; Venieri, D.; Diamadopoulos, E. Detection and Fate of Antibiotic Resistant Bacteria in Wastewater Treatment Plants: A Review. Ecotoxicol. Environ. Saf. 2013, 91, 1–9. [Google Scholar] [CrossRef]
- Chu, B.T.T.; Petrovich, M.L.; Chaudhary, A.; Wright, D.; Murphy, B.; Wells, G.; Poretsky, R. Metagenomics Reveals the Impact of Wastewater Treatment Plants on the Dispersal of Microorganisms and Genes in Aquatic Sediments. Appl. Environ. Microbiol. 2018, 84. [Google Scholar] [CrossRef]
- Iwane, T.; Urase, T.; Yamamoto, K. Possible Impact of Treated Wastewater Discharge on Incidence of Antibiotic Resistant Bacteria in River Water. Water Sci. Technol. 2001, 43, 91–99. [Google Scholar] [CrossRef]
- Jindal, A.; Kocherginskaya, S.; Mehboob, A.; Robert, M.; Mackie, R.I.; Raskin, L.; Zilles, J.L. Antimicrobial Use and Resistance in Swine Waste Treatment Systems. Appl. Environ. Microbiol. 2006, 72, 7813–7820. [Google Scholar] [CrossRef]
- Korzeniewska, E.; Korzeniewska, A.; Harnisz, M. Antibiotic Resistant Escherichia coli in Hospital and Municipal Sewage and Their Emission to the Environment. Ecotoxicol. Environ. Saf. 2013, 91, 96–102. [Google Scholar] [CrossRef] [PubMed]

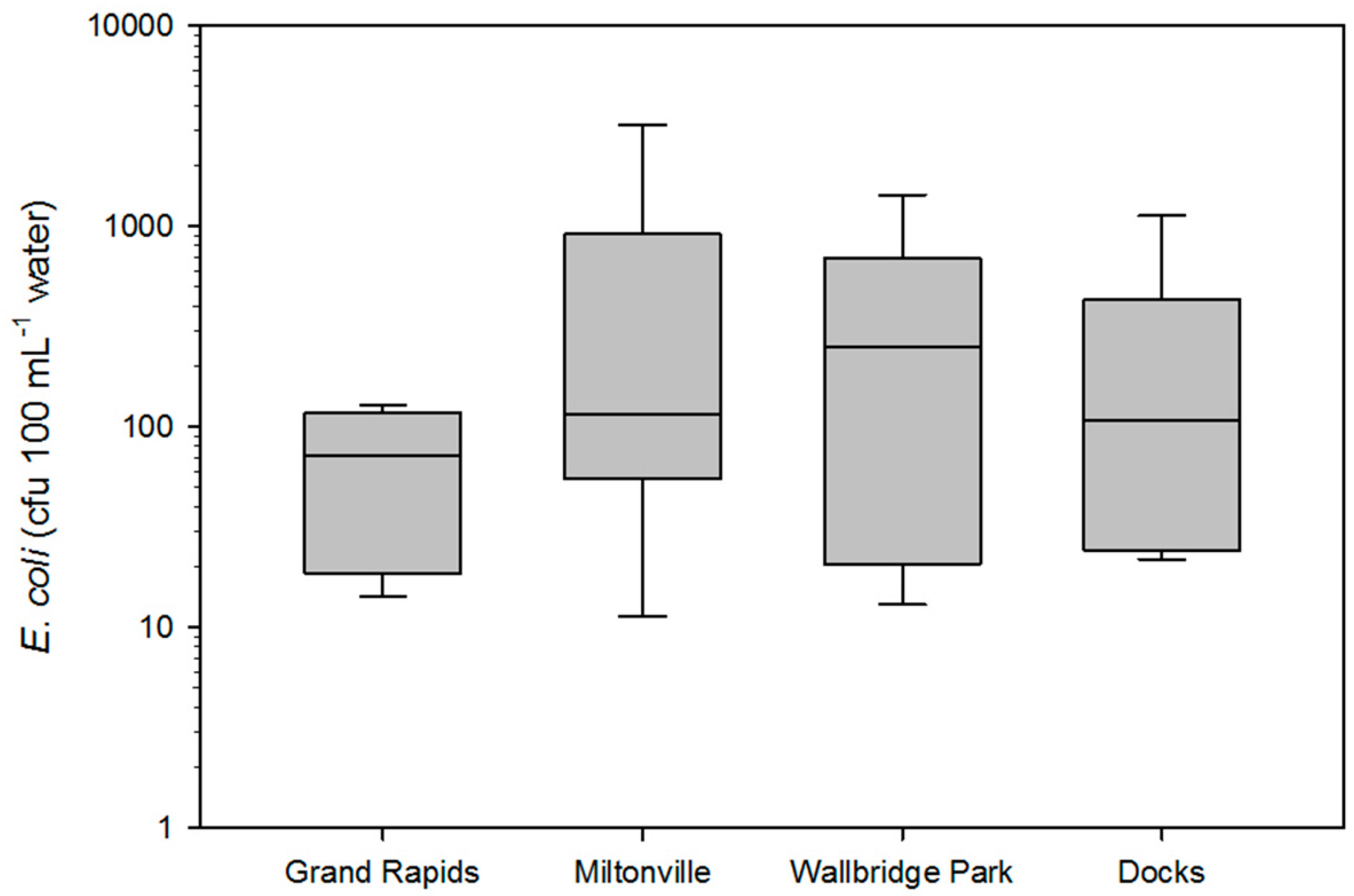
| Location ID | October-16 | November-16 | December-16 | May-17 | June-17 | July-17 | Mean |
|---|---|---|---|---|---|---|---|
| Grand Rapids | 20 | 14.3 | 113 | 76.2 | 68.8 | 128.8 | 60.7 |
| Miltonville | 3230 | 11.4 | 140 | 90.9 | 69.2 | 138.9 | 526.1 |
| Walbridge Park | 1440 | 257 | 247 | 140.9 | 23 | 446.6 | 425.7 |
| Docks | 1130 | 56 | 160 | 25.1 | 21.8 | 196.4 | 264.9 |
| Antibiotic | Grand Rapids (n = 69) | Miltonville (n = 78) | Walbridge Park (n = 97) | Docks (n = 85) | Total (n = 329) |
|---|---|---|---|---|---|
| Tetracycline | 6 (8.7%) | 5 (6.4%) | 12 (12.3%) | 6 (7%) | 29 (8.81%) |
| Imipenem | 0 (0%) | 0 (0%) | 0 (0%) | 1 (1.2%) | 1 (0.3%) |
| Cephalothin | 59 (85.5%) | 72 (92.3%) | 92 (94.8%) | 81 (95.3%) | 309 (95.3%) |
| Gentamicin | 2 (3%) | 7 (9%) | 9 (9.3%) | 9 (10.6%) | 27 (8.2%) |
| Sulfamethoxazole | 1 (1.4%) | 1 (1.3%) | 0 (0%) | 3 (3.5%) | 5 (1.5%) |
| Ciprofloxacin | 2 (2.9%) | 2 (2.5%) | 6 (6.2%) | 4 (4.7%) | 14 (4.2%) |
| Cefoperazone | 3 (4.3%) | 2 (2.5%) | 2 (2%) | 6 (7%) | 13 (4%) |
| Ampicillin | 17 (24.6%) | 27 (34%) | 42 (43.3%) | 40 (47%) | 126 (38.3%) |
| MDR Categories | Grand Rapids (n = 69) | Miltonville (n = 78) | Walbridge Park (n = 97) | Docks (n = 85) | Total (n = 329) |
|---|---|---|---|---|---|
| ≥1 antibiotic | 62 (89.8%) | 75 (96%) | 94 (96.9%) | 82 (96.5%) | 313 (95%) |
| ≥2 antibiotic | 23 (33.3%) | 32 (41%) | 43 (44.3%) | 46 (54.1%) | 144 (43%) |
| ≥3 antibiotic | 7 (10%) | 6 (7.7%) | 23 (23.7%) | 14 (16.5%) | 50 (15%) |
| ≥4 antibiotic | 2 (2.9%) | 2 (2.5%) | 6 (6.2%) | 6 (7%) | 16 (4.9%) |
| ≥5 antibiotic | 1 (1.4%) | 1 (1.3%) | 0 (0%) | 2 (2.3%) | 4 (1.2%) |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mukherjee, M.; Marie, L.; Liles, C.; Mustafa, N.; Bullerjahn, G.; Gentry, T.J.; Brooks, J.P. Elevated Incidences of Antimicrobial Resistance and Multidrug Resistance in the Maumee River (Ohio, USA), a Major Tributary of Lake Erie. Microorganisms 2021, 9, 911. https://doi.org/10.3390/microorganisms9050911
Mukherjee M, Marie L, Liles C, Mustafa N, Bullerjahn G, Gentry TJ, Brooks JP. Elevated Incidences of Antimicrobial Resistance and Multidrug Resistance in the Maumee River (Ohio, USA), a Major Tributary of Lake Erie. Microorganisms. 2021; 9(5):911. https://doi.org/10.3390/microorganisms9050911
Chicago/Turabian StyleMukherjee, Maitreyee, Leah Marie, Cheyenne Liles, Nadia Mustafa, George Bullerjahn, Terry J. Gentry, and John P. Brooks. 2021. "Elevated Incidences of Antimicrobial Resistance and Multidrug Resistance in the Maumee River (Ohio, USA), a Major Tributary of Lake Erie" Microorganisms 9, no. 5: 911. https://doi.org/10.3390/microorganisms9050911
APA StyleMukherjee, M., Marie, L., Liles, C., Mustafa, N., Bullerjahn, G., Gentry, T. J., & Brooks, J. P. (2021). Elevated Incidences of Antimicrobial Resistance and Multidrug Resistance in the Maumee River (Ohio, USA), a Major Tributary of Lake Erie. Microorganisms, 9(5), 911. https://doi.org/10.3390/microorganisms9050911








