The Influence of Diet and Sex on the Gut Microbiota of Lean and Obese JCR:LA-cp Rats
Abstract
1. Introduction
2. Materials and Methods
2.1. Animal Model and Experimental Protocol
2.2. Biological Sample Collection
2.3. Illumina MiSeq Sequencing of 16S rRNA Gene V4 Region of Gut Microbiota
2.4. Quality Control
2.5. Short Chain Fatty Acid Measurement
2.6. Sequence Metrics and Taxonomic Composition
2.7. Statistical Analysis
3. Results
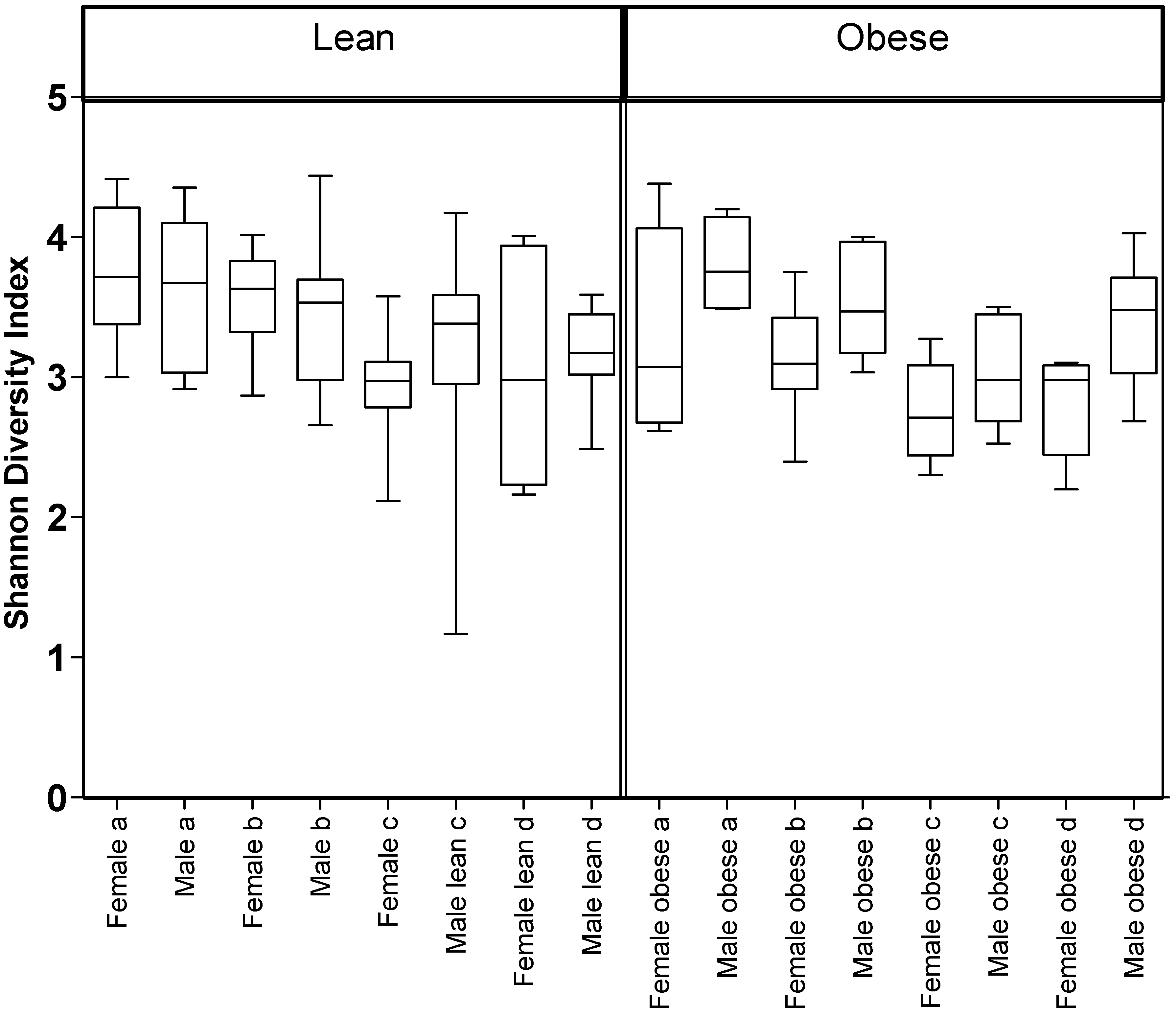
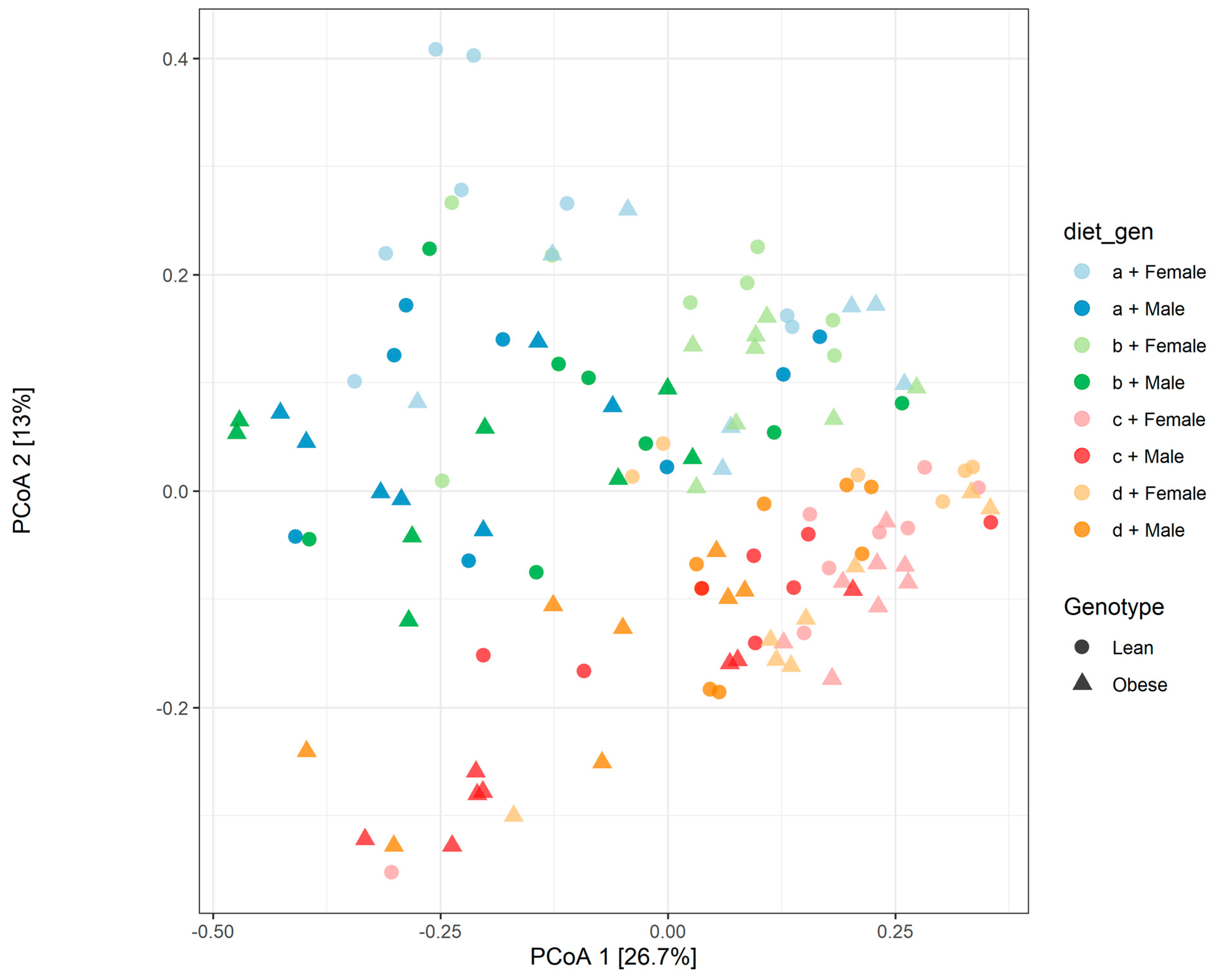
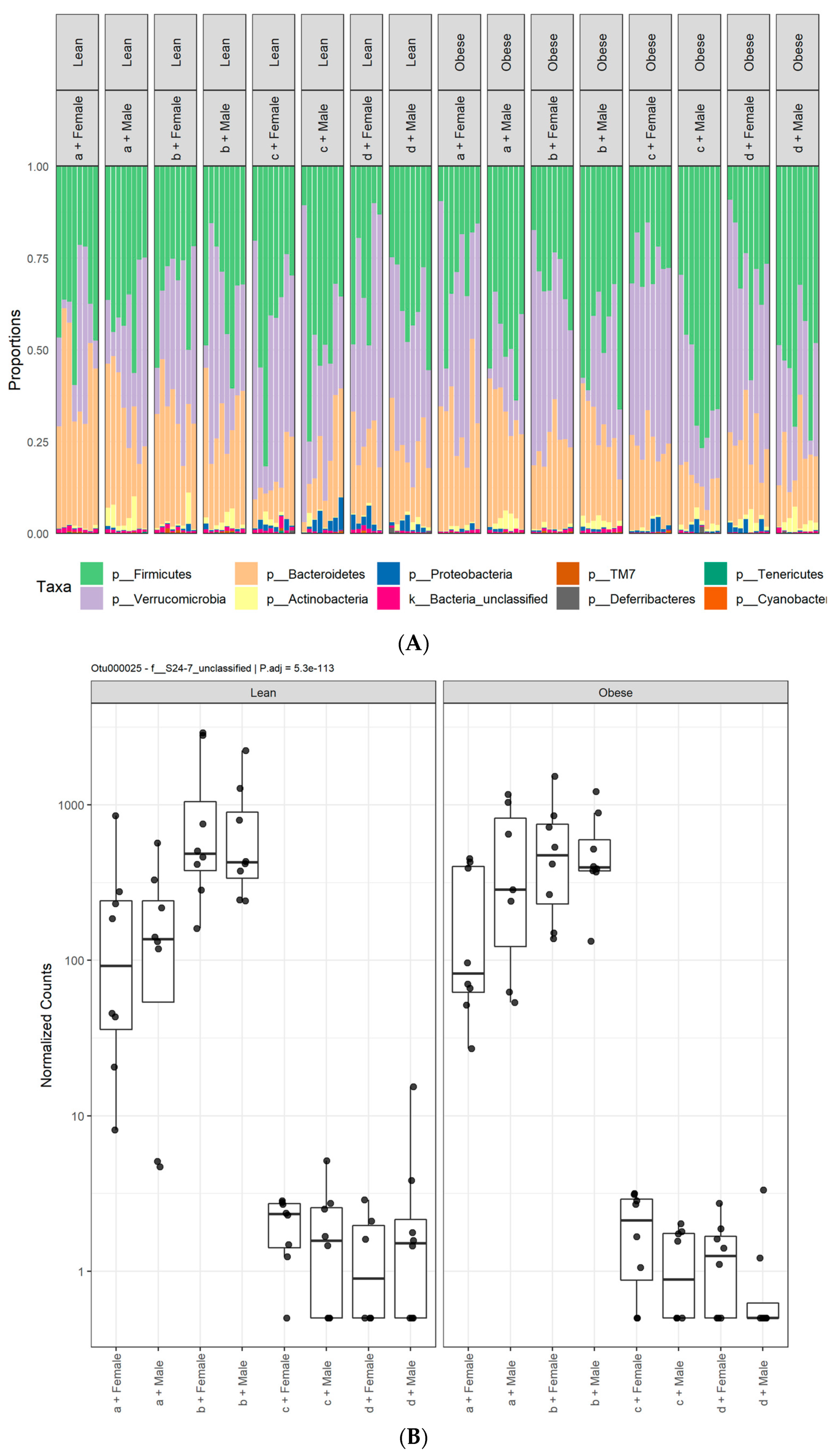
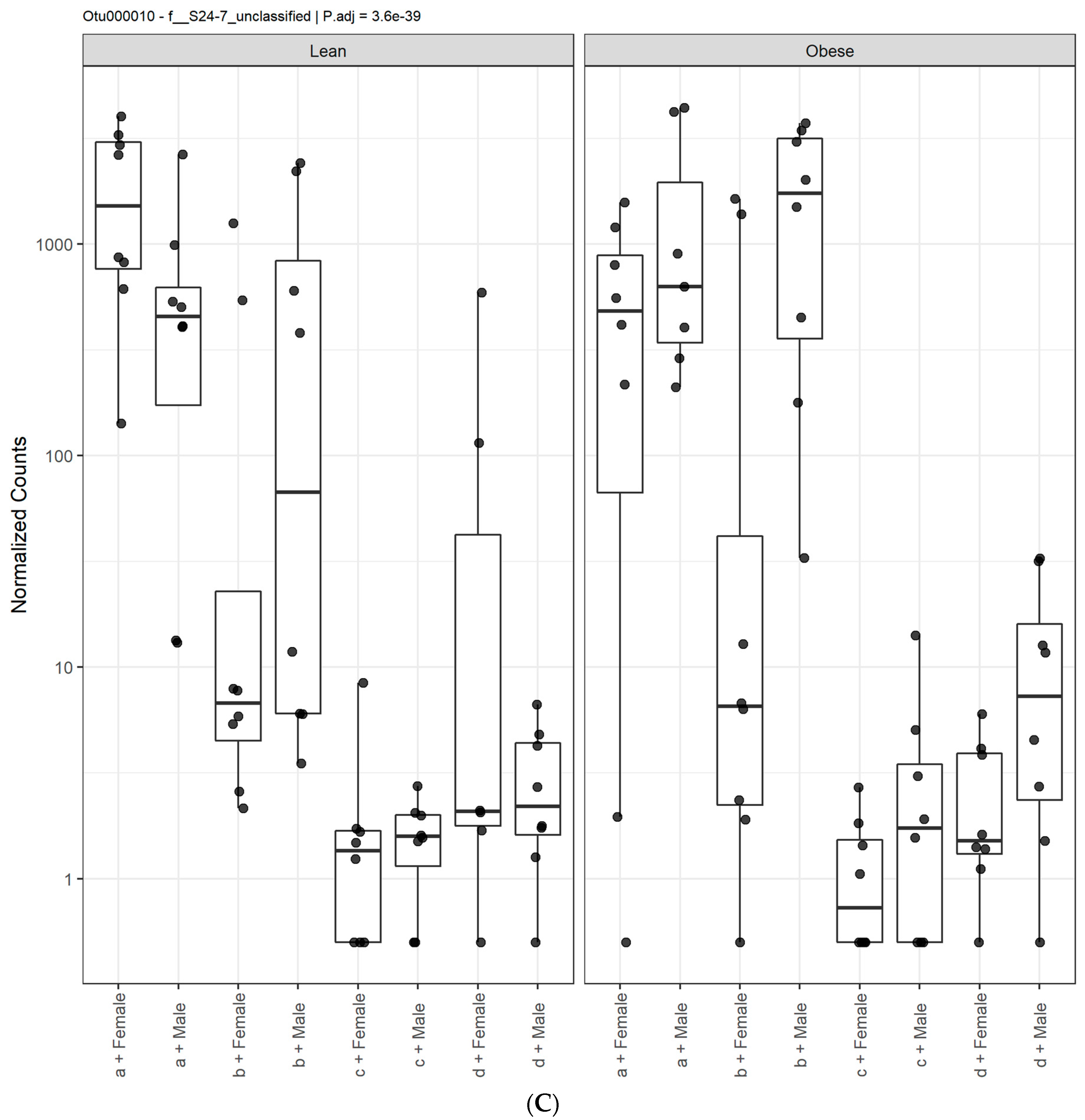
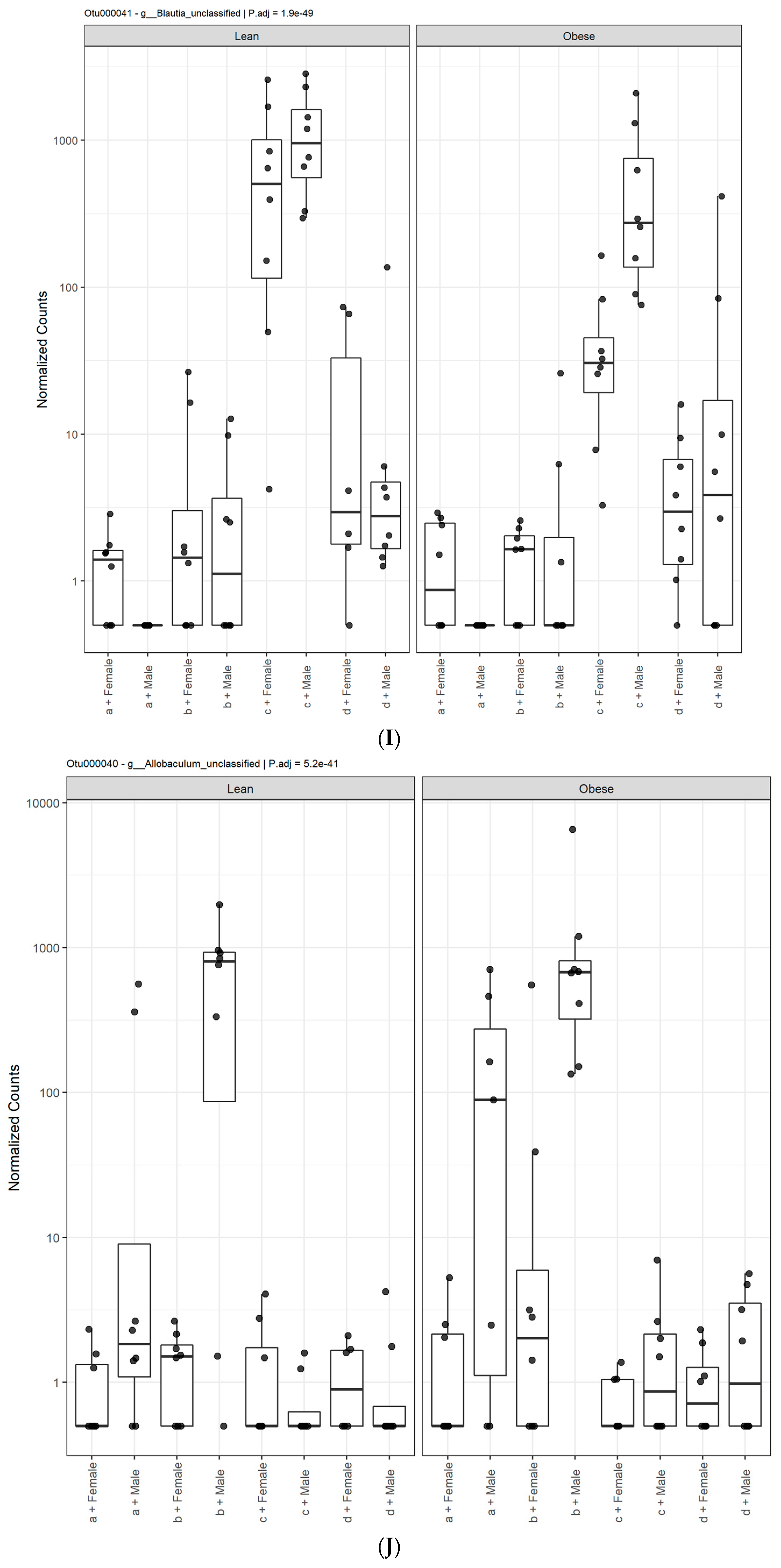
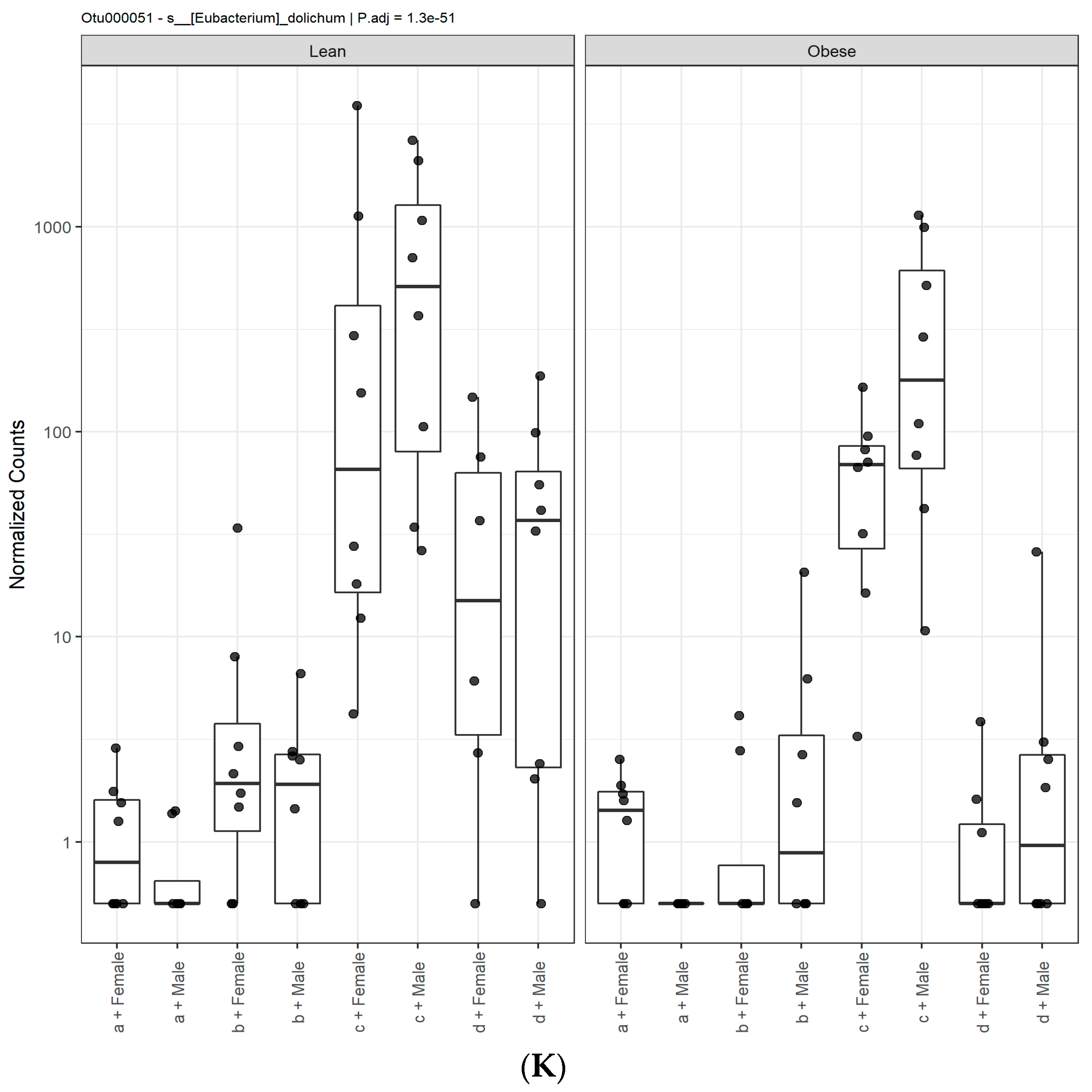
3.1. Diet, Sex, and Genotype Alters the Composition and Diversity of Gut Microbial Ecology
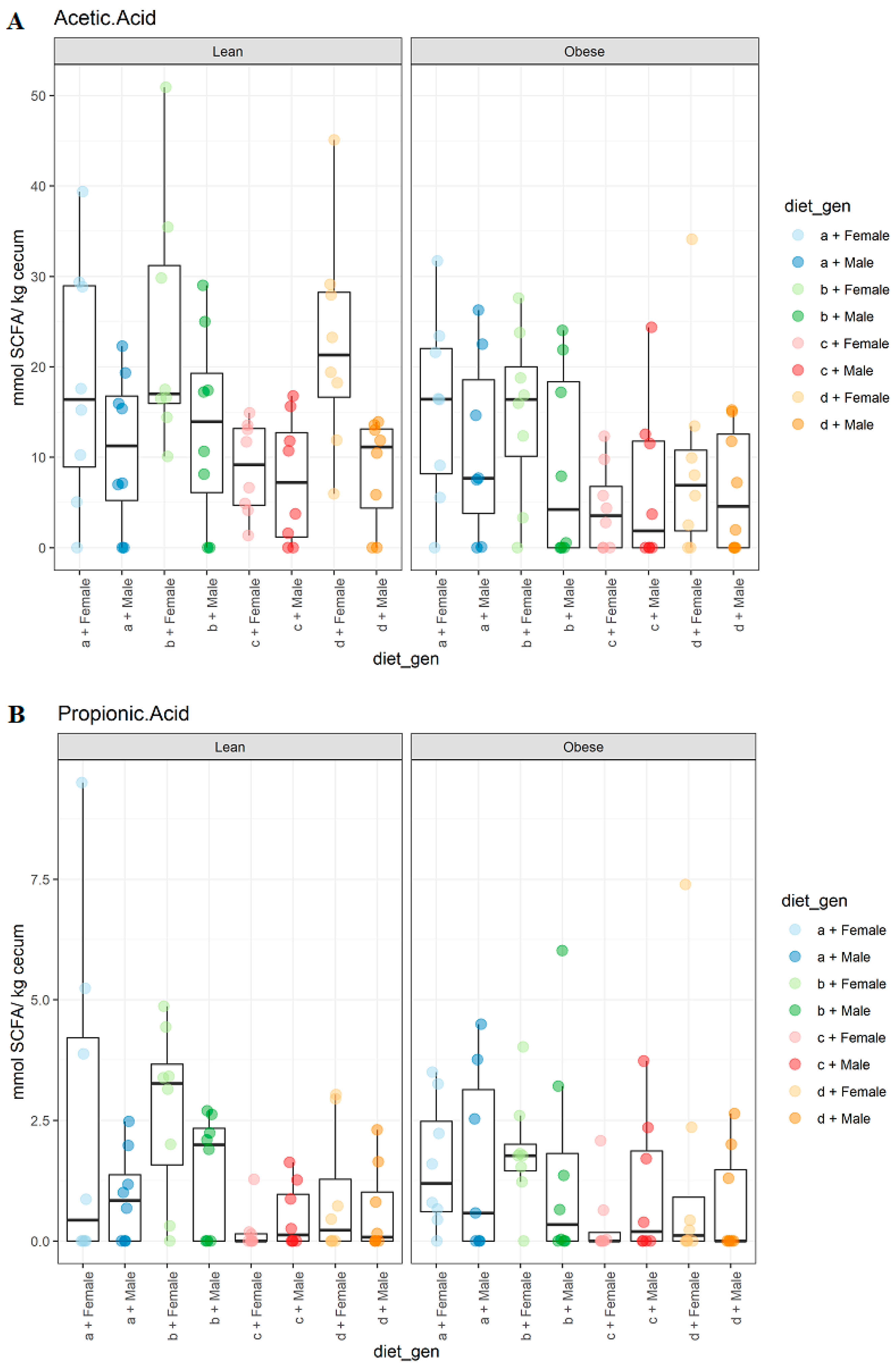
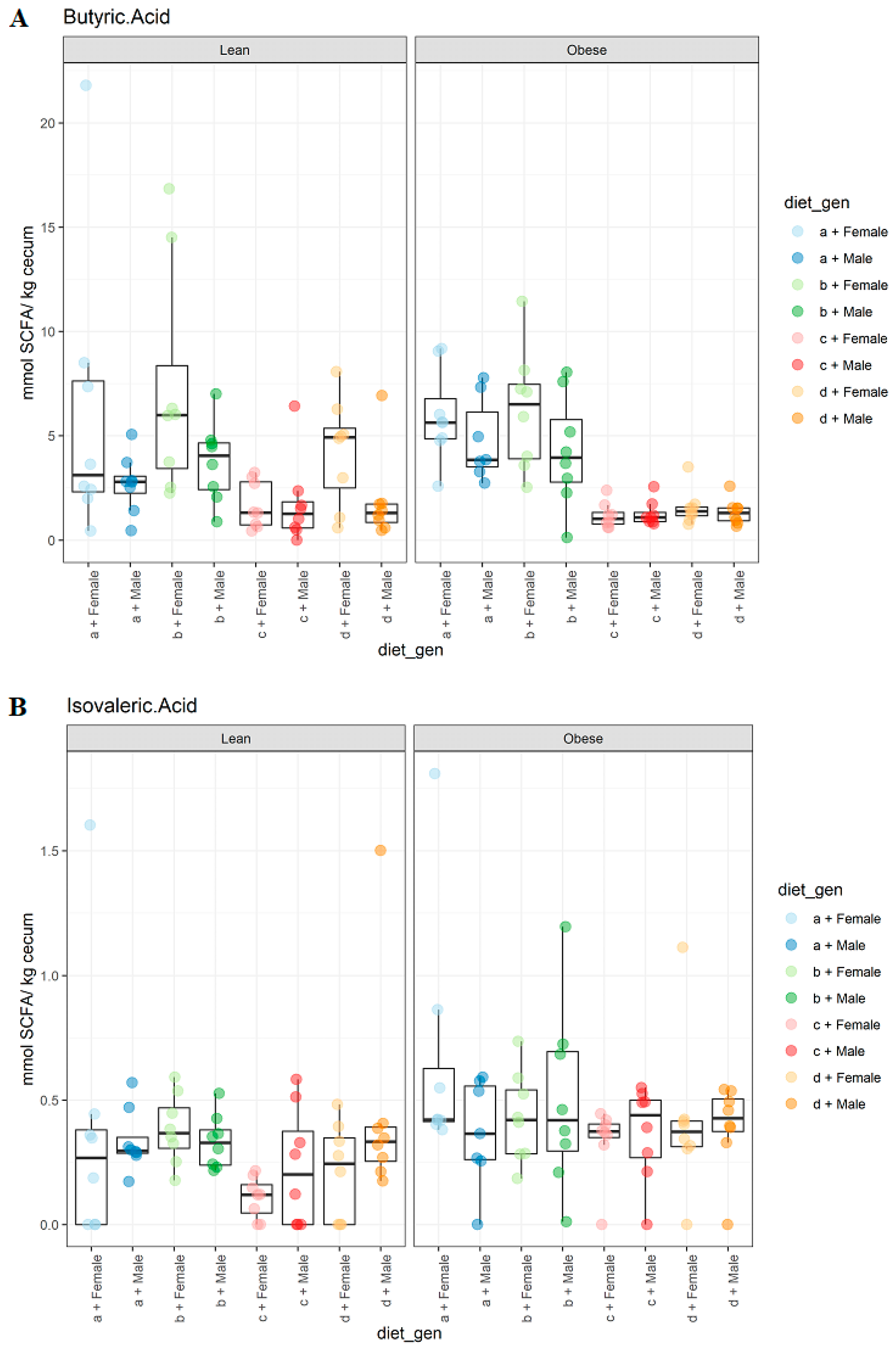
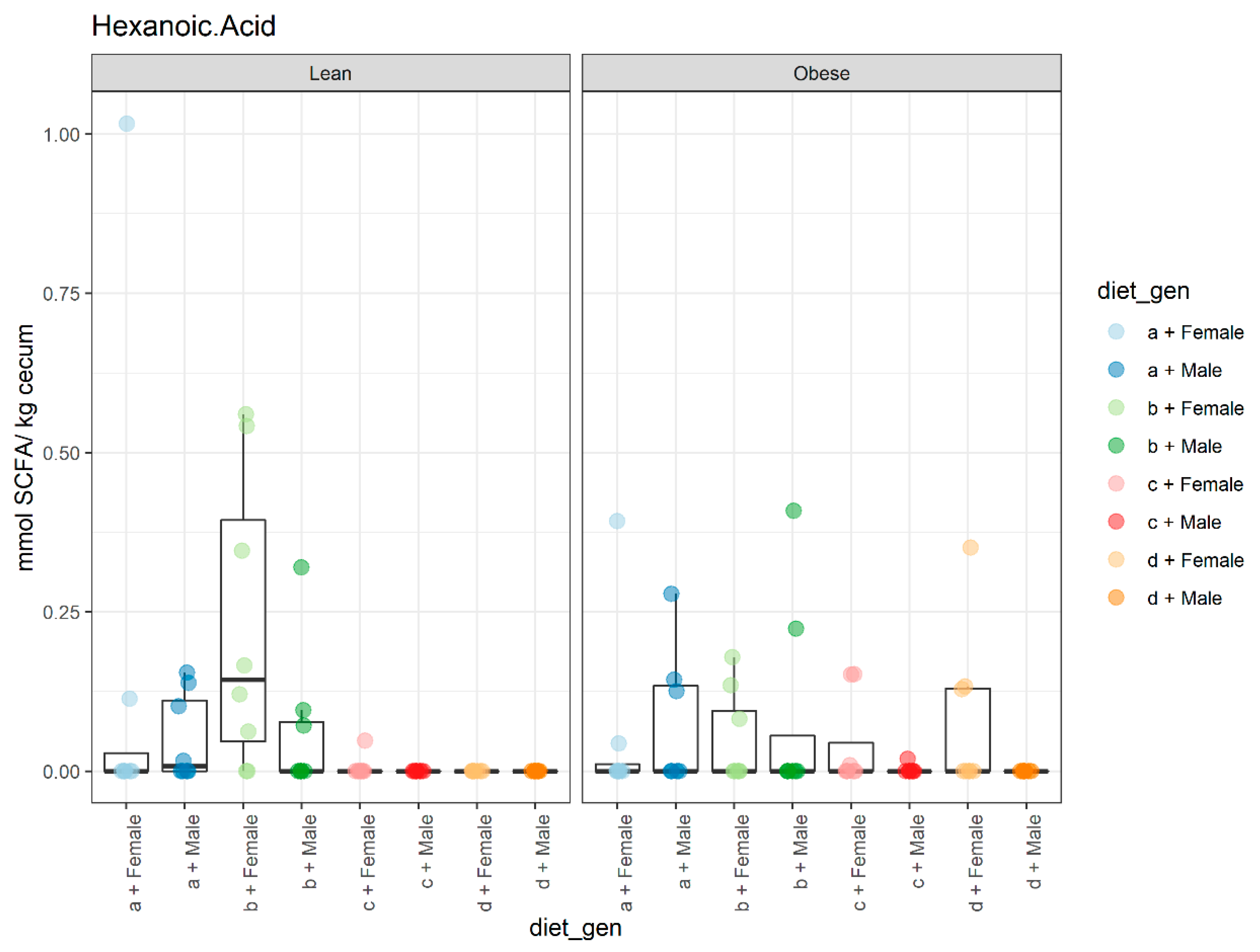
3.2. Diet, Sex, and Genotype Impacts the Gut SCFA Composition
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Lynch, S.V.; Pedersen, O. The human intestinal microbiome in health and disease. N. Engl. J. Med. 2016, 375, 2369–2379. [Google Scholar] [CrossRef] [PubMed]
- Power, S.E.; O’Toole, P.W.; Stanton, C.; Ross, R.P.; Fitzgerald, G.F. Intestinal microbiota, diet and health. Br. J. Nutr. 2014, 111, 387–402. [Google Scholar] [CrossRef] [PubMed]
- Turner, J.R. Intestinal mucosal barrier function in health and disease. Nat. Rev. Immunol. 2009, 9, 799–809. [Google Scholar] [CrossRef] [PubMed]
- Turnbaugh, P.J.; Ley, R.E.; Mahowald, M.A.; Magrini, V.; Mardis, E.R.; Gordon, J.I. An obesity-associated gut microbiome with increased capacity for energy harvest. Nature 2006, 444, 1027–1031. [Google Scholar] [CrossRef] [PubMed]
- Vandeputte, D.; Falony, G.; Vieira-Silva, S.; Tito, R.Y.; Joossens, M.; Raes, J. Stool consistency is strongly associated with gut microbiota richness and composition, enterotypes and bacterial growth rates. Gut 2016, 65, 57–62. [Google Scholar] [CrossRef] [PubMed]
- Hakkak, R.; Korourian, S.; Foley, S.L.; Erickson, B.D. Assessment of gut microbiota populations in lean and obese zucker rats. PLoS ONE 2017, 12, e0181451. [Google Scholar] [CrossRef]
- Turnbaugh, P.J.; Hamady, M.; Yatsunenko, T.; Cantarel, B.L.; Duncan, A.; Ley, R.E.; Sogin, M.L.; Jones, W.J.; Roe, B.A.; Affourtit, J.P.; et al. A core gut microbiome in obese and lean twins. Nature 2009, 457, 480–484. [Google Scholar] [CrossRef]
- Ley, R.E.; Turnbaugh, P.J.; Klein, S.; Gordon, J.I. Human gut microbes associated with obesity. Nature 2006, 444, 1022–1023. [Google Scholar] [CrossRef]
- Lee, C.J.; Sears, C.L.; Maruthur, N. Gut microbiome and its role in obesity and insulin resistance. Ann. N. Y. Acad. Sci. 2020, 1461, 37–52. [Google Scholar] [CrossRef]
- Le Chatelier, E.; Nielsen, T.; Qin, J.; Prifti, E.; Hildebrand, F.; Falony, G.; Almeida, M.; Arumugam, M.; Batto, J.M.; Kennedy, S.; et al. Richness of human gut microbiome correlates with metabolic markers. Nature 2013, 500, 541–546. [Google Scholar] [CrossRef]
- Zhao, L.; Zhang, Q.; Ma, W.; Tian, F.; Shen, H.; Zhou, M. A combination of quercetin and resveratrol reduces obesity in high-fat diet-fed rats by modulation of gut microbiota. J. Food Funct. 2017, 8, 4644–4656. [Google Scholar] [CrossRef] [PubMed]
- Xiao, S.; Fei, N.; Pang, X.; Shen, J.; Wang, L.; Zhang, B.; Zhang, M.; Zhang, X.; Zhang, C.; Li, M.; et al. A gut microbiota-targeted dietary intervention for amelioration of chronic inflammation underlying metabolic syndrome. FEMS Microbiol. Ecol. 2014, 87, 357–367. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Tang, H.; Zhang, C.; Zhao, Y.; Derrien, M.; Rocher, E.; van-Hylckama Vlieg, J.E.T.; Strissel, K.; Zhao, L.; Obin, M.; et al. Modulation of gut microbiota during probiotic-mediated attenuation of metabolic syndrome in high fat diet-fed mice. ISME J. 2015, 9, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Delzenne, N.M.; Neyrinck, A.M.; Bäckhed, F.; Cani, P.D. Targeting gut microbiota in obesity: Effects of prebiotics and probiotics. Nat. Rev. Endocrinol. 2011, 7, 639–646. [Google Scholar] [CrossRef]
- Shin, N.R.; Lee, J.C.; Lee, H.Y.; Kim, M.S.; Whon, T.W.; Lee, M.S.; Bae, J.W. An increase in the akkermansia spp. Population induced by metformin treatment improves glucose homeostasis in diet-induced obese mice. Gut 2014, 63, 727–735. [Google Scholar] [CrossRef]
- Hildebrandt, M.A.; Hoffmann, C.; Sherrill-Mix, S.A.; Keilbaugh, S.A.; Hamady, M.; Chen, Y.Y.; Knight, R.; Ahima, R.S.; Bushman, F.; Wu, G.D. High-fat diet determines the composition of the murine gut microbiome independently of obesity. J. Gastroenterol 2009, 137, 1641–1648. [Google Scholar] [CrossRef]
- David, L.A.; Maurice, C.F.; Carmody, R.N.; Gootenberg, D.B.; Button, J.E.; Wolfe, B.E.; Ling, A.V.; Devlin, A.S.; Varma, Y.; Fischbach, M.A.; et al. Diet rapidly and reproducibly alters the human gut microbiome. Nature 2014, 505, 559–563. [Google Scholar] [CrossRef]
- De La Serre, C.B.; Ellis, C.L.; Lee, J.; Hartman, A.L.; Rutledge, J.C.; Raybould, H.E. Propensity to high-fat diet-induced obesity in rats is associated with changes in the gut microbiota and gut inflammation. Am. J. Physiol. Gastrointest Liver Physiol. 2010, 299, G440–G448. [Google Scholar] [CrossRef]
- Turnbaugh, P.J.; Bäckhed, F.; Fulton, L.; Gordon, J.I. Diet-induced obesity is linked to marked but reversible alterations in the mouse distal gut microbiome. Cell Host Microbe 2008, 3, 213–223. [Google Scholar] [CrossRef]
- Wang, J.; Jia, H. Metagenome-wide association studies: Fine-mining the microbiome. Nat. Rev. Microbiol. 2016, 14, 508–522. [Google Scholar] [CrossRef]
- Parks, B.W.; Nam, E.; Org, E.; Kostem, E.; Norheim, F.; Hui, S.T.; Pan, C.; Civelek, M.; Rau, C.D.; Bennett, B.J.; et al. Genetic control of obesity and gut microbiota composition in response to high-fat, high-sucrose diet in mice. Cell Metab. 2013, 17, 141–152. [Google Scholar] [CrossRef] [PubMed]
- Carmody, R.N.; Gerber, G.K.; Luevano, J.M., Jr.; Gatti, D.M.; Somes, L.; Svenson, K.L.; Turnbaugh, P.J. Diet dominates host genotype in shaping the murine gut microbiota. Cell Host Microbe 2015, 17, 72–84. [Google Scholar] [CrossRef]
- Parikh, M.; Netticadan, T.; Pierce, G.N. Flaxseed: Its bioactive components and their cardiovascular benefits. Am. J. Physiol. Heart Circ. Physiol. 2017, 314, H146–H159. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Wang, H.; Yin, P.; Fan, H.; Sun, L.; Liu, Y. Flaxseed oil ameliorates alcoholic liver disease via anti-inflammation and modulating gut microbiota in mice. Lipids Health Dis. 2017, 16, 44. [Google Scholar] [CrossRef] [PubMed]
- Gomides, A.F.; Gonçalves, R.V.; de Paula, S.O.; Ferreira, C.L.; Comastri, D.S.; Peluzio Mdo, C. Defatted flaxseed meal prevents the appearance of aberrant crypt foci in the colon of mice increasing the gene expression of p53. Nutr. Hosp. 2015, 31, 1675–1681. [Google Scholar] [CrossRef] [PubMed]
- Brahe, L.K.; Le Chatelier, E.; Prifti, E.; Pons, N.; Kennedy, S.; Blædel, T.; Håkansson, J.; Dalsgaard, T.K.; Hansen, T.; Pedersen, O.; et al. Dietary modulation of the gut microbiota-a randomised controlled trial in obese postmenopausal women. Br. J. Nutr. 2015, 114, 406–417. [Google Scholar] [CrossRef]
- Parikh, M.; Maddaford, T.G.; Austria, J.A.; Aliani, M.; Netticadan, T.; Pierce, G.N. Dietary flaxseed as a strategy for improving human health. Nutrients 2019, 11, 1171. [Google Scholar] [CrossRef]
- Edel, A.L.; Aliani, M.; Pierce, G.N. Supported liquid extraction in the quantitation of plasma enterolignans using isotope dilution GC?MS with application to flaxseed consumption in healthy adults. J. Chromatog. B Analyt. Technol. Biomed. Life Sci. 2013, 912, 24–32. [Google Scholar] [CrossRef] [PubMed]
- Pulkrabek, M.; Rhee, Y.; Gibbs, P.; Hall, C. Flaxseed- and buckwheat-supplemented diets altered enterobacteriaceae diversity and prevalence in the cecum and feces of obese mice. J. Diet Suppl. 2017, 14, 667–678. [Google Scholar] [CrossRef]
- Power, K.A.; Lepp, D.; Zarepoor, L.; Monk, J.M.; Wu, W.; Tsao, R.; Liu, R. Dietary flaxseed modulates the colonic microenvironment in healthy c57bl/6 male mice which may alter susceptibility to gut-associated diseases. J. Nutr. Biochem. 2016, 28, 61–69. [Google Scholar] [CrossRef]
- Patterson, E.; RM, O.D.; Murphy, E.F.; Wall, R.; O’Sullivan, O.S.; Nilaweera, K.; Fitzgerald, G.F.; Cotter, P.D.; Ross, R.P.; Stanton, C. Impact of dietary fatty acids on metabolic activity and host intestinal microbiota composition in c57bl/6j mice. Br. J. Nutr. 2014, 111, 1905–1917. [Google Scholar] [CrossRef] [PubMed]
- Diane, A.; Pierce, W.D.; Kelly, S.E.; Sokolik, S.; Borthwick, F.; Jacome-Sosa, M.; Mangat, R.; Pradillo, J.M.; Allan, S.M.; Ruth, M.R.; et al. Mechanisms of comorbidities associated with the metabolic syndrome: Insights from the JCR:LA-cp corpulent rat strain. Front Nutr. 2016, 3. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Russell, J.C.; Koeslag, D.G. Jcr:La-corpulent rat: A strain with spontaneous vascular and myocardial disease. ILAR J. 1990, 32, 27–32. [Google Scholar] [CrossRef]
- Russell, J.C.; Proctor, S.D. Small animal models of cardiovascular disease: Tools for the study of the roles of metabolic syndrome, dyslipidemia, and atherosclerosis. Cardiovasc. Pathol. 2006, 15, 318–330. [Google Scholar] [CrossRef] [PubMed]
- Maddaford, T.G.; Russell, J.C.; Pierce, G.N. Post-ischemic cardiac performance in the insulin-resistant JCR:LA-cp rat. Am. J. Physiol. 1997, 273, H1187–H1192. [Google Scholar] [CrossRef] [PubMed]
- Kozich, J.J.; Westcott, S.L.; Baxter, N.T.; Highlander, S.K.; Schloss, P.D. Development of a dual-index sequencing strategy and curation pipeline for analyzing amplicon sequence data on the miseq illumina sequencing platform. Appl. Environ. Microbiol. 2013, 79, 5112–5120. [Google Scholar] [CrossRef] [PubMed]
- Schloss, P.D.; Westcott, S.L.; Ryabin, T.; Hall, J.R.; Hartmann, M.; Hollister, E.B.; Lesniewski, R.A.; Oakley, B.B.; Parks, D.H.; Robinson, C.J.; et al. Introducing mothur: Open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl. Environ. Microbiol. 2009, 75, 7537–7541. [Google Scholar] [CrossRef]
- Zhao, G.; Nyman, M.; Jönsson, J.A. Rapid determination of short-chain fatty acids in colonic contents and faeces of humans and rats by acidified water-extraction and direct-injection gas chromatography. Biomed. Chromatog. BMC 2006, 20, 674–682. [Google Scholar] [CrossRef] [PubMed]
- Ericsson, A.C.; Franklin, C.L. Manipulating the gut microbiota: Methods and challenges. ILAR J. 2015, 56, 205–217. [Google Scholar] [CrossRef]
- Henke, M.T.; Kenny, D.J.; Cassilly, C.D.; Vlamakis, H.; Xavier, R.J.; Clardy, J. Ruminococcus gnavus, a member of the human gut microbiome associated with crohn’s disease, produces an inflammatory polysaccharide. Proc. Natl. Acad. Sci. USA 2019, 116, 12672–12677. [Google Scholar] [CrossRef]
- Xie, G.; Wang, X.; Liu, P.; Wei, R.; Chen, W.; Rajani, C.; Hernandez, B.Y.; Alegado, R.; Dong, B.; Li, D.; et al. Distinctly altered gut microbiota in the progression of liver disease. Oncotarget 2016, 7, 19355–19366. [Google Scholar] [CrossRef]
- Chen, Y.-R.; Zheng, H.-M.; Zhang, G.-X.; Chen, F.-L.; Chen, L.-D.; Yang, Z.-C. High oscillospira abundance indicates constipation and low bmi in the guangdong gut microbiome project. Sci. Rep. 2020, 10, 9364. [Google Scholar] [CrossRef]
- Pessione, E. Lactic acid bacteria contribution to gut microbiota complexity: Lights and shadows. Front Cell Infect Microbiol. 2012, 2, 86. [Google Scholar] [CrossRef]
- Ozato, N.; Saito, S.; Yamaguchi, T.; Katashima, M.; Tokuda, I.; Sawada, K.; Katsuragi, Y.; Kakuta, M.; Imoto, S.; Ihara, K.; et al. Blautia genus associated with visceral fat accumulation in adults 20–76 years of age. NPJ Biofilms Microbiomes. 2019, 5, 28. [Google Scholar] [CrossRef]
- Pallister, T.; Jackson, M.A.; Martin, T.C.; Glastonbury, C.A.; Jennings, A.; Beaumont, M.; Mohney, R.P.; Small, K.S.; MacGregor, A.; Steves, C.J.; et al. Untangling the relationship between diet and visceral fat mass through blood metabolomics and gut microbiome profiling. Int. Obes. 2017, 41, 1106–1113. [Google Scholar] [CrossRef] [PubMed]
- Matsuzawa, Y. Establishment of a concept of visceral fat syndrome and discovery of adiponectin. Proc. Japan Acad. Ser. B Phys. Biol. Sci. 2010, 86, 131–141. [Google Scholar] [CrossRef] [PubMed]
- Ravussin, Y.; Koren, O.; Spor, A.; LeDuc, C.; Gutman, R.; Stombaugh, J.; Knight, R.; Ley, R.E.; Leibel, R.L. Responses of gut microbiota to diet composition and weight loss in lean and obese mice. Obesity 2012, 20, 738–747. [Google Scholar] [CrossRef] [PubMed]
- Han, K.; Jin, W.; Mao, Z.; Dong, S.; Zhang, Q.; Yang, Y.; Chen, B.; Wu, H.; Zeng, M. Microbiome and butyrate production are altered in the gut of rats fed a glycated fish protein diet. J. Funct. Foods 2018, 47, 423–433. [Google Scholar] [CrossRef]
- Tachon, S.; Zhou, J.; Keenan, M.; Martin, R.; Marco, M.L. The intestinal microbiota in aged mice is modulated by dietary resistant starch and correlated with improvements in host responses. FEMS Microbiol. Ecol. 2013, 83, 299–309. [Google Scholar] [CrossRef] [PubMed]
- Leclercq, S.; Matamoros, S.; Cani, P.D.; Neyrinck, A.M.; Jamar, F.; Stärkel, P.; Windey, K.; Tremaroli, V.; Bäckhed, F.; Verbeke, K.; et al. Intestinal permeability, gut-bacterial dysbiosis, and behavioral markers of alcohol-dependence severity. Proc. Natl. Acad. Sci. USA 2014, 111, E4485–E4493. [Google Scholar] [CrossRef]
- Flint, H.J.; Bayer, E.A.; Rincon, M.T.; Lamed, R.; White, B.A. Polysaccharide utilization by gut bacteria: Potential for new insights from genomic analysis. Nat. Rev. Microbiol. 2008, 6, 121–131. [Google Scholar] [CrossRef]
- Riley, L.W.; Raphael, E.; Faerstein, E. Obesity in the united states—Dysbiosis from exposure to low-dose antibiotics? Front Public Health 2013, 1, 69. [Google Scholar] [CrossRef]
- Diamant, M.; Blaak, E.E.; de Vos, W.M. Do nutrient-gut-microbiota interactions play a role in human obesity, insulin resistance and type 2 diabetes? Obes. Rev. 2011, 12, 272–281. [Google Scholar] [CrossRef] [PubMed]
- Priyadarshini, M.; Kotlo, K.U.; Dudeja, P.K.; Layden, B.T. Role of short chain fatty acid receptors in intestinal physiology and pathophysiology. Compr. Physiol. 2018, 8, 1091–1115. [Google Scholar] [CrossRef] [PubMed]
- Arora, T.; Sharma, R.; Frost, G. Propionate. Anti-obesity and satiety enhancing factor? Appetite 2011, 56, 511–515. [Google Scholar] [CrossRef] [PubMed]
- Diraison, F.; Dusserre, E.; Vidal, H.; Sothier, M.; Beylot, M. Increased hepatic lipogenesis but decreased expression of lipogenic gene in adipose tissue in human obesity. Am. J. Physiol. Endocrinol. Metab. 2002, 282, E46–E51. [Google Scholar] [CrossRef]
- Kasubuchi, M.; Hasegawa, S.; Hiramatsu, T.; Ichimura, A.; Kimura, I. Dietary gut microbial metabolites, short-chain fatty acids, and host metabolic regulation. Nutrients 2015, 7, 2839–2849. [Google Scholar] [CrossRef]
- De la Cuesta-Zuluaga, J.; Mueller, N.T.; Alvarez-Quintero, R.; Velasquez-Mejia, E.P.; Sierra, J.A.; Corrales-Agudelo, V.; Carmona, J.A.; Abad, J.M.; Escobar, J.S. Higher fecal short-chain fatty acid levels are associated with gut microbiome dysbiosis, obesity, hypertension and cardiometabolic disease risk factors. Nutrients 2018, 11, 51. [Google Scholar] [CrossRef]
- Holmes, Z.C.; Silverman, J.D.; Dressman, H.K.; Wei, Z.; Dallow, E.P.; Armstrong, S.C.; Seed, P.C.; Rawls, J.F.; David, L.A. Short-chain fatty acid production by gut microbiota from children with obesity differs according to prebiotic choice and bacterial community composition. mBio 2020, 11, e00914-20. [Google Scholar] [CrossRef] [PubMed]
- Morrison, D.J.; Preston, T. Formation of short chain fatty acids by the gut microbiota and their impact on human metabolism. Gut Microbes 2016, 7, 189–200. [Google Scholar] [CrossRef] [PubMed]
- Murphy, E.F.; Cotter, P.D.; Healy, S.; Marques, T.M.; O’Sullivan, O.; Fouhy, F.; Clarke, S.F.; O’Toole, P.W.; Quigley, E.M.; Stanton, C.; et al. Composition and energy harvesting capacity of the gut microbiota: Relationship to diet, obesity and time in mouse models. Gut 2010, 59, 1635–1642. [Google Scholar] [CrossRef] [PubMed]
- Frost, G.; Sleeth, M.L.; Sahuri-Arisoylu, M.; Lizarbe, B.; Cerdan, S.; Brody, L.; Anastasovska, J.; Ghourab, S.; Hankir, M.; Zhang, S.; et al. The short-chain fatty acid acetate reduces appetite via a central homeostatic mechanism. Nat. Commun. 2014, 5, 3611. [Google Scholar] [CrossRef]
- Chambers, E.S.; Viardot, A.; Psichas, A.; Morrison, D.J.; Murphy, K.G.; Zac-Varghese, S.E.; MacDougall, K.; Preston, T.; Tedford, C.; Finlayson, G.S.; et al. Effects of targeted delivery of propionate to the human colon on appetite regulation, body weight maintenance and adiposity in overweight adults. Gut 2015, 64, 1744–1754. [Google Scholar] [CrossRef] [PubMed]
- Cani, P.D.; Bibiloni, R.; Knauf, C.; Waget, A.; Neyrinck, A.M.; Delzenne, N.M.; Burcelin, R. Changes in gut microbiota control metabolic endotoxemia-induced inflammation in high-fat diet-induced obesity and diabetes in mice. Diabetes 2008, 57, 1470–1481. [Google Scholar] [CrossRef] [PubMed]
- Levy, M.; Thaiss, C.A.; Elinav, E. Metabolites: Messengers between the microbiota and the immune system. Genes Dev. 2016, 30, 1589–1597. [Google Scholar] [CrossRef] [PubMed]
- Smith, B.J.; Miller, R.A.; Ericsson, A.C.; Harrison, D.C.; Strong, R.; Schmidt, T.M. Changes in the gut microbiome and fermentation products concurrent with enhanced longevity in acarbose-treated mice. BMC Microbiol. 2019, 19, 130. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Resch, C.; Parikh, M.; Austria, J.A.; Proctor, S.D.; Netticadan, T.; Blewett, H.; Pierce, G.N. The Influence of Diet and Sex on the Gut Microbiota of Lean and Obese JCR:LA-cp Rats. Microorganisms 2021, 9, 1037. https://doi.org/10.3390/microorganisms9051037
Resch C, Parikh M, Austria JA, Proctor SD, Netticadan T, Blewett H, Pierce GN. The Influence of Diet and Sex on the Gut Microbiota of Lean and Obese JCR:LA-cp Rats. Microorganisms. 2021; 9(5):1037. https://doi.org/10.3390/microorganisms9051037
Chicago/Turabian StyleResch, Craig, Mihir Parikh, J. Alejandro Austria, Spencer D. Proctor, Thomas Netticadan, Heather Blewett, and Grant N. Pierce. 2021. "The Influence of Diet and Sex on the Gut Microbiota of Lean and Obese JCR:LA-cp Rats" Microorganisms 9, no. 5: 1037. https://doi.org/10.3390/microorganisms9051037
APA StyleResch, C., Parikh, M., Austria, J. A., Proctor, S. D., Netticadan, T., Blewett, H., & Pierce, G. N. (2021). The Influence of Diet and Sex on the Gut Microbiota of Lean and Obese JCR:LA-cp Rats. Microorganisms, 9(5), 1037. https://doi.org/10.3390/microorganisms9051037





