SARS-CoV-2 IgG Antibodies Seroprevalence and Sera Neutralizing Activity in MEXICO: A National Cross-Sectional Study during 2020
Abstract
1. Introduction
2. Materials and Methods
2.1. Study Design
2.2. Sample Shipping, Treatment, and Storage
2.3. Determination of Neutralizing Antibodies Against the RBD Antigen of the SARS-CoV-2 Virus by ELISA
2.4. Statistical Analysis
2.5. Role of the Funding Source
3. Results
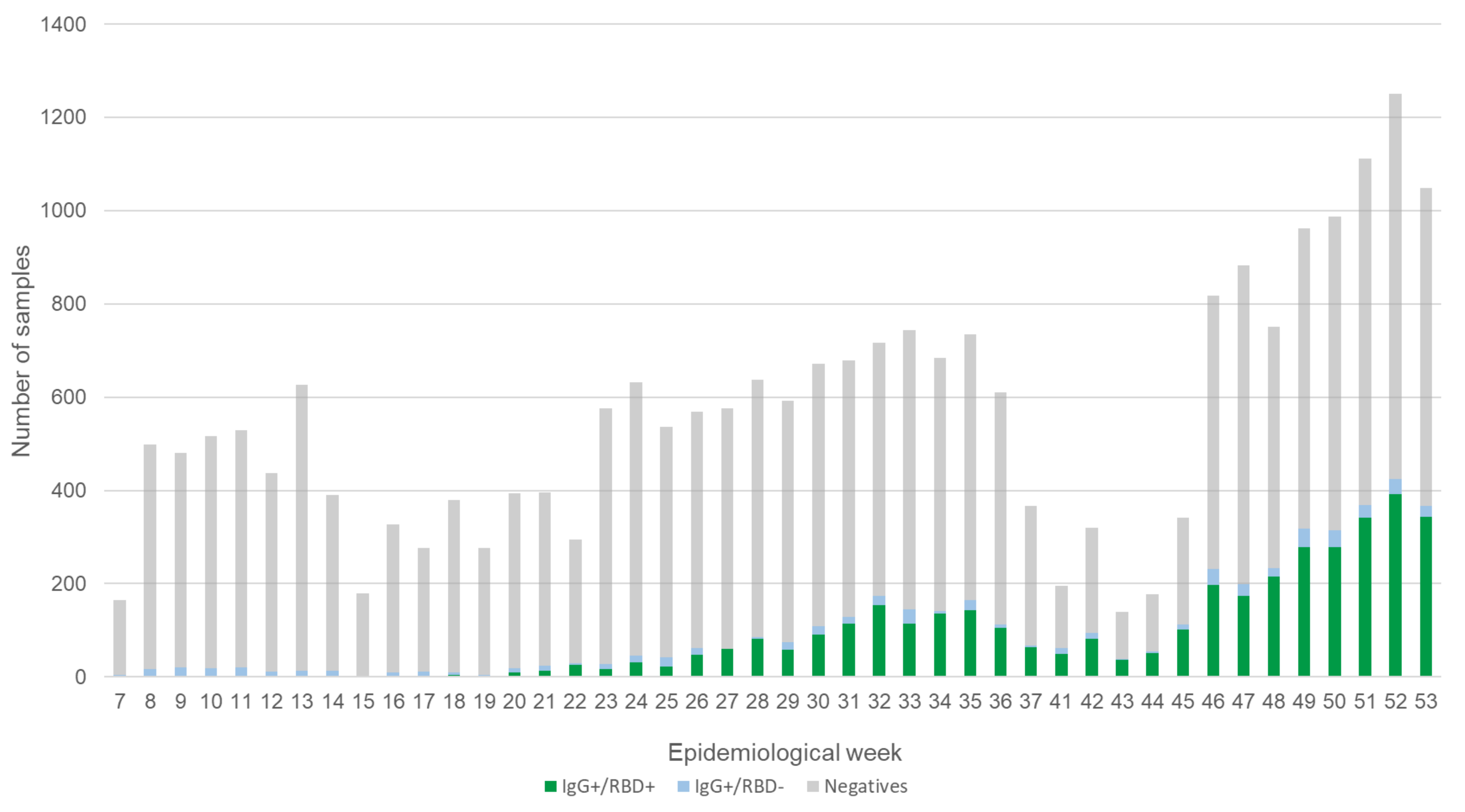
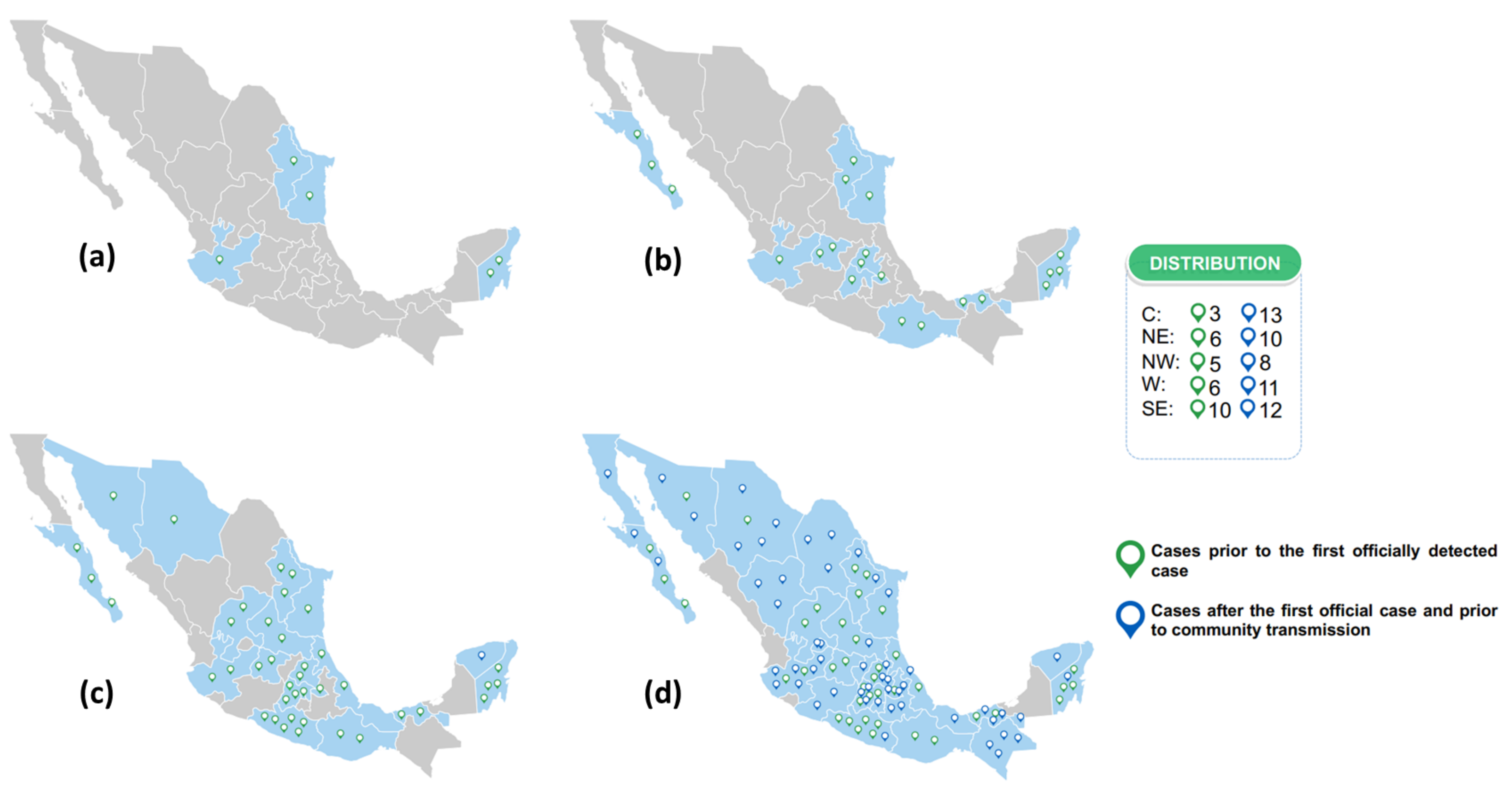
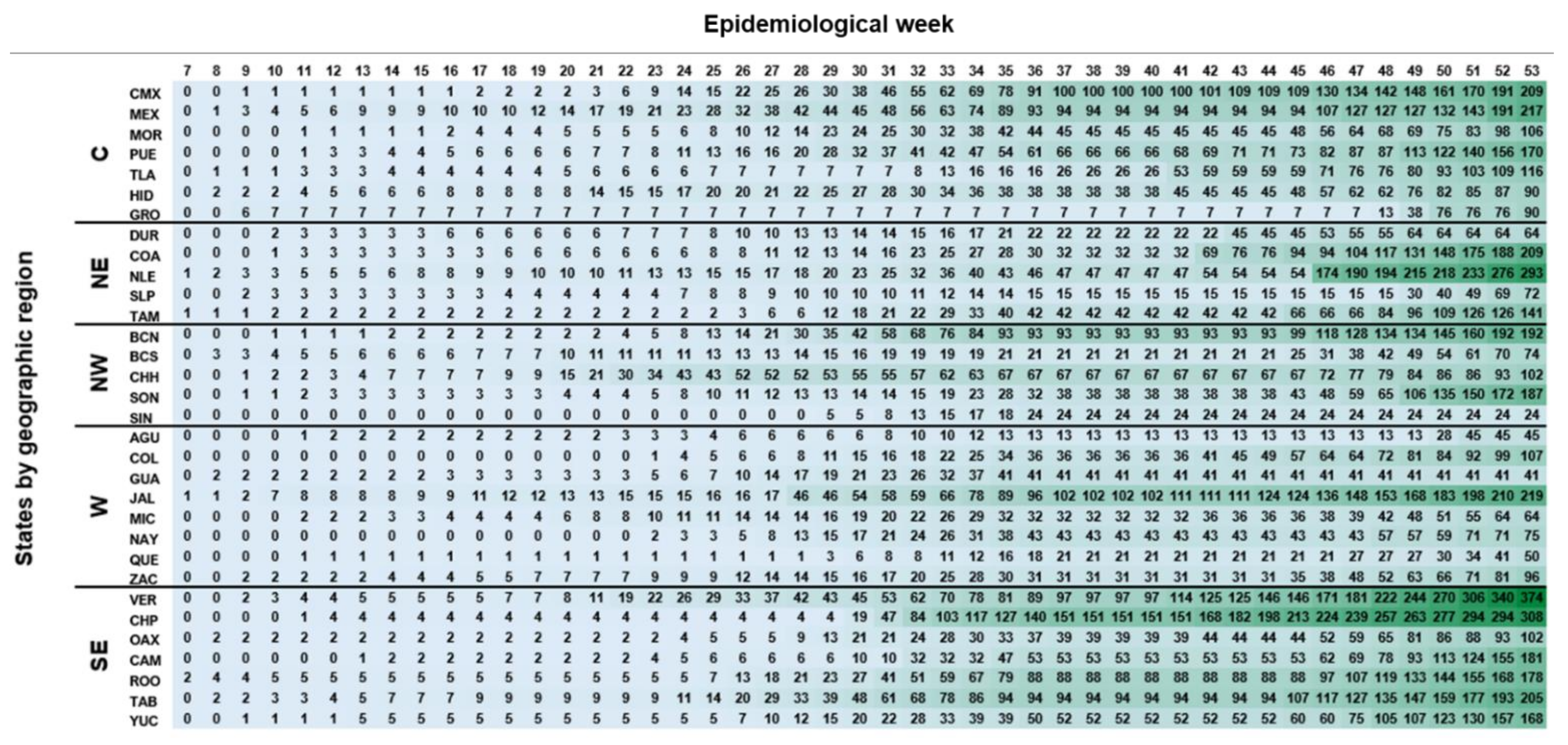
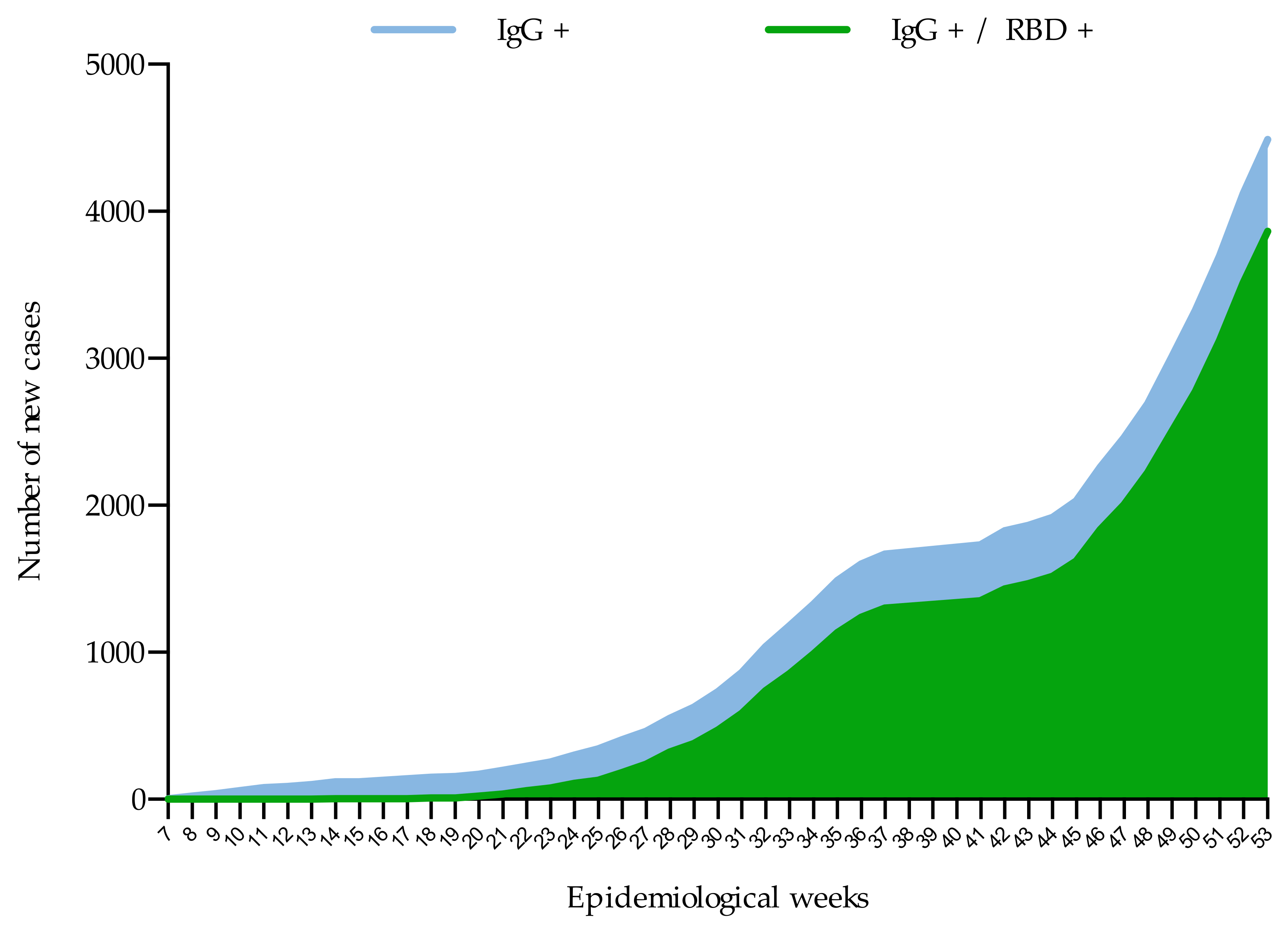
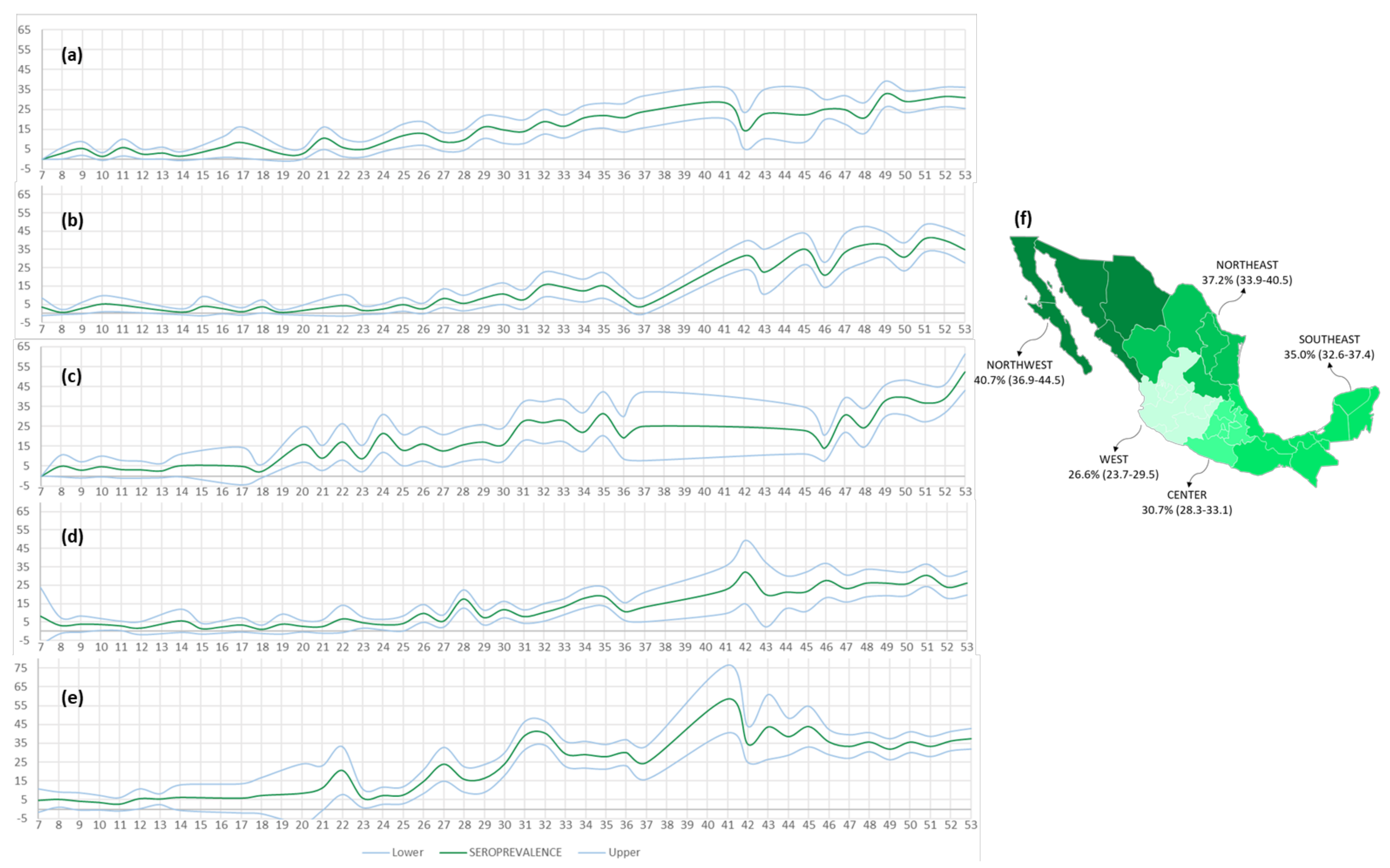
Dynamics and Extent of Infection (Cumulative Results from 44 Weeks)
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Acknowledgments
Conflicts of Interest
References
- Zhu, N.; Zhang, D.; Wang, W.; Li, X.; Yang, B.; Song, J.; Zhao, X.; Huang, B.; Shi, W.; Lu, R.; et al. A Novel Coronavirus from Patients with Pneumonia in China 2019. N. Engl. J. Med. 2020, 382, 727–733. [Google Scholar] [CrossRef] [PubMed]
- Huang, C.; Wang, Y.; Li, X.; Ren, L.; Zhao, J.; Hu, Y.; Zhang, L.; Fan, G.; Xu, J.; Gu, X.; et al. Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Lancet 2020, 395, 497–506. [Google Scholar] [CrossRef]
- World Health Organization. Coronavirus (COVID-19). Cases by Country, Territory or Area. 2020. Available online: https://who.sprinklr.com (accessed on 22 March 2021).
- Dirección General de Epidemiología. Informe Técnico Diario COVID-19 México. 2021. Available online: https://www.gob.mx/cms/uploads/attachment/file/603816/Comunicado_Tecnico_Diario_COVID-19_2020.12.31.pdf (accessed on 19 March 2021).
- Feng, W.; Newbigging, A.M.; Le, C.; Pang, B.; Peng, H.; Cao, Y.; Wu, J.; Abbas, G.; Song, J.; Wang, D.B.; et al. Molecular Diagnosis of COVID-19: Challenges and Research Needs. Anal. Chem. 2020, 92, 10196–10209. [Google Scholar] [CrossRef] [PubMed]
- Gobierno de México, Dirección General de Epidemiología. Lineamiento Estandarizado Para la Vigilancia Epidemiológica y Por Laboratorio de la Enfermedad Respiratoria Viral. 2020; Available online: Lineamiento_VE_Lab_enfermedad_respiratoria_viral-_20052020.pdf (coronavirus.gob.mx); (accessed on 5 April 2021). [Google Scholar]
- Ghaffari, A.; Meurant, R.; Ardakani, A. COVID-19 Serological Tests: How Well Do They Actually Perform? Diagnostics 2020, 10, 453. [Google Scholar] [CrossRef]
- Kubina, R.; Dziedzic, A. Molecular and Serological Tests for COVID-19. A Comparative Review of SARS-CoV-2 Coronavirus Laboratory and Point-of-Care Diagnostics. Diagnostics 2020, 10, 434. [Google Scholar] [CrossRef]
- World Health Organization. Population-Based Age-Stratified Seroepidemiological Investigation Protocol for Coronavirus 2019 (COVID-19) Infection; World Health Organization: Geneva, Switzerland, 2020. [Google Scholar]
- Centers for Disease Control and Prevention. Interim Guidelines for COVID-19 Antibody Testing. 2021. Available online: https://www.cdc.gov/coronavirus/2019-ncov/lab/resources/antibody-tests-guidelines.html#print. (accessed on 5 April 2021).
- Berlin, D.A.; Gulick, R.M.; Martinez, F.J. Severe Covid-19. N. Engl. J. Med. 2020, 383, 2451–2460. [Google Scholar] [CrossRef]
- Azer, S.A. COVID-19: Pathophysiology, diagnosis, complications and investigational therapeutics. New Microbes New Infect. 2020, 37, 100738. [Google Scholar] [CrossRef]
- Norma Oficial Mexicana. NOM-253-SSA-2012. Para la Disposición de Sangre Humana y sus Componentes con Fines Terapéuticos. 2012. Available online: https://www.gob.mx/cms/uploads/attachment/file/219069/NOM-253-SSA1-2012_Para_la_disposicion_de_sangre_humana_y_sus_componentes_con_fines_terapeuticos.pdf (accessed on 11 April 2021).
- Organización Mundial de la Salud. Guía Sobre la Reglamentación Relativa al Transporte de Sustancias Infecciosas 2019–2020; Organización Mundial de la Salud: Geneva, Switzerland, 2019. [Google Scholar]
- Bonelli, F.; Sarasini, A.; Zierold, C.; Calleri, M.; Bonetti, A.; Vismara, C.; Blocki, F.; Pallavicini, L.; Chinali, A.; Campisi, D.; et al. Clinical and analytical performance of an automated serological test that identifies S1/S2-neutralizing IgG in COVID-19 patients semiquantitatively. J. Clin. Microbiol. 2020, 58, 9. [Google Scholar] [CrossRef]
- Tan, C.W.; Chia, W.N.; Qin, X.; Liu, P.; Chen, M.I.C.; Tiu, C.; Hu, Z.; Chen, V.C.W.; Young, B.E.; Sia, W.R.; et al. A SARS-CoV-2 surrogate virus neutralization test based on antibody-mediated blockage of ACE2–spike protein–protein interaction. Nat. Biotechnol. 2020, 38, 1073–1078. [Google Scholar]
- Perico, L.; Tomasoni, S.; Peracchi, T.; Perna, A.; Pezzotta, A.; Remuzzi, G.; Benigni, A. COVID-19 and lombardy: TESTing the impact of the first wave of the pandemic. EBioMedicine. 2020, 61, 103069. [Google Scholar] [CrossRef]
- Apolone, G.; Montomoli, E.; Manenti, A.; Boeri, M.; Sabia, F.; Hyseni, I.; Mazzini, L.; Martinuzzi, D.; Cantone, L.; Milanese, G.; et al. Unexpected detection of SARS-CoV-2 antibodies in the prepandemic period in Italy. Tumori J. 2020, 11, 755. [Google Scholar] [CrossRef]
- Basavajaru, S.V.; Patton, M.E.; Grimm, K.; Rasheed, M.A.U.; Lester, S.; Mills, L.; Stumpf, M.; Freeman, B.; Tamin, A.; Harcourt, J.; et al. Serologic testing of U.S. blood donations to identify SARS-CoV-2-reactive antibodies: December 2019–January 2020. Clin. Infect. Dis. 2020, 30, 1785. [Google Scholar] [CrossRef]
- Stringhini, S.; Wisniak, A.; Piumatti, G.; Azman, A.S.; Lauer, S.A.; Baysson, H.; De Ridder, D.; Petrovic, D.; Schrempft, S.; Marcus, K.; et al. Seroprevalence of anti-SARS-CoV-2 IgG antibodies in Geneva, Switzerland (SEROCoV-POP): A population-based study. Lancet 2020, 396, 313–319. [Google Scholar] [CrossRef]
- Xu, X.; Sun, J.; Nie, S.; Li, H.; Kong, Y.; Liang, M.; Hou, J.; Huang, X.; Li, D.; Ma, T.; et al. Seroprevalence of immunoglobulin M and G antibodies against SARS-CoV-2 in China. Nat. Med. 2020, 26, 1193–1195. [Google Scholar] [CrossRef] [PubMed]
- Bryan, A.; Pepper, G.; Wener, M.H.; Fink, S.L.; Morishima, C.; Chaudhary, A.; Jerome, K.R.; Mathias, P.C.; Greninger, A.L. Performance characteristics of the Abbott Architect SARS-CoV-2 IgG assay and seroprevalence in Boise, Idaho. J. Clin. Microbiol. 2020, 58, e00941. [Google Scholar] [CrossRef]
- Dirección General de Epidemiología. Aviso Epidemiológico CONAVE/09/2020/COVID-19. 2020. Available online: https://www.gob.mx/cms/uploads/attachment/file/545445/AE_Enfermedad_COVID-19_SARS-CoV-2_2020.04.06.pdf (accessed on 25 November 2020).
- Taboada, B.; Vázquez-Pérez, J.A.; Muñoz-Medina, J.E.; Ramos-Cervantes, P.; Escalera-Zamudio, M.; Boukadida, C.; Sánchez-Flores, A.; Isa, P.; Mendieta-Condado, E.; Martínez-Orozco, J.A.; et al. Genomic Analysis of Early SARS-CoV-2 Variants Introduced in México. J. Virol. 2020, 94, e01056-20. [Google Scholar] [CrossRef]
- Remes-Troje, J.M.; Ramos-De-la-Medina, A.; Manríquez-Reyes, M.; Martínez-Pérez-Maldonado, L.; Solis-González, A.; Hernández-Flores, K.G.; Vivanco-Cid, H. Prevalence of SARS-CoV-2 IgG antibodies in a population from Veracruz (Southeastern Mexico). medRxiv 2020. [Google Scholar] [CrossRef]
- Secretaría de Salud. Available online: https://www.gob.mx/salud/prensa/255-secretaria-de-salud-presenta-resultados-preliminares-de-la-encuesta-nacional-de-salud-y-nutricion-covid-19?idiom=es (accessed on 22 March 2021).
- Havers, F.P.; Reed, C.; Lim, T.; Montgomery, J.M.; Klena, J.D.; Hall, A.J.; Fry, A.M.; Cannon, D.L.; Chiang, C.F.; Gibbons, A.; et al. Seroprevalence of antibodies to SARS-CoV-2 in 10 sites in the United States, March 23–May 12, 2020. JAMA Inter. Med. 2020, 21, 220. [Google Scholar] [CrossRef]
- Bajema, K.L.; Wiegand, R.E.; Cuffe, K.; Patel, S.V.; Iachan, R.; Lim, T.; Lee, A.; Moyse, D.; Havers, F.P.; Harding, L.; et al. Estimated SARS-CoV-2 Seroprevalence in the US as of September 2020. JAMA Intern. Med. 2020, 24, 460. [Google Scholar]
- Aguas, R.; Corder, R.M.; King, J.G.; Goncalves, G.; Ferreira, M.U.; Gomes, M.G.M. Herd immunity thresholds for SARS-CoV-2 estimated from unfolding epidemics. medRxiv 2020. [Google Scholar] [CrossRef]
- Gomes, M.G.M.; Corder, R.M.; King, J.G.; Langwig, K.E.; Souto-Maior, C.; Carneiro, J.; Goncalves, G.; Penha-Goncalves, C.; Ferreira, M.U.; Aguas, R. Individual variation in susceptibility or exposure to SARS-CoV-2 lowers the herd immunity threshold. medRxiv 2020. [Google Scholar] [CrossRef]
- Premkumar, L.; Segovia-Chumbez, B.; Jadi, R.; Martinez, D.R.; Raut, R.; Markmann, A.; Cornaby, C.; Bartelt, L.; Weiss, S.; Park, Y.; et al. The receptor-binding domain of the viral spike protein is an immunodominant and highly specific target of antibodies in SARS-CoV-2 patients. Sci. Immunol. 2020, 5, 8413. [Google Scholar] [CrossRef] [PubMed]
- Weisberg, S.P.; Connors, T.J.; Zhu, Y.; Baldwin, M.R.; Lin, W.H.; Wontakal, S.; Szabo, P.A.; Wells, S.B.; Dogra, P.; Gray, J.; et al. Distinct antibody responses to SARS-CoV-2 in children and adults across the COVID-19 clinical spectrum. Nat. Immunol. 2020, 22, 25–31. [Google Scholar] [CrossRef] [PubMed]
- Bosnjak, B.; Stein, S.C.; Willenzon, S.; Cordes, A.K.; Puppe, W.; Bernhardt, G.; Ravens, I.; Ritter, C.; Schultze-Florey, C.R.; Gödecke, N.; et al. Low serum neutralizing anti-SARS-CoV-2 S antibody levels in mild affected COVID-19 convalescent patients revealed by two different detection methods. Cell. Mol. Immunol. 2020, 19, 1–9. [Google Scholar]
| Region | Age Group | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Total | 0–20 Years | 21–40 Years | 41–60 Years | 61–80 Years | 81–100 Years | |||||||
| Pos/Total | % (CI) | Pos/Total | % (CI) | Pos/Total | % (CI) | Pos/Total | % (CI) | Pos/Total | % (CI) | Pos/Total | % (CI) | |
| C | 438/1428 | 30.7 (28.3–33.1) | 43/163 | 26.4 (19.6–33.1) | 166/430 | 38.6 (34.0–43.2) | 152/500 | 30.4 (26.4–34.4) | 61/248 | 24.6 (19.2–30.0) | 16/87 | 18.4 (10.3–26.5) |
| NE | 314/844 | 37.2 (33.9–40.5) | 41/121 | 33.9 (25.5–42.3) | 98/254 | 38.6 (32.6–44.6) | 105/263 | 39.9 (34.0–45.8) | 48/143 | 33.6 (25.8–41.3) | 22/63 | 34.9 (23.1–46.7) |
| NW | 267/656 | 40.7 (36.9–44.5) | 29/59 | 49.2 (36.4–61.9) | 92/229 | 40.2 (33.8–46.5) | 97/228 | 42.5 (36.1–49.0) | 37/107 | 34.6 (25.6–43.6) | 12/33 | 36.4 (20.0–52.8) |
| W | 240/902 | 26.6 (23.7–29.5) | 35/128 | 27.3 (19.6–35.1) | 91/306 | 29.7 (24.6–34.9) | 82/271 | 30.3 (24.8–35.7) | 27/139 | 19.4 (12.8–26.0) | 5/58 | 8.6 (1.4–15.8) |
| SE | 535/1529 | 35.0 (32.6–37.4) | 77/213 | 36.2 (29.7–42.6) | 216/540 | 40.0 (35.9–44.1) | 145/447 | 32.4 (28.1–36.8) | 67/220 | 30.5 (24.4–36.5) | 30/109 | 27.5 (19.1–35.9) |
| Total | 1794/5359 | 33.5 (32.2–34.7) | 225/684 | 32.9 (29.4–36.4) | 663/1759 | 37.7 (35.4–40.0) | 581/1709 | 34.0 (31.8–36.2) | 240/857 | 28.0 (25.0–31.0) | 85/350 | 24.3 (19.8–28.8) |
| Region | Age Group | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Total | 0–20 Years | 21–40 Years | 41–60 Years | 61–80 Years | 81–100 Years | |||||||
| Pos/Total | % (CI) | Pos/Total | % (CI) | Pos/Total | % (CI) | Pos/Total | % (CI) | Pos/Total | % (CI) | Pos/Total | % (CI) | |
| C | 401/1428 | 28.1 (25.8–30.4) | 40/163 | 24.5 (17.9–31.1) | 152/430 | 35.3 (30.8–39.9) | 139/500 | 27.8 (23.9–31.7) | 57/248 | 23.0 (17.7–28.2) | 13/86 | 15.1 (7.5–22.7) |
| NE | 292/844 | 34.6 (31.4–37.9) | 35/121 | 28.9 (20.8–37.0) | 92/254 | 36.2 (30.3–42.1) | 100/262 | 38.2 (32.3–44.1) | 47/143 | 32.9 (25.2–40.6) | 18/63 | 28.6 (17.4–39.7) |
| NW | 243/656 | 37.0 (32.6–40.0) | 26/59 | 44.1 (31.4–56.7) | 81/226 | 35.8 (29.6–42.1) | 86/225 | 38.2 (31.9–44.6) | 33/107 | 30.8 (22.1–39.6) | 10/33 | 30.3 (14.6–46.0) |
| W | 211/902 | 23.4 (20.6–26.2) | 32/128 | 25.0 (17.5–32.5) | 81/306 | 26.5 (21.5–31.4) | 72/271 | 26.6 (21.3–31.8) | 21/139 | 15.1 (9.2–21.1) | 5/58 | 8.6 (1.4–15.8) |
| SE | 494/1529 | 32.3 (30.0–34.7) | 70/212 | 33.0 (26.7–39.3) | 203/540 | 37.6 (33.5–41.7) | 131/446 | 29.4 (25.1–33.6) | 64/220 | 29.1 (23.1–35.1) | 26/109 | 23.9 (15.9–31.9) |
| Total | 1641/5359 | 30.6 (29.3–31.8) | 203/683 | 29.7 (26.3–33.1) | 609/1756 | 34.7 (32.5–36.9) | 528/1704 | 31.1 (28.8–33.2) | 222/857 | 25.9 (23.0–28.8) | 72/349 | 20.6 (16.4–24.9) |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Muñoz-Medina, J.E.; Grajales-Muñiz, C.; Salas-Lais, A.G.; Fernandes-Matano, L.; López-Macías, C.; Monroy-Muñoz, I.E.; Santos Coy-Arechavaleta, A.; Palomec-Nava, I.D.; Duque-Molina, C.; Madera-Sandoval, R.L.; et al. SARS-CoV-2 IgG Antibodies Seroprevalence and Sera Neutralizing Activity in MEXICO: A National Cross-Sectional Study during 2020. Microorganisms 2021, 9, 850. https://doi.org/10.3390/microorganisms9040850
Muñoz-Medina JE, Grajales-Muñiz C, Salas-Lais AG, Fernandes-Matano L, López-Macías C, Monroy-Muñoz IE, Santos Coy-Arechavaleta A, Palomec-Nava ID, Duque-Molina C, Madera-Sandoval RL, et al. SARS-CoV-2 IgG Antibodies Seroprevalence and Sera Neutralizing Activity in MEXICO: A National Cross-Sectional Study during 2020. Microorganisms. 2021; 9(4):850. https://doi.org/10.3390/microorganisms9040850
Chicago/Turabian StyleMuñoz-Medina, José Esteban, Concepción Grajales-Muñiz, Angel Gustavo Salas-Lais, Larissa Fernandes-Matano, Constantino López-Macías, Irma Eloísa Monroy-Muñoz, Andrea Santos Coy-Arechavaleta, Iliana Donají Palomec-Nava, Célida Duque-Molina, Ruth Lizzeth Madera-Sandoval, and et al. 2021. "SARS-CoV-2 IgG Antibodies Seroprevalence and Sera Neutralizing Activity in MEXICO: A National Cross-Sectional Study during 2020" Microorganisms 9, no. 4: 850. https://doi.org/10.3390/microorganisms9040850
APA StyleMuñoz-Medina, J. E., Grajales-Muñiz, C., Salas-Lais, A. G., Fernandes-Matano, L., López-Macías, C., Monroy-Muñoz, I. E., Santos Coy-Arechavaleta, A., Palomec-Nava, I. D., Duque-Molina, C., Madera-Sandoval, R. L., Rivero-Arredondo, V., González-Ibarra, J., Alvarado-Yaah, J. E., Rojas-Mendoza, T., Santacruz-Tinoco, C. E., González-Bonilla, C. R., & Borja-Aburto, V. H. (2021). SARS-CoV-2 IgG Antibodies Seroprevalence and Sera Neutralizing Activity in MEXICO: A National Cross-Sectional Study during 2020. Microorganisms, 9(4), 850. https://doi.org/10.3390/microorganisms9040850









